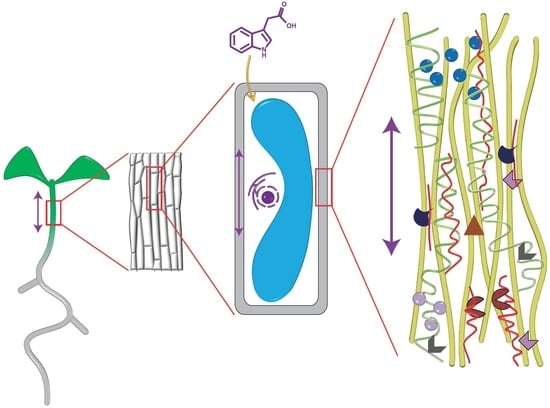
The Role of Auxin in Cell Wall Expansion
Abstract
1. Introduction
2. Plant Cell Walls
2.1. Cellulose Microfibrils (CMFs)
2.2. Hemicelluloses and Pectins
2.3. Structural Proteins
2.4. Interactions within the Cell Wall
3. The Role of Auxin in Wall Extension
3.1. Auxin Signaling Stimulates Cell Elongation
3.2. Auxin and Cell Wall-Related Genes
3.3. Auxin Induces Acid Growth
3.4. EXPANSINs Mediate Acid Growth
3.5. Cellulose and Xyloglucan Modification during Wall Expansion
3.6. Pectin Methylesterification and Its Consequences in Wall Loosening
3.7. Crosslinking of the Wall Polysaccharides
4. Conclusions
Acknowledgements
Author Contributions
Conflicts of Interest
References
- Burton, R.A.; Gidley, M.J.; Fincher, G.B. Heterogeneity in the chemistry, structure and function of plant cell walls. Nat. Chem. Biol. 2010, 6, 724–732. [Google Scholar] [CrossRef] [PubMed]
- Wolf, S.; Hématy, K.; Höfte, H. Growth Control and Cell Wall Signaling in Plants. Annu. Rev. Plant Biol. 2012, 63, 381–407. [Google Scholar] [CrossRef] [PubMed]
- Cosgrove, D.J. Growth of the plant cell wall. Nat. Rev. Mol. Cell Biol. 2005, 6, 850–861. [Google Scholar] [CrossRef] [PubMed]
- Fry, S.C.; Love, J.; Bryant, J.A. Plant Cell Wall Polymers. In Biofuels and Bioenergy; John Wiley & Sons, Ltd.: Hoboken, NJ, USA, 2017; pp. 59–87. ISBN 978-1-11-835055-3. [Google Scholar]
- Carpita, N.C.; Gibeaut, D.M. Structural models of primary cell walls in flowering plants: Consistency of molecular structure with the physical properties of the walls during growth. Plant J. 1993, 3, 1–30. [Google Scholar] [CrossRef] [PubMed]
- Cosgrove, D.J. Wall Extensiblity: Its Nature, Measurement and Relationship to Plant Cell Growth. New Phytol. 1993, 124, 1–23. [Google Scholar] [CrossRef] [PubMed]
- Cosgrove, D.J. Diffuse growth of plant cell walls. Plant Physiol. 2018, 176, 16–27. [Google Scholar] [CrossRef] [PubMed]
- Cosgrove, D.J. Plant cell wall extensibility: Connecting plant cell growth with cell wall structure, mechanics, and the action of wall-modifying enzymes. J. Exp. Bot. 2016, 67, 463–476. [Google Scholar] [CrossRef] [PubMed]
- Guerriero, G.; Hausman, J.F.; Cai, G. No stress! Relax! Mechanisms governing growth and shape in plant cells. Int. J. Mol. Sci. 2014, 15, 5094–5114. [Google Scholar] [CrossRef] [PubMed]
- Wei, C.; Lintilhac, P.M. Loss of stability—A new model for stress relaxation in plant cell walls. J. Theor. Biol. 2003, 224, 305–312. [Google Scholar] [CrossRef]
- Wei, C.; Lintilhac, P.M. Loss of Stability: A New Look at the Physics of Cell Wall Behavior during Plant Cell Growth. Plant Physiol. 2007, 145, 763–772. [Google Scholar] [CrossRef] [PubMed]
- Kroeger, J.H.; Zerzour, R.; Geitmann, A. Regulator or driving force? The role of turgor pressure in oscillatory plant cell growth. PLoS ONE 2011, 6, e18549. [Google Scholar] [CrossRef] [PubMed]
- Green, P.B. Mechanism for Plant Cellular Morphogenesis. Science 1962, 138, 1404–1405. [Google Scholar] [CrossRef] [PubMed]
- Kutschera, U. The current status of the acid-growth hypothesis. New Phytol. 1994, 126, 549–569. [Google Scholar] [CrossRef]
- Rayle, D.L.; Cleland, R.E. Evidence that Auxin-induced Growth of Soybean Hypocotyls Involves Proton Excretion. Plant Physiol. 1980, 66, 433–437. [Google Scholar] [CrossRef] [PubMed]
- Rayle, D.L.; Cleland, R.E. The Acid Growth Theory of auxin-induced cell elongation is alive and well. Plant Physiol. 1992, 99, 1271–1274. [Google Scholar] [CrossRef] [PubMed]
- Ray, P.M. Cell wall synthesis and cell elongation in oat coleoptile tissue. Am. J. Bot. 1962, 49, 928–939. [Google Scholar] [CrossRef]
- Ray, P.M.; Ruesink, A.W. Kinetics experiments on the nature of the growth mechanism in oat coleoptile cellls. Dev. Biol. 1962, 4, 377–397. [Google Scholar] [CrossRef]
- Derbyshire, P.; Findlay, K.; McCann, M.C.; Roberts, K. Cell elongation in Arabidopsis hypocotyls involves dynamic changes in cell wall thickness. J. Exp. Bot. 2007, 58, 2079–2089. [Google Scholar] [CrossRef] [PubMed]
- Freshour, G.; Clay, R.P.; Fuller, M.S.; Albersheim, P.; Darvill, A.G.; Hahn, M.G. Developmental and Tissue-Specific Structural Alterations of the Cell-Wall Polysaccharides of Arabidopsis thaliana Roots. Plant Physiol. 1996, 110, 1413–1429. [Google Scholar] [CrossRef] [PubMed]
- Phyo, P.; Wang, T.; Kiemle, S.N.; O’Neill, H.; Pingali, S.V.; Hong, M.; Cosgrove, D.J. Gradients in wall mechanics and polysaccharides along growing inflorescence stems. Plant Physiol. 2017. [Google Scholar] [CrossRef] [PubMed]
- Majda, M.; Grones, P.; Sintorn, I.M.; Vain, T.; Milani, P.; Krupinski, P.; Zagórska-Marek, B.; Viotti, C.; Jönsson, H.; Mellerowicz, E.J.; et al. Mechanochemical Polarization of Contiguous Cell Walls Shapes Plant Pavement Cells. Dev. Cell 2017, 43, 290–304. [Google Scholar] [CrossRef] [PubMed]
- Refrégier, G.; Pelletier, S.; Jaillard, D.; Höfte, H. Interaction between wall deposition and cell elongation in dark-grown hypocotyl cells in Arabidopsis. Plant Physiol. 2004, 135, 959–968. [Google Scholar] [CrossRef] [PubMed]
- Majda, M. Role of the Cell Wall in Cell Shape Acquisition; Swedish University of Agricultural Sciences: Umeå, Sweden, 2018; pp. 1652–6880. ISBN 978-9-17-760160-9. [Google Scholar]
- Doblin, M. Cellulose Biosynthesis in Plants: From Genes to Rosettes. Plant Cell Physiol. 2002, 43, 1407–1420. [Google Scholar] [CrossRef] [PubMed]
- Somerville, C. Cellulose Synthesis in Higher Plants. Annu. Rev. Cell Dev. Biol. 2006, 22, 53–78. [Google Scholar] [CrossRef] [PubMed]
- Baskin, T.I. Anisotropic Expansion of the Plant Cell Wall. Annu. Rev. Cell Dev. Biol. 2005, 21, 203–222. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Ehrhardt, D.W.; Somerville, C.R. Mutations of cellulose synthase (CESA1) phosphorylation sites modulate anisotropic cell expansion and bidirectional mobility of cellulose synthase. Proc. Natl. Acad. Sci. USA 2010, 107, 17188–17193. [Google Scholar] [CrossRef] [PubMed]
- Fagard, M.; Desnos, T.; Desprez, T.; Goubet, F.; Refregier, G.; Mouille, G.; McCann, M.; Rayon, C.; Vernhettes, S.; Hofte, H. PROCUSTE1 Encodes a Cellulose Synthase Required for Normal Cell Elongation Specifically in Roots and Dark-Grown Hypocotyls of Arabidopsis. Plant Cell 2000, 12, 2409–2424. [Google Scholar] [CrossRef] [PubMed]
- Robert, S.; Bichet, A.; Grandjean, O.; Kierzkowski, D.; Satiat-Jeaunemaitre, B.; Pelletier, S.; Hauser, M.-T.; Höfte, H.; Vernhettes, S. An Arabidopsis Endo-1,4-β-d-Glucanase Involved in Cellulose Synthesis Undergoes Regulated Intracellular Cycling. Plant Cell 2005, 17, 3378–3389. [Google Scholar] [CrossRef] [PubMed]
- Mansoori, N.; Timmers, J.; Desprez, T.; Kamei, C.L.A.; Dees, D.C.T.; Vincken, J.P.; Visser, R.G.F.; Fte, H.H.; Vernhettes, S.; Trindade, L.M. KORRIGAN1 interacts specifically with integral components of the Cellulose synthase machinery. PLoS ONE 2014, 9, e112387. [Google Scholar] [CrossRef] [PubMed]
- Paredez, A.R.; Somerville, C.; Ehrhardt, D. Visualization of Cellulose Synthase with Microtubules. Science 2006, 312, 1491–1495. [Google Scholar] [CrossRef] [PubMed]
- Höfte, H.; Voxeur, A. Plant cell walls. Curr. Biol. 2017, 27, R865–R870. [Google Scholar] [CrossRef] [PubMed]
- Wasteneys, G.O.; Galway, M.E. Remodeling the cytoskeleton for growth and form: An overview with some new views. Annu. Rev. Plant Biol. 2003, 54, 691–722. [Google Scholar] [CrossRef] [PubMed]
- Wasteneys, G.O. Progress in understanding the role of microtubules in plant cells. Curr. Opin. Plant Biol. 2004, 7, 651–660. [Google Scholar] [CrossRef] [PubMed]
- Gutierrez, R.; Lindeboom, J.J.; Paredez, A.R.; Emons, A.M.C.; Ehrhardt, D.W. Arabidopsis cortical microtubules position cellulose synthase delivery to the plasma membrane and interact with cellulose synthase trafficking compartments. Nat. Cell Biol. 2009, 11, 797–806. [Google Scholar] [CrossRef] [PubMed]
- Crowell, E.F.; Bischoff, V.; Desprez, T.; Rolland, A.; Stierhof, Y.-D.; Schumacher, K.; Gonneau, M.; Hofte, H.; Vernhettes, S. Pausing of Golgi Bodies on Microtubules Regulates Secretion of Cellulose Synthase Complexes in Arabidopsis. Plant Cell 2009, 21, 1141–1154. [Google Scholar] [CrossRef] [PubMed]
- Eckardt, N.A. Role of Xyloglucan in Primary Cell Walls. Plant Cell 2008, 20, 1421–1422. [Google Scholar] [CrossRef] [PubMed]
- Takeda, T.; Furuta, Y.; Awano, T.; Mizuno, K.; Mitsuishi, Y.; Hayashi, T. Suppression and acceleration of cell elongation by integration of xyloglucans in pea stem segments. Proc. Natl. Acad. Sci. USA 2002, 99, 9055–9060. [Google Scholar] [CrossRef] [PubMed]
- Cavalier, D.M.; Lerouxel, O.; Neumetzler, L.; Yamauchi, K.; Reinecke, A.; Freshour, G.; Zabotina, O.A.; Hahn, M.G.; Burgert, I.; Pauly, M.; et al. Disrupting Two Arabidopsis thaliana Xylosyltransferase Genes Results in Plants Deficient in Xyloglucan, a Major Primary Cell Wall Component. Plant Cell 2008, 20, 1519–1537. [Google Scholar] [CrossRef] [PubMed]
- Hayashi, T. Xyloglucans in the primary cell wall. Annu. Rev. Plant Physiol. Plant Mol. Biol. 1989, 20, 1519–1537. [Google Scholar] [CrossRef]
- Scheller, H.V.; Ulvskov, P. Hemicelluloses. Annu. Rev. Plant Biol. 2010, 61, 263–289. [Google Scholar] [CrossRef] [PubMed]
- Xiao, C.; Zhang, T.; Zheng, Y.; Cosgrove, D.J.; Anderson, C.T. Xyloglucan Deficiency Disrupts Microtubule Stability and Cellulose Biosynthesis in Arabidopsis, Altering Cell Growth and Morphogenesis. Plant Physiol. 2016, 170, 234–249. [Google Scholar] [CrossRef] [PubMed]
- Moller, I.; Sørensen, I.; Bernal, A.J.; Blaukopf, C.; Lee, K.; Øbro, J.; Pettolino, F.; Roberts, A.; Mikkelsen, J.D.; Knox, J.P.; et al. High-throughput mapping of cell-wall polymers within and between plants using novel microarrays. Plant J. 2007, 50, 1118–1128. [Google Scholar] [CrossRef] [PubMed]
- Willats, W.G.T.; Mccartney, L.; Mackie, W.; Knox, J.P. Pectin: Cell biology and prospects for functional analysis. Plant Mol. Biol. 2001, 47, 9–27. [Google Scholar] [CrossRef] [PubMed]
- Iwai, H.; Masaoka, N.; Ishii, T.; Satoh, S. A pectin glucuronyltransferase gene is essential for intercellular attachment in the plant meristem. Proc. Natl. Acad. Sci. USA 2002, 99, 16319–16324. [Google Scholar] [CrossRef] [PubMed]
- Ridley, B.L.; O’Neill, M.A.; Mohnen, D. Pectins: Structure, biosynthesis, and oligogalacturonide-related signaling. Phytochemistry 2001, 57, 929–967. [Google Scholar] [CrossRef]
- Verger, S.; Chabout, S.; Gineau, E.; Mouille, G. Cell adhesion in plants is under the control of putative O-fucosyltransferases. Development 2016, 143, 2536–2540. [Google Scholar] [CrossRef] [PubMed]
- Vincken, J.-P. If Homogalacturonan Were a Side Chain of Rhamnogalacturonan I. Implications for Cell Wall Architecture. Plant Physiol. 2003, 132, 1781–1789. [Google Scholar] [CrossRef] [PubMed]
- Vincken, J.P.; Schols, H.A.; Oomen, R.J.F.J.; Beldman, G.; Visser, R.G.F.; Voragen, A.G.J. Pectin : The hairy thing : Evidence that homogalacturonan is a side chain of rhamnogalacturonan I. In Advances in Pectin and Pectinase Research; Voragen, A.G.J., Voragen, F., Schols, H., Visser, R., Eds.; Kluwer Academic Publishers: Dordrecht, The Netherlands, 2003; pp. 47–61. [Google Scholar]
- Caffall, K.H.; Mohnen, D. The structure, function, and biosynthesis of plant cell wall pectic polysaccharides. Carbohydr. Res. 2009, 344, 1879–1900. [Google Scholar] [CrossRef] [PubMed]
- Round, A.N.; Rigby, N.M.; MacDougall, A.J.; Morris, V.J. A new view of pectin structure revealed by acid hydrolysis and atomic force microscopy. Carbohydr. Res. 2010, 345, 487–497. [Google Scholar] [CrossRef] [PubMed]
- Cornuault, V.; Posé, S.; Knox, J.P. Disentangling pectic homogalacturonan and rhamnogalacturonan-I polysaccharides: Evidence for sub-populations in fruit parenchyma systems. Food Chem. 2018, 246, 275–285. [Google Scholar] [CrossRef] [PubMed]
- Matsunaga, T.; Ishii, T.; Matsumoto, S.; Higuchi, M.; Darvill, A.; Albersheim, P.; O’Neill, M.A. Occurrence of the primary cell wall polysaccharide rhamnogalacturonan II in pteridophytes, lycophytes, and bryophytes. Implications for the evolution of vascular plants. Plant Physiol. 2004, 134, 339–351. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.J.; Brandizzi, F. The plant secretory pathway for the trafficking of cell wall polysaccharides and glycoproteins. Glycobiology 2016, 26, 940–949. [Google Scholar] [CrossRef] [PubMed]
- Zhu, C.; Ganguly, A.; Baskin, T.I.; McClosky, D.D.; Anderson, C.T.; Foster, C.; Meunier, K.A.; Okamoto, R.; Berg, H.; Dixit, R. The Fragile Fiber1 Kinesin Contributes to Cortical Microtubule-Mediated Trafficking of Cell Wall Components. Plant Physiol. 2015, 167, 780–792. [Google Scholar] [CrossRef] [PubMed]
- Rose, J.K.C.; Lee, S.J. Straying off the Highway: Trafficking of Secreted Plant Proteins and Complexity in the Plant Cell Wall Proteome. Plant Physiol. 2010, 153, 433–436. [Google Scholar] [CrossRef] [PubMed]
- Toyooka, K.; Goto, Y.; Asatsuma, S.; Koizumi, M.; Mitsui, T.; Matsuoka, K. A Mobile Secretory Vesicle Cluster Involved in Mass Transport from the Golgi to the Plant Cell Exterior. Plant Cell 2009, 21, 1212–1229. [Google Scholar] [CrossRef] [PubMed]
- Proseus, T.E.; Boyer, J.S. Turgor pressure moves polysaccharides into growing cell walls of Chara corallina. Ann. Bot. 2005, 95, 967–979. [Google Scholar] [CrossRef] [PubMed]
- Scheible, W.R.; Pauly, M. Glycosyltransferases and cell wall biosynthesis: Novel players and insights. Curr. Opin. Plant Biol. 2004, 7, 285–295. [Google Scholar] [CrossRef] [PubMed]
- Cassab, G.I. Plant cell wall proteins. Annu. Rev. Plant Physiol. Plant Mol. Biol. 1998, 49, 281–309. [Google Scholar] [CrossRef] [PubMed]
- Carpita, N.C. Structure and Biogenesis of the Cell Walls of Grasses. Annu. Rev. Plant Physiol. Plant Mol. Biol. 1996, 47, 445–476. [Google Scholar] [CrossRef] [PubMed]
- Re, A.; Plant, P.; Bioi, M.; Cassab, G.I.; Varner, J.E. Cell wall proteins. Annu. Rev. Plant Physiol. Plant Mol. Biol. 1988, 39, 321–353. [Google Scholar]
- Cosgrove, D.J. Loosening of plant cell walls by expansins. Nature 2000, 407, 321–326. [Google Scholar] [CrossRef] [PubMed]
- Kieliszewski, M.J.; Lamport, D.T.A. Extensin: Repetitive motifs, functional sites, post-translational codes, and phylogeny. Plant J. 1994, 5, 157–172. [Google Scholar] [CrossRef] [PubMed]
- Lamport, D.T.A.; Kieliszewski, M.J.; Chen, Y.; Cannon, M.C. Role of the Extensin Superfamily in Primary Cell Wall Architecture. Plant Physiol. 2011, 156, 11–19. [Google Scholar] [CrossRef] [PubMed]
- Lamport, D.T. Oxygen Fixation into Hydroxyproline of Oxygen Fixation into Hydroxyproline Plant Cell Wall Protein. J. Biol. Chem. 1963, 238, 1438–1440. [Google Scholar] [PubMed]
- Showalter, A.M. Structure and function of plant cell wall proteins. Plant Cell 1993, 5, 9–23. [Google Scholar] [CrossRef] [PubMed]
- Cannon, M.C.; Terneus, K.; Hall, Q.; Tan, L.; Wang, Y.; Wegenhart, B.L.; Chen, L.; Lamport, D.T.A.; Chen, Y.; Kieliszewski, M.J. Self-assembly of the plant cell wall requires an extensin scaffold. Proc. Natl. Acad. Sci. USA 2008, 105, 2226–2231. [Google Scholar] [CrossRef] [PubMed]
- Kreuger, M.; Van Hoist, G.J. Arabinogalactan proteins and plant differentiation. Plant Mol. Biol. 1996, 30, 1077–1086. [Google Scholar] [CrossRef] [PubMed]
- Kreuger, M.; Van Hoist, G. Arabinogalactan proteins are essential in somatic embryogenesis of Daucus carota L. Planta 1993, 189, 243–248. [Google Scholar] [CrossRef]
- Veytsman, B.A.; Cosgrove, D.J. A model of cell wall expansion based on thermodynamics of polymer networks. Biophys. J. 1998, 75, 2240–2250. [Google Scholar] [CrossRef]
- Zhang, T.; Zheng, Y.; Cosgrove, D.J. Spatial organization of cellulose microfibrils and matrix polysaccharides in primary plant cell walls as imaged by multichannel atomic force microscopy. Plant J. 2016, 85, 179–192. [Google Scholar] [CrossRef] [PubMed]
- Cosgrove, D.J. Re-constructing our models of cellulose and primary cell wall assembly. Curr. Opin. Plant Biol. 2014, 22, 122–131. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Hong, M. Solid-state NMR investigations of cellulose structure and interactions with matrix polysaccharides in plant primary cell walls. J. Exp. Bot. 2016, 67, 503–514. [Google Scholar] [CrossRef] [PubMed]
- Keegstra, K.; Kenneth, W.; Bauer, W.D.; Albersheim, P. The Structure of Plant Cell Walls. Plant Physiol. 1973, 188–196. [Google Scholar] [CrossRef]
- Whitney, S.E.C.; Wilson, E.; Webster, J.; Bacic, A.; Grant Reid, J.S.; Gidley, M.J. Effects of structural variation in xyloglucan polymers on interactions with bacterial cellulose. Am. J. Bot. 2006, 93, 1402–1414. [Google Scholar] [CrossRef] [PubMed]
- Talbott, L.D.; Ray, P.M. Molecular size and separability features of pea cell wall polysaccharides: Implications for models of primary wall structure. Plant Physiol. 1992, 98, 357–368. [Google Scholar] [CrossRef] [PubMed]
- Park, Y.B.; Cosgrove, D.J. Xyloglucan and its interactions with other components of the growing cell wall. Plant Cell Physiol. 2015, 56, 180–194. [Google Scholar] [CrossRef] [PubMed]
- Hrmova, M.; Farkas, V.; Lahnstein, J.; Fincher, G.B. A barley xyloglucan xyloglucosyl transferase covalently links xyloglucan, cellulosic substrates, and (1,3;1,4)-β-d-glucans. J. Biol. Chem. 2007, 282, 12951–12962. [Google Scholar] [CrossRef] [PubMed]
- Hanus, J.; Mazeau, K. The Xyloglucan-Cellulose Assembly at the Atomic Scale. Biopolymers 2006, 82, 59–73. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Z.; Crespi, V.H.; Kubicki, J.D.; Cosgrove, D.J.; Zhong, L. Molecular dynamics simulation study of xyloglucan adsorption on cellulose surfaces: Effects of surface hydrophobicity and side-chain variation. Cellulose 2014, 21, 1025–1039. [Google Scholar] [CrossRef]
- Vissenberg, K.; Fry, S.C.; Pauly, M.; Höfte, H.; Verbelen, J.P. XTH acts at the microfibril-matrix interface during cell elongation. J. Exp. Bot. 2005, 56, 673–683. [Google Scholar] [CrossRef] [PubMed]
- Fry, S.C.; Smith, R.C.; Renwick, K.F.; Martin, D.J.; Hodge, S.K.; Matthews, K.J. Xyloglucan endotransglycosylase, a new wall-loosening enzyme activity from plants. Biochem. J. 1992, 282, 821–828. [Google Scholar] [CrossRef] [PubMed]
- Steele, N.M.; Sulová, Z.; Campbell, P.; Braam, J.; Farkas, V.; Fry, S.C. Ten isoenzymes of xyloglucan endotransglycosylase from plant cell walls select and cleave the donor substrate stochastically. Biochem. J. 2001, 355, 671–679. [Google Scholar] [CrossRef] [PubMed]
- Rose, J.K.C.; Braam, J.; Fry, S.C.; Nishitani, K. The XTH family of enzymes involved in xyloglucan endotransglucosylation and endohydrolysis: Current perspectives and a new unifying nomenclature. Plant Cell Physiol. 2002, 43, 1421–1435. [Google Scholar] [CrossRef] [PubMed]
- Nishitani, K.; Tominaga, R. Endo-xyloglucan transferase, a novel class of glycosyltransferase that catalyzes transfer of a segment of xyloglucan molecule to another xyloglucan molecule. J. Biol. Chem. 1992, 267, 21058–21064. [Google Scholar] [PubMed]
- Thompson, J.E.; Fry, S.C. Restructuring of wall-bound xyloglucan by transglycosylation in living plant cells. Plant J. 2001, 26, 23–34. [Google Scholar] [CrossRef] [PubMed]
- Strohmeier, M.; Hrmova, M.; Fischer, M.; Harvey, A.J.; Fincher, G.B.; Pleiss, J. Molecular modeling of family GH16 glycoside hydrolases: Potential roles for xyloglucan transglucosylases/hydrolases in cell wall modification in the poaceae. Protein Sci. 2004, 13, 3200–3213. [Google Scholar] [CrossRef] [PubMed]
- Thompson, J.E.; Fry, S.C. Trimming and solubilization of xyloglucan after deposition in the walls of cultured rose cells. J. Exp. Bot. 1997, 48, 297–305. [Google Scholar] [CrossRef]
- Baluška, F.; Hlavacka, A.; Šamaj, J.; Palme, K.; Robinson, D.G.; Matoh, T.; Mccurdy, D.W.; Menzel, D.; Volkmann, D. F-Actin-Dependent Endocytosis of Cell Wall Pectins in Meristematic Root Cells. Insights from Brefeldin A-Induced Compartments. Plant Physiol. 2002, 130, 422–431. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Zabotina, O.; Hong, M. Pectin-cellulose interactions in the Arabidopsis primary cell wall from two-dimensional magic-angle-spinning solid-state nuclear magnetic resonance. Biochemistry 2012, 51, 9846–9856. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Park, Y.B.; Cosgrove, D.J.; Hong, M. Cellulose-Pectin Spatial Contacts Are Inherent to Never-Dried Arabidopsis Primary Cell Walls: Evidence from Solid-State Nuclear Magnetic Resonance. Plant Physiol. 2015, 168, 871–884. [Google Scholar] [CrossRef] [PubMed]
- Yoneda, A.; Ito, T.; Higaki, T.; Kutsuna, N.; Saito, T.; Ishimizu, T.; Osada, H.; Hasezawa, S.; Matsui, M.; Demura, T. Cobtorin target analysis reveals that pectin functions in the deposition of cellulose microfibrils in parallel with cortical microtubules. Plant J. 2010, 64, 657–667. [Google Scholar] [CrossRef] [PubMed]
- Dick-Pérez, M.; Zhang, Y.; Hayes, J.; Salazar, A.; Zabotina, O.A.; Hong, M. Structure and interactions of plant cell-wall polysaccharides by two- and three-dimensional magic-angle-spinning solid-state NMR. Biochemistry 2011, 50, 989–1000. [Google Scholar] [CrossRef] [PubMed]
- Chanliaud, E.; Gidley, M.J. In vitro synthesis and properties of pectin/Acetobacter xylinus cellulose composites. Plant J. 1999, 20, 25–35. [Google Scholar] [CrossRef] [PubMed]
- Virk, S.S.; Cleland, R.E. The role of wall calcium in the extension of cell walls of soybean hypocotyls. Planta 1990, 182, 559–564. [Google Scholar] [CrossRef] [PubMed]
- Hocq, L.; Pelloux, J.; Lefebvre, V. Connecting Homogalacturonan-Type Pectin Remodeling to Acid Growth. Trends Plant Sci. 2017, 22, 20–29. [Google Scholar] [CrossRef] [PubMed]
- Braybrook, S.A.; Peaucelle, A. Mechano-Chemical Aspects of Organ Formation in Arabidopsis thaliana: The Relationship between Auxin and Pectin. PLoS ONE 2013, 8, e57813. [Google Scholar] [CrossRef] [PubMed]
- Zykwinska, A.; Thibault, J.F.; Ralet, M.C. Organization of pectic arabinan and galactan side chains in association with cellulose microfibrils in primary cell walls and related models envisaged. J. Exp. Bot. 2007, 58, 1795–1802. [Google Scholar] [CrossRef] [PubMed]
- Zykwinska, A.W.; Ralet, M.J.; Garnier, C.D.; Thibault, J.-F.J. Evidence for in vitro binding of pectin side chains to cellulose. Plant Physiol. 2005, 139, 397–407. [Google Scholar] [CrossRef] [PubMed]
- Proseus, T.E.; Boyer, J.S. Periplasm turgor pressure controls wall deposition and assembly in growing Chara corallina cells. Ann. Bot. 2006, 98, 93–105. [Google Scholar] [CrossRef] [PubMed]
- Schopfer, P. Biomechanics of plant growth. Am. J. Bot. 2006, 93, 1415–1425. [Google Scholar] [CrossRef] [PubMed]
- Kutschera, U.; Bergfeld, R.; Schopfer, P. Cooperation of epidermis and inner tissues in auxin-mediated growth of maize coleoptiles. Planta 1987, 170, 168–180. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, R. Stress relaxation property of the cell wall and auxin-induced cell elongation. J. Plant Res. 1996, 109, 75–84. [Google Scholar] [CrossRef]
- Perrot-Rechenmann, C. Cellular responses to auxin: Division versus expansion. Cold Spring Harb. Perspect. Biol. 2010, 2, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Masuda, Y. Auxin-induced cell elongation and cell wall changes. Bot. Mag. 1990, 103, 345–370. [Google Scholar] [CrossRef]
- Velasquez, S.M.; Barbez, E.; Kleine-Vehn, J.; Estevez, J. Auxin and cellular elongation. Plant Physiol. 2016, 170. [Google Scholar] [CrossRef] [PubMed]
- Del Pozo, J.C. The Balance between Cell Division and Endoreplication Depends on E2FC-DPB, Transcription Factors Regulated by the Ubiquitin-SCFSKP2A Pathway in Arabidopsis. Plant Cell 2006, 18, 2224–2235. [Google Scholar] [CrossRef] [PubMed]
- Nishitani, K.; Masuda, Y. Auxin-induced changes in the cell wall structure: Changes in the sugar compositions, intrinsic viscosity and molecular weight distributions of matrix polysaccharides. Physiol. Plant. 1981, 52, 482–494. [Google Scholar] [CrossRef]
- Gendreau, E.; Traas, J.; Desnos, T.; Grandjean, O.; Caboche, M.; Höfte, H. Cellular basis of hypocotyl growth in Arabidopsis thaliana. Plant Physiol. 1997, 114, 295–305. [Google Scholar] [CrossRef] [PubMed]
- Ren, H.; Gray, W.M. SAUR Proteins as Effectors of Hormonal and Environmental Signals in Plant Growth. Mol. Plant 2015, 8, 1153–1164. [Google Scholar] [CrossRef] [PubMed]
- Blancaflor, E.B.; Hasenstein, K.H. Time course and auxin sensitivity of cortical microtubule reorientation in maize roots. Protoplasma 1995, 185, 72–82. [Google Scholar] [CrossRef] [PubMed]
- Takesue, K.; Shibaoka, H. Auxin-induced longitudinal-to-transverse reorientation of cortical microtubules in nonelongating epidermal cells of azuki bean epicotyls. Protoplasma 1999, 206, 27–30. [Google Scholar] [CrossRef]
- Fu, Y.; Li, H.; Yang, Z. The ROP2 GTPase Controls the Formation of Cortical Fine F-Actin and the Early Phase of Directional Cell Expansion during Arabidopsis Organogenesis. Plant Cell 2002, 14, 777–794. [Google Scholar] [CrossRef] [PubMed]
- Gu, Y.; Wang, Z.; Yang, Z. ROP/RAC GTPase: An old new master regulator for plant signaling. Curr. Opin. Plant Biol. 2004, 7, 527–536. [Google Scholar] [CrossRef] [PubMed]
- Chapman, E.J.; Greenham, K.; Castillejo, C.; Sartor, R.; Bialy, A.; Sun, T.; Estelle, M. Hypocotyl transcriptome reveals auxin regulation of growth-promoting genes through GA-dependent and -independent pathways. PLoS ONE 2012, 7. [Google Scholar] [CrossRef] [PubMed]
- Dharmasiri, N.; Dharmasiri, S.; Weijers, D.; Lechner, E.; Yamada, M.; Hobbie, L.; Ehrismann, J.S.; Jürgens, G.; Estelle, M. Plant development is regulated by a family of auxin receptor F box proteins. Dev. Cell 2005, 9, 109–119. [Google Scholar] [CrossRef] [PubMed]
- Berleth, T.; Krogan, N.T.; Scarpella, E. Auxin signals—Turning genes on and turning cells around. Curr. Opin. Plant Biol. 2004, 7, 553–563. [Google Scholar] [CrossRef] [PubMed]
- Reed, J.W. Roles and activities of Aux/IAA proteins in Arabidopsis. Trends Plant Sci. 2001, 6, 420–425. [Google Scholar] [CrossRef]
- Ramos, J.A.; Zenser, N.; Leyser, O.; Callis, J. Rapid degradation of auxin/indoleacetic acid proteins requires conserved amino acids of domain II and is proteasome dependent. Plant Cell 2001, 13, 2349–2360. [Google Scholar] [CrossRef] [PubMed]
- Overvoorde, P.J.P.; Okushima, Y.; Alonso, J.J.M.; Chan, A.; Chang, C.; Ecker, J.R.; Hughes, B.; Liu, A.; Onodera, C.; Quach, H.; et al. Functional genomic analysis of the AUXIN/INDOLE-3-ACETIC ACID gene family members in Arabidopsis thaliana. Plant Cell 2005, 17, 3282–3300. [Google Scholar] [CrossRef] [PubMed]
- Sato, A.; Yamamoto, K.T. Overexpression of the non-canonical Aux/IAA genes causes auxin-related aberrant phenotypes in Arabidopsis. Physiol. Plant. 2008, 133, 397–405. [Google Scholar] [CrossRef] [PubMed]
- Liscum, E.; Reed, J.W. Genetics of Aux/IAA and ARF action in plant growth and development. Plant Mol. Biol. 2002, 49, 387–400. [Google Scholar] [CrossRef] [PubMed]
- Mockaitis, K.; Estelle, M. Auxin Receptors and Plant Development: A New Signaling Paradigm. Annu. Rev. Cell Dev. Biol. 2008, 24, 55–80. [Google Scholar] [CrossRef] [PubMed]
- Ulmasov, T.; Hagen, G.; Guilfoyle, T.J. Activation and repression of transcription by auxin-response factors. Proc. Natl. Acad. Sci. USA 1999, 96, 5844–5849. [Google Scholar] [CrossRef] [PubMed]
- Tiwari, S.B.; Hagen, G.; Guilfoyle, T. The roles of auxin response factor domains in auxin-responsive transcription. Plant Cell 2003, 15, 533–543. [Google Scholar] [CrossRef] [PubMed]
- Hagen, G.; Guilfoyle, T. Auxin-responsive gene expression: Genes, promoters and regulatory factors. Plant Mol. Biol. 2002, 49, 373–385. [Google Scholar] [CrossRef] [PubMed]
- Esmon, C.A.; Tinsley, A.G.; Ljung, K.; Sandberg, G.; Hearne, L.B.; Liscum, E. A gradient of auxin and auxin-dependent transcription precedes tropic growth responses. Proc. Natl. Acad. Sci. USA 2006, 103, 236–241. [Google Scholar] [CrossRef] [PubMed]
- Pelletier, S.; Van Orden, J.; Wolf, S.; Vissenberg, K.; Delacourt, J.; Ndong, Y.A.; Pelloux, J.; Bischoff, V.; Urbain, A.; Mouille, G.; et al. A role for pectin de-methylesterification in a developmentally regulated growth acceleration in dark-grown Arabidopsis hypocotyls. New Phytol. 2010, 188, 726–739. [Google Scholar] [CrossRef] [PubMed]
- Nemhauser, J.L.; Hong, F.; Chory, J. Different Plant Hormones Regulate Similar Processes through Largely Nonoverlapping Transcriptional Responses. Cell 2006, 126, 467–475. [Google Scholar] [CrossRef] [PubMed]
- Cosgrove, D.J. Enzymes and Other Agents That Enhance Cell Wall Extensibility. Annu. Rev. Plant Physiol. Plant Mol. Biol. 1999, 50, 391–417. [Google Scholar] [CrossRef] [PubMed]
- Eklof, J.M.; Brumer, H. The XTH Gene Family: An Update on Enzyme Structure, Function, and Phylogeny in Xyloglucan Remodeling. Plant Physiol. 2010, 153, 456–466. [Google Scholar] [CrossRef] [PubMed]
- Motose, H.; Sugiyama, M.; Fukuda, H. A proteoglycan mediates inductive interaction during plant vascular development. Nature 2004, 429, 873–878. [Google Scholar] [CrossRef] [PubMed]
- Seifert, G.J.; Roberts, K. The Biology of Arabinogalactan Proteins. Annu. Rev. Plant Biol. 2007, 58, 137–161. [Google Scholar] [CrossRef] [PubMed]
- Peaucelle, A.; Braybrook, S.A.; Le Guillou, L.; Bron, E.; Kuhlemeier, C.; Höfte, H. Pectin-induced changes in cell wall mechanics underlie organ initiation in Arabidopsis. Curr. Biol. 2011, 21, 1720–1726. [Google Scholar] [CrossRef] [PubMed]
- Savaldi-Goldstein, S.; Baiga, T.J.; Pojer, F.; Dabi, T.; Butterfield, C.; Parry, G.; Santner, A.; Dharmasiri, N.; Tao, Y.; Estelle, M.; et al. New auxin analogs with growth-promoting effects in intact plants reveal a chemical strategy to improve hormone delivery. Proc. Natl. Acad. Sci. USA 2008, 105, 15190–15195. [Google Scholar] [CrossRef] [PubMed]
- Hong, F.; Breitling, R.; McEntee, C.W.; Wittner, B.S.; Nemhauser, J.L.; Chory, J. A bioconductor package for detecting differentially expressed genes in meta-analysis. Bioinformatics 2006, 22, 2825–2827. [Google Scholar] [CrossRef] [PubMed]
- Breitling, R.; Armengaud, P.; Amtmann, A.; Herzyk, P. Rank products: A simple, yet powerful, new method to detect differentially regulated genes in replicated microarray experiments. FEBS Lett. 2004, 573, 83–92. [Google Scholar] [CrossRef] [PubMed]
- Hager, A.; Menzel, H.; Krauss, A. Versuche und Hypothese zur Primärwirkung des Auxins beim Streckungswachstum. Planta 1971, 100, 47–75. [Google Scholar] [CrossRef] [PubMed]
- Rayle, D.L.; Cleland, R. Enhancement of wall loosening and elongation by Acid solutions. Plant Physiol. 1970, 46, 250–253. [Google Scholar] [CrossRef] [PubMed]
- Lüthen, H.; Bigdon, M.; Böttger, M. Reexamination of the Acid growth theory of auxin action. Plant Physiol. 1990, 93, 931–939. [Google Scholar] [CrossRef] [PubMed]
- Cleland, R.E.; Buckley, G.; Nowbar, S.; Lew, N.M.; Stinemetz, C.; Evans, M.L.; Rayle, D.L. The pH profile for acid-induced elongation of coleoptile and epicotyl sections is consistent with the acid-growth theory. Planta 1991, 186, 70–74. [Google Scholar] [CrossRef] [PubMed]
- Karcz, W.; Stolarek, J.; Pietruszka, M.; Malkowski, E. The dose-response curves for IAA induced elongation growth and acidification of the incubation medium of Zea mays coleoptile segments. Physiol. Plant. 1990, 80, 257–261. [Google Scholar] [CrossRef]
- Arsuffi, G.; Braybrook, S.A. Acid growth: An ongoing trip. J. Exp. Bot. 2017, 2, 137–146. [Google Scholar] [CrossRef] [PubMed]
- Fendrych, M.; Leung, J.; Friml, J. Tir1/AFB-Aux/IAA auxin perception mediates rapid cell wall acidification and growth of Arabidopsis hypocotyls. Elife 2016, 5, 1–19. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, K.; Hayashi, K.; Kinoshita, T. Auxin Activates the Plasma Membrane H+-ATPase by Phosphorylation during Hypocotyl Elongation in Arabidopsis. Plant Physiol. 2012, 159, 632–641. [Google Scholar] [CrossRef] [PubMed]
- Spartz, A.K.; Ren, H.; Park, M.Y.; Grandt, K.N.; Lee, S.H.; Murphy, A.S.; Sussman, M.R.; Overvoorde, P.J.; Gray, W.M. SAUR Inhibition of PP2C-D Phosphatases Activates Plasma Membrane H+-ATPases to Promote Cell Expansion in Arabidopsis. Plant Cell 2014, 26, 2129–2142. [Google Scholar] [CrossRef] [PubMed]
- Hager, A.; Debus, G.; Edel, H.G.; Stransky, H.; Serrano, R. Auxin induces exocytosis and the rapid synthesis of a high-turnover pool of plasma-membrane H+-ATPase. Planta 1991, 185, 527–537. [Google Scholar] [CrossRef] [PubMed]
- Thiel, G.; Weise, R. Auxin augments conductance of K+ inward rectifier in maize coleoptile protoplasts. Planta 1999, 208, 38–45. [Google Scholar] [CrossRef]
- Philippar, K.; Ivashikina, N.; Ache, P.; Christian, M.; Lüthen, H.; Palme, K.; Hedrich, R. Auxin activates KAT1 and KAT2, two K+-channel genes expressed in seedlings of Arabidopsis thaliana. Plant J. 2004, 37, 815–827. [Google Scholar] [CrossRef] [PubMed]
- Frías, I.; Caldeira, M.T.; Pérez-Castiñeira, J.R.; Navarro-Aviñó, J.P.; Culiañez-Maciá, F.; Kuppinger, O.; Stransky, H.; Pagés, M.; Hager, A.; Serrano, R. A major isoform of the maize plasma membrane H+-ATPase: Characterization and induction by auxin in coleoptiles. Plant Cell 1996, 8, 1533–1544. [Google Scholar] [PubMed]
- Kutschera, U.; Niklas, K.J. The epidermal-growth-control theory of stem elongation: An old and a new perspective. J. Plant Physiol. 2007, 164, 1395–1409. [Google Scholar] [CrossRef] [PubMed]
- Marcotrigiano, M. A role for leaf epidermis in the control of leaf size and the rate and extent of mesophyll cell division. Am. J. Bot. 2010, 97, 224–233. [Google Scholar] [CrossRef] [PubMed]
- Philippar, K.; Fuchs, I.; Luthen, H.; Hoth, S.; Bauer, C.S.; Haga, K.; Thiel, G.; Ljung, K.; Sandberg, G.; Bottger, M.; et al. Auxin-induced K+ channel expression represents an essential step in coleoptile growth and gravitropism. Proc. Natl. Acad. Sci. USA 1999, 96, 12186–12191. [Google Scholar] [CrossRef] [PubMed]
- Savaldi-Goldstein, S.; Chory, J. Growth coordination and the shoot epidermis. Curr. Opin. Plant Biol. 2008, 11, 42–48. [Google Scholar] [CrossRef] [PubMed]
- Van Volkenburgh, E.; Schmidt, M.G.; Cleland, R.E. Loss of capacity for acid-induced wall loosening as the principal cause of the cessation of cell enlargement in light-grown bean leaves. Planta 1985, 163, 500–505. [Google Scholar] [CrossRef] [PubMed]
- Winch, S.; Pritchard, J. Acid-induced wall loosening is confined to the accelerating region of the root growing zone. J. Exp. Bot. 1999, 50, 1481–1487. [Google Scholar] [CrossRef]
- McQueen-Mason, S.; Durachko, D.M.; Cosgrove, D.J. Two Endogenous Proteins That Induce Cell Wall Extension in Plants. Plant Cell 1992, 4, 1425. [Google Scholar] [CrossRef] [PubMed]
- McQueen-Mason, S.; Cosgrove, D.J. Disruption of hydrogen bonding between plant cell wall polymers by proteins that induce wall extension. Proc. Natl. Acad. Sci. USA 1994, 91, 6574–6578. [Google Scholar] [CrossRef] [PubMed]
- McQueen-Mason, S.J.; Cosgrove, D.J. Expansin Mode of Action on Cell Walls. Plant Physiol. 1995, 107, 87–100. [Google Scholar] [CrossRef] [PubMed]
- Link, B.M.; Cosgrove, D.J. Acid-growth response and α-expansins in suspension cultures of bright yellow 2 tobacco. Plant Physiol. 1998, 118, 907–916. [Google Scholar] [CrossRef] [PubMed]
- Yennawar, N.H.; Li, L.-C.; Dudzinski, D.M.; Tabuchi, A.; Cosgrove, D.J. Crystal structure and activities of EXPB1 (Zea m 1), a β-expansin and group-1 pollen allergen from maize. Proc. Natl. Acad. Sci. USA 2006, 103, 14664–14671. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Ruan, Y.-L. Regulation of cell division and expansion by sugar and auxin signaling. Front. Plant Sci. 2013, 4, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Purugganan, M.M.; Braam, J.; Fry, S.C. The Arabidopsis TCH4 xyloglucan endotransglycosylase. Substrate specificity, pH optimum, and cold tolerance. Plant Physiol. 1997, 115, 181–190. [Google Scholar] [CrossRef] [PubMed]
- Shipp, M.; Nadella, R.; Gao, H.; Farkas, V.; Sigrist, H.; Faik, A. Glyco-array technology for efficient monitoring of plant cell wall glycosyltransferase activities. Glycoconj. J. 2008, 25, 49–58. [Google Scholar] [CrossRef] [PubMed]
- Kotake, T.; Nakagawa, N.; Takeda, K.; Sakurai, N. Auxin-Induced Elongation Growth and Expressions of Cell Wall-Bound Exo- and Endo-β-Glucanases in Barley Coleoptiles. Plant Cell Physiol. 2000, 41, 1272–1278. [Google Scholar] [CrossRef] [PubMed]
- Yokoyama, R.; Nishitani, K. A comprehensive expression analysis of all members of a gene family encoding cell-wall enzymes allowed us to predict cis-regulatory regions involved in cell-wall construction in specific organs of Arabidopsis. Plant Cell Physiol. 2001, 42, 1025–1033. [Google Scholar] [CrossRef] [PubMed]
- Swarup, K.; Benková, E.; Swarup, R.; Casimiro, I.; Péret, B.; Yang, Y.; Parry, G.; Nielsen, E.; De Smet, I.; Vanneste, S.; et al. The auxin influx carrier LAX3 promotes lateral root emergence. Nat. Cell Biol. 2008, 10, 946–954. [Google Scholar] [CrossRef] [PubMed]
- Xu, W. Arabidopsis TCH4, Regulated by Hormones and the Environment, Encodes a Xyloglucan Endotransglycosylase. Plant Cell 1995, 7, 1555–1567. [Google Scholar] [CrossRef] [PubMed]
- Xu, W.; Campbell, P.; Vargheese, A.K.; Braam, J. The Arabidopsis XET-related gene family: Environmental and hormonal regulation of expression. Plant J. 1996, 9, 879–889. [Google Scholar] [CrossRef] [PubMed]
- Ohmiya, Y.; Samejima, M.; Shiroishi, M.; Amano, Y.; Kanda, T.; Sakai, F.; Hayashi, T. Evidence that endo-1,4-β-glucanases act on cellulose in suspension- cultured poplar cells. Plant J. 2000, 24, 147–158. [Google Scholar] [CrossRef] [PubMed]
- Verma, D.P.S.; Maclachlan, G.A.; Byrne, H.; Ewings, D. Regulation In Vitro Translation of Messenger Ribonucleic Acid for Cellulase from Auxin-Treated Pea Epicotyls. J. Biol. Chem. 1975, 250, 1019–1026. [Google Scholar] [PubMed]
- Fry, S.C. Cellulases, hemicelluloses and auxin-stimulated growth: A possible relationship. Physiol. Plant. 1989, 75, 532–536. [Google Scholar] [CrossRef]
- Catala, C.; Rose, J.K.; Bennett, A.B. Auxin regulation and spatial localization of an endo-1,4-β-d-glucanase and a xyloglucan endotransglycosylase in expanding tomato hypocotyls. Plant J. 1997, 12, 417–426. [Google Scholar] [CrossRef] [PubMed]
- Antosiewicz, D.M.; Purugganan, M.M.; Polisensky, D.H.; Braam, J. Cellular localization of Arabidopsis xyloglucan endotransglycosylase-related proteins during development and after wind stimulation. Plant Physiol. 1997, 115, 1319–1328. [Google Scholar] [CrossRef] [PubMed]
- Van Sandt, V.S.T.; Suslov, D.; Verbelen, J.P.; Vissenberg, K. Xyloglucan endotransglucosylase activity loosens a plant cell wall. Ann. Bot. 2007, 100, 1467–1473. [Google Scholar] [CrossRef] [PubMed]
- McDougall, G.J.; Fry, S.C. Inhibition of auxin-stimulated growth of pea stem segments by a specific nonasaccharide of xyloglucan. Planta 1988, 175, 412–416. [Google Scholar] [CrossRef] [PubMed]
- Paque, S.; Mouille, G.; Grandont, L.; Alabadi, D.; Gaertner, C.; Goyallon, A.; Muller, P.; Primard-Brisset, C.; Sormani, R.; Blazquez, M.A.; et al. AUXIN BINDING PROTEIN1 Links Cell Wall Remodeling, Auxin Signaling, and Cell Expansion in Arabidopsis. Plant Cell 2014, 26, 280–295. [Google Scholar] [CrossRef] [PubMed]
- York, W.S.; Darvill, A.G.; Albersheim, P. Inhibition of 2,4-dichlorophenoxyacetic Acid-stimulated elongation of pea stem segments by a xyloglucan oligosaccharide. Plant Physiol. 1984, 75, 295–297. [Google Scholar] [CrossRef] [PubMed]
- Hyodo, H.; Yamakawa, S.; Takeda, Y.; Tsuduki, M.; Yokota, A.; Nishitani, K.; Kohchi, T. Active gene expression of a xyloglucan endotransglucosylase/hydrolase gene, XTH9, in inflorescence apices is related to cell elongation in Arabidopsis thaliana. Plant Mol. Biol. 2003, 52, 473–482. [Google Scholar] [CrossRef] [PubMed]
- Vissenberg, K.; Fry, S.C.; Verbelen, J.P. Root hair initiation is coupled to a highly localized increase of xyloglucan endotransglycosylase action in Arabidopsis roots. Plant Physiol. 2001, 127, 1125–1135. [Google Scholar] [CrossRef] [PubMed]
- Fry, S.C. The Structure and Functions of Xyloglucan. J. Exp. Bot. 1989, 40, 1–11. [Google Scholar] [CrossRef]
- Marin-Rodriguez, M.C.; Orchard, J.; Seymour, G.B. Pectate lyases, cell wall degradation and fruit softening. J. Exp. Bot. 2002, 53, 2115–2119. [Google Scholar] [CrossRef] [PubMed]
- Wakabayashi, K.; Chun, J.P.; Huber, D.J. Extensive solubilization and depolymerization of cell wall polysaccharides during avocado (Persea americana) ripening involves concerted action of polygalacturonase and pectinmethylesterase. Physiol. Plant. 2000, 108, 345–352. [Google Scholar] [CrossRef]
- Wakabayashi, K.; Hoson, T.; Huber, D.J. Methyl de-esterification as a major factor regulating the extent of pectin depolymerization during fruit ripening: A comparison of the action of avocado (Persea americana) and tomato (Lycopersicon esculentum) polygalacturonases. J. Plant Physiol. 2003, 160, 667–673. [Google Scholar] [CrossRef] [PubMed]
- Osorio, S.; Castillejo, C.; Quesada, M.A.; Medina-Escobar, N.; Brownsey, G.J.; Suau, R.; Heredia, A.; Botella, M.A.; Valpuesta, V. Partial demethylation of oligogalacturonides by pectin methyl esterase 1 is required for eliciting defence responses in wild strawberry (Fragaria vesca). Plant J. 2008, 54, 43–55. [Google Scholar] [CrossRef] [PubMed]
- Ferrari, S. Oligogalacturonides: Plant damage-associated molecular patterns and regulators of growth and development. Front. Plant Sci. 2013, 4, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Savatin, D.V.; Ferrari, S.; Sicilia, F.; De Lorenzo, G. Oligogalacturonide-Auxin Antagonism Does Not Require Posttranscriptional Gene Silencing or Stabilization of Auxin Response Repressors in Arabidopsis. Plant Physiol. 2011, 157, 1163–1174. [Google Scholar] [CrossRef] [PubMed]
- Branca, C.; De Lorenzo, G.; Cervone, F. Competitive inhibition of the auxin-induced elongation by α-d-oligogalacturonides in pea stem segments. Physiol. Plant. 1988, 72, 499–504. [Google Scholar] [CrossRef]
- Bellincampi, D.; Cardarelli, M.; Zaghi, D.; Serino, G.; Salvi, G.; Gatz, C.; Cervone, F.; Altamura, M.M.; Costantino, P.; Lorenzoai, G.D. Oligogalacturonides Prevent Rhizogenesis in rolB-Transformed Tobacco Explants by lnhibiting Auxin-lnduced Expression of the rolB Gene. Plant Cell 1996, 8, 477–487. [Google Scholar] [CrossRef] [PubMed]
- Ferrari, S.; Galletti, R.; Pontiggia, D.; Manfredini, C.; Lionetti, V.; Bellincampi, D.; Cervone, F.; De Lorenzo, G. Transgenic expression of a fungal endo-polygalacturonase increases plant resistance to pathogens and reduces auxin sensitivity. Plant Physiol. 2008, 146, 669–681. [Google Scholar] [CrossRef] [PubMed]
- Galletti, R.; Denoux, C.; Gambetta, S.; Dewdney, J.; Ausubel, F.M.; De Lorenzo, G.; Ferrari, S. The AtrbohD-Mediated Oxidative Burst Elicited by Oligogalacturonides in Arabidopsis Is Dispensable for the Activation of Defense Responses Effective against Botrytis cinerea. Plant Physiol. 2008, 148, 1695–1706. [Google Scholar] [CrossRef] [PubMed]
- Noctor, G.; Reichheld, J.P.; Foyer, C.H. ROS-related redox regulation and signaling in plants. Semin. Cell Dev. Biol. 2017. [Google Scholar] [CrossRef] [PubMed]
- Tenhaken, R. Cell wall remodeling under abiotic stress. Front. Plant Sci. 2015, 5, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Francoz, E.; Ranocha, P.; Nguyen-Kim, H.; Jamet, E.; Burlat, V.; Dunand, C. Roles of cell wall peroxidases in plant development. Phytochemistry 2015, 112, 15–21. [Google Scholar] [CrossRef] [PubMed]
- Bailey-Serres, J.; Mittler, R. The roles of reactive oxygen species in plant cells. Plant Physiol. 2006, 141, 900191. [Google Scholar] [CrossRef] [PubMed]
- Schopfer, P.; Liszkay, A.; Bechtold, M.; Frahry, G.; Wagner, A. Evidence that hydroxyl radicals mediate auxin-induced extension growth. Planta 2002, 214, 821–828. [Google Scholar] [CrossRef] [PubMed]
- Marga, F.; Grandbois, M.; Cosgrove, D.J.; Baskin, T.I. Cell wall extension results in the coordinate separation of parallel microfibrils: Evidence from scanning electron microscopy and atomic force microscopy. Plant J. 2005, 43, 181–190. [Google Scholar] [CrossRef] [PubMed]
- Thompson, D.S. Space and time in the plant cell wall: Relationships between cell type, cell wall rheology and cell function. Ann. Bot. 2008, 101, 203–211. [Google Scholar] [CrossRef] [PubMed]
- Lockhart, J.A. An analysis of irreversible plant cell elongation. J. Theor. Biol. 1965, 8, 264–275. [Google Scholar] [CrossRef]
- Nakagawa, Y.; Katagiri, T.; Shinozaki, K.; Qi, Z.; Tatsumi, H.; Furuichi, T.; Kishigami, A.; Sokabe, M.; Kojima, I.; Sato, S.; et al. Arabidopsis plasma membrane protein crucial for Ca2+ influx and touch sensing in roots. Proc. Natl. Acad. Sci. USA 2007, 104, 3639–3644. [Google Scholar] [CrossRef] [PubMed]
- Monshausen, G.B.; Bibikova, T.N.; Weisenseel, M.H.; Gilroy, S. Ca2+ Regulates Reactive Oxygen Species Production and pH during Mechanosensing in Arabidopsis Roots. Plant Cell 2009, 21, 2341–2356. [Google Scholar] [CrossRef] [PubMed]
- Knight, M.R.; Campbell, A.K.; Smith, S.M.; Trewavas, A.J. Transgenic plant aequorin reports the effects of touch and cold-shock and elicitors on cytoplasmic calcium. Nature 1991, 352, 524–526. [Google Scholar] [CrossRef] [PubMed]
- Valentin, R.; Cerclier, C.; Geneix, N.; Aguié-Béghin, V.; Gaillard, C.; Ralet, M.C.; Cathala, B. Elaboration of extensin-pectin thin film model of primary plant cell wall. Langmuir 2010, 26, 9891–9898. [Google Scholar] [CrossRef] [PubMed]
- Wolf, S.; Mouille, G.; Pelloux, J. Homogalacturonan methyl-esterification and plant development. Mol. Plant 2009, 2, 851–860. [Google Scholar] [CrossRef] [PubMed]
- Willats, W.G.T.; Orfila, C.; Limberg, G.; Buchholt, H.C.; Van Alebeek, G.J.W.M.; Voragen, A.G.J.; Marcus, S.E.; Christensen, T.M.I.E.; Mikkelsen, J.D.; Murray, B.S.; et al. Modulation of the degree and pattern of methyl-esterification of pectic homogalacturonan in plant cell walls: Implications for pectin methyl esterase action, matrix properties, and cell adhesion. J. Biol. Chem. 2001, 276, 19404–19413. [Google Scholar] [CrossRef] [PubMed]
- Cameron, R.G.; Luzio, G.A.; Goodner, K.; Williams, M.A.K. Demethylation of a model homogalacturonan with a salt-independent pectin methylesterase from citrus: I. Effect of pH on demethylated block size, block number and enzyme mode of action. Carbohydr. Polym. 2008, 71, 287–299. [Google Scholar] [CrossRef]
- Catoire, L.; Pierron, M.; Morvan, C.; Du Penhoat, C.H.; Goldberg, R. Investigation of the action patterns of pectinmethylesterase isoforms through kinetic analyses and NMR spectroscopy: Implications in cell wall expansion. J. Biol. Chem. 1998, 273, 33150–33156. [Google Scholar] [CrossRef] [PubMed]
- Denès, J.M.; Baron, A.; Renard, C.M.G.C.; Péan, C.; Drilleau, J.F. Different action patterns for apple pectin methylesterase at pH 7.0 and 4.5. Carbohydr. Res. 2000, 327, 385–393. [Google Scholar] [CrossRef]
- Jolie, R.P.; Duvetter, T.; Van Loey, A.M.; Hendrickx, M.E. Pectin methylesterase and its proteinaceous inhibitor: A review. Carbohydr. Res. 2010, 345, 2583–2595. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.; Teng, Q.; Wicker, L. Action pattern of Valencia orange PME de-esterification of high methoxyl pectin and characterization of modified pectins. Carbohydr. Res. 2005, 340, 2620–2629. [Google Scholar] [CrossRef] [PubMed]
- Wolf, S.; Rausch, T.; Greiner, S. The N-terminal pro region mediates retention of unprocessed type-I PME in the Golgi apparatus. Plant J. 2009, 58, 361–375. [Google Scholar] [CrossRef] [PubMed]
- Western, T.L.; Burn, J.; Tan, W.L.; Skinner, D.J.; Martin-McCaffrey, L.; Moffatt, B.A.; Haughn, G.W. Isolation and Characterization of Mutants Defective in Seed Coat Mucilage Secretory Cell Development in Arabidopsis. Plant Physiol. 2001, 127, 998–1011. [Google Scholar] [CrossRef] [PubMed]
- Griffiths, J.S.; Tsai, A.Y.-L.; Xue, H.; Voiniciuc, C.; Ola, K.; Seifert, G.J.; Mansfield, S.D.; Haughn, G.W. SALT-OVERLY SENSITIVE5 Mediates Arabidopsis Seed Coat Mucilage Adherence and Organization through Pectins. Plant Physiol. 2014, 165, 991–1004. [Google Scholar] [CrossRef] [PubMed]
- Griffiths, J.S.; Crepeau, M.-J.; Ralet, M.-C.; Seifert, G.J.; North, H.M. Dissecting Seed Mucilage Adherence Mediated by FEI2 and SOS5. Front. Plant Sci. 2016, 7, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Raggi, S.; Ferrarini, A.; Delledonne, M.; Dunand, C.; Ranocha, P.; De Lorenzo, G.; Cervone, F.; Ferrari, S. The Arabidopsis thaliana Class III Peroxidase AtPRX71 Negatively Regulates Growth under Physiological Conditions and in Response to Cell Wall Damage. Plant Physiol. 2015, 169, 2513–2525. [Google Scholar] [CrossRef] [PubMed]
- Pereira, C.S.; Ribeiro, J.M.L.; Vatulescu, A.D.; Findlay, K.; MacDougall, A.J.; Jackson, P.A.P. Extensin network formation in Vitis vinifera callus cells is an essential and causal event in rapid and H2O2-induced reduction in primary cell wall hydration. BMC Plant Biol. 2011, 11, 106. [Google Scholar] [CrossRef] [PubMed]
- Swanson, S.; Gilroy, S. ROS in plant development. Physiol. Plant. 2010, 138, 384–392. [Google Scholar] [CrossRef] [PubMed]
- Fry, S.C. Oxidative scission of plant cell wall polysaccharides by ascorbate-induced hydroxyl radicals. Biochem. J. 1998, 332, 507–515. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Krens, S.F.G.; Fendrych, M.; Friml, J. Real-time Analysis of Auxin Response, Cell Wall pH and Elongation in Arabidopsis thaliana Hypocotyls. Bio-Protocol 2018, 7, 1–13. [Google Scholar] [CrossRef] [PubMed]
| Cellulose Related | |
|---|---|
| CELLULOSE SYNTHASE-LIKE | CSLC4; CSLC5 |
| EXPANSIN related | |
| EXPANSIN | EXPA4; EXPA11 |
| EXPANSIN-LIKE | EXLA3 |
| XTH related | |
| XYLOGLUCAN ENDOTRANSGLUCOSYLASE/HYDROLASE | XTH18; XTH19; XTH23; XTH33 |
| TOUCH | TCH2; TCH3; TCH4 |
| XYLOSYLTRANSFERASE | XT1 |
| ACT DOMAIN REPEAT 7 | ACR7 |
| Pectin related | |
| PECTIN METHYLESTERASE | PME1; PME34 |
| PLANT INVERTASE/PECTIN METHYLESTERASE INHIBITOR SUPERFAMILY | |
| PECTIN ACETYLESTERASE | PAE11 |
| GALACTURONOSYLTRANSFERASE-LIKE 10 | GATL10 |
| GALACTURONOSYLTRANSFERASE-LIKE | GATL3 |
| GALACTAN SYNTHASE | GALS3 |
| POLYGALACTURONASE INHIBITING PROTEIN 1 | PGIP1 |
| PEROXIDASE related | |
| PEROXIDASE SUPERFAMILY PROTEINS | |
| Secondary cell wall related | |
| OVATE FAMILY PROTEIN 1 | OFP1 |
| REDUCED IN LATERAL GROWTH 1 | RUL1 |
| Other/biosynthesis related | |
| EXORDIUM LIKE 2 | EXL2 |
| Cellulose Related | |
|---|---|
| CELLULOSE SYNTHASE-LIKE | CSLC04; CSLC12; CSLD2; CSLD3 |
| CELLULOSE SYNTHASE-INTERACTIVE PROTEIN 1 | CSI1 |
| EXPANSIN related | |
| EXPANSIN | EXPA1; XPA7; EXPA10; EXPA12; EXPA18 |
| EXPANSIN-LIKE | EXLA1; EXLA2; EXLA3 |
| XTH related | |
| XYLOGLUCAN ENDOTRANSGLUCOSYLASE/HYDROLASE | XTH8; XTH17; XTH18; XTH19 XTH23; XTH33 |
| TOUCH 3 | TCH3 |
| XYLOSYLTRANSFERASE 1 | XT1 |
| Pectin related | |
| PECTIN METHYLESTERASE | PME2; PME41 |
| PLANT INVERTASE/PECTIN METHYLESTERASE INHIBITOR SUPERFAMILY | |
| PECTIN METHYLESTERASE INIHIBITOR 7 | PMEI7 |
| PECTIN ACETYLESTERASE 11 | PAE9; PAE11 |
| PECTIN LYASE-LIKE SUPERFAMILY PROTEIN | |
| POLYGALACTURONASE INHIBITING PROTEIN 1 | PGIP1; PGIP2 |
| ARABINOXYLAN PECTIN ARABINOGALACTAN PROTEIN 1 | APAP1 |
| FRA8 HOMOLOG | F8H |
| GALACTAN SYNTHASE | GALS2; GALS3 |
| BETA-GALACTOSIDASE | BGAL10; BGAL12 |
| GALACTURONOSYLTRANSFERASE-LIKE | GATL7; GATL10 |
| MALE GAMETOPHYTE DEFECTIVE 4 | MGP4 |
| Hemicelluloserelated | |
| ENDO-BETA-MANNASE 7 | MAN7 |
| ALPHA-XYLOSIDASE 1 | XYL1 |
| GLYCOSYLTRANSFERASE 18 | GT18 |
| MURUS 3 | MUR3 |
| AGPrelated | |
| ARABINOGALACTAN PROTEIN | AGP2; AGP9 |
| PROLINE/LEUCIN-RICH PROTEIN related | |
| PROLINE-RICH PROTEIN 1 | PRP1; PRP2 |
| PROLINE-RICH PROTEIN-LIKE 1 | PRPL1 |
| LEUCINE-RICH REPEAT/EXTENSIN 2 | LRX2 |
| LEUCINE-RICH REPEAT PROTEIN | LRR1 |
| Peroxidase related | |
| PEROXIDASE SUPERFAMILY PROTEIN | |
| PEROXIDASE 7 | PER7; PRX25; PRX33 |
| Secondary cell wall related | |
| PARVUS | PARVUS |
| TRANSPARENT TESTA 8 | TT8 |
| GLABRA 2 | GL2 |
| Signal Perception | |
| FORMIN HOMOLOGY | FH1; FH5 |
| WALL ASSIOCIATED KINASE | WAK |
| THESEUS 1 | THE1 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Majda, M.; Robert, S. The Role of Auxin in Cell Wall Expansion. Int. J. Mol. Sci. 2018, 19, 951. https://doi.org/10.3390/ijms19040951
Majda M, Robert S. The Role of Auxin in Cell Wall Expansion. International Journal of Molecular Sciences. 2018; 19(4):951. https://doi.org/10.3390/ijms19040951
Chicago/Turabian StyleMajda, Mateusz, and Stéphanie Robert. 2018. "The Role of Auxin in Cell Wall Expansion" International Journal of Molecular Sciences 19, no. 4: 951. https://doi.org/10.3390/ijms19040951
APA StyleMajda, M., & Robert, S. (2018). The Role of Auxin in Cell Wall Expansion. International Journal of Molecular Sciences, 19(4), 951. https://doi.org/10.3390/ijms19040951