Why Two? On the Role of (A-)Symmetry in Negative Supercoiling of DNA by Gyrase
Abstract
1. Introduction
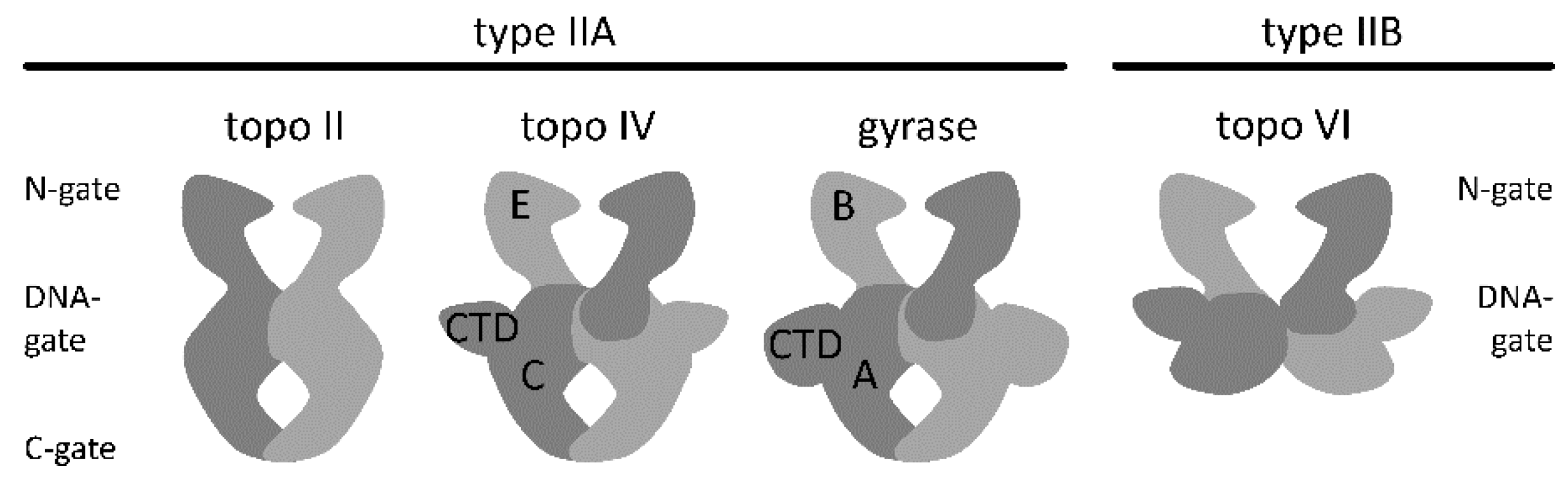
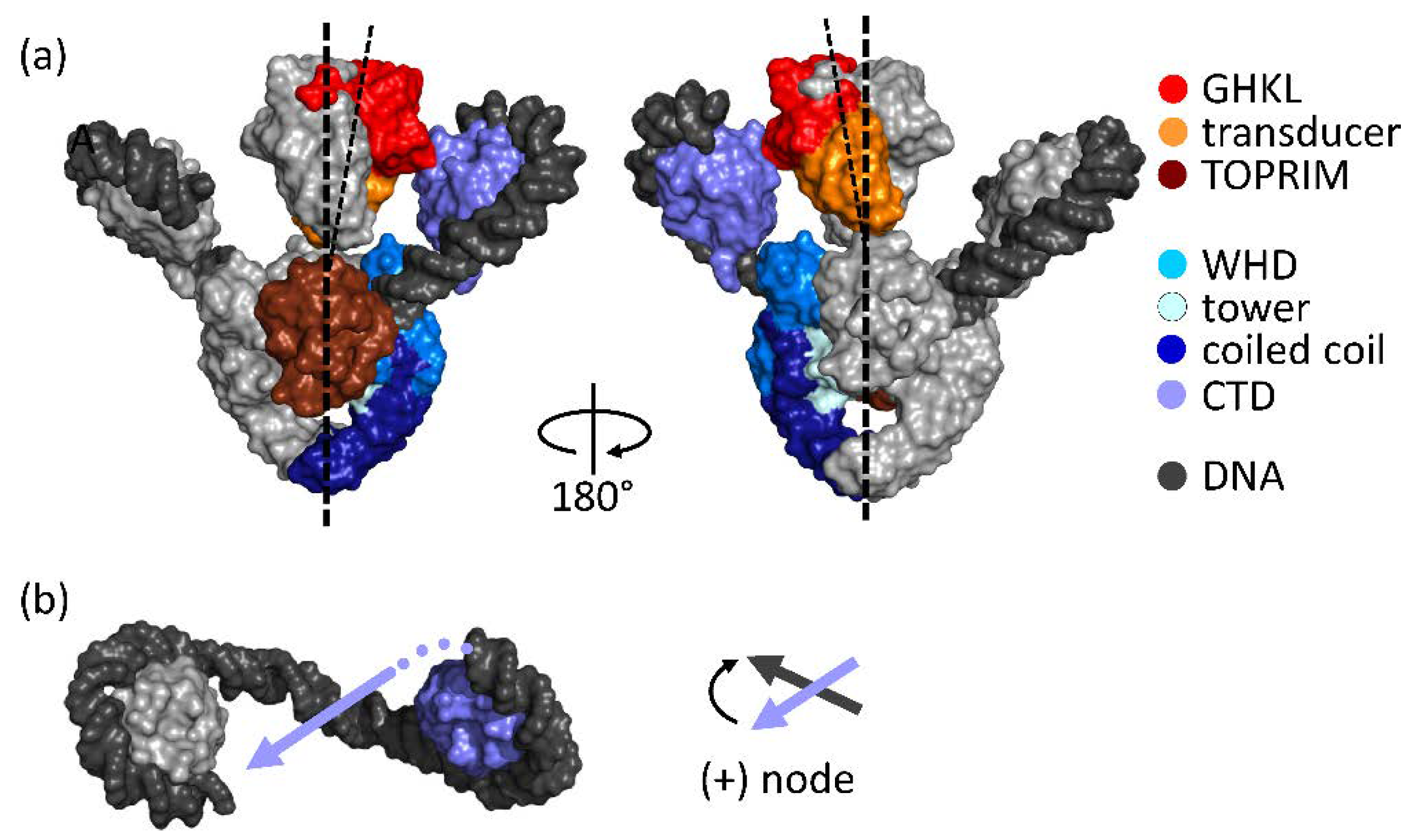
2. Architecture of Gyrase
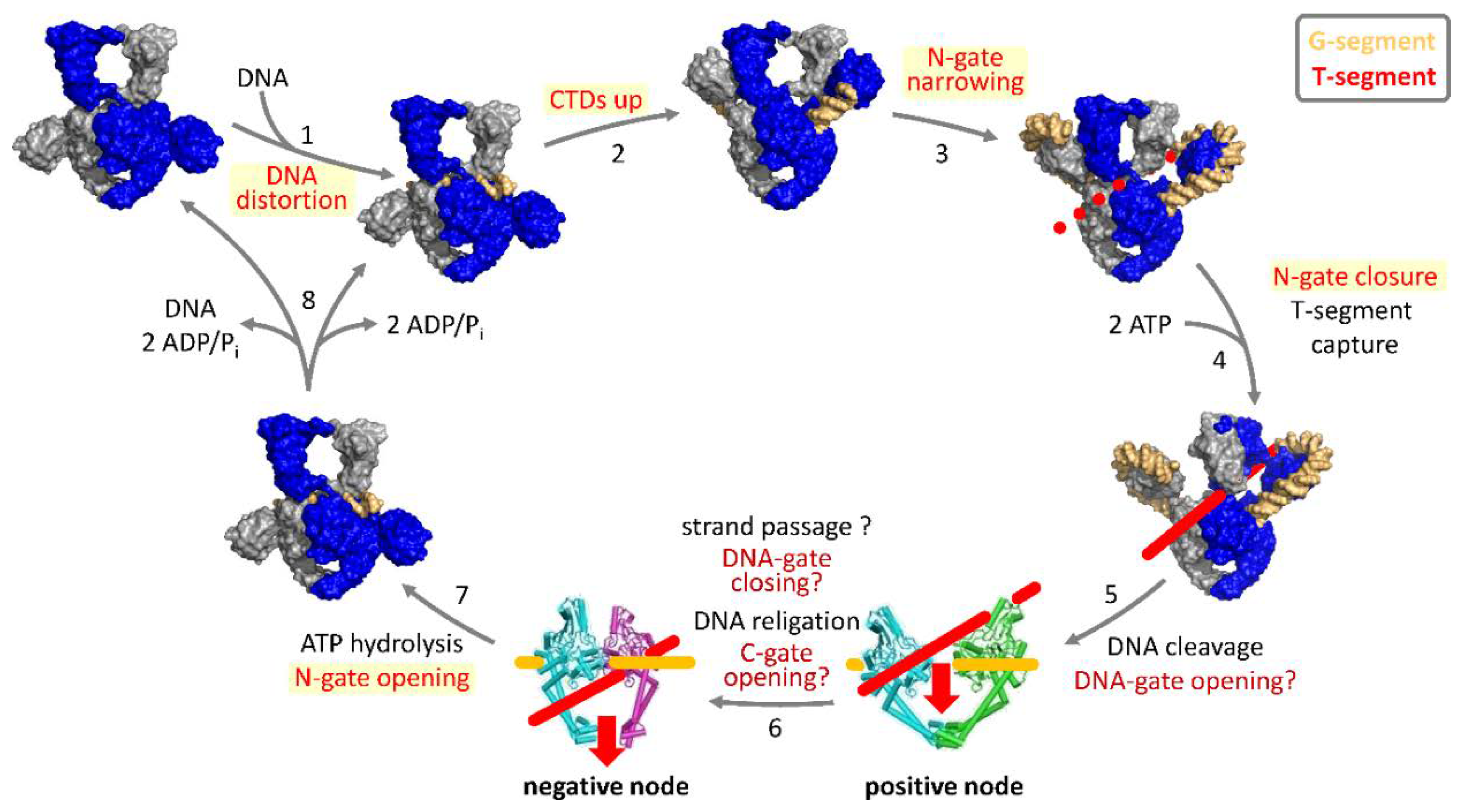
3. Strand-Passage Mechanism for DNA Supercoiling
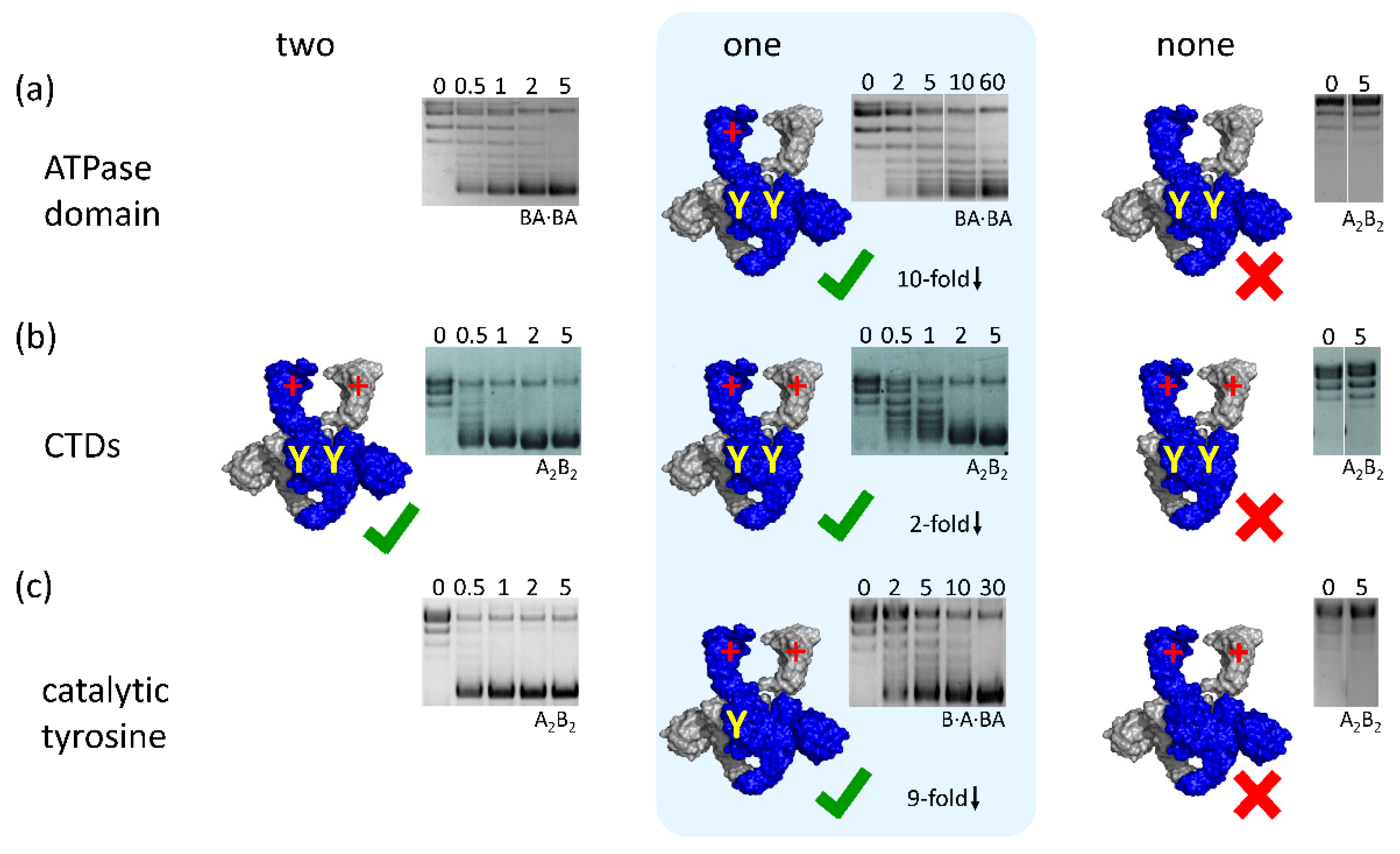
4. Symmetry, Asymmetry, and Inter-Subunit Communication in Gyrase
4.1. Binding and Hydrolysis of a Single ATP is Sufficient for N-Gate Closure, Trapping of a T-Segment, and DNA Supercoiling
4.2. Gyrase with a Single CTD Catalyzes Negative Supercoiling of DNA in Steps of Two
4.3. Gyrase with a Single Catalytic Tyrosine: DNA Supercoiling in the Absence of Strand Passage
4.4. A Minimal, Non-Redundant Gyrase? Implications for Inter-Domain Communication
5. Conclusions
Acknowledgments
Conflicts of Interest
Abbreviations
| ADPNP | 5′-adenylyl-β,γ-imidotriphosphate |
| CTD | C-terminal domain (of GyrA) |
| FRET | Förster resonance energy transfer |
| GHKL | GyrB-Hsp90-histidine/serine protein kinases-MutL |
| G-segment | gate segment |
| GyrA | GyrA subunit of gyrase |
| GyrB | GyrB subunit of gyrase |
| GyrBA | GyrB-GyrA fusion protein |
| NTD | N-terminal domain (of GyrA) |
| sc | supercoil |
| topo | topoisomerase |
| TOPRIM | topoisomerase-primase |
| T-segment | transport segment |
| WHD | winged-helix domain |
References
- Schoeffler, A.J.; Berger, J.M. DNA topoisomerases: Harnessing and constraining energy to govern chromosome topology. Q. Rev. Biophys. 2008, 41, 41–101. [Google Scholar] [CrossRef] [PubMed]
- Kouzine, F.; Wojtowicz, D.; Yamane, A.; Resch, W.; Kieffer-Kwon, K.R.; Bandle, R.; Nelson, S.; Nakahashi, H.; Awasthi, P.; Feigenbaum, L.; et al. Global regulation of promoter melting in naive lymphocytes. Cell 2013, 153, 988–999. [Google Scholar] [CrossRef] [PubMed]
- Kouzine, F.; Gupta, A.; Baranello, L.; Wojtowicz, D.; Ben-Aissa, K.; Liu, J.; Przytycka, T.M.; Levens, D. Transcription-dependent dynamic supercoiling is a short-range genomic force. Nat. Struct. Mol. Biol. 2013, 20, 396–403. [Google Scholar] [CrossRef] [PubMed]
- Vos, S.M.; Tretter, E.M.; Schmidt, B.H.; Berger, J.M. All tangled up: How cells direct, manage and exploit topoisomerase function. Nat. Rev. Mol. Cell Biol. 2011, 12, 827–841. [Google Scholar] [CrossRef] [PubMed]
- Schoeffler, A.J.; Berger, J.M. Recent advances in understanding structure-function relationships in the type II topoisomerase mechanism. Biochem. Soc. Trans. 2005, 33, 1465–1470. [Google Scholar] [CrossRef] [PubMed]
- Horowitz, D.S.; Wang, J.C. Mapping the active site tyrosine of Escherichia coli DNA gyrase. J. Biol. Chem. 1987, 262, 5339–5344. [Google Scholar] [PubMed]
- Goto, T.; Wang, J.C. Yeast DNA topoisomerase II. An ATP-dependent type II topoisomerase that catalyzes the catenation, decatenation, unknotting, and relaxation of double-stranded DNA rings. J. Biol. Chem. 1982, 257, 5866–5872. [Google Scholar] [PubMed]
- Gellert, M.; Mizuuchi, K.; O’Dea, M.H.; Nash, H.A. DNA gyrase: An enzyme that introduces superhelical turns into DNA. Proc. Natl. Acad. Sci. USA 1976, 73, 3872–3876. [Google Scholar] [CrossRef] [PubMed]
- Peng, H.; Marians, K.J. Decatenation activity of topoisomerase IV during oriC and pBR322 DNA replication in vitro. Proc. Natl. Acad. Sci. USA 1993, 90, 8571–8575. [Google Scholar] [CrossRef] [PubMed]
- Buhler, C.; Lebbink, J.H.; Bocs, C.; Ladenstein, R.; Forterre, P. DNA topoisomerase VI generates ATP-dependent double-strand breaks with two-nucleotide overhangs. J. Biol. Chem. 2001, 276, 37215–37222. [Google Scholar] [CrossRef] [PubMed]
- Corbett, K.D.; Benedetti, P.; Berger, J.M. Holoenzyme assembly and ATP-mediated conformational dynamics of topoisomerase VI. Nat. Struct. Mol. Biol. 2007, 14, 611–619. [Google Scholar] [CrossRef] [PubMed]
- Roca, J.; Berger, J.M.; Harrison, S.C.; Wang, J.C. DNA transport by a type II topoisomerase: Direct evidence for a two-gate mechanism. Proc. Natl. Acad. Sci. USA 1996, 93, 4057–4062. [Google Scholar] [CrossRef] [PubMed]
- Williams, N.L.; Maxwell, A. Probing the two-gate mechanism of DNA gyrase using cysteine cross-linking. Biochemistry 1999, 38, 13502–13511. [Google Scholar] [CrossRef] [PubMed]
- Williams, N.L.; Maxwell, A. Locking the DNA gate of DNA gyrase: Investigating the effects on DNA cleavage and ATP hydrolysis. Biochemistry 1999, 38, 14157–14164. [Google Scholar] [CrossRef] [PubMed]
- Sissi, C.; Palumbo, M. In front of and behind the replication fork: Bacterial type IIA topoisomerases. Cell. Mol. Life Sci. 2010, 67, 2001–2024. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, W.; Sala, C.; Hegde, S.R.; Jha, R.K.; Cole, S.T.; Nagaraja, V. Transcription facilitated genome-wide recruitment of topoisomerase I and DNA gyrase. PLoS Genet. 2017, 13, e1006754. [Google Scholar] [CrossRef] [PubMed]
- Collin, F.; Karkare, S.; Maxwell, A. Exploiting bacterial DNA gyrase as a drug target: Current state and perspectives. Appl. Microbiol. Biotechnol. 2011, 92, 479–497. [Google Scholar] [CrossRef] [PubMed]
- Sugino, A.; Higgins, N.P.; Cozzarelli, N.R. DNA gyrase subunit stoichiometry and the covalent attachment of subunit A to DNA during DNA cleavage. Nucleic Acids Res. 1980, 8, 3865–3874. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Morais Cabral, J.H.; Jackson, A.P.; Smith, C.V.; Shikotra, N.; Maxwell, A.; Liddington, R.C. Crystal structure of the breakage-reunion domain of DNA gyrase. Nature 1997, 388, 903–906. [Google Scholar] [CrossRef] [PubMed]
- Papillon, J.; Menetret, J.F.; Batisse, C.; Helye, R.; Schultz, P.; Potier, N.; Lamour, V. Structural insight into negative DNA supercoiling by DNA gyrase, a bacterial type 2A DNA topoisomerase. Nucleic Acids Res. 2013, 41, 7815–7827. [Google Scholar] [CrossRef] [PubMed]
- Gubaev, A.; Klostermeier, D. DNA-induced narrowing of the gyrase N-gate coordinates T-segment capture and strand passage. Proc. Natl. Acad. Sci. USA 2011, 108, 14085–14090. [Google Scholar] [CrossRef] [PubMed]
- Gubaev, A.; Klostermeier, D. Potassium ions are required for nucleotide-induced closure of gyrase N-gate. J. Biol. Chem. 2012, 287, 10916–10921. [Google Scholar] [CrossRef] [PubMed]
- Wigley, D.B.; Davies, G.J.; Dodson, E.J.; Maxwell, A.; Dodson, G. Crystal structure of an N-terminal fragment of the DNA gyrase B protein. Nature 1991, 351, 624–629. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Zhang, Z.; Mitchenall, L.A.; Maxwell, A.; Deng, J.; Zhang, H.; Zhou, Y.; Chen, Y.Y.; Wang, D.C.; Zhang, X.E.; et al. The dimer state of GyrB is an active form: Implications for the initial complex assembly and processive strand passage. Nucleic Acids Res. 2011, 39, 8488–8502. [Google Scholar] [CrossRef] [PubMed]
- Gottler, T.; Klostermeier, D. Dissection of the Nucleotide Cycle of B. subtilis DNA Gyrase and its Modulation by DNA. J. Mol. Biol. 2007, 367, 1392–1404. [Google Scholar] [CrossRef] [PubMed]
- Costenaro, L.; Grossmann, J.G.; Ebel, C.; Maxwell, A. Small-angle X-ray scattering reveals the solution structure of the full-length DNA gyrase a subunit. Structure 2005, 13, 287–296. [Google Scholar] [CrossRef] [PubMed]
- Lanz, M.A.; Klostermeier, D. Guiding strand passage: DNA-induced movement of the gyrase C-terminal domains defines an early step in the supercoiling cycle. Nucleic Acids Res. 2011, 39, 9681–9694. [Google Scholar] [CrossRef] [PubMed]
- Corbett, K.D.; Shultzaberger, R.K.; Berger, J.M. The C-terminal domain of DNA gyrase A adopts a DNA-bending beta-pinwheel fold. Proc. Natl. Acad. Sci. USA 2004, 101, 7293–7298. [Google Scholar] [CrossRef] [PubMed]
- Reece, R.J.; Maxwell, A. The C-terminal domain of the Escherichia coli DNA gyrase A subunit is a DNA-binding protein. Nucleic Acids Res. 1991, 19, 1399–1405. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.F.; Wang, J.C. DNA-DNA gyrase complex: The wrapping of the DNA duplex outside the enzyme. Cell 1978, 15, 979–984. [Google Scholar] [CrossRef]
- Orphanides, G.; Maxwell, A. Evidence for a conformational change in the DNA gyrase-DNA complex from hydroxyl radical footprinting. Nucleic Acids Res. 1994, 22, 1567–1575. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Nollmann, M.; Stone, M.D.; Bryant, Z.; Gore, J.; Crisona, N.J.; Hong, S.C.; Mitelheiser, S.; Maxwell, A.; Bustamante, C.; Cozzarelli, N.R. Multiple modes of Escherichia coli DNA gyrase activity revealed by force and torque. Nat. Struct. Mol. Biol. 2007, 14, 264–271. [Google Scholar] [CrossRef] [PubMed]
- Brown, P.O.; Cozzarelli, N.R. A sign inversion mechanism for enzymatic supercoiling of DNA. Science 1979, 206, 1081–1083. [Google Scholar] [CrossRef] [PubMed]
- Mizuuchi, K.; Fisher, L.M.; O’Dea, M.H.; Gellert, M. DNA gyrase action involves the introduction of transient double-strand breaks into DNA. Proc. Natl. Acad. Sci. USA 1980, 77, 1847–1851. [Google Scholar] [CrossRef] [PubMed]
- Hartmann, S.; Weidlich, D.; Klostermeier, D. Single-Molecule Confocal FRET Microscopy to Dissect Conformational Changes in the Catalytic Cycle of DNA Topoisomerases. Methods Enzymol. 2016, 581, 317–351. [Google Scholar] [CrossRef]
- Gubaev, A.; Klostermeier, D. The mechanism of negative DNA supercoiling: A cascade of DNA-induced conformational changes prepares gyrase for strand passage. DNA Repair 2014, 16, 23–34. [Google Scholar] [CrossRef] [PubMed]
- Gubaev, A.; Hilbert, M.; Klostermeier, D. The DNA gate of Bacillus subtilis gyrase is predominantly in the closed conformation during the DNA supercoiling reaction. Proc. Natl. Acad. Sci. USA 2009, 106, 13278–13283. [Google Scholar] [CrossRef] [PubMed]
- Lanz, M.A.; Klostermeier, D. The GyrA-box determines the geometry of DNA bound to gyrase and couples DNA binding to the nucleotide cycle. Nucleic Acids Res. 2012, 40, 10893–10903. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Rudolph, M.G.; Klostermeier, D. Mapping the Spectrum of Conformational States of the DNA- and C-Gates in Bacillus subtilis Gyrase. J. Mol. Biol. 2013, 425, 2632–2640. [Google Scholar] [CrossRef] [PubMed]
- Gubaev, A.; Weidlich, D.; Klostermeier, D. DNA gyrase with a single catalytic tyrosine can catalyze DNA supercoiling by a nicking-closing mechanism. Nucleic Acids Res. 2016, 44, 10354–10366. [Google Scholar] [CrossRef] [PubMed]
- Hartmann, S.; Gubaev, A.; Klostermeier, D. Binding and Hydrolysis of a Single ATP Is Sufficient for N-Gate Closure and DNA Supercoiling by Gyrase. J. Mol. Biol. 2017, 429, 3717–3729. [Google Scholar] [CrossRef] [PubMed]
- Stelljes, J.T.; Weidlich, D.; Gubaev, A.; Klostermeier, D. Gyrase containing a single C-terminal domain catalyzes negative supercoiling of DNA by decreasing the linking number in steps of two. Nucleic Acids Res. 2018. [Google Scholar] [CrossRef]
- Harkins, T.T.; Lewis, T.J.; Lindsley, J.E. Pre-steady-state analysis of ATP hydrolysis by Saccharomyces cerevisiae DNA topoisomerase II. 2. Kinetic mechanism for the sequential hydrolysis of two ATP. Biochemistry 1998, 37, 7299–7312. [Google Scholar] [CrossRef] [PubMed]
- Harkins, T.T.; Lindsley, J.E. Pre-steady-state analysis of ATP hydrolysis by Saccharomyces cerevisiae DNA topoisomerase II. 1. A DNA-dependent burst in ATP hydrolysis. Biochemistry 1998, 37, 7292–7298. [Google Scholar] [CrossRef] [PubMed]
- Baird, C.L.; Harkins, T.T.; Morris, S.K.; Lindsley, J.E. Topoisomerase II drives DNA transport by hydrolyzing one ATP. Proc. Natl. Acad. Sci. USA 1999, 96, 13685–13690. [Google Scholar] [CrossRef] [PubMed]
- Lindsley, J.E.; Wang, J.C. Study of allosteric communication between protomers by immunotagging. Nature 1993, 361, 749–750. [Google Scholar] [CrossRef] [PubMed]
- Kampranis, S.C.; Maxwell, A. Hydrolysis of ATP at only one GyrB subunit is sufficient to promote supercoiling by DNA gyrase. J. Biol. Chem. 1998, 273, 26305–26309. [Google Scholar] [CrossRef] [PubMed]
- Jackson, A.P.; Maxwell, A. Identifying the catalytic residue of the ATPase reaction of DNA gyrase. Proc. Natl. Acad. Sci. USA 1993, 90, 11232–11236. [Google Scholar] [CrossRef] [PubMed]
- Gross, C.H.; Parsons, J.D.; Grossman, T.H.; Charifson, P.S.; Bellon, S.; Jernee, J.; Dwyer, M.; Chambers, S.P.; Markland, W.; Botfield, M.; et al. Active-site residues of Escherichia coli DNA gyrase required in coupling ATP hydrolysis to DNA supercoiling and amino acid substitutions leading to novobiocin resistance. Antimicrob. Agents Chemother. 2003, 47, 1037–1046. [Google Scholar] [CrossRef] [PubMed]
- Kampranis, S.C.; Gormley, N.A.; Tranter, R.; Orphanides, G.; Maxwell, A. Probing the binding of coumarins and cyclothialidines to DNA gyrase. Biochemistry 1999, 38, 1967–1976. [Google Scholar] [CrossRef] [PubMed]
- Sugino, A.; Higgins, N.P.; Brown, P.O.; Peebles, C.L.; Cozzarelli, N.R. Energy coupling in DNA gyrase and the mechanism of action of novobiocin. Proc. Natl. Acad. Sci. USA 1978, 75, 4838–4842. [Google Scholar] [CrossRef] [PubMed]
- Ruthenburg, A.J.; Graybosch, D.M.; Huetsch, J.C.; Verdine, G.L. A superhelical spiral in the Escherichia coli DNA gyrase A C-terminal domain imparts unidirectional supercoiling bias. J. Biol. Chem. 2005, 280, 26177–26184. [Google Scholar] [CrossRef] [PubMed]
- Tretter, E.M.; Berger, J.M. Mechanisms For Defining Supercoiling Setpoint By DNA Gyrase Orthologs II. The shape of the GyrA CTD is not a sole determinant for controlling supercoiling efficiency. J. Biol. Chem. 2012, 287, 18645–18654. [Google Scholar] [CrossRef] [PubMed]
- Tretter, E.M.; Berger, J.M. Mechanisms for defining the supercoiling setpoint of DNA gyrase orthologs I. A non-conserved acidic C-terminal tail modulates E. coli gyrase activity. J. Biol. Chem. 2012, 287, 18636–18644. [Google Scholar] [CrossRef] [PubMed]
- Lanz, M.A.; Farhat, M.; Klostermeier, D. The acidic C-terminal tail of the GyrA subunit moderates the DNA supercoiling activity of Bacillus subtilis gyrase. J. Biol. Chem. 2014, 289, 12275–12285. [Google Scholar] [CrossRef] [PubMed]
- Kampranis, S.C.; Maxwell, A. Conversion of DNA gyrase into a conventional type II topoisomerase. Proc. Natl. Acad. Sci. USA 1996, 93, 14416–14421. [Google Scholar] [CrossRef] [PubMed]
- Critchlow, S.E.; Maxwell, A. DNA cleavage is not required for the binding of quinolone drugs to the DNA gyrase-DNA complex. Biochemistry 1996, 35, 7387–7393. [Google Scholar] [CrossRef] [PubMed]
- Sugino, A.; Peebles, C.L.; Kreuzer, K.N.; Cozzarelli, N.R. Mechanism of action of nalidixic acid: Purification of Escherichia coli nalA gene product and its relationship to DNA gyrase and a novel nicking-closing enzyme. Proc. Natl. Acad. Sci. USA 1977, 74, 4767–4771. [Google Scholar] [CrossRef] [PubMed]
- Mizuuchi, K.; O’Dea, M.H.; Gellert, M. DNA gyrase: Subunit structure and ATPase activity of the purified enzyme. Proc. Natl. Acad. Sci. USA 1978, 75, 5960–5963. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.F.; Wang, J.C. Micrococcus luteus DNA gyrase: Active components and a model for its supercoiling of DNA. Proc. Natl. Acad. Sci. USA 1978, 75, 2098–2102. [Google Scholar] [CrossRef] [PubMed]
- Kampranis, S.C.; Maxwell, A. Conformational changes in DNA gyrase revealed by limited proteolysis. J. Biol. Chem. 1998, 273, 22606–22614. [Google Scholar] [CrossRef] [PubMed]
- Morrison, A.; Cozzarelli, N.R. Contacts between DNA gyrase and its binding site on DNA: Features of symmetry and asymmetry revealed by protection from nucleases. Proc. Natl. Acad. Sci. USA 1981, 78, 1416–1420. [Google Scholar] [CrossRef] [PubMed]
- Ali, J.A.; Orphanides, G.; Maxwell, A. Nucleotide binding to the 43-kilodalton N-terminal fragment of the DNA gyrase B protein. Biochemistry 1995, 34, 9801–9808. [Google Scholar] [CrossRef] [PubMed]

© 2018 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Klostermeier, D. Why Two? On the Role of (A-)Symmetry in Negative Supercoiling of DNA by Gyrase. Int. J. Mol. Sci. 2018, 19, 1489. https://doi.org/10.3390/ijms19051489
Klostermeier D. Why Two? On the Role of (A-)Symmetry in Negative Supercoiling of DNA by Gyrase. International Journal of Molecular Sciences. 2018; 19(5):1489. https://doi.org/10.3390/ijms19051489
Chicago/Turabian StyleKlostermeier, Dagmar. 2018. "Why Two? On the Role of (A-)Symmetry in Negative Supercoiling of DNA by Gyrase" International Journal of Molecular Sciences 19, no. 5: 1489. https://doi.org/10.3390/ijms19051489
APA StyleKlostermeier, D. (2018). Why Two? On the Role of (A-)Symmetry in Negative Supercoiling of DNA by Gyrase. International Journal of Molecular Sciences, 19(5), 1489. https://doi.org/10.3390/ijms19051489




