The Expensive-Tissue Hypothesis in Vertebrates: Gut Microbiota Effect, a Review
Abstract
1. Introduction
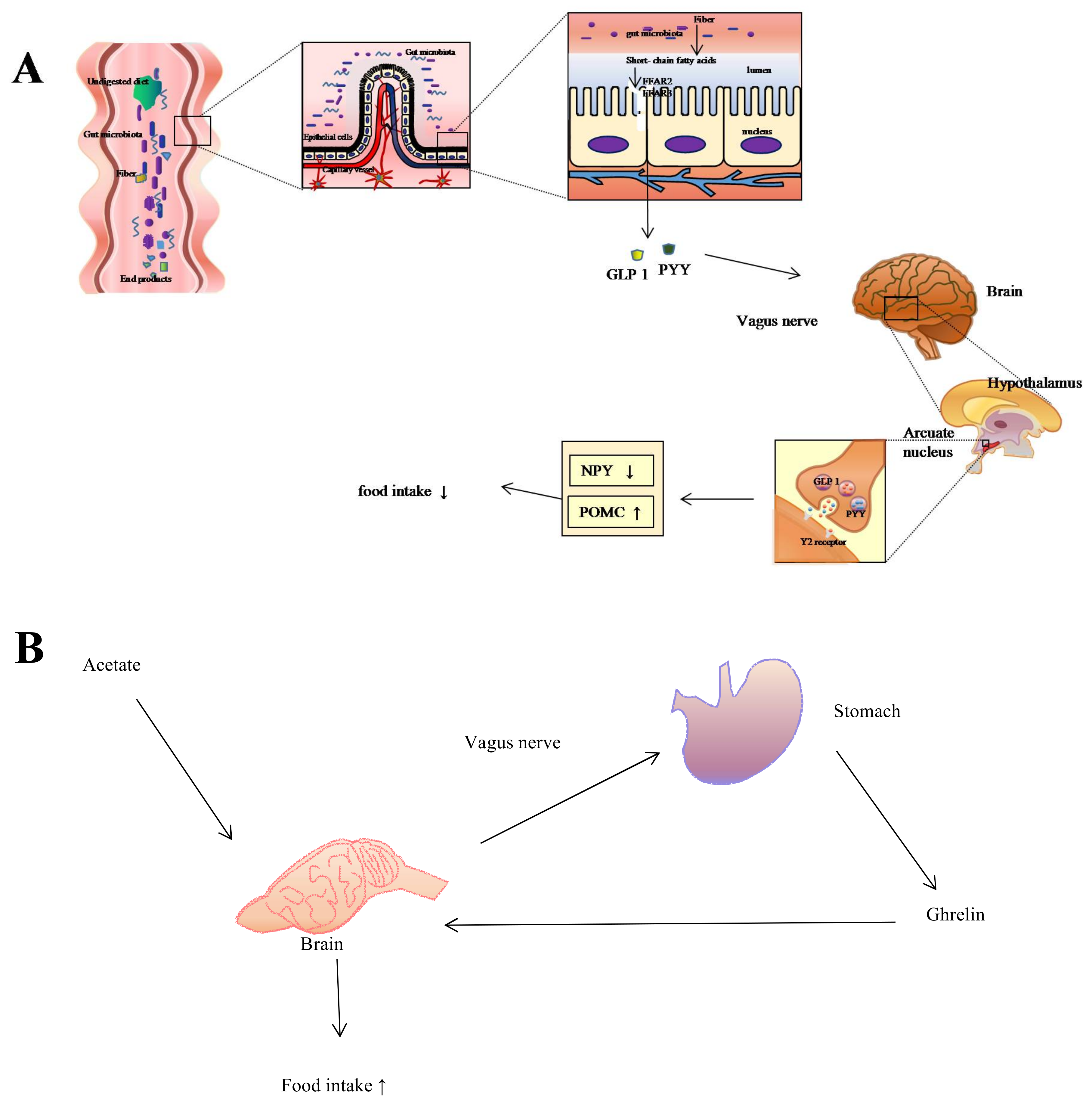
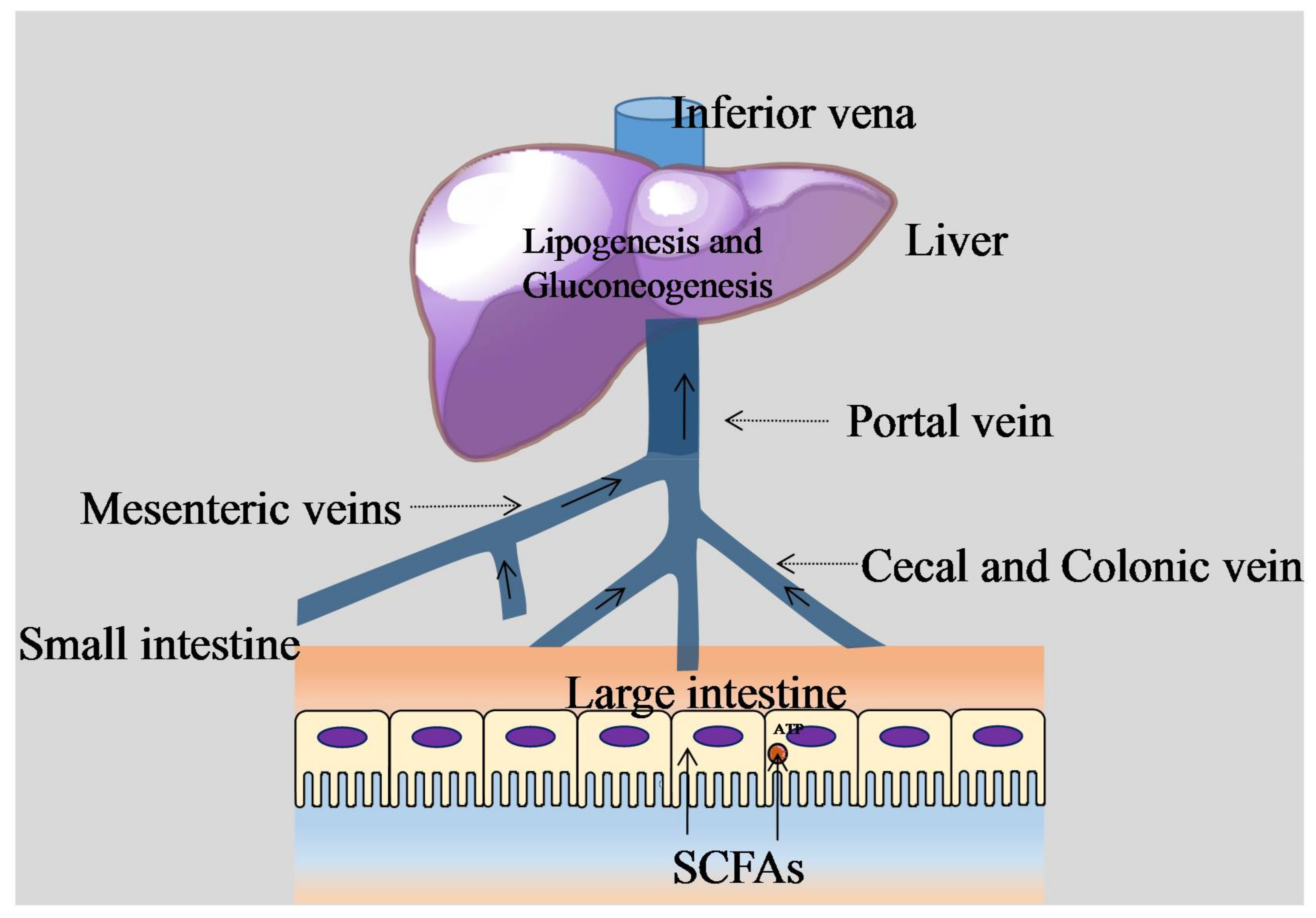
2. Influences of Gut Microbiota on Host Digestive System
3. Gut Microbiota Associated with Diet Affects Gut Size
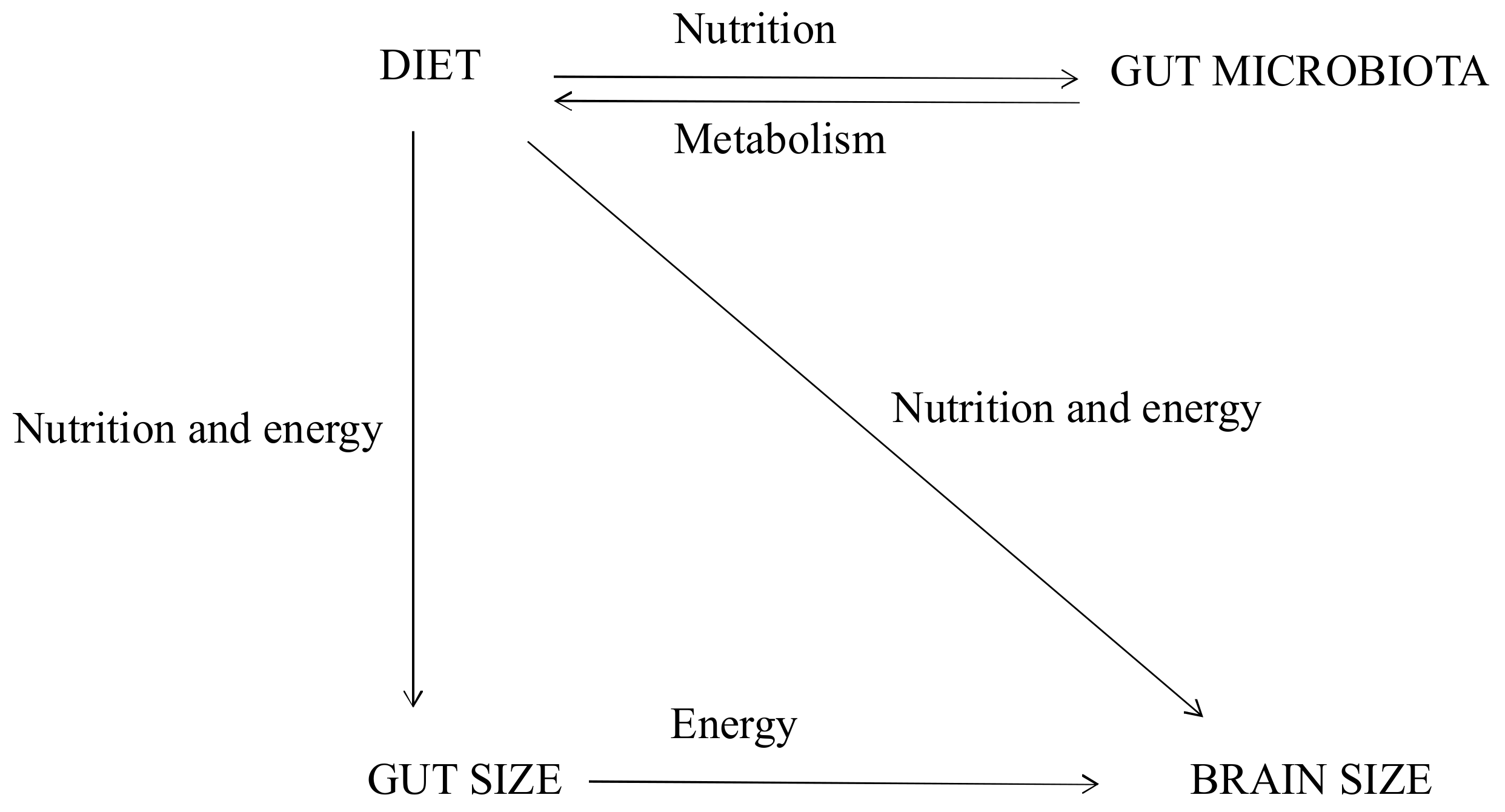
4. The Expensive-Tissue Hypothesis (ETH): Gut Size and Brain Size
5. Diet, Gut Microbiota, and the Trade-Off between Gut and Brain Size
6. Future Perspectives
7. Conclusions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| ETH | the Expensive-Tissue Hypothesis |
| SCFAs | short-chain fatty acids |
| FFAR2 | free fatty acid receptor 2 |
| FFAR3 | free fatty acid receptor 3 |
| PYY | peptide YY |
| NPY | neuropeptide Y |
| GLP1 | glucagon-like peptide 1 |
| POMC | anorexigenic pro-opiomelanocortin |
| HDAC | histone deacetylase |
| CK2 | casein kinase II |
| Sp1 | specificity proteins 1 |
| Sp3 | specificity proteins 3 |
| GF | germ free |
| CV | conventional |
References
- Gordon, J.I. Honor thy gut symbionts redux. Science 2012, 336, 1251–1253. [Google Scholar] [CrossRef] [PubMed]
- Smith, P.A. The tantalizing links between gut microbes and the brain. Nature 2015, 526, 312–314. [Google Scholar] [CrossRef] [PubMed]
- Hosokawa, T.; Kikuchi, Y.; Nikoh, N.T.; Shimada, M.; Fukatsu, T. Strict host-symbiont cospeciation and reductive genome evolution in insect gut bacteria. PLoS Biol. 2006, 4, 1841–1851. [Google Scholar] [CrossRef] [PubMed]
- Dominguez-Bello, M.G.; Gordon, J.I. Delivery mode shapes the acquisition and structure of the initial microbiota across multiple body habitats. Proc. Natl. Acad. Sci. USA 2010, 107, 11971–11975. [Google Scholar] [CrossRef] [PubMed]
- Nicholson, J.K.; Holmes, E.; Kinross, J.; Burcelin, R.; Gibson, G.; Jia, W.; Pettersson, S. Host-gut microbiota metabolic interactions. Science 2012, 336, 1262–1267. [Google Scholar] [CrossRef] [PubMed]
- Trajkovski, M.; Wollheim, C.B. Physiology: Microbial signals to the brain control weight. Nature 2016, 534, 185–187. [Google Scholar] [CrossRef] [PubMed]
- Sonnenburg, J.L.; Bäckhed, F. Diet-microbiota interactions as moderators of human metabolism. Nature 2016, 535, 56–64. [Google Scholar] [CrossRef] [PubMed]
- De Filippo, C.; Cavalieri, D.; Di Paola, M.; Ramazzotti, M.; Poullet, J.B.; Massart, S.; Collini, S.; Pieraccini, G.; lionetti, P. Impact of diet in shaping gut microbiota revealed by a comparative study in children from Europe and rural Africa. Proc. Natl. Acad. Sci. USA 2010, 107, 14691–14696. [Google Scholar] [CrossRef] [PubMed]
- Wu, G.D.; Chen, J.; Hoffmann, C.; Bittinger, K.; Chen, Y.Y.; Keilbaugh, S.A.; Bewtra, M.; Kinghts, D.; Walters, W.A.; Knight, R.; et al. Linking long-term dietary patterns with gut microbial enterotypes. Science 2011, 334, 105–108. [Google Scholar] [CrossRef] [PubMed]
- Zhu, L.; Wu, Q.; Dai, J.; Zhang, S.; Wei, F. Evidence of cellulose metabolism by the giant panda gut microbiome. Proc. Natl. Acad. Sci. USA 2011, 108, 17714–17719. [Google Scholar] [CrossRef] [PubMed]
- David, L.A.; Maurice, C.F.; Carmody, R.N.; Gootenberg, D.B.; Button, J.E.; Wolfe, B.E.; Ling, A.V.; Devlin, A.S.; Varman, Y.; Fischbach, M.A.; et al. Diet rapidly and reproducibly alters the human gut microbiome. Nature 2014, 505, 559–563. [Google Scholar] [CrossRef] [PubMed]
- Bletz, M.C.; Goedbloed, D.J; Sanchez, E.; Reinhardt, T.; Tebbe, C.C.; Bhuju, S.; Geffers, R.; Jarek, M.; Vences, M.; Steinfartz, S. Amphibian gut microbiota shifts differentially in community structure but converges on habitat-specific predicted functions. Nat. Commun. 2016, 7, 13699. [Google Scholar] [CrossRef] [PubMed]
- Chivers, D.J.; Hladik, C.M. Morphology of the gastrointestinal tract in primates: Comparisons with other mammals in relation to diet. J. Morphol. 1980, 166, 337–386. [Google Scholar] [CrossRef] [PubMed]
- Moss, R. Effects of captivity on gut lengths in red grouse. J. Wildl. Manag. 1972, 36, 99–104. [Google Scholar] [CrossRef]
- Aiello, L.C.; Wheeler, P. The expensive-tissue hypothesis: The brain and the digestive system in human and primate evolution. Curr. Anthropol. 1995, 36, 199–221. [Google Scholar] [CrossRef]
- Hladik, C.M.; Chivers, D.J.; Pasquet, P. On Diet and Gut Size in Non-human Primates and Humans: Is There a Relationship to Brain Size? Curr. Anthropol. 1999, 40, 695–697. [Google Scholar] [PubMed]
- Savory, C.J.; Gentle, M.J. Changes in food intake and gut size in Japanese quail in response to manipulation of dietary fibre content. Br. Poult. Sci. 1976, 17, 571–580. [Google Scholar] [CrossRef] [PubMed]
- Sullam, K.E.; Dalton, C.M.; Russell, J.A.; Kilham, S.S.; EI-Sabaawi, R.; German, D.P.; Flecker, A.S. Changes in digestive traits and body nutritional composition accommodate a trophic niche shift in Trinidadian guppies. Oecologia 2015, 177, 245–257. [Google Scholar] [CrossRef] [PubMed]
- Xi, X.X. Correlations between the Gut Microbiota of Newborns and Microbial Communities of Multiple Body Habitats in Mothers; Inner Mongolia Agricultural University: Hohhot, China, 2017. [Google Scholar]
- Gordon, H.A.; Bruckner-Kardoss, E. Effect of normal microbial flora on intestinal surface area. Am. J. Physiol. 1961, 201, 175–178. [Google Scholar] [CrossRef] [PubMed]
- Alam, M.; Midtvedt, T.; Uribe, A. Differential cell kinetics in the ileum and colon of germfree rats. Scand. J. Gastroenterol. 1994, 29, 445–451. [Google Scholar] [CrossRef] [PubMed]
- Wostmann, B.S. The germfree animal in nutritional studies. Annu. Rev. Nutr. 1981, 1, 257–279. [Google Scholar] [CrossRef] [PubMed]
- Sommer, F.; Bäckhed, F. The gut microbiota—Masters of host development and physiology. Nat. Rev. Microbiol. 2013, 11, 227–238. [Google Scholar] [CrossRef] [PubMed]
- Perry, R.J.; Peng, L.; Barry, N.A.; Cline, G.W.; Zhang, D.; Cardone, R.L.; Petersen, K.F.; Kibbey, R.G.; Goodman, A.L.; Shulman, G.I. Acetate mediates a microbiome-brain-β-cell axis to promote metabolic syndrome. Nature 2016, 534, 213–217. [Google Scholar] [CrossRef] [PubMed]
- Cummings, J.H; Pomare, E.W.; Branch, W.J.; Naylor, C.P.; Macfarlane, G.T. Short chain fatty acids in human large intestine, portal, hepatic and venous blood. Gut 1987, 28, 1221–1227. [Google Scholar] [CrossRef] [PubMed]
- Demigne, C.; Remesy, C.; Morand, C. Short chain fatty acids. In Colonic Microbiota, Nutrition and Health; Gibson, G.R., Roberfroid, M.B., Eds.; Springer: Hiderberg, Germany, 1999; pp. 55–69. [Google Scholar]
- Scheppach, W. Effects of short chain fatty acids on gut morphology and function. Gut 1994, 35, S35–S38. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.H.; Shen, R.-L.; Li, W.Q. Production and function of short chain fatty acids. Anim. Husb. Vet. Sci. Technol. Inf. 2007, 2, 12–13. [Google Scholar]
- Xu, Y.J.; Fang, R.J. The nutritional physiology of short chain fatty acids. Fodd. Res. 2007, 8, 26–28. [Google Scholar]
- Eckburg, P.B.; Bik, E.M.; Bernstein, C.N.; Purdom, E.; Dethlefsen, L.; Sargent, M.; Gill, S.R.; Nelson, K.E.; Relman, D.A. Diversity of the human intestinal microbial flora. Science 2005, 308, 1635–1638. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Bjursell, M.K.; Himrod, J.; Deng, S.; Carmichael, L.K.; Chiang, H.C.; Hooper, L.V.; Gordon, J.I. A genomic view of the human-Bacteroides thetaiotaomicron symbiosis. Science 2003, 299, 2074–2076. [Google Scholar] [CrossRef] [PubMed]
- Martens, E.C.; Lowe, E.C.; Chiang, H.; Pudlo, N.A.; Wu, M.; McNulty, N.P.; Abbott, D.W.; Henrissat, B.; Gilbert, H.J.; Bolam, D.N.; et al. Recognition and degradation of plant cell wall polysaccharides by two human gut symbionts. PLoS Biol. 2011, 9, e1001221. [Google Scholar] [CrossRef] [PubMed]
- Tremaroli, V.; Bäckhed, F. Functional interactions between the gut microbiota and host metabolism. Nature 2012, 489, 242–249. [Google Scholar] [CrossRef] [PubMed]
- Van Wey, A.S.; Cookson, A.L.; Roy, N.C.; McNabb, W.C.; Soboleva, T.K.; Shorten, P.R. Bacterial biofilms associated with food particles in the human large bowel. Mol. Nutr. Food Res. 2011, 55, 969–978. [Google Scholar] [CrossRef] [PubMed]
- Ze, X.; Duncan, S.H.; Louis, P.; Flint, H.J. Ruminococcus bromii is a keystone species for the degradation of resistant starch in the human colon. ISME J. 2012, 6, 1535–1543. [Google Scholar] [CrossRef] [PubMed]
- Liao, W.B.; Lou, S.L.; Zeng, Y.; Kotrschal, A. Large brains, small guts: The expensive tissue hypothesis supported in anurans. Am. Nat. 2016, 188, 693–700. [Google Scholar] [CrossRef] [PubMed]
- Kaufman, J.A.; Hladik, C.M.; Pasquet, P. On the expensive tissue hypothesis: Independent support from highly encephalized fish. Curr. Anthropol. 2003, 44, 705–707. [Google Scholar] [CrossRef]
- Tsuboi, M.; Husby, A.; Kotrschal, A.; Hayward, A.; Buechel, S.D.; Zidar, J.; Løvlie, H.; Kolm, N. Comparative support for the expensive tissue hypothesis: Big brains are correlated with smaller gut and greater parental investment in Lake Tanganyika cichlids. Evolution 2015, 69, 190–200. [Google Scholar] [CrossRef] [PubMed]
- Aiello, L.C.; Bates, N.; Joffe, T. Defense of the Expensive Tissue Hypothesis: Ontogeny, Maternal Care and Organ Size. In Evolutionary Anatomy of the Primate Cerebral Cortex; Falk, D., Gibson, K.R., Eds.; Cambridge University Press: Cambridge, UK, 2001; pp. 57–78. [Google Scholar]
- Jin, L.; Liu, W.C.; Li, Y.H.; Zeng, Y.; Liao, W.B. Evidence for the expensive-tissue hypothesis in the Omei Wood Frog (Rana omeimontis). Herpetol. J. 2015, 25, 127–130. [Google Scholar]
- Kotrschal, A.; Corral-Lopez, A.; Szidat, S.; Kolm, N. The effect of brain size evolution on feeding propensity, digestive efficiency, and juvenile growth. Evolution 2015, 69, 3013–3020. [Google Scholar] [CrossRef] [PubMed]
- Aiello, L.C.; Wells, J.C.K. Energetics and the evolution of the genus Homo. Annu. Rev. Anthropol. 2002, 31, 323–338. [Google Scholar] [CrossRef]
- Isler, K.; van Schaik, C.P. The Expensive Brain: A framework for explaining evolutionary changes in brain size. J. Hum. Evol. 2009, 57, 392–400. [Google Scholar] [CrossRef] [PubMed]
- Isler, K.; van Schaik, C.P. Metabolic costs of brain size evolution. Biol. Lett. 2006, 2, 557–560. [Google Scholar] [CrossRef] [PubMed]
- Isler, K.; van Schaik, C.P. Costs of encephalization: The energy trade-off hypothesis tested on birds. J. Hum. Evol. 2006, 51, 228–243. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Zhou, C.Q.; Liao, W.B. Evidence for neither the compensation hypothesis nor the expensive-tissue hypothesis in Carassius auratus. Anim. Biol. 2014, 64, 177–187. [Google Scholar] [CrossRef]
- Navarrete, A.; van Schaik, C.P.; Isler, K. Energetics and the evolution of human brain size. Nature 2011, 480, 91–93. [Google Scholar] [CrossRef] [PubMed]
- Yu, X.; Zhong, M.J.; Li, D.Y.; Jin, L.; Liao, W.B.; Kotrschal, A. Large-brained frogs mature later and live longer. Evolution 2018. [Google Scholar] [CrossRef] [PubMed]
- Pitnick, S.; Jones, K.E.; Wilkinson, G.S. Mating system and brain size in bats. Proc. R. Soc. B 2006, 273, 719–724. [Google Scholar] [CrossRef] [PubMed]
- Kotrschal, A.; Rogell, B.; Bundsen, A.; Svensson, B.; Zajitschek, S.; Brännström, I.; Immler, S.; Maklakov, A.A.; Kolm, N. Artificial selection on relative brain size in the guppy reveals costs and benefits of evolving a larger brain. Curr. Biol. 2013, 23, 168–171. [Google Scholar] [CrossRef] [PubMed]
- Jiang, A.; Zhong, M.J.; Xie, M.; Lou, S.L.; Jin, L.; Robert, J.; Liao, W.B. Seasonality and age is positively related to brain size in Andrew’s toad (Bufo andrewsi). Evol. Biol. 2015, 42, 339–348. [Google Scholar] [CrossRef]
- Crawford, M.A. The role of dietary fatty acids in biology: Their place in the evolution of the human brain. Nutr. Rev. 1992, 50, 3–11. [Google Scholar] [CrossRef] [PubMed]
- Goyal, M.S.; Venkatesh, S.; Milbrandt, J.; Gordon, J.I; Raichle, M.E. Feeding the brain and nurturing the mind: Linking nutrition and the gut microbiota to brain development. Proc. Natl. Acad. Sci. USA 2015, 112, 14105–14112. [Google Scholar] [CrossRef] [PubMed]
- Sender, R.; Fuchs, S.; Milo, R. Are we really vastly outnumbered? Revisiting the ratio of bacterial to host cells in humans. Cell 2016, 164, 337–340. [Google Scholar] [CrossRef] [PubMed]
- Prado-Irwin, S.R.; Bird, A.K.; Zink, A.G.; Vredenburg, V.T. Intraspecific variation in the skin-associated microbiome of a terrestrial salamander. Microb. Ecol. 2017, 74, 45–756. [Google Scholar] [CrossRef] [PubMed]
- Ley, R.E.; Lozupone, C.A.; Hamady, M.; Knight, R.; Gordon, J.I. Worlds within worlds: Evolution of the vertebrate gut microbiota. Nat. Rev. Microbiol. 2008, 6, 776–788. [Google Scholar] [CrossRef] [PubMed]
- Bestion, E.; Jacob, S.; Zinger, L.; Gesu, L.D.; Richard, M.; White, J.; Cote, J. Climate warming reduces gut microbiota diversity in a vertebrate ectotherm. Nat. Ecol. Evol. 2017, 1, 016. [Google Scholar] [CrossRef] [PubMed]
- Flint, H.J.; Scott, K.P.; Louis, P.; Duncan, S.H. The role of the gut microbiota in nutrition and health. Nat. Rev. Gastroenterol. Hepatol. 2012, 9, 577–589. [Google Scholar] [CrossRef] [PubMed]
- Eģģesbø, M.; Moen, B.; Peddada, S.; Baird, D.; Rugtveit, J.; Midtvedt, T.; Bushel, P.R.; Sekelja, M.; Rudi, K. Development of gut microbiota in infants not exposed to medical interventions. Apmis 2011, 119, 17–35. [Google Scholar] [CrossRef] [PubMed]
- Schiffrin, E.J.; Blum, S. Interactions between the microbiota and the intestinal mucosa. Eur. J. Clin. Nutr. 2002, 56, S60–S64. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Zhang, M.; Wang, S.; Han, R.; Cao, Y.; Hua, W.; Mao, Y.; Zhang, X.; Pang, X.; Wei, C.; et al. Interactions between gut microbiota, host genetics and diet relevant to development of metabolic syndromes in mice. ISME J. 2010, 4, 232–241. [Google Scholar] [CrossRef] [PubMed]
- McFall-Ngai, M.; Hadfield, M.G.; Bosch, T.C.G.; Carey, H.V.; Domazet-Lošo, T.; Douglas, A.E.; Dubilier, N.; Eberl, G.; Fukami, T.; Gilbert, S.F.; et al. Animals in a bacterial world, a new imperative for the life sciences. Proc. Natl. Acad. Sci. USA 2013, 110, 3229–3236. [Google Scholar] [CrossRef] [PubMed]
- Martens, E.C.; Kelly, A.G.; Tauzin, A.S.; Brumer, H. The devil lies in the details: How variations in polysaccharide fine-structure impact the physiology and evolution of gut microbes. J. Mol. Biol. 2014, 426, 3851–3865. [Google Scholar] [CrossRef] [PubMed]
- Wei, F.; Wang, X.; Wu, Q. The giant panda gut microbiome. Trends Microbiol. 2015, 23, 450–452. [Google Scholar] [CrossRef] [PubMed]
- Sleeth, M.L.; Thompson, E.L.; Ford, H.E.; Zac-Varghese, S.E.K. Free fatty acid receptor 2 and nutrient sensing: A proposed role for fibre, fermentable carbohydrates and short-chain fatty acids in appetite regulation. Nutr. Res. Rev. 2010, 23, 135–145. [Google Scholar] [CrossRef] [PubMed]
- Mortensen, F.V.; Nielsen, H.; Aalkjær, C.; Mulvany, M.J.; Hessov, I. Short Chain Fatty Acids Relax Isolated Resistance Arteries from the Human Ileum by a Mechanism Dependent on Anion-Exchange. Basic Clin. Pharmacol. 1994, 75, 181–185. [Google Scholar] [CrossRef]
- Høverstad, T.; Midtvedt, T. Short-chain fatty acids in germfree mice and rats. J. Nutr. 1986, 116, 1772–1776. [Google Scholar] [CrossRef] [PubMed]
- Wostmann, B.S.; Larkin, C.; Moriarty, A.; Bruckner-Kardoss, E. Dietary intake, energy metabolism, and excretory losses of adult male germfree Wistar rats. Lab. Anim. Sci. 1983, 33, 46–50. [Google Scholar] [PubMed]
- Stappenbeck, T.S.; Hooper, L.V.; Gordon, J.I. Developmental regulation of intestinal angiogenesis by indigenous microbes via Paneth cells. Proc. Natl. Acad. Sci. USA 2002, 99, 15451–15455. [Google Scholar] [CrossRef] [PubMed]
- Reinhardt, C.; Bergentall, M.; Greiner, T.U.; Schaffner, F.; Östergren-Lundén, G.; Petersen, L.C.; Ruf, W.; Bäckhed, F. Tissue factor and PAR1 promote microbiota-induced intestinal vascular remodelling. Nature 2012, 483, 627–631. [Google Scholar] [CrossRef] [PubMed]
- Lewis, S.J.; Heaton, K.W. Increasing butyrate concentration in the distal colon by accelerating intestinal transit. Gut 1997, 41, 245–251. [Google Scholar] [CrossRef] [PubMed]
- Davie, J.R. Inhibition of histone deacetylase activity by butyrate. J. Nutr. 2003, 133, 2485S–2493S. [Google Scholar] [CrossRef] [PubMed]
- Louis, P.; Scott, K.P.; Duncan, S.H.; Flint, H.J. Understanding the effects of diet on bacterial metabolism in the large intestine. J. Appl. Microbiol. 2007, 102, 1197–1208. [Google Scholar] [CrossRef] [PubMed]
- Samuel, B.S.; Gordon, J.I. A humanized gnotobiotic mouse model of host-archaeal-bacterial mutualism. Proc. Natl. Acad. Sci. USA 2006, 103, 10011–10016. [Google Scholar] [CrossRef] [PubMed]
- Macfarlane, S.; Macfarlane, G.T. Regulation of short-chain fatty acid production. Proc. Nutr. Soc. 2003, 62, 67–72. [Google Scholar] [CrossRef] [PubMed]
- Rey, F.; Faith, J.J.; Bain, J.; Muehlbauer, M.J.; Stevens, R.D.P.; Newgard, C.B.; Gordon, J.I. Dissecting the in vivo metabolic potential of two human gut acetogens. J. Biol. Chem. 2010, 285, 22082–22090. [Google Scholar] [CrossRef] [PubMed]
- El Oufir, L.; Flourié, B.; des Varannes, S.B.; Barry, J.L.; Cloarec, D.; Bornet, F.; Galmiche, J.P. Relations between transit time, fermentation products, and hydrogen consuming flora in healthy humans. Gut 1996, 38, 870–877. [Google Scholar] [CrossRef] [PubMed]
- Sahakian, A.B.; Jee, S.R.; Pimentel, M. Methane and the gastrointestinal tract. Dig. Dis. Sci. 2010, 55, 2135–2143. [Google Scholar] [CrossRef] [PubMed]
- Derting, T.L.; Bogue, B.A. Responses of the gut to moderate energy demands in a small herbivore (Microtus pennsylvanicus). J. Mammal. 1993, 74, 59–68. [Google Scholar] [CrossRef]
- Franks, A.H.; Harmsen, H.J.; Raangs, G.C.; Jansen, G.J.; Schut, F.; Welling, G.W. Variations of bacterial populations in human feces measured by fluorescent in situ hybridization with group-specific 16S rRNA-targeted oligonucleotide probes. Appl. Environ. Microb. 1998, 64, 3336–3345. [Google Scholar]
- Zoetendal, E.G.; Akkermans, A.D.L.; De Vos, W.M. Temperature gradient gel electrophoresis analysis of 16S rRNA from human fecal samples reveals stable and host-specific communities of active bacteria. Appl. Environ. Microb. 1998, 64, 3854–3859. [Google Scholar]
- Costello, E.K.; Lauber, C.L.; Hamady, M.; Fierer, N.; Gordon, J.I.; Knight, R. Bacterial community variation in human body habitats across space and time. Science 2009, 326, 1694–1697. [Google Scholar] [CrossRef] [PubMed]
- Duncan, S.H.; Louis, P.; Thomson, J.M.; Flint, H.J. The role of pH in determining the species composition of the human colonic microbiota. Environ. Microbiol. 2009, 11, 2112–2122. [Google Scholar] [CrossRef] [PubMed]
- Lauraeus, M.; Wikstrom, M. The terminal quinol oxidases of Bacillus subtilis have different energy conservation properties. J. Biol. Chem. 1993, 268, 11470–11473. [Google Scholar] [PubMed]
- Lemma, E.; Simon, J.; Schagger, H.; Kroger, A. Properties of the menaquinol oxidase (Qox) and of qox deletion mutants of Bacillus subtilis. Arch. Microbiol. 1995, 163, 432–438. [Google Scholar] [CrossRef] [PubMed]
- Yi, S.M.; Narasimhulu, K.V.; Samoilova, R.I.; Gennis, R.B.; Dikanov, S.A. Characterization of the semiquinone radical stabilized by the cytochrome aa3-600 menaquinol oxidase of Bacillus subtilis. J. Biol. Chem. 2010, 285, 18241–18251. [Google Scholar] [CrossRef] [PubMed]
- Baughn, A.D.; Malamy, M.H. The strict anaerobe Bacteroides fragilis grows in and benefits from nanomolar concentrations of oxygen. Nature 2004, 427, 441–444. [Google Scholar] [CrossRef] [PubMed]
- Flint, H.J.; Duncan, S.H.; Scott, K.P.; Louis, P. Interactions and competition within the microbial community of the human colon: Links between diet and health. Environ. Microbiol. 2007, 9, 1101–1111. [Google Scholar] [CrossRef] [PubMed]
- Booijink, C.C.; El-Aidy, S.; Rajilić-Stojanović, M.; Heilig, H.G.; Troost, F.J.; Smidt, H.; Kleerebezem, M.; De Vos, W.M.; Zoetendal, E.G. High temporal and inter-individual variation detected in the human ileal microbiota. Environ. Microbiol. 2010, 12, 3213–3227. [Google Scholar] [CrossRef] [PubMed]
- Zoetendal, E.G.; Raes, J.; Van Den Bogert, B.; Arumugam, M.; Booijink, C.C.; Troost, F.J.; Bork, P.; Wels, M.; de Vos, W.M.; Kleerebezem, M. The human small intestinal microbiota is driven by rapid uptake and conversion of simple carbohydrates. ISME J. 2012, 6, 1415–1426. [Google Scholar] [CrossRef] [PubMed]
- Ley, R.E.; Hamady, M.; Lozupone, C.; Tozupone, C.; Turnbaugh, P.J.; Ramey, R.R.; Bircher, J.S.; Schlegel, M.L.; Tucker, T.A.; Schrenzel, M.D.; et al. Evolution of mammals and their gut microbes. Science 2008, 320, 1647–1651. [Google Scholar] [CrossRef] [PubMed]
- Walker, A.W.; Duncan, S.H.; Harmsen, H.J.M.; Holtrop, G.; Welling, G.W.; Flint, H.J. The species composition of the human intestinal microbiota differs between particle-associated and liquid phase communities. Environ. Microbiol. 2008, 10, 3275–3283. [Google Scholar] [CrossRef] [PubMed]
- Weng, F.C.H.; Shaw, G.T.W.; Weng, C.Y.; Yang, Y.J.; Wang, D. Inferring microbial interactions in the Gut of the Hong Kong Whipping Frog (Polypedates megacephalus) and a validation using probiotics. Front. Microbiol. 2017, 8, 525. [Google Scholar] [CrossRef] [PubMed]
- Stevens, C.E.; Hume, I.D. Comparative Physiology of the Vertebrate Digestive System; Cambridge University Press: Cambridge, UK, 2004. [Google Scholar]
- Grant, P.R.; Grant, B.R. Evolution of character displacement in Darwin’s finches. Science 2006, 31, 224–226. [Google Scholar] [CrossRef] [PubMed]
- Sibly, R.M. Strategies of digestion and defecation. Saude Soc. 1981, 24, 129–140. [Google Scholar]
- Kramer, D.L.; Bryant, M.J. Intestine length in the fishes of a tropical stream: 2. Relationships to diet—The long and short of a convoluted issue. Environ. Biol. Fish. 1995, 42, 129–141. [Google Scholar] [CrossRef]
- German, D.P.; Michael, H.H. Gut length and mass in herbivorous and carnivorous prickleback fishes (Teleostei: Stichaeidae): Ontogenetic, dietary, and phylogenetic effects. Mar. Biol. 2006, 148, 1123–1134. [Google Scholar] [CrossRef]
- German, D.P.; Gawlicka, A.K.; Horn, M.H. Evolution of ontogenetic dietary shifts and associated gut features in prickleback fishes (Teleostei: Stichaeidae). Comp. Biochem. Physiol. Part B 2014, 168, 12–18. [Google Scholar] [CrossRef] [PubMed]
- Reddy, C.V.; Jensen, L.S.; Merrill, L.H.; Mcginnis, J. Influence of mechanical alteration of dietary density on energy available for chick growth. J. Nutr. 1962, 77, 428–432. [Google Scholar] [CrossRef] [PubMed]
- Karasov, W.H.; del Rio, C.M.; Caviedes-Vidal, E. Ecological physiology of diet and digestive systems. Annu. Rev. Physiol. 2011, 73, 69–93. [Google Scholar] [CrossRef] [PubMed]
- Cant, J.P.; McBride, B.W.; Croom, W.J. The regulation of intestinal metabolism and its impact on whole animal energetics. J. Anim. Sci. 1996, 74, 2541–2553. [Google Scholar] [CrossRef] [PubMed]
- Heneghan, J.B. Enterocyte kinetics, mucosal surface area and mucus in gnotobiotes. Clin. Exp. Gnotobiot. 1979, 6, 19–27. [Google Scholar]
- Heneghan, J.B.; Gordon, H.A.; Miniats, O.P. Intestinal mucosal surface area and goblet cells in germfree and conventional piglets. Zentralblatt fur Bakteriologie, Parasitenkunde, Infekstionskrankheiten und Hygiene. I. Abt.: Supplemente 1979, 6, 107–111. [Google Scholar]
- Baez, S.; Waldemar, Y.; Bruckner, G.; Miniats, O.P.; Gordon, H.A. Vascular smooth muscle depressant substance in germfree piglets. In Clinical and Experimental Gnotobiotics, zbl Bakt. Suppl 7; Fliedner, T., Heit, H., Niethammer, D., Pflieger, H., Eds.; Fischer: Stuttgart, Germany, 1979; pp. 29–34. [Google Scholar]
- Gordon, H.A.; Bruckner-Kardoss, E.; Staley, T.E.; Wagner, M.; Wostmann, B.S. Characteristics of the germfree rat. Acta Anat. 1966, 64, 301–323. [Google Scholar] [CrossRef]
- Meshn, J.C.; Sacquet, E.; Guenet, J.L. Action de la flore bacterienne sur la morphologie et la surface de la muguese de I'intestine grele du rat. Ann. Biol. Anim. Biochim. Biophys. 1973, 13, 203–214. [Google Scholar]
- Abrams, G.D.; Bauer, H.; Sprinz, H. Influence of the normal flora on mucosal morphology and cellular renewal in the ileum. A comparison of germfree and conventional mice. Lab. Investig. 1963, 12, 355–364. [Google Scholar] [PubMed]
- Mink, J.W.; Blumenschine, R.J.; Adams, D.B. Ratio of central nervous system to body metabolism in vertebrates: Its constancy and functional basis. Am. J. Physiol. Reg. Integr. 1981, 241, R203–R212. [Google Scholar] [CrossRef] [PubMed]
- Kety, S.S. Circulation and energy metabolism of the brain. Clin. Neurosurg. 1963, 9, 56–66. [Google Scholar] [CrossRef] [PubMed]
- Raichle, M.E.; Mintun, M.A. Brain work and brain imaging. Annu. Rev. Neurosci. 2006, 29, 449–476. [Google Scholar] [CrossRef] [PubMed]
- Wrangham, R.W. Catching fire: How cooking made us human. Hum. Nat. 2014, 20, 447–449. [Google Scholar]
- Wrangham, R. Catching Fire: How Cooking Made Us Human; Basic Books: New York, NY, USA, 2009. [Google Scholar]
- Van Woerden, J.T.; van Schaik, C.P.; Isler, K. Effects of seasonality on brain size evolution: Evidence from strepsirrhine primates. Am. Nat. 2010, 176, 758–767. [Google Scholar] [CrossRef] [PubMed]
- Bruhn, J.M.; Benedict, F.G. The respiratory metabolism of the chimpanzee. Proc. Am. Acad. Arts. Sci. 1936, 71, 259–326. [Google Scholar] [CrossRef]
- Whiten, A.; Byrne, R. Machiavellian Intelligence: Social Expertise and the Evolution of Intellect in Monkeys, Apes and Humans; Clarendon Press/Oxford University Press: New York, NY, USA, 1988; pp. 1–9. [Google Scholar]
- DeCasien, A.R.; Williams, S.A.; Higham, J.P. Primate brain size is predicted by diet but not sociality. Nat. Ecol. Evol. 2017, 1, 0112. [Google Scholar] [CrossRef] [PubMed]
- Goyal, M.S.; Hawrylycz, M.; Miller, J.A.; Snyder, A.Z.; Raichle, M.E. Aerobic glycolysis in the human brain is associated with development and neotenous gene expression. Cell Metab. 2014, 19, 4–5. [Google Scholar] [CrossRef] [PubMed]
- Bélanger, M.; Allaman, I.; Magistretti, P.J. Brain energy metabolism: Focus on astrocyte-neuron metabolic cooperation. Cell Metab. 2011, 14, 724–738. [Google Scholar] [CrossRef] [PubMed]
- Fünfschilling, U.; Supplie, L.M.; Mahad, D.; Boretius, S.; Saab, A.S.; Edgar, J.; Brinkmann, B.G.; Kassmann, C.M.; Tzvetanova, I.D.; Möbius, W.; et al. Glycolytic oligodendrocytes maintain myelin and long-term axonal integrity. Nature 2012, 485, 517–521. [Google Scholar] [CrossRef] [PubMed]
- Lu, J.; Lu, L.; Yu, Y.; Cluette-Brown, J.; Martin, C.R.; Claud, E.C. Effects of Intestinal Microbiota on Brain Development in Humanized Gnotobiotic Mice. Sci. Rep. 2018, 8, 5443. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, C. Mental health: Thinking from the gut. Nature 2015, 518, S12–S15. [Google Scholar] [CrossRef] [PubMed]
- Erny, D.; Prinz, M. Microbiology: Gut microbes augment neurodegeneration. Nature 2017, 544, 304–305. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.K.; Menezes, J.S.; Umesaki, Y.; Mazmanian, S.K. Proinflammatory T-cell responses to gut microbiota promote experimental autoimmune encephalomyelitis. Proc. Natl. Acad. Sci. USA 2011, 108, 4615–4622. [Google Scholar] [CrossRef] [PubMed]
- Heijtz, R.D.; Wang, S.; Anuar, F.; Qian, Y.; Björkholm, B.; Samuelsson, A.; Hibberd, M.L.; Forssberg, H.; Pettersson, S. Normal gut microbiota modulates brain development and behavior. Proc. Natl. Acad. Sci. USA 2011, 108, 3047–3052. [Google Scholar] [CrossRef] [PubMed]
- Borre, Y.E.; Moloney, R.D.; Clarke, G.; Dinan, T.G.; Cryan, J.F. The impact of microbiota on brain and behavior: Mechanisms & therapeutic potential. Adv. Exp. Med. Biol. 2014, 817, 373–403. [Google Scholar] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Huang, C.H.; Yu, X.; Liao, W.B. The Expensive-Tissue Hypothesis in Vertebrates: Gut Microbiota Effect, a Review. Int. J. Mol. Sci. 2018, 19, 1792. https://doi.org/10.3390/ijms19061792
Huang CH, Yu X, Liao WB. The Expensive-Tissue Hypothesis in Vertebrates: Gut Microbiota Effect, a Review. International Journal of Molecular Sciences. 2018; 19(6):1792. https://doi.org/10.3390/ijms19061792
Chicago/Turabian StyleHuang, Chun Hua, Xin Yu, and Wen Bo Liao. 2018. "The Expensive-Tissue Hypothesis in Vertebrates: Gut Microbiota Effect, a Review" International Journal of Molecular Sciences 19, no. 6: 1792. https://doi.org/10.3390/ijms19061792
APA StyleHuang, C. H., Yu, X., & Liao, W. B. (2018). The Expensive-Tissue Hypothesis in Vertebrates: Gut Microbiota Effect, a Review. International Journal of Molecular Sciences, 19(6), 1792. https://doi.org/10.3390/ijms19061792



