Epigenomic Control of Thermogenic Adipocyte Differentiation and Function
Abstract
1. Introduction
2. Genome-Wide Studies on Histone Modifications during Thermogenic Adipogenesis
3. Genome-Wide Studies on Chromatin Remodeling during Thermogenic Adipogenesis
4. Genome-Wide Studies on DNA Methylation during Thermogenic Adipogenesis
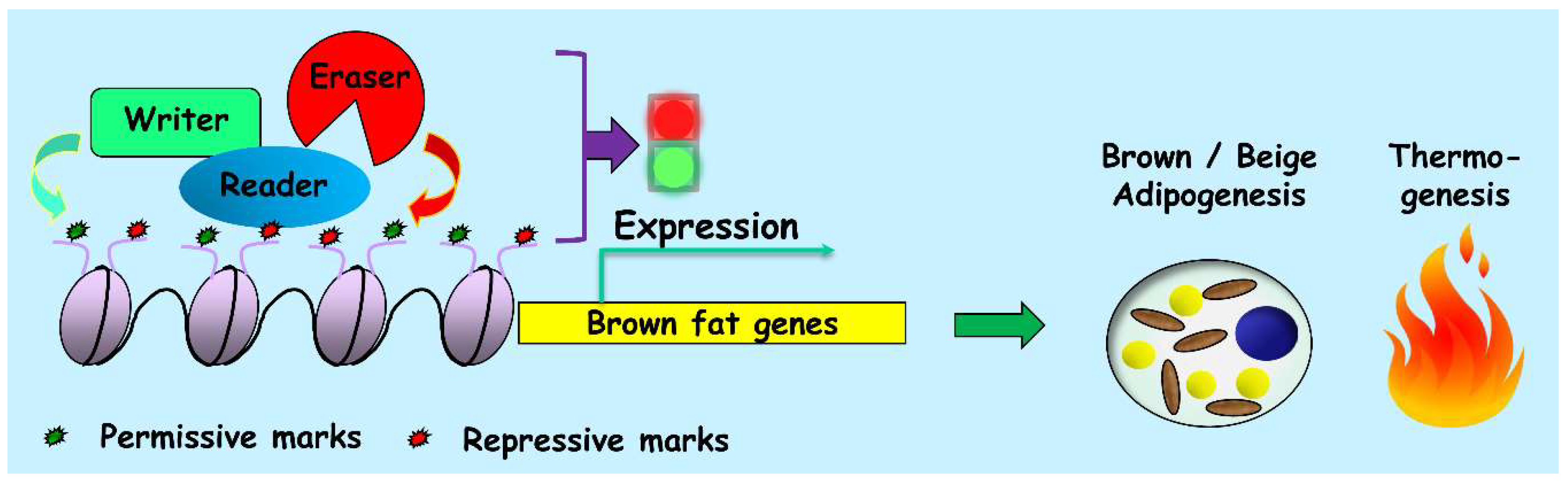
5. Genome-Wide Studies on the Writers, Erasers and Readers of the Epigenome during Thermogenic Adipogenesis
6. Novel Regulators of Thermogenic Adipogenesis Identified through Epigenomic Studies
7. Genome-Wide Studies on Chromatin Modifications and Interactions in Disease Models
8. Concluding Remarks
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| BAT | Brown adipose tissue |
| WAT | White adipose tissue |
| eWAT | Epididymal white adipose tissue |
| PET-CT | Positron emission tomography-computed tomography |
| MSC | Mesenchymal stem cell |
| PTM | Post-translational modification |
| PCA | Principal component analysis |
| hMADS | Human multipotent adipose-derived stem |
| ChIP-seq | Chromatin immunoprecipitation-sequencing |
| FAIRE-seq | Formaldehyde-assisted isolation of regulatory elements-sequencing |
| ATAC-seq | Assays for transposase-accessible chromatin-sequencing |
| 4C-seq | Circular chromosome conformation capture-sequencing |
| RRBS | Reduced representation bisulfite sequencing |
| FACS | Fluorescence-activated cell sorting |
| SNP | Single-nucleotide polymorphism |
| GWAS | Genome-wide association study |
| TF | Transcription factor |
| UCP1 | Uncoupling protein 1 |
| PPARγ | Peroxisome Proliferator Activated Receptor Gamma |
| EBF2 | Early B cell factor-2 |
| HOXC10 | Homeobox C10 |
| CBP | CREB-Binding protein |
| MLL4 | Mixed-lineage leukemia 4 |
| PRDM16 | PR domain containing 16 |
| CPT1b | Carnitine palmitoyltransferase IB |
| NFIA | Nuclear factor I/A |
| IL-10 | Interleukin 10 |
| KLF11 | Kruppel-like factor 11 |
| LSD1 | Lysine-specific demethylase 1 |
| IRX3 | Iroquois homeobox 3 |
| JMJD3 | Jumonji domain containing protein 3 |
| SWI/SNF | SWItch/sucrose non-fermentable |
| BRD4 | Bromodomain-containing protein 4 |
| PKA | Protein kinase A |
| GR | Glucocorticoid receptor |
| SIX1 | SIX Homeobox 1 |
| RREB1 | Ras-responsive element-binding protein 1 |
| PIM1 | Pim-1 Proto-Oncogene, Serine/Threonine Kinase |
| DLC1 | Deleted in liver cancer 1 |
| FTO | Fat mass and obesity-associated |
| FGF21 | Fibroblast growth factor 21 |
| CoREST | Co-repressor to RE1 silencing transcription factor |
| NRF1 | Nuclear respiratory factor 1 |
References
- Seale, P.; Bjork, B.; Yang, W.; Kajimura, S.; Chin, S.; Kuang, S.; Scime, A.; Devarakonda, S.; Conroe, H.M.; Erdjument-Bromage, H.; et al. PRDM16 controls a brown fat/skeletal muscle switch. Nature 2008, 454, 961–967. [Google Scholar] [CrossRef] [PubMed]
- Sanchez-Gurmaches, J.; Guertin, D.A. Adipocytes arise from multiple lineages that are heterogeneously and dynamically distributed. Nat. Commun. 2014, 5, 4099. [Google Scholar] [CrossRef] [PubMed]
- Peirce, V.; Carobbio, S.; Vidal-Puig, A. The different shades of fat. Nature 2014, 510, 76–83. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Bostrom, P.; Sparks, L.M.; Ye, L.; Choi, J.H.; Giang, A.H.; Khandekar, M.; Virtanen, K.A.; Nuutila, P.; Schaart, G.; et al. Beige adipocytes are a distinct type of thermogenic fat cell in mouse and human. Cell 2012, 150, 366–376. [Google Scholar] [CrossRef] [PubMed]
- Ikeda, K.; Maretich, P.; Kajimura, S. The common and distinct features of brown and beige adipocytes. Trends Endocrinol. Metab. 2018, 29, 191–200. [Google Scholar] [CrossRef] [PubMed]
- Van Marken Lichtenbelt, W.D.; Vanhommerig, J.W.; Smulders, N.M.; Drossaerts, J.M.; Kemerink, G.J.; Bouvy, N.D.; Schrauwen, P.; Teule, G.J. Cold-activated brown adipose tissue in healthy men. N. Engl. J. Med. 2009, 360, 1500–1508. [Google Scholar] [CrossRef] [PubMed]
- Cypess, A.M.; Lehman, S.; Williams, G.; Tal, I.; Rodman, D.; Goldfine, A.B.; Kuo, F.C.; Palmer, E.L.; Tseng, Y.H.; Doria, A.; et al. Identification and importance of brown adipose tissue in adult humans. N. Engl. J. Med. 2009, 360, 1509–1517. [Google Scholar] [CrossRef] [PubMed]
- Virtanen, K.A.; Lidell, M.E.; Orava, J.; Heglind, M.; Westergren, R.; Niemi, T.; Taittonen, M.; Laine, J.; Savisto, N.J.; Enerback, S.; et al. Functional brown adipose tissue in healthy adults. N. Engl. J. Med. 2009, 360, 1518–1525. [Google Scholar] [CrossRef] [PubMed]
- Saito, M.; Okamatsu-Ogura, Y.; Matsushita, M.; Watanabe, K.; Yoneshiro, T.; Nio-Kobayashi, J.; Iwanaga, T.; Miyagawa, M.; Kameya, T.; Nakada, K.; et al. High incidence of metabolically active brown adipose tissue in healthy adult humans: Effects of cold exposure and adiposity. Diabetes 2009, 58, 1526–1531. [Google Scholar] [CrossRef] [PubMed]
- Bartelt, A.; Bruns, O.T.; Reimer, R.; Hohenberg, H.; Ittrich, H.; Peldschus, K.; Kaul, M.G.; Tromsdorf, U.I.; Weller, H.; Waurisch, C.; et al. Brown adipose tissue activity controls triglyceride clearance. Nat. Med. 2011, 17, 200–205. [Google Scholar] [CrossRef] [PubMed]
- Stanford, K.I.; Middelbeek, R.J.; Townsend, K.L.; An, D.; Nygaard, E.B.; Hitchcox, K.M.; Markan, K.R.; Nakano, K.; Hirshman, M.F.; Tseng, Y.H.; et al. Brown adipose tissue regulates glucose homeostasis and insulin sensitivity. J. Clin. Investig. 2013, 123, 215–223. [Google Scholar] [CrossRef] [PubMed]
- Connolly, E.; Morrisey, R.D.; Carnie, J.A. The effect of interscapular brown adipose tissue removal on body-weight and cold response in the mouse. Br. J. Nutr. 1982, 47, 653–658. [Google Scholar] [CrossRef] [PubMed]
- Hamann, A.; Flier, J.S.; Lowell, B.B. Decreased brown fat markedly enhances susceptibility to diet-induced obesity, diabetes, and hyperlipidemia. Endocrinology 1996, 137, 21–29. [Google Scholar] [CrossRef] [PubMed]
- Lowell, B.B.; V, S.S.; Hamann, A.; Lawitts, J.A.; Himms-Hagen, J.; Boyer, B.B.; Kozak, L.P.; Flier, J.S. Development of obesity in transgenic mice after genetic ablation of brown adipose tissue. Nature 1993, 366, 740–742. [Google Scholar] [CrossRef] [PubMed]
- Feldmann, H.M.; Golozoubova, V.; Cannon, B.; Nedergaard, J. Ucp1 ablation induces obesity and abolishes diet-induced thermogenesis in mice exempt from thermal stress by living at thermoneutrality. Cell Metab. 2009, 9, 203–209. [Google Scholar] [CrossRef] [PubMed]
- Tontonoz, P.; Hu, E.; Spiegelman, B.M. Stimulation of adipogenesis in fibroblasts by ppar gamma 2, a lipid-activated transcription factor. Cell 1994, 79, 1147–1156. [Google Scholar] [CrossRef]
- Farmer, S.R. Transcriptional control of adipocyte formation. Cell Metab. 2006, 4, 263–273. [Google Scholar] [CrossRef] [PubMed]
- Bracken, A.P.; Dietrich, N.; Pasini, D.; Hansen, K.H.; Helin, K. Genome-wide mapping of polycomb target genes unravels their roles in cell fate transitions. Genes Dev. 2006, 20, 1123–1136. [Google Scholar] [CrossRef] [PubMed]
- Bowers, R.R.; Kim, J.W.; Otto, T.C.; Lane, M.D. Stable stem cell commitment to the adipocyte lineage by inhibition of DNA methylation: Role of the BMP-4 gene. Proc. Natl. Acad. Sci. USA 2006, 103, 13022–13027. [Google Scholar] [CrossRef] [PubMed]
- Heintzman, N.D.; Hon, G.C.; Hawkins, R.D.; Kheradpour, P.; Stark, A.; Harp, L.F.; Ye, Z.; Lee, L.K.; Stuart, R.K.; Ching, C.W.; et al. Histone modifications at human enhancers reflect global cell-type-specific gene expression. Nature 2009, 459, 108–112. [Google Scholar] [CrossRef] [PubMed]
- Heintzman, N.D.; Stuart, R.K.; Hon, G.; Fu, Y.; Ching, C.W.; Hawkins, R.D.; Barrera, L.O.; Van Calcar, S.; Qu, C.; Ching, K.A.; et al. Distinct and predictive chromatin signatures of transcriptional promoters and enhancers in the human genome. Nat. Genet. 2007, 39, 311–318. [Google Scholar] [CrossRef] [PubMed]
- Rada-Iglesias, A.; Bajpai, R.; Swigut, T.; Brugmann, S.A.; Flynn, R.A.; Wysocka, J. A unique chromatin signature uncovers early developmental enhancers in humans. Nature 2011, 470, 279–283. [Google Scholar] [CrossRef] [PubMed]
- Creyghton, M.P.; Cheng, A.W.; Welstead, G.G.; Kooistra, T.; Carey, B.W.; Steine, E.J.; Hanna, J.; Lodato, M.A.; Frampton, G.M.; Sharp, P.A.; et al. Histone H3K27ac separates active from poised enhancers and predicts developmental state. Proc. Natl. Acad. Sci. USA 2010, 107, 21931–21936. [Google Scholar] [CrossRef] [PubMed]
- Sun, L.; Xie, H.; Mori, M.A.; Alexander, R.; Yuan, B.; Hattangadi, S.M.; Liu, Q.; Kahn, C.R.; Lodish, H.F. Mir193b-365 is essential for brown fat differentiation. Nat. Cell Biol. 2011, 13, 958–965. [Google Scholar] [CrossRef] [PubMed]
- Adaikalakoteswari, A.; Vatish, M.; Alam, M.T.; Ott, S.; Kumar, S.; Saravanan, P. Low vitamin B12 in pregnancy is associated with adipose-derived circulating mirs targeting ppargamma and insulin resistance. J. Clin. Endocrinol. Metab. 2017, 102, 4200–4209. [Google Scholar] [CrossRef] [PubMed]
- Oldenburg, A.; Briand, N.; Sorensen, A.L.; Cahyani, I.; Shah, A.; Moskaug, J.O.; Collas, P. A lipodystrophy-causing lamin a mutant alters conformation and epigenetic regulation of the anti-adipogenic MIR335 locus. J. Cell Biol. 2017, 216, 2731–2743. [Google Scholar] [CrossRef] [PubMed]
- Nardelli, C.; Iaffaldano, L.; Pilone, V.; Labruna, G.; Ferrigno, M.; Carlomagno, N.; Dodaro, C.A.; Forestieri, P.; Buono, P.; Salvatore, F.; et al. Changes in the microrna profile observed in the subcutaneous adipose tissue of obese patients after laparoscopic adjustable gastric banding. J. Obes. 2017, 2017, 6754734. [Google Scholar] [CrossRef] [PubMed]
- Alexander, R.; Lodish, H.; Sun, L. Micrornas in adipogenesis and as therapeutic targets for obesity. Expert Opin. Ther. Targets 2011, 15, 623–636. [Google Scholar] [CrossRef] [PubMed]
- Shamsi, F.; Zhang, H.; Tseng, Y.H. Microrna regulation of brown adipogenesis and thermogenic energy expenditure. Front. Endocrinol. 2017, 8, 205. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.; Sabari, B.R.; Garcia, B.A.; Allis, C.D.; Zhao, Y. Snapshot: Histone modifications. Cell 2014, 159, 458–458 e451. [Google Scholar] [CrossRef] [PubMed]
- Pan, D.; Huang, L.; Zhu, L.J.; Zou, T.; Ou, J.; Zhou, W.; Wang, Y.X. Jmjd3-mediated H3K27me3 dynamics orchestrate brown fat development and regulate white fat plasticity. Dev. Cell 2015, 35, 568–583. [Google Scholar] [CrossRef] [PubMed]
- Brunmeir, R.; Wu, J.; Peng, X.; Kim, S.Y.; Julien, S.G.; Zhang, Q.; Xie, W.; Xu, F. Comparative transcriptomic and epigenomic analyses reveal new regulators of murine brown adipogenesis. PLoS Genet. 2016, 12, e1006474. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.E.; Wang, C.; Xu, S.; Cho, Y.W.; Wang, L.; Feng, X.; Baldridge, A.; Sartorelli, V.; Zhuang, L.; Peng, W.; et al. H3K4 mono- and di-methyltransferase MLL4 is required for enhancer activation during cell differentiation. eLife 2013, 2, e01503. [Google Scholar] [CrossRef] [PubMed]
- Lai, B.; Lee, J.E.; Jang, Y.; Wang, L.; Peng, W.; Ge, K. MLL3/MLL4 are required for CBP/p300 binding on enhancers and super-enhancer formation in brown adipogenesis. Nucleic Acids Res. 2017, 45, 6388–6403. [Google Scholar] [CrossRef] [PubMed]
- Shi, Y.; Lan, F.; Matson, C.; Mulligan, P.; Whetstine, J.R.; Cole, P.A.; Casero, R.A. Histone demethylation mediated by the nuclear amine oxidase homolog lsd1. Cell 2004, 119, 941–953. [Google Scholar] [CrossRef] [PubMed]
- Zeng, X.; Jedrychowski, M.P.; Chen, Y.; Serag, S.; Lavery, G.G.; Gygi, S.P.; Spiegelman, B.M. Lysine-specific demethylase 1 promotes brown adipose tissue thermogenesis via repressing glucocorticoid activation. Genes Dev. 2016, 30, 1822–1836. [Google Scholar] [CrossRef] [PubMed]
- Roh, H.C.; Tsai, L.T.Y.; Shao, M.; Tenen, D.; Shen, Y.; Kumari, M.; Lyubetskaya, A.; Jacobs, C.; Dawes, B.; Gupta, R.K.; et al. Warming induces significant reprogramming of beige, but not brown, adipocyte cellular identity. Cell Metab. 2018, 27, 1121–1137. [Google Scholar] [CrossRef] [PubMed]
- Abe, Y.; Rozqie, R.; Matsumura, Y.; Kawamura, T.; Nakaki, R.; Tsurutani, Y.; Tanimura-Inagaki, K.; Shiono, A.; Magoori, K.; Nakamura, K.; et al. JMJD1A is a signal-sensing scaffold that regulates acute chromatin dynamics via SWI/SNF association for thermogenesis. Nat. Commun. 2015, 6, 7052. [Google Scholar] [CrossRef] [PubMed]
- Tateishi, K.; Okada, Y.; Kallin, E.M.; Zhang, Y. Role of Jhdm2a in regulating metabolic gene expression and obesity resistance. Nature 2009, 458, 757–761. [Google Scholar] [CrossRef] [PubMed]
- Loft, A.; Forss, I.; Siersbaek, M.S.; Schmidt, S.F.; Larsen, A.S.; Madsen, J.G.; Pisani, D.F.; Nielsen, R.; Aagaard, M.M.; Mathison, A.; et al. Browning of human adipocytes requires KLF11 and reprogramming of PPARgamma superenhancers. Genes Dev. 2015, 29, 7–22. [Google Scholar] [CrossRef] [PubMed]
- Waki, H.; Nakamura, M.; Yamauchi, T.; Wakabayashi, K.; Yu, J.; Hirose-Yotsuya, L.; Take, K.; Sun, W.; Iwabu, M.; Okada-Iwabu, M.; et al. Global mapping of cell type-specific open chromatin by faire-seq reveals the regulatory role of the nfi family in adipocyte differentiation. PLoS Genet. 2011, 7, e1002311. [Google Scholar] [CrossRef] [PubMed]
- Hiraike, Y.; Waki, H.; Yu, J.; Nakamura, M.; Miyake, K.; Nagano, G.; Nakaki, R.; Suzuki, K.; Kobayashi, H.; Yamamoto, S.; et al. NFIA co-localizes with PPARgamma and transcriptionally controls the brown fat gene program. Nat. Cell Biol. 2017, 19, 1081–1092. [Google Scholar] [CrossRef] [PubMed]
- Rajbhandari, P.; Thomas, B.J.; Feng, A.C.; Hong, C.; Wang, J.; Vergnes, L.; Sallam, T.; Wang, B.; Sandhu, J.; Seldin, M.M.; et al. IL-10 signaling remodels adipose chromatin architecture to limit thermogenesis and energy expenditure. Cell 2018, 172, 218–233.e217. [Google Scholar] [CrossRef] [PubMed]
- Kiskinis, E.; Hallberg, M.; Christian, M.; Olofsson, M.; Dilworth, S.M.; White, R.; Parker, M.G. RIP140 directs histone and DNA methylation to silence Ucp1 expression in white adipocytes. EMBO J. 2007, 26, 4831–4840. [Google Scholar] [CrossRef] [PubMed]
- Shore, A.; Karamitri, A.; Kemp, P.; Speakman, J.R.; Lomax, M.A. Role of Ucp1 enhancer methylation and chromatin remodelling in the control of Ucp1 expression in murine adipose tissue. Diabetologia 2010, 53, 1164–1173. [Google Scholar] [CrossRef] [PubMed]
- Derecka, M.; Gornicka, A.; Koralov, S.B.; Szczepanek, K.; Morgan, M.; Raje, V.; Sisler, J.; Zhang, Q.; Otero, D.; Cichy, J.; et al. Tyk2 and Stat3 regulate brown adipose tissue differentiation and obesity. Cell Metab. 2012, 16, 814–824. [Google Scholar] [CrossRef] [PubMed]
- Lim, Y.C.; Chia, S.Y.; Jin, S.; Han, W.; Ding, C.; Sun, L. Dynamic DNA methylation landscape defines brown and white cell specificity during adipogenesis. Mol. Metab. 2016, 5, 1033–1041. [Google Scholar] [CrossRef] [PubMed]
- Yue, F.; Cheng, Y.; Breschi, A.; Vierstra, J.; Wu, W.; Ryba, T.; Sandstrom, R.; Ma, Z.; Davis, C.; Pope, B.D.; et al. A comparative encyclopedia of DNA elements in the mouse genome. Nature 2014, 515, 355–364. [Google Scholar] [CrossRef] [PubMed]
- Park, Y.K.; Ge, K. Glucocorticoid receptor accelerates, but is dispensable for, adipogenesis. Mol. Cell. Biol. 2017, 37. [Google Scholar] [CrossRef] [PubMed]
- Roh, H.C.; Tsai, L.T.; Lyubetskaya, A.; Tenen, D.; Kumari, M.; Rosen, E.D. Simultaneous transcriptional and epigenomic profiling from specific cell types within heterogeneous tissues in vivo. Cell Rep. 2017, 18, 1048–1061. [Google Scholar] [CrossRef] [PubMed]
- Harms, M.J.; Lim, H.W.; Ho, Y.; Shapira, S.N.; Ishibashi, J.; Rajakumari, S.; Steger, D.J.; Lazar, M.A.; Won, K.J.; Seale, P. PRDM16 binds MED1 and controls chromatin architecture to determine a brown fat transcriptional program. Genes Dev. 2015, 29, 298–307. [Google Scholar] [CrossRef] [PubMed]
- Pradhan, R.N.; Bues, J.J.; Gardeux, V.; Schwalie, P.C.; Alpern, D.; Chen, W.; Russeil, J.; Raghav, S.K.; Deplancke, B. Dissecting the brown adipogenic regulatory network using integrative genomics. Sci. Rep. 2017, 7, 42130. [Google Scholar] [CrossRef] [PubMed]
- Kurdistani, S.K.; Grunstein, M. Histone acetylation and deacetylation in yeast. Nat. Rev. Mol. Cell Biol. 2003, 4, 276–284. [Google Scholar] [CrossRef] [PubMed]
- Shahbazian, M.D.; Grunstein, M. Functions of site-specific histone acetylation and deacetylation. Annu. Rev. Biochem. 2007, 76, 75–100. [Google Scholar] [CrossRef] [PubMed]
- Bannister, A.J.; Kouzarides, T. Regulation of chromatin by histone modifications. Cell Res. 2011, 21, 381–395. [Google Scholar] [CrossRef] [PubMed]
- Jenuwein, T.; Allis, C.D. Translating the histone code. Science 2001, 293, 1074–1080. [Google Scholar] [CrossRef] [PubMed]
- Owen, D.J.; Ornaghi, P.; Yang, J.C.; Lowe, N.; Evans, P.R.; Ballario, P.; Neuhaus, D.; Filetici, P.; Travers, A.A. The structural basis for the recognition of acetylated histone H4 by the bromodomain of histone acetyltransferase gcn5p. EMBO J. 2000, 19, 6141–6149. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, P.R.; Nietlispach, D.; Mott, H.R.; Callaghan, J.; Bannister, A.; Kouzarides, T.; Murzin, A.G.; Murzina, N.V.; Laue, E.D. Structure of the HP1 chromodomain bound to histone H3 methylated at lysine 9. Nature 2002, 416, 103–107. [Google Scholar] [CrossRef] [PubMed]
- Jacobs, S.A.; Khorasanizadeh, S. Structure of HP1 chromodomain bound to a lysine 9-methylated histone H3 tail. Science 2002, 295, 2080–2083. [Google Scholar] [CrossRef] [PubMed]
- Bannister, A.J.; Zegerman, P.; Partridge, J.F.; Miska, E.A.; Thomas, J.O.; Allshire, R.C.; Kouzarides, T. Selective recognition of methylated lysine 9 on histone H3 by the HP1 chromo domain. Nature 2001, 410, 120–124. [Google Scholar] [CrossRef] [PubMed]
- Duteil, D.; Tosic, M.; Lausecker, F.; Nenseth, H.Z.; Muller, J.M.; Urban, S.; Willmann, D.; Petroll, K.; Messaddeq, N.; Arrigoni, L.; et al. Lsd1 ablation triggers metabolic reprogramming of brown adipose tissue. Cell Rep. 2016, 17, 1008–1021. [Google Scholar] [CrossRef] [PubMed]
- Abe, Y.; Fujiwara, Y.; Takahashi, H.; Matsumura, Y.; Sawada, T.; Jiang, S.; Nakaki, R.; Uchida, A.; Nagao, N.; Naito, M.; et al. Histone demethylase JMJD1A coordinates acute and chronic adaptation to cold stress via thermogenic phospho-switch. Nat. Commun. 2018, 9, 1566. [Google Scholar] [CrossRef] [PubMed]
- Loven, J.; Hoke, H.A.; Lin, C.Y.; Lau, A.; Orlando, D.A.; Vakoc, C.R.; Bradner, J.E.; Lee, T.I.; Young, R.A. Selective inhibition of tumor oncogenes by disruption of super-enhancers. Cell 2013, 153, 320–334. [Google Scholar] [CrossRef] [PubMed]
- Roe, J.S.; Mercan, F.; Rivera, K.; Pappin, D.J.; Vakoc, C.R. Bet bromodomain inhibition suppresses the function of hematopoietic transcription factors in acute myeloid leukemia. Mol. Cell 2015, 58, 1028–1039. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.E.; Park, Y.K.; Park, S.; Jang, Y.; Waring, N.; Dey, A.; Ozato, K.; Lai, B.; Peng, W.; Ge, K. Brd4 binds to active enhancers to control cell identity gene induction in adipogenesis and myogenesis. Nat. Commun. 2017, 8, 2217. [Google Scholar] [CrossRef] [PubMed]
- Rajakumari, S.; Wu, J.; Ishibashi, J.; Lim, H.W.; Giang, A.H.; Won, K.J.; Reed, R.R.; Seale, P. Ebf2 determines and maintains brown adipocyte identity. Cell Metab. 2013, 17, 562–574. [Google Scholar] [CrossRef] [PubMed]
- Whyte, W.A.; Orlando, D.A.; Hnisz, D.; Abraham, B.J.; Lin, C.Y.; Kagey, M.H.; Rahl, P.B.; Lee, T.I.; Young, R.A. Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell 2013, 153, 307–319. [Google Scholar] [CrossRef] [PubMed]
- Sim, C.K.; Kim, S.Y.; Brunmeir, R.; Zhang, Q.; Li, H.; Dharmasegaran, D.; Leong, C.; Lim, Y.Y.; Han, W.; Xu, F. Regulation of white and brown adipocyte differentiation by RhoGAP DLC1. PLoS ONE 2017, 12, e0174761. [Google Scholar] [CrossRef] [PubMed]
- Ng, R.; Hussain, N.A.; Zhang, Q.; Chang, C.; Li, H.; Fu, Y.; Cao, L.; Han, W.; Stunkel, W.; Xu, F. Mirna-32 drives brown fat thermogenesis and trans-activates subcutaneous white fat browning in mice. Cell Rep. 2017, 19, 1229–1246. [Google Scholar] [CrossRef] [PubMed]
- Smemo, S.; Tena, J.J.; Kim, K.H.; Gamazon, E.R.; Sakabe, N.J.; Gomez-Marin, C.; Aneas, I.; Credidio, F.L.; Sobreira, D.R.; Wasserman, N.F.; et al. Obesity-associated variants within FTO form long-range functional connections with IRX3. Nature 2014, 507, 371–375. [Google Scholar] [CrossRef] [PubMed]
- Claussnitzer, M.; Dankel, S.N.; Kim, K.H.; Quon, G.; Meuleman, W.; Haugen, C.; Glunk, V.; Sousa, I.S.; Beaudry, J.L.; Puviindran, V.; et al. Fto obesity variant circuitry and adipocyte browning in humans. N. Engl. J. Med. 2015, 373, 895–907. [Google Scholar] [CrossRef] [PubMed]
- Ng, Y.; Tan, S.X.; Chia, S.Y.; Tan, H.Y.; Gun, S.Y.; Sun, L.; Hong, W.; Han, W. Hoxc10 suppresses browning of white adipose tissues. Exp. Mol. Med. 2017, 49, e292. [Google Scholar] [CrossRef] [PubMed]
- Kang, S.; Tsai, L.T.; Zhou, Y.; Evertts, A.; Xu, S.; Griffin, M.J.; Issner, R.; Whitton, H.J.; Garcia, B.A.; Epstein, C.B.; et al. Identification of nuclear hormone receptor pathways causing insulin resistance by transcriptional and epigenomic analysis. Nat. Cell Biol. 2015, 17, 44–56. [Google Scholar] [CrossRef] [PubMed]

| Chromatin Markers/Regions | Function | Systems | Refs |
|---|---|---|---|
| H3K4me1 | Enhancer priming | Immortalized primary brown pre-adipocytes; C3H10T1/2 mesenchymal stem cells (MSCs); brown adipose tissue (BAT); FACS sorted cell type-specific nuclei from cold/warm beige and brown adipocytes | [32,33,34,36,37,48,49,50] |
| H3K4me2 | Gene activation | Immortalized primary brown pre-adipocytes; BAT | [33,34,36] |
| H3K4me3 | Promoter activation | Immortalized primary brown preadipocytes; C3H10T1/2 MSCs; BAT | [32,33,34,38,48,51] |
| H3K9me2 | Gene repression | Immortalized primary brown pre-adipocytes; | [34] |
| H3K27me3 | Gene repression | Immortalized primary brown pre-adipocytes; C3H10T1/2 MSCs; | [31,32,34] |
| H3K36me3 | Transcriptional elongation | Immortalized primary brown pre-adipocytes; | [34] |
| H3K9ac | Gene activation | C3H10T1/2 MSCs | [32] |
| H3K27ac | Enhancer activation | Immortalized primary brown pre-adipocytes; BAT; C3H10T1/2 MSCs; FACS sorted cell type-specific nuclei from cold/warm beige and brown adipocytes; human multipotent adipose-derived stem (hMADS) cells | [32,33,34,37,38,40,42,48,49,50,51,52] |
| Open chromatin region (FAIRE-seq (formaldehyde-assisted isolation of regulatory elements sequencing) and ATAC-seq (assays for transposase-accessible chromatin sequencing)) | Active promoters and enhancers | Immortalized primary brown preadipocytes; BAT; Immortalized beige pre-adipocytes | [34,42,43] |
| DNA methylation | Gene repression | Primary brown pre-adipocytes | [47] |
| Chromatin Factors | Function | Systems | Refs |
|---|---|---|---|
| MLL4 (mixed-lineage leukemia 4) | H3K4me1/2 methylase (writer) | Immortalized primary brown pre-adipocytes | [33,34] |
| LSD1 (lysine-specific demethylase 1) | H3K4me1/2 demethylase (eraser) | Brown adipocytes; BAT | [36,61] |
| JMJD1A (Jumonji domain containing 3) | H3K9me2 demethylase (eraser) | Immortalized primary brown pre-adipocytes | [38] |
| CBP (Carnitine palmitoyltransferase) | H3K27ac acetylase (writer) | Immortalized primary brown pre-adipocytes; hMADS cells | [34,40,49] |
| BRD4 | Reader of lysine acetylation | Immortalized primary brown pre-adipocytes | [65] |
| Regulators | Function | Approaches | Refs |
|---|---|---|---|
| SIX1 | Promotes brown adipogenesis | Binding motif search in enhancers | [32] |
| RREB1 | Promotes brown adipogenesis | Binding motif search in H3K27me3 peaks; Super-enhancer association analysis | [31,32] |
| KLF11 | Promotes browning in human brite adipocytes | Super-enhancer association analysis | [40] |
| PIM1 | Promotes brown adipogenesis | Super-enhancer association analysis | [32] |
| miR-32 | Promotes brown fat thermogenesis and white fat browning | Super-enhancer association analysis | [69] |
| IRX3 | Represses white fat browning | Long-range chromatin interaction analysis | [70,71] |
| NFIA | Promotes brown adipogenesis | Open chromatin region analysis | [42] |
| HOXC10 | Represses white fat browning | DNA methylation analysis | [47] |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Peng, X.; Zhang, Q.; Liao, C.; Han, W.; Xu, F. Epigenomic Control of Thermogenic Adipocyte Differentiation and Function. Int. J. Mol. Sci. 2018, 19, 1793. https://doi.org/10.3390/ijms19061793
Peng X, Zhang Q, Liao C, Han W, Xu F. Epigenomic Control of Thermogenic Adipocyte Differentiation and Function. International Journal of Molecular Sciences. 2018; 19(6):1793. https://doi.org/10.3390/ijms19061793
Chicago/Turabian StylePeng, Xu, Qiongyi Zhang, Cheng Liao, Weiping Han, and Feng Xu. 2018. "Epigenomic Control of Thermogenic Adipocyte Differentiation and Function" International Journal of Molecular Sciences 19, no. 6: 1793. https://doi.org/10.3390/ijms19061793
APA StylePeng, X., Zhang, Q., Liao, C., Han, W., & Xu, F. (2018). Epigenomic Control of Thermogenic Adipocyte Differentiation and Function. International Journal of Molecular Sciences, 19(6), 1793. https://doi.org/10.3390/ijms19061793






