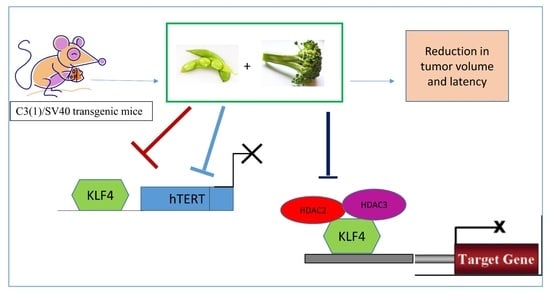
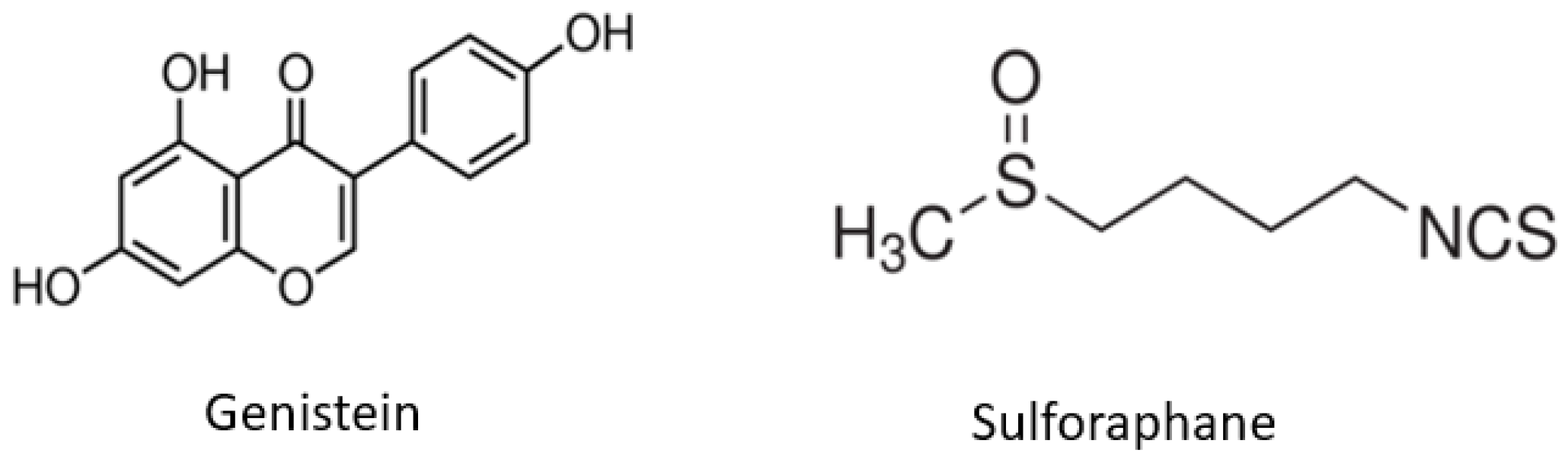
The Effects of Combinatorial Genistein and Sulforaphane in Breast Tumor Inhibition: Role in Epigenetic Regulation
Abstract
1. Introduction
2. Results
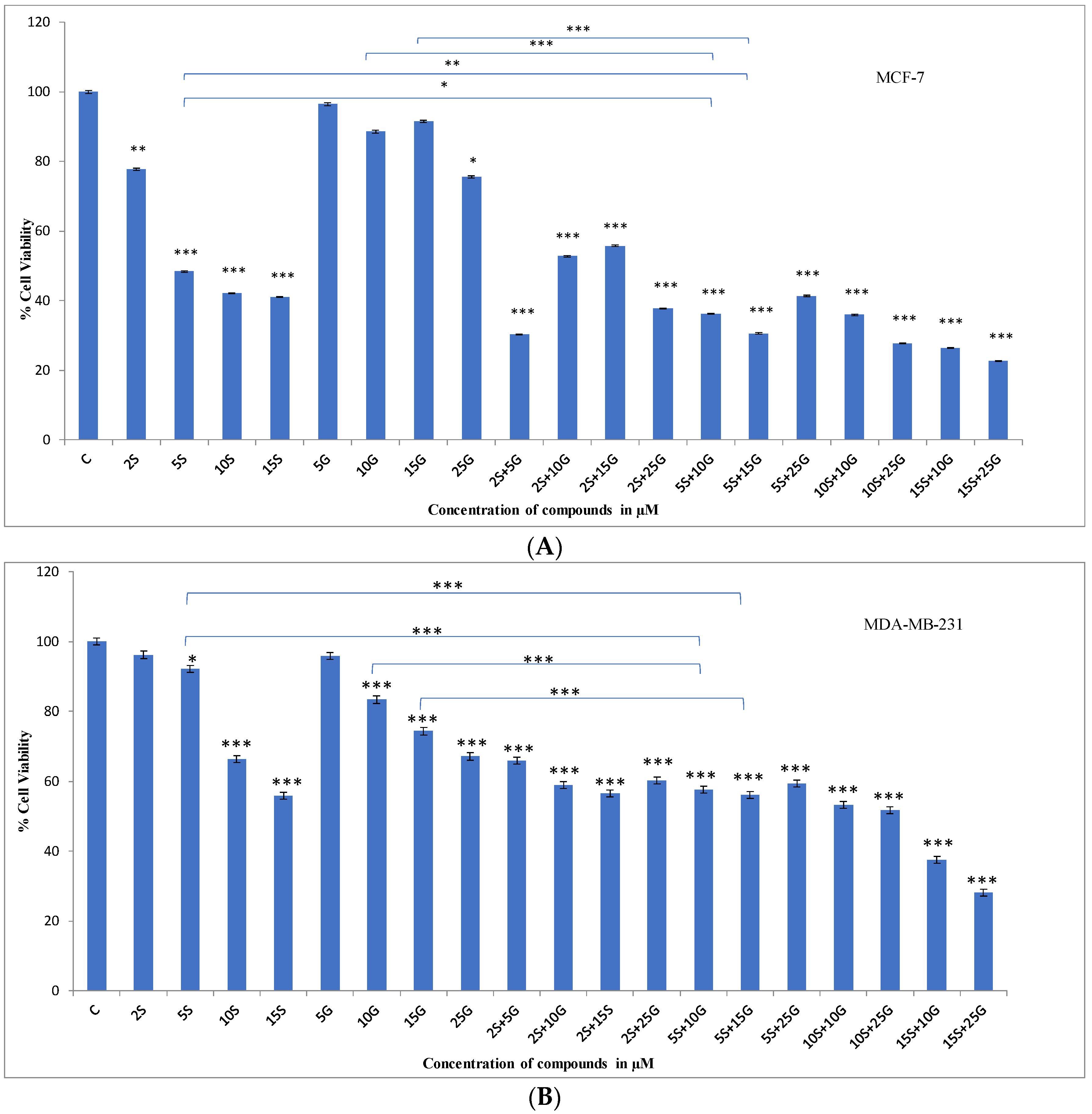
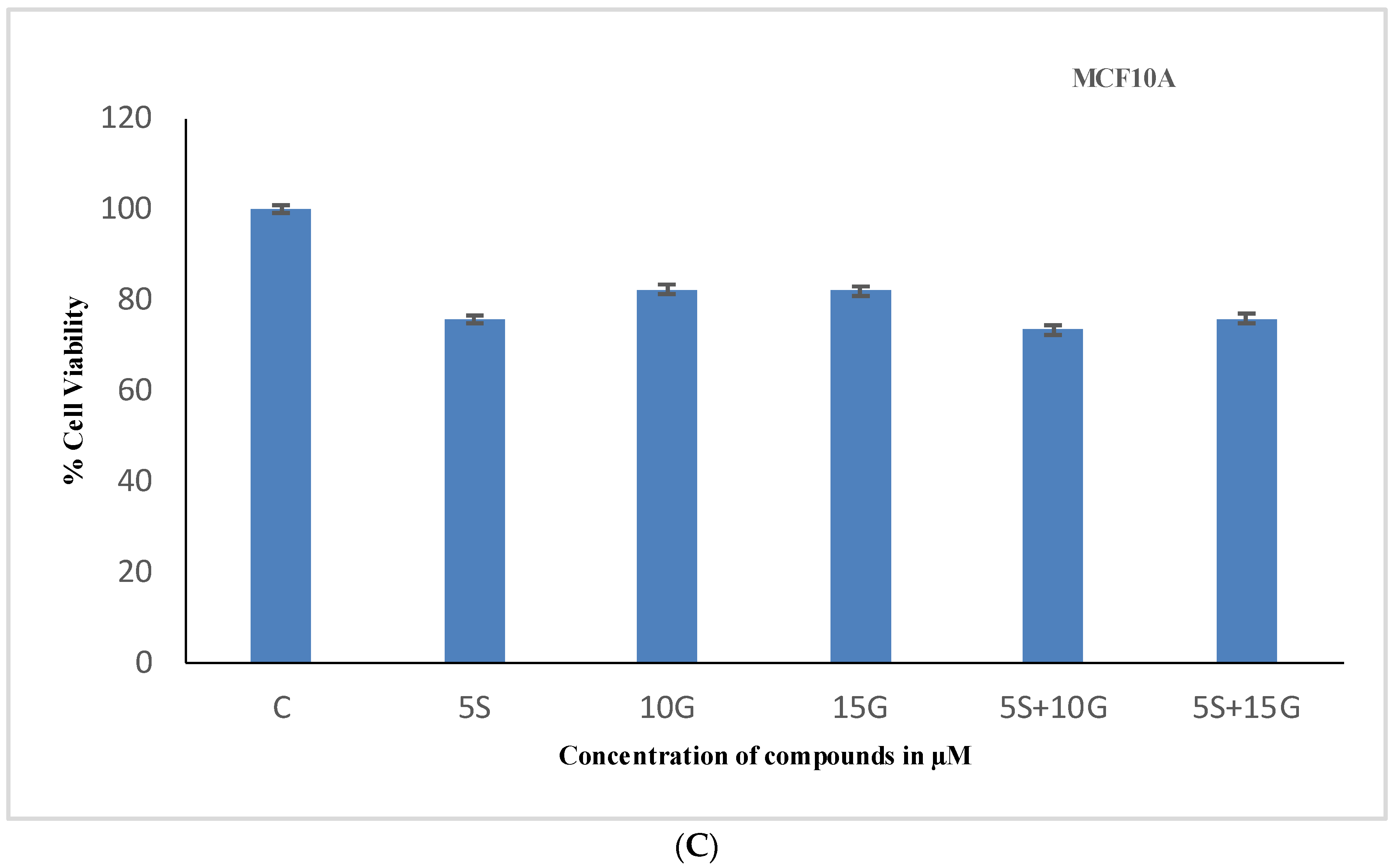
2.1. Effect of the Combination Treatment on the Cellular Viability of MCF-7 and MDA-MB-231 Breast Cancer Cells
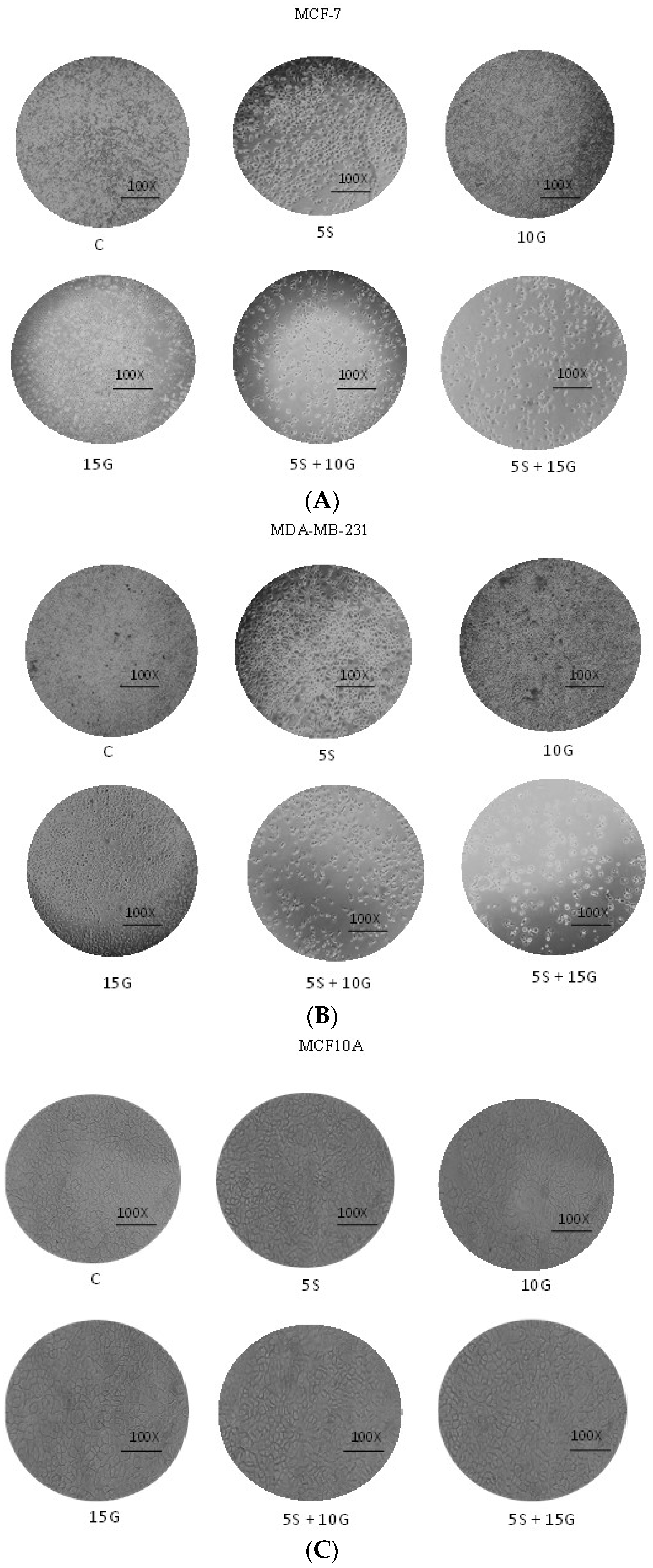
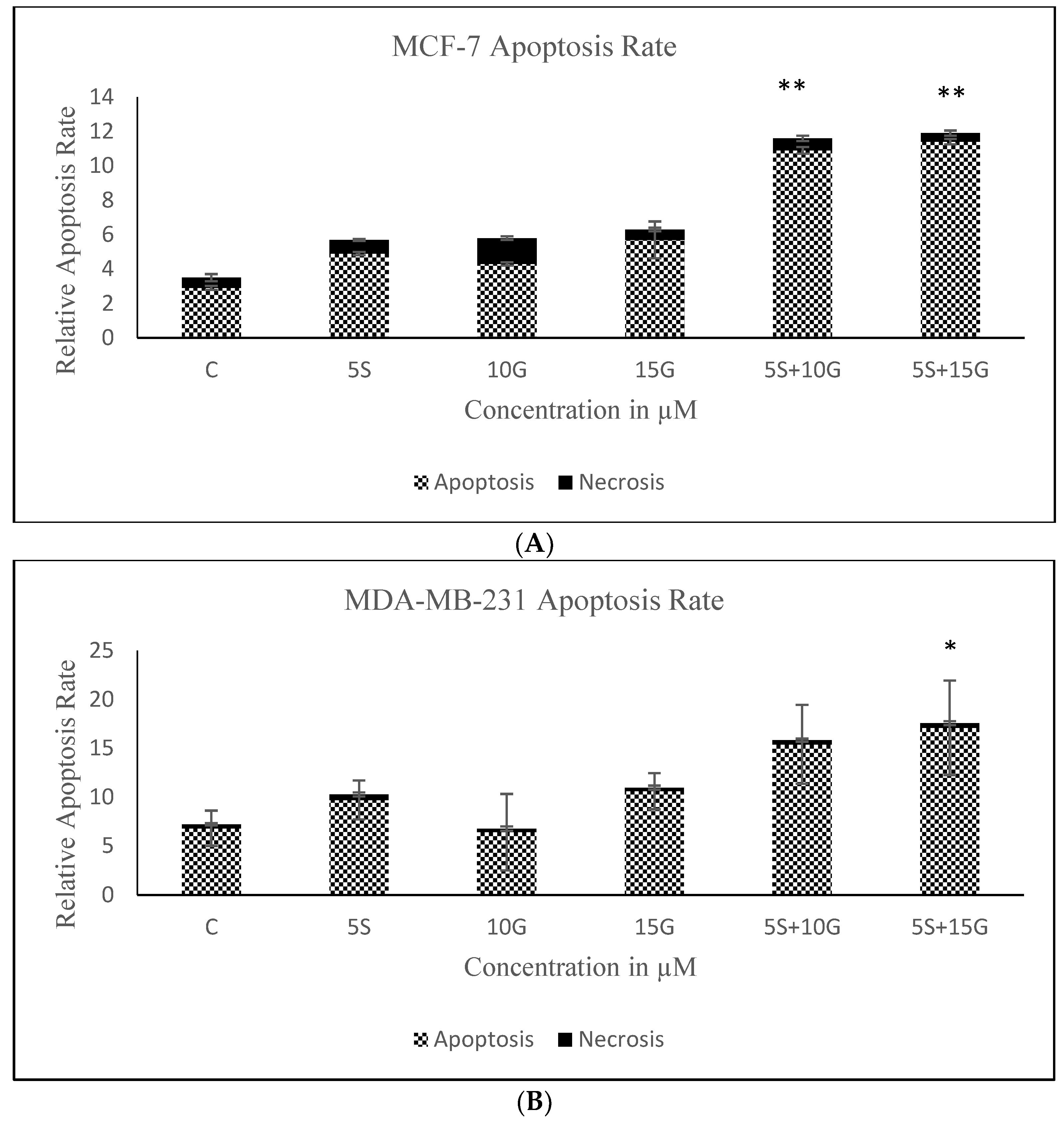
2.2. Combination of GEN and SFN Promote Cell Death as Evidenced through Cell Density Analysis and Apoptosis Assay
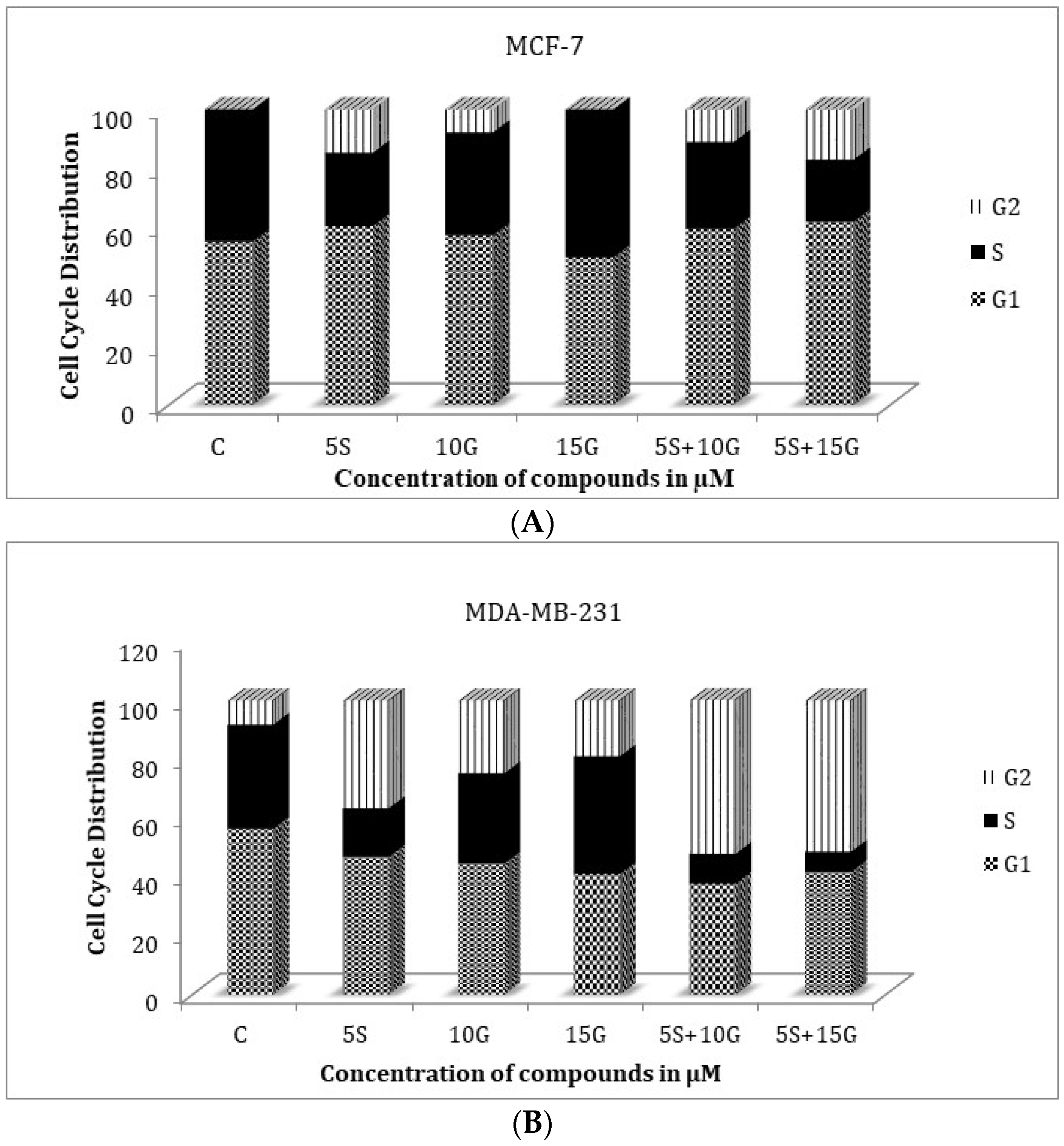
2.3. Effect of the Combination on Treatment Cell Cycle Arrest
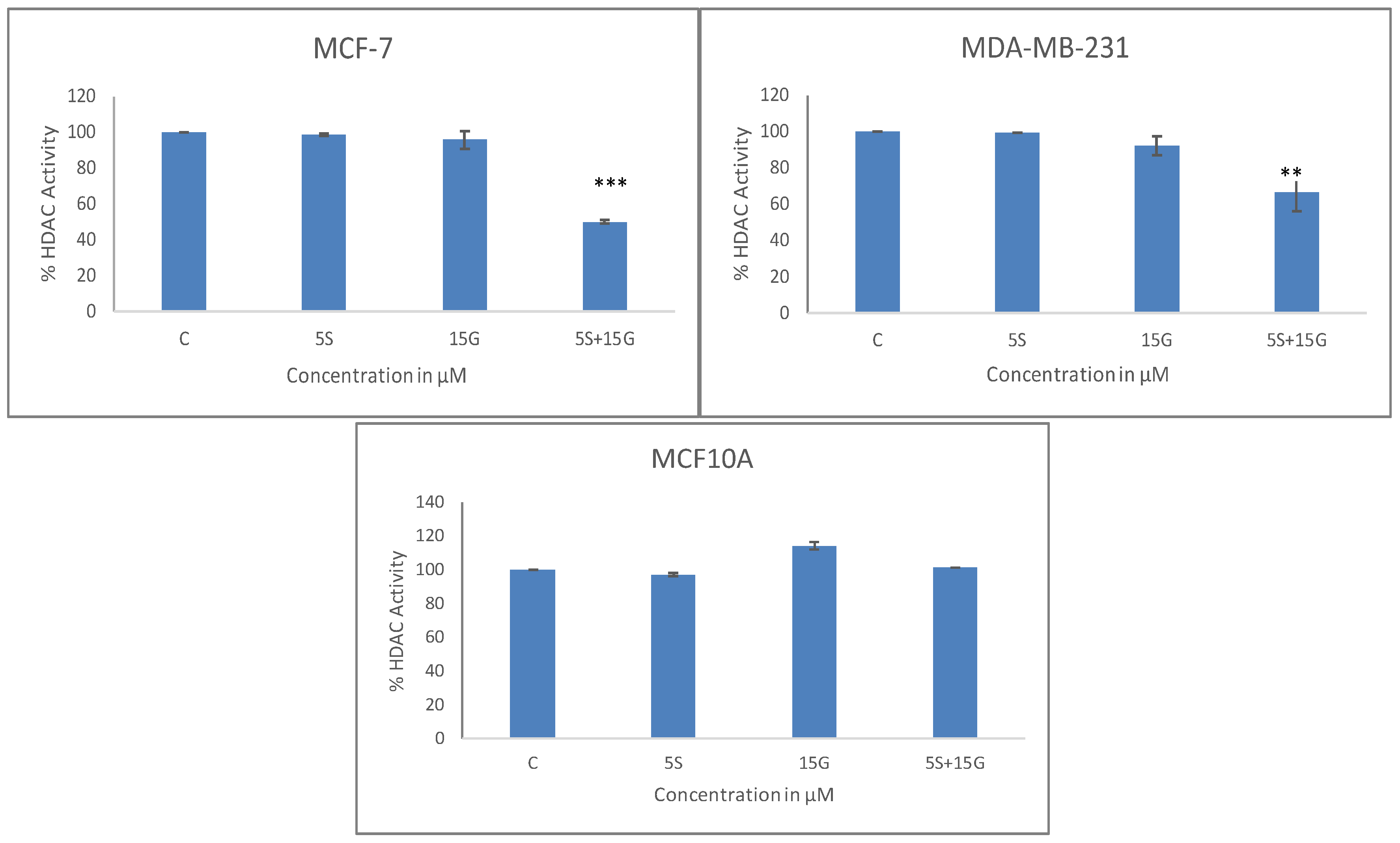
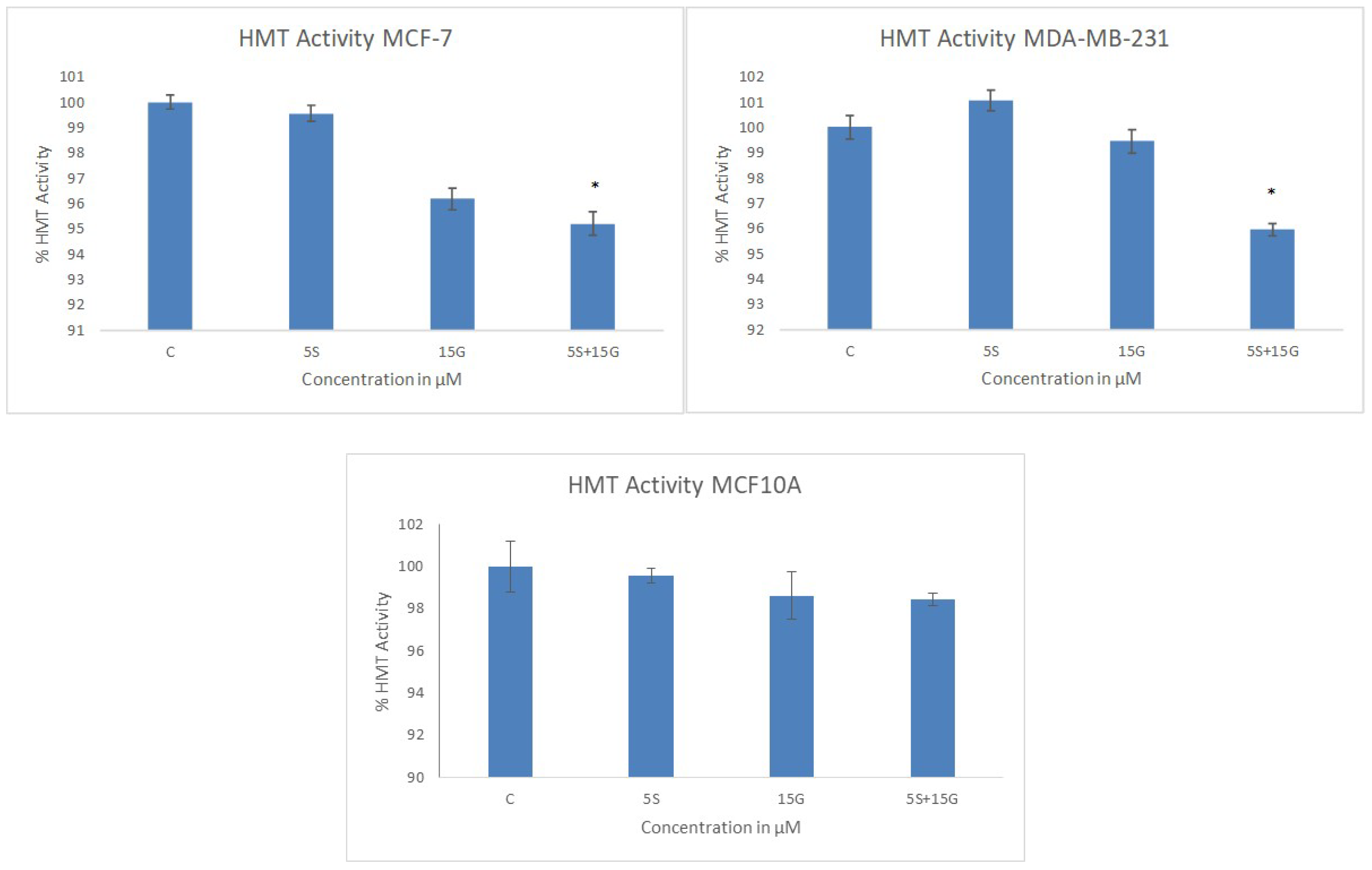
2.4. Combinatorial GEN and SFN Administration Decreases HDAC and HMT Expression
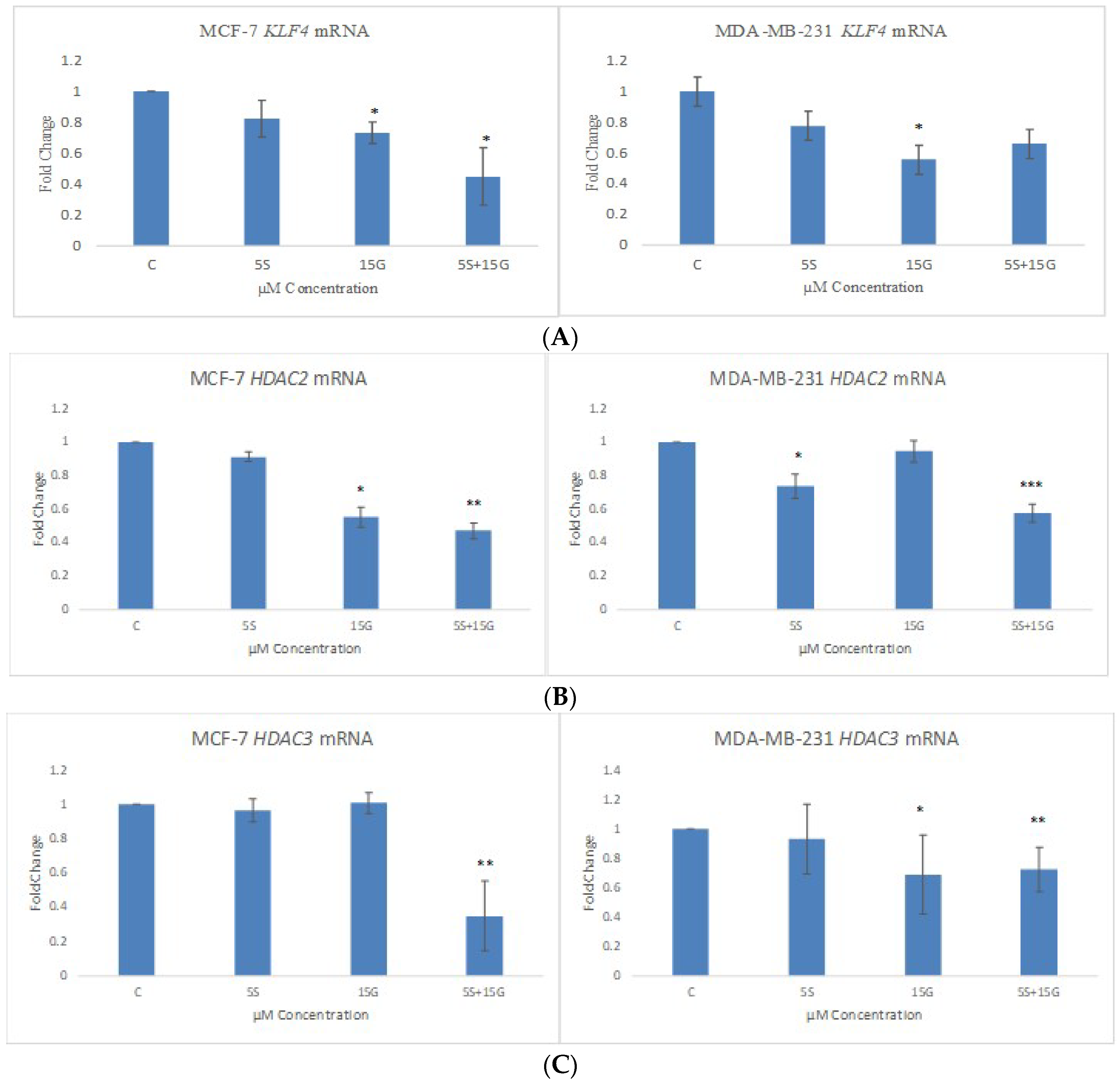
2.5. GEN and SFN in Combination Induces Changes in KLF4, HDAC2 and HDAC3 mRNA Levels
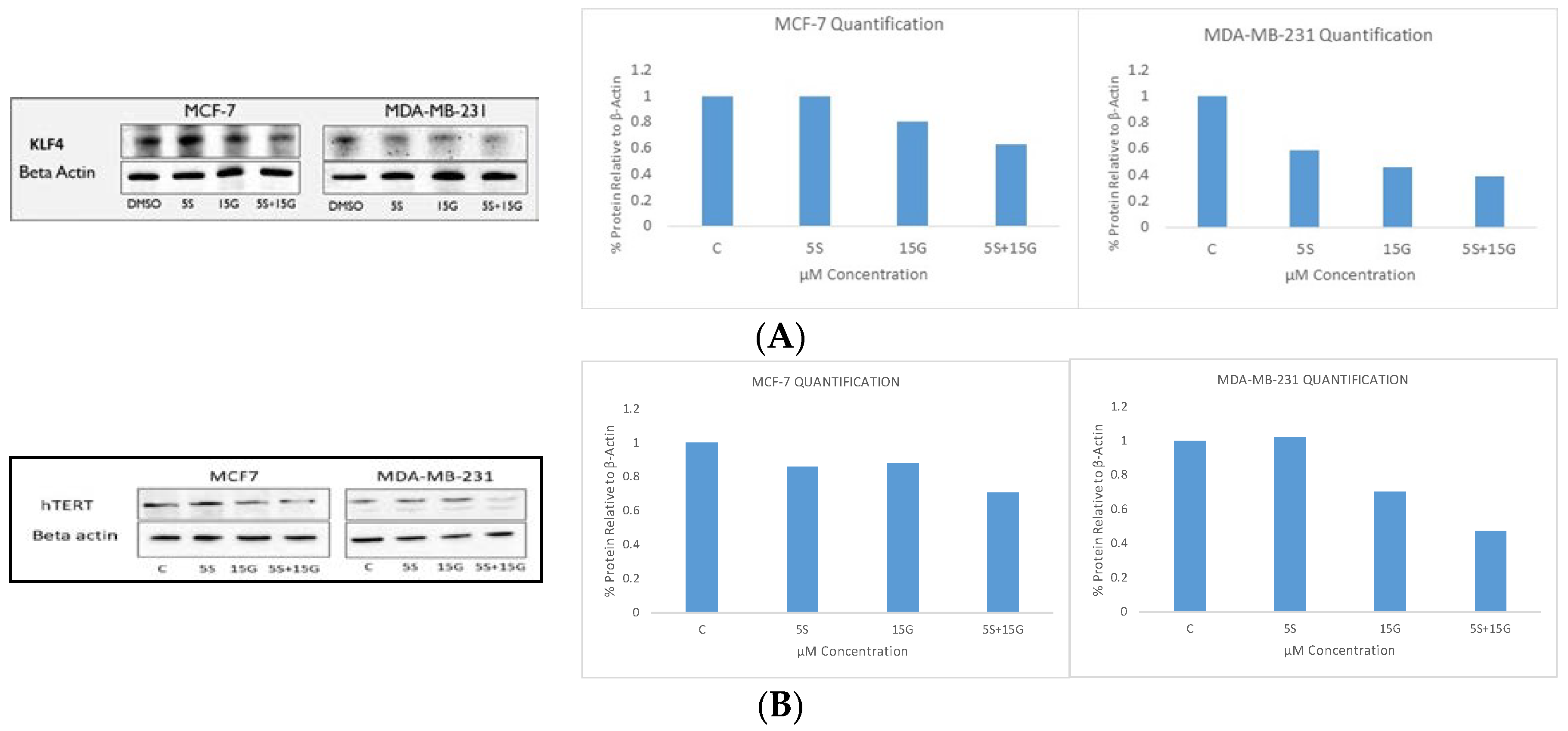
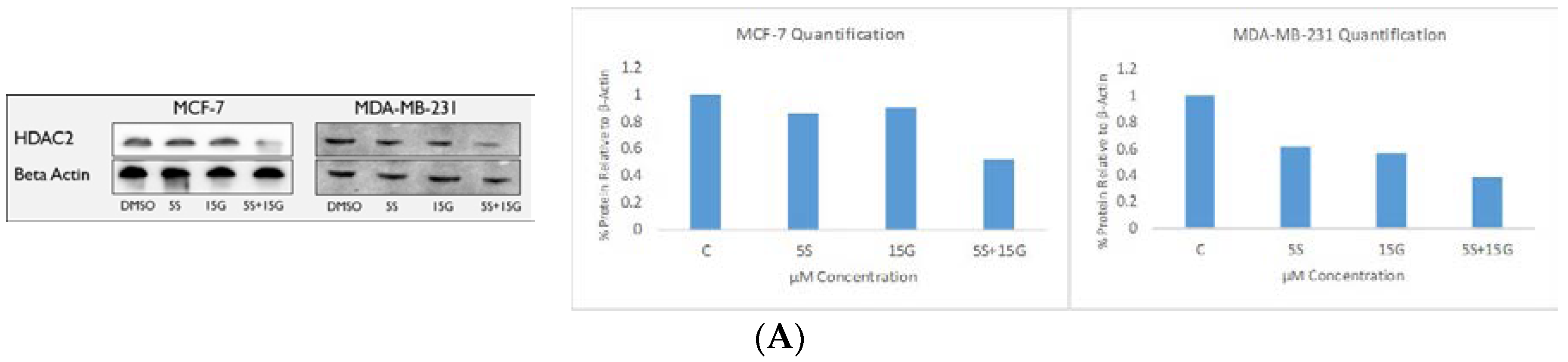
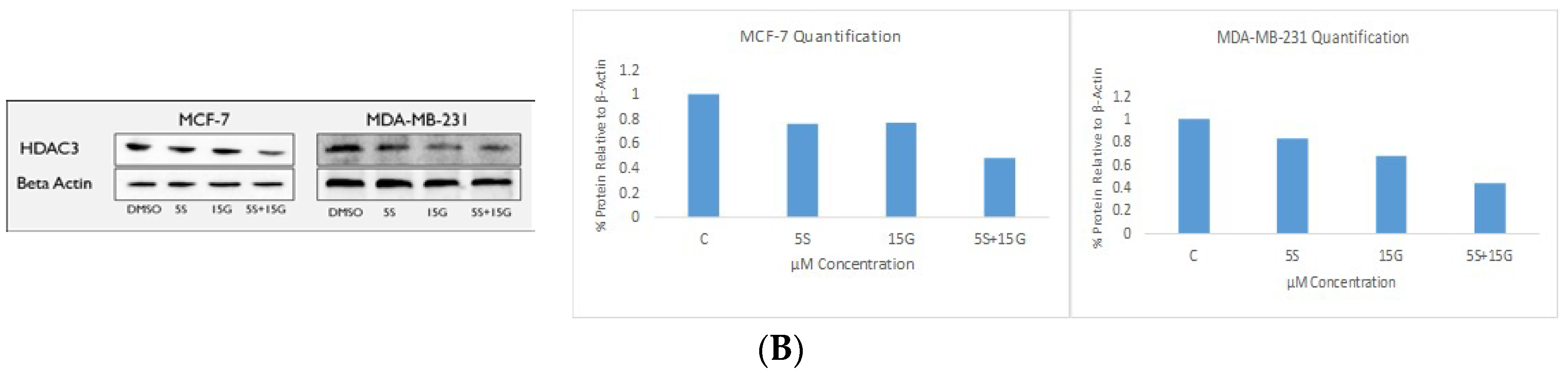
2.6. Combination Treatment Induces Changes in KLF4, hTERT, HDAC2 AND HDAC3 Protein Levels
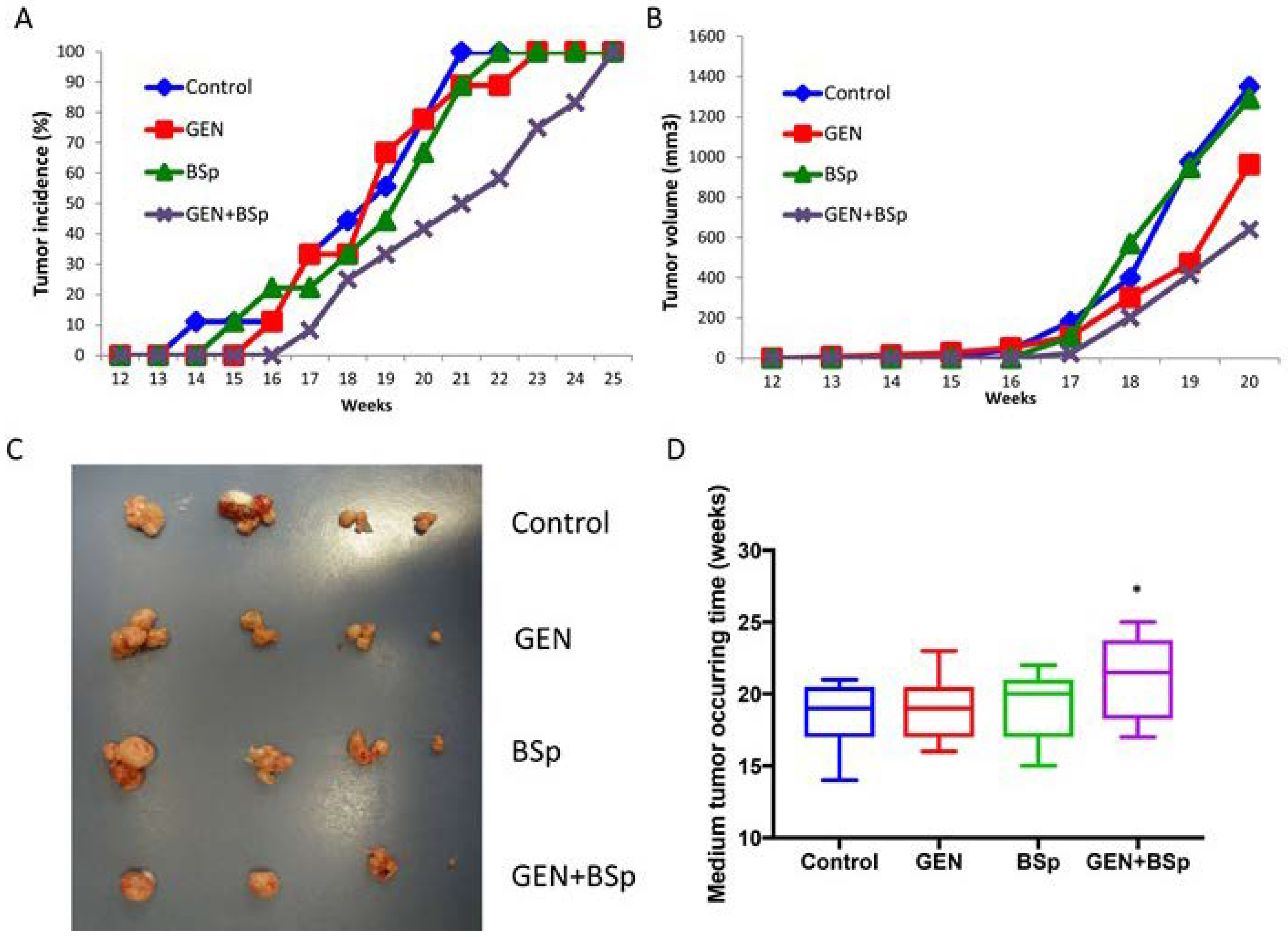
2.7. Combinatorial Dietary Regimen of GEN and Broccoli Sprouts (BSp) Is Effective in Reducing Mammary Tumor Incidence and Delaying Tumor Latency in a Spontaneous Breast Cancer Mouse Model
3. Discussion
4. Materials and Methods
5. Conclusions
Supplementary Materials
Author Contributions
Acknowledgments
Conflicts of Interest
References
- DeSantis, C.E.; Ma, J.; Goding Sauer, A.; Newman, L.A.; Jemal, A. Breast cancer statistics, 2017, racial disparity in mortality by state. CA Cancer J. Clin. 2017, 67, 439–448. [Google Scholar] [CrossRef] [PubMed]
- Hardy, T.M.; Tollefsbol, T.O. Epigenetic diet: Impact on the epigenome and cancer. Epigenomics 2011, 3, 503–518. [Google Scholar] [CrossRef] [PubMed]
- Herceg, Z. Epigenetics and cancer: Towards an evaluation of the impact of environmental and dietary factors. Mutagenesis 2007, 22, 91–103. [Google Scholar] [CrossRef] [PubMed]
- Russo, G.L.; Vastolo, V.; Ciccarelli, M.; Albano, L.; Macchia, P.E.; Ungaro, P. Dietary polyphenols and chromatin remodeling. Crit. Rev. Food Sci. Nutr. 2017, 57, 2589–2599. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Meeran, S.M.; Tollefsbol, T.O. Combinatorial bioactive botanicals re-sensitize tamoxifen treatment in ER-negative breast cancer via epigenetic reactivation of ERα expression. Sci. Rep. 2017, 7, 9345. [Google Scholar] [CrossRef] [PubMed]
- Murrell, A. What lies beneath the epigenetic signatures associated with breast cancer and how do we find out? In Proceedings of the UK Breast Cancer Research Symposium, London, UK, 22–23 July 2016. [Google Scholar]
- Park, J.-H.; Kim, S.-H.; Lee, M.S.; Kim, M.-S. Epigenetic modification by dietary factors: Implications in metabolic syndrome. Mol. Aspects Med. 2017, 54, 58–70. [Google Scholar] [CrossRef] [PubMed]
- Tahir, A.A.; Sani, N.F.A.; Murad, N.A.; Makpol, S.; Ngah, W.Z.W.; Yusof, Y.A.M. Combined ginger extract & Gelam honey modulate Ras/ERK and PI3K/AKT pathway genes in colon cancer HT29 cells. Nutr. J. 2015, 14, 31. [Google Scholar] [PubMed]
- Kotecha, R.; Takami, A.; Espinoza, J.L. Dietary phytochemicals and cancer chemoprevention: A review of the clinical evidence. Oncotarget 2016, 7, 52517. [Google Scholar] [CrossRef] [PubMed]
- Chou, T.-C. Drug combination studies and their synergy quantification using the chou-talalay method. Cancer Res. 2010, 70, 440–446. [Google Scholar] [CrossRef] [PubMed]
- Xie, Q.; Bai, Q.; Zou, L.Y.; Zhang, Q.Y.; Zhou, Y.; Chang, H.; Yi, L.; Zhu, J.D.; Mi, M.T. Genistein inhibits DNA methylation and increases expression of tumor suppressor genes in human breast cancer cells. Genes Chromosomes Cancer 2014, 53, 422–431. [Google Scholar] [CrossRef] [PubMed]
- Tortorella, S.M.; Royce, S.G.; Licciardi, P.V.; Karagiannis, T.C. Dietary sulforaphane in cancer chemoprevention: The role of epigenetic regulation and HDAC inhibition. Antioxid. Redox Signal. 2015, 22, 1382–1424. [Google Scholar] [CrossRef] [PubMed]
- Meeran, S.M.; Ahmed, A.; Tollefsbol, T.O. Epigenetic targets of bioactive dietary components for cancer prevention and therapy. Clin. Epigenet. 2010, 1, 101–116. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Yuan, Y.-Y.; Meeran, S.M.; Tollefsbol, T.O. Synergistic epigenetic reactivation of estrogen receptor-α (ERα) by combined green tea polyphenol and histone deacetylase inhibitor in ERα-negative breast cancer cells. Mol. Cancer 2010, 9, 274. [Google Scholar] [CrossRef] [PubMed]
- Royston, K.J.; Udayakumar, N.; Lewis, K.; Tollefsbol, T.O. A novel combination of withaferin A and sulforaphane inhibits epigenetic machinery, cellular viability and induces apoptosis of breast cancer cells. Int. J. Mol. Sci. 2017, 18, 1092. [Google Scholar] [CrossRef] [PubMed]
- Dashwood, R.H.; Ho, E. Dietary agents as histone deacetylase inhibitors: Sulforaphane and structurally related isothiocyanates. Nutr. Rev. 2008, 66. [Google Scholar] [CrossRef] [PubMed]
- Traka, M.H.; Chambers, K.F.; Lund, E.K.; Goodlad, R.A.; Johnson, I.T.; Mithen, R.F. Involvement of KLF4 in sulforaphane-and iberin-mediated induction of p21 waf1/cip1. Nutr. Cancer 2008, 61, 137–145. [Google Scholar] [CrossRef] [PubMed]
- Rowland, B.D.; Peeper, D.S. KLF4, p21 and context-dependent opposing forces in cancer. Nat. Rev. Cancer 2006, 6, 11. [Google Scholar] [CrossRef] [PubMed]
- Wei, D.; Kanai, M.; Huang, S.; Xie, K. Emerging role of KLF4 in human gastrointestinal cancer. Carcinogenesis 2005, 27, 23–31. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Goldstein, B.G.; Chao, H.-H.; Katz, J. KLF4 and KLF5 regulate proliferation, apoptosis and invasion in esophageal cancer cells. Cancer Biol. Ther. 2005, 4, 1216–1221. [Google Scholar] [CrossRef] [PubMed]
- Yu, F.; Li, J.; Chen, H.; Fu, J.; Ray, S.; Huang, S.; Zheng, H.; Ai, W. Kruppel-like factor 4 (KLF4) is required for maintenance of breast cancer stem cells and for cell migration and invasion. Oncogene 2011, 30, 2161. [Google Scholar] [CrossRef] [PubMed]
- Okuda, H.; Xing, F.; Pandey, P.R.; Sharma, S.; Watabe, M.; Pai, S.K.; Mo, Y.-Y.; Iiizumi-Gairani, M.; Hirota, S.; Liu, Y. Mir-7 suppresses brain metastasis of breast cancer stem-like cells by modulating KLF4. Cancer Res. 2013, 73, 1434–1444. [Google Scholar] [CrossRef] [PubMed]
- Pandya, A.Y.; Talley, L.I.; Frost, A.R.; Fitzgerald, T.J.; Trivedi, V.; Chakravarthy, M.; Chhieng, D.C.; Grizzle, W.E.; Engler, J.A.; Krontiras, H. Nuclear localization of KLF4 is associated with an aggressive phenotype in early-stage breast cancer. Clin. Cancer. Res. 2004, 10, 2709–2719. [Google Scholar] [CrossRef] [PubMed]
- Li, R.; Liang, J.; Ni, S.; Zhou, T.; Qing, X.; Li, H.; He, W.; Chen, J.; Li, F.; Zhuang, Q. A mesenchymal-to-epithelial transition initiates and is required for the nuclear reprogramming of mouse fibroblasts. Cell Stem Cell 2010, 7, 51–63. [Google Scholar] [CrossRef] [PubMed]
- Ray, A.; Alalem, M.; Ray, B.K. Loss of epigenetic kruppel-like factor 4 histone deacetylase (KLF-4-HDAC)-mediated transcriptional suppression is crucial in increasing vascular endothelial growth factor (VEGF) expression in breast cancer. J. Biol. Chem. 2013, 288, 27232–27242. [Google Scholar] [CrossRef] [PubMed]
- Hsieh, M.-H.; Chen, Y.-T.; Chen, Y.-T.; Lee, Y.-H.; Lu, J.; Chien, C.-L.; Chen, H.-F.; Ho, H.-N.; Yu, C.-J.; Wang, Z.-Q. Parp1 controls KLF4-mediated telomerase expression in stem cells and cancer cells. Nucleic Acids Res. 2017, 45, 10492–10503. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.-H.; Wan, Y.; Singh, S.V. Breast cancer stem cell inhibition by benzyl isothiocyanate is hampered by induction of kruppel-like factor 4. In Proceedings of the AACR Annual Meeting, Washington, DC, USA, 1–5 April 2017. [Google Scholar]
- Chou, T.; Martin, N. A computer program for quantitation of synergism and antagonism in drug combinations, and the determination of IC50 and ED50 and LD50 values. In CompuSyn for Drug Combinations: PC Software and User’s Guide; ComboSyn: Paramus, NJ, USA, 2005. [Google Scholar]
- Fu, J.; Chou, T.-C. Abstract 4554A: Simple, efficient, and quantitative approach for determination of synergism, additive effect, and antagonism of drugs in vivo using combination index method: A proposition for clinical protocol design and regulatory synergy claims. In Proceedings of the AACR Annual Meeting, Washington, DC, USA, 1–5 April 2017. [Google Scholar]
- Chou, T.-C. Preclinical versus clinical drug combination studies. Leuk. Lymphoma 2008, 49, 2059–2080. [Google Scholar] [CrossRef] [PubMed]
- Hu, W.; Jia, Y.; Xiao, X.; Lv, K.; Chen, Y.; Wang, L.; Luo, X.; Liu, T.; Li, W.; Li, Y. KLF4 downregulates hTERT expression and telomerase activity to inhibit lung carcinoma growth. Oncotarget 2016, 7, 52870. [Google Scholar] [CrossRef] [PubMed]
- Key, T.J.; Schatzkin, A.; Willett, W.C.; Allen, N.E.; Spencer, E.A.; Travis, R.C. Diet, nutrition and the prevention of cancer. Public Health Nutr. 2004, 7, 187–200. [Google Scholar] [CrossRef] [PubMed]
- Esteller, M.; Herman, J.G. Cancer as an epigenetic disease: DNA methylation and chromatin alterations in human tumours. J. Pathol. 2002, 196, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Paul, B.; Barnes, S.; Demark-Wahnefried, W.; Morrow, C.; Salvador, C.; Skibola, C.; Tollefsbol, T.O. Influences of diet and the gut microbiome on epigenetic modulation in cancer and other diseases. Clin. Epigenet. 2015, 7, 112. [Google Scholar] [CrossRef] [PubMed]
- Youlden, D.R.; Cramb, S.M.; Yip, C.H.; Baade, P.D. Incidence and mortality of female breast cancer in the Asia-pacific region. Cancer Biol. Med. 2014, 11, 101. [Google Scholar] [PubMed]
- Ferrini, K.; Ghelfi, F.; Mannucci, R.; Titta, L. Lifestyle, nutrition and breast cancer: Facts and presumptions for consideration. Ecancermedicalscience 2015, 9, 557. [Google Scholar] [CrossRef] [PubMed]
- Grosso, G.; Bella, F.; Godos, J.; Sciacca, S.; Del Rio, D.; Ray, S.; Galvano, F.; Giovannucci, E.L. Possible role of diet in cancer: Systematic review and multiple meta-analyses of dietary patterns, lifestyle factors, and cancer risk. Nutr. Rev. 2017, 75, 405–419. [Google Scholar] [CrossRef] [PubMed]
- Meeran, S.M.; Patel, S.N.; Li, Y.; Shukla, S.; Tollefsbol, T.O. Bioactive dietary supplements reactivate ER expression in ER-negative breast cancer cells by active chromatin modifications. PLoS ONE 2012, 7, e37748. [Google Scholar] [CrossRef] [PubMed]
- Sinha, S.; Shukla, S.; Khan, S.; Tollefsbol, T.O.; Meeran, S.M. Epigenetic reactivation of p21CIP1/WAF1 and KLOTHO by a combination of bioactive dietary supplements is partially ERα-dependent in ERα-negative human breast cancer cells. Mol. Cell. Endocrinol. 2015, 406, 102–114. [Google Scholar] [CrossRef] [PubMed]
- Kikuno, N.; Shiina, H.; Urakami, S.; Kawamoto, K.; Hirata, H.; Tanaka, Y.; Majid, S.; Igawa, M.; Dahiya, R. Genistein mediated histone acetylation and demethylation activates tumor suppressor genes in prostate cancer cells. Int. J. Cancer 2008, 123, 552–560. [Google Scholar] [CrossRef] [PubMed]
- Majid, S.; Dar, A.A.; Ahmad, A.E.; Hirata, H.; Kawakami, K.; Shahryari, V.; Saini, S.; Tanaka, Y.; Dahiya, A.V.; Khatri, G. BTG3 tumor suppressor gene promoter demethylation, histone modification and cell cycle arrest by genistein in renal cancer. Carcinogenesis 2009, 30, 662–670. [Google Scholar] [CrossRef] [PubMed]
- Paul, B.; Royston, K.J.; Li, Y.; Stoll, M.L.; Skibola, C.F.; Wilson, L.S.; Barnes, S.; Morrow, C.D.; Tollefsbol, T.O. Impact of genistein on the gut microbiome of humanized mice and its role in breast tumor inhibition. PLoS ONE 2017, 12, e0189756. [Google Scholar] [CrossRef] [PubMed]
- Ganai, S.A. Histone deacetylase inhibitor sulforaphane: The phytochemical with vibrant activity against prostate cancer. Biomed. Pharmacother. 2016, 81, 250–257. [Google Scholar] [CrossRef] [PubMed]
- Tarapore, R.S.; Siddiqui, I.A.; Mukhtar, H. Modulation of wnt/β-catenin signaling pathway by bioactive food components. Carcinogenesis 2011, 33, 483–491. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Zhang, T.; Korkaya, H.; Liu, S.; Lee, H.-F.; Newman, B.; Yu, Y.; Clouthier, S.G.; Schwartz, S.J.; Wicha, M.S. Sulforaphane, a dietary component of broccoli/broccoli sprouts, inhibits breast cancer stem cells. Clin. Cancer. Res. 2010, 16, 2580–2590. [Google Scholar] [CrossRef] [PubMed]
- Meeran, S.M.; Patel, S.N.; Tollefsbol, T.O. Sulforaphane causes epigenetic repression of hTERT expression in human breast cancer cell lines. PLoS ONE 2010, 5, e11457. [Google Scholar] [CrossRef] [PubMed]
- West, A.C.; Johnstone, R.W. New and emerging HDAC inhibitors for cancer treatment. J. Clin. Investig. 2014, 124, 30–39. [Google Scholar] [CrossRef] [PubMed]
- Myzak, M.C.; Karplus, P.A.; Chung, F.-L.; Dashwood, R.H. A novel mechanism of chemoprotection by sulforaphane: Inhibition of histone deacetylase. Cancer Res. 2004, 64, 5767–5774. [Google Scholar] [CrossRef] [PubMed]
- Sasamura, H.; Takahashi, A.; Yuan, J.; Kitamura, H.; Masumori, N.; Miyao, N.; Itoh, N.; Tsukamoto, T. Antiproliferative and antiangiogenic activities of genistein in human renal cell carcinoma. Urology 2004, 64, 389–393. [Google Scholar] [CrossRef] [PubMed]
- Majid, S.; Kikuno, N.; Nelles, J.; Noonan, E.; Tanaka, Y.; Kawamoto, K.; Hirata, H.; Li, L.C.; Zhao, H.; Okino, S.T. Genistein induces the p21WAF1/CIP1 and p16INK4a tumor suppressor genes in prostate cancer cells by epigenetic mechanisms involving active chromatin modification. Cancer Res. 2008, 68, 2736–2744. [Google Scholar] [CrossRef] [PubMed]
- Fahey, J.W.; Zhang, Y.; Talalay, P. Broccoli sprouts: An exceptionally rich source of inducers of enzymes that protect against chemical carcinogens. Proc. Natl. Acad. Sci. USA 1997, 94, 10367–10372. [Google Scholar] [CrossRef] [PubMed]
- Atwell, L.L.; Hsu, A.; Wong, C.P.; Stevens, J.F.; Bella, D.; Yu, T.-W.; Pereira, C.B.; Löhr, C.V.; Christensen, J.M.; Dashwood, R.H.; et al. Absorption and chemopreventive targets of sulforaphane in humans following consumption of broccoli sprouts or a myrosinase-treated broccoli sprout extract. Mol. Nutr. Food Res. 2015, 59, 424–433. [Google Scholar] [CrossRef] [PubMed]
- Hu, R.; Khor, T.O.; Shen, G.; Jeong, W.-S.; Hebbar, V.; Chen, C.; Xu, C.; Reddy, B.; Chada, K.; Kong, A.-N.T. Cancer chemoprevention of intestinal polyposis in ApcMin/+ mice by sulforaphane, a natural product derived from cruciferous vegetable. Carcinogenesis 2006, 27, 2038–2046. [Google Scholar] [CrossRef] [PubMed]
- Lamartiniere, C.A.; Cotroneo, M.S.; Fritz, W.A.; Wang, J.; Mentor-Marcel, R.; Elgavish, A. Genistein chemoprevention: Timing and mechanisms of action in murine mammary and prostate. J. Nutr. 2002, 132, 552S–558S. [Google Scholar] [CrossRef] [PubMed]
- Jin, Z.; MacDonald, R.S. Soy isoflavones increase latency of spontaneous mammary tumors in mice. J. Nutr. 2002, 132, 3186–3190. [Google Scholar] [CrossRef] [PubMed]
- Ferralli, J.; Chiquet-Ehrismann, R.; Degen, M. KLF4A stimulates breast cancer cell proliferation by acting as a KLF4 antagonist. Oncotarget 2016, 7, 45608. [Google Scholar] [CrossRef] [PubMed]
- Ema, M.; Mori, D.; Niwa, H.; Hasegawa, Y.; Yamanaka, Y.; Hitoshi, S.; Mimura, J.; Kawabe, Y.-I.; Hosoya, T.; Morita, M. Krüppel-like factor 5 is essential for blastocyst development and the normal self-renewal of mouse ESCs. Cell Stem Cell 2008, 3, 555–567. [Google Scholar] [CrossRef] [PubMed]
- Wong, C.W.; Hou, P.S.; Tseng, S.F.; Chien, C.L.; Wu, K.J.; Chen, H.F.; Ho, H.N.; Kyo, S.; Teng, S.C. Krüppel-like transcription factor 4 contributes to maintenance of telomerase activity in stem cells. Stem Cells 2010, 28, 1510–1517. [Google Scholar] [CrossRef] [PubMed]
- Pons, D.G.; Nadal-Serrano, M.; Blanquer-Rossello, M.; Sastre-Serra, J.; Oliver, J.; Roca, P. Genistein modulates proliferation and mitochondrial functionality in breast cancer cells depending on ERalpha/ERbeta ratio. J. Cell. Biochem. 2014, 115, 949–958. [Google Scholar] [CrossRef] [PubMed]
- Yori, J.L.; Seachrist, D.D.; Johnson, E.; Lozada, K.L.; Abdul-Karim, F.W.; Chodosh, L.A.; Schiemann, W.P.; Keri, R.A. Krüppel-like factor 4 inhibits tumorigenic progression and metastasis in a mouse model of breast cancer. Neoplasia 2011, 13, 601–610. [Google Scholar] [CrossRef] [PubMed]
- Su, Y.; Simmen, R.C. Soy isoflavone genistein upregulates epithelial adhesion molecule E-cadherin expression and attenuates β-catenin signaling in mammary epithelial cells. Carcinogenesis 2008, 30, 331–339. [Google Scholar] [CrossRef] [PubMed]
- Singh, S.V.; Warin, R.; Xiao, D.; Powolny, A.A.; Stan, S.D.; Arlotti, J.A.; Zeng, Y.; Hahm, E.-R.; Marynowski, S.W.; Bommareddy, A. Sulforaphane inhibits prostate carcinogenesis and pulmonary metastasis in tramp mice in association with increased cytotoxicity of natural killer cells. Cancer Res. 2009, 69, 2117–2125. [Google Scholar] [CrossRef] [PubMed]
- Shen, G.; Khor, T.O.; Hu, R.; Yu, S.; Nair, S.; Ho, C.-T.; Reddy, B.S.; Huang, M.-T.; Newmark, H.L.; Kong, A.-N.T. Chemoprevention of familial adenomatous polyposis by natural dietary compounds sulforaphane and dibenzoylmethane alone and in combination in ApcMin/+ mouse. Cancer Res. 2007, 67, 9937–9944. [Google Scholar] [CrossRef] [PubMed]
- Fritz, W.A.; Wang, J.; Eltoum, I.-E.; Lamartiniere, C.A. Dietary genistein downregulates androgen and estrogen receptor expression in the rat prostate. Mol. Cell. Endocrinol. 2002, 186, 89–99. [Google Scholar] [CrossRef]
- Lee, A.V.; Oesterreich, S.; Davidson, N.E. Mcf-7 cells—Changing the course of breast cancer research and care for 45 years. JNCI J. Natl. Cancer Inst. 2015, 107. [Google Scholar] [CrossRef] [PubMed]
- Lacroix, M.; Leclercq, G. Relevance of breast cancer cell lines as models for breast tumours: An update. Breast Cancer Res. Treat. 2004, 83, 249–289. [Google Scholar] [CrossRef] [PubMed]
- Neve, R.M.; Chin, K.; Fridlyand, J.; Yeh, J.; Baehner, F.L.; Fevr, T.; Clark, L.; Bayani, N.; Coppe, J.-P.; Tong, F. A collection of breast cancer cell lines for the study of functionally distinct cancer subtypes. Cancer Cell 2006, 10, 515–527. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Landen, C.N.; Li, Y.; Alvarez, R.D.; Tollefsbol, T.O. Enhancement of cisplatin-mediated apoptosis in ovarian cancer cells through potentiating G2/M arrest and p21 upregulation by combinatorial epigallocatechin gallate and sulforaphane. J. Oncol. 2013, 2013, 872957. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Meeran, S.M.; Patel, S.N.; Chen, H.; Hardy, T.M.; Tollefsbol, T.O. Epigenetic reactivation of estrogen receptor-α (ERα) by genistein enhances hormonal therapy sensitivity in ERα-negative breast cancer. Mol. Cancer 2013, 12, 9. [Google Scholar] [CrossRef] [PubMed]

| Cell Lines | SFN | GEN | Average CI |
|---|---|---|---|
| MCF-7 | 5.0 µM | 10.0 µM | 0.40468 |
| MCF-7 | 5.0 µM | 15.0 µM | 0.29520 |
| MDA-MB-231 | 5.0 µM | 10.0 µM | 0.45254 |
| MDA-MB-231 | 5.0 µM | 15.0 µM | 0.56832 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Paul, B.; Li, Y.; Tollefsbol, T.O. The Effects of Combinatorial Genistein and Sulforaphane in Breast Tumor Inhibition: Role in Epigenetic Regulation. Int. J. Mol. Sci. 2018, 19, 1754. https://doi.org/10.3390/ijms19061754
Paul B, Li Y, Tollefsbol TO. The Effects of Combinatorial Genistein and Sulforaphane in Breast Tumor Inhibition: Role in Epigenetic Regulation. International Journal of Molecular Sciences. 2018; 19(6):1754. https://doi.org/10.3390/ijms19061754
Chicago/Turabian StylePaul, Bidisha, Yuanyuan Li, and Trygve O. Tollefsbol. 2018. "The Effects of Combinatorial Genistein and Sulforaphane in Breast Tumor Inhibition: Role in Epigenetic Regulation" International Journal of Molecular Sciences 19, no. 6: 1754. https://doi.org/10.3390/ijms19061754
APA StylePaul, B., Li, Y., & Tollefsbol, T. O. (2018). The Effects of Combinatorial Genistein and Sulforaphane in Breast Tumor Inhibition: Role in Epigenetic Regulation. International Journal of Molecular Sciences, 19(6), 1754. https://doi.org/10.3390/ijms19061754