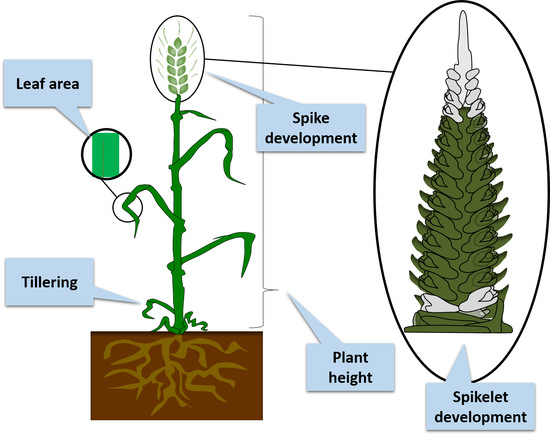
Key Hormonal Components Regulate Agronomically Important Traits in Barley
Abstract
1. Introduction
2. Hormonal Regulation of Agronomically Important Traits in Barley
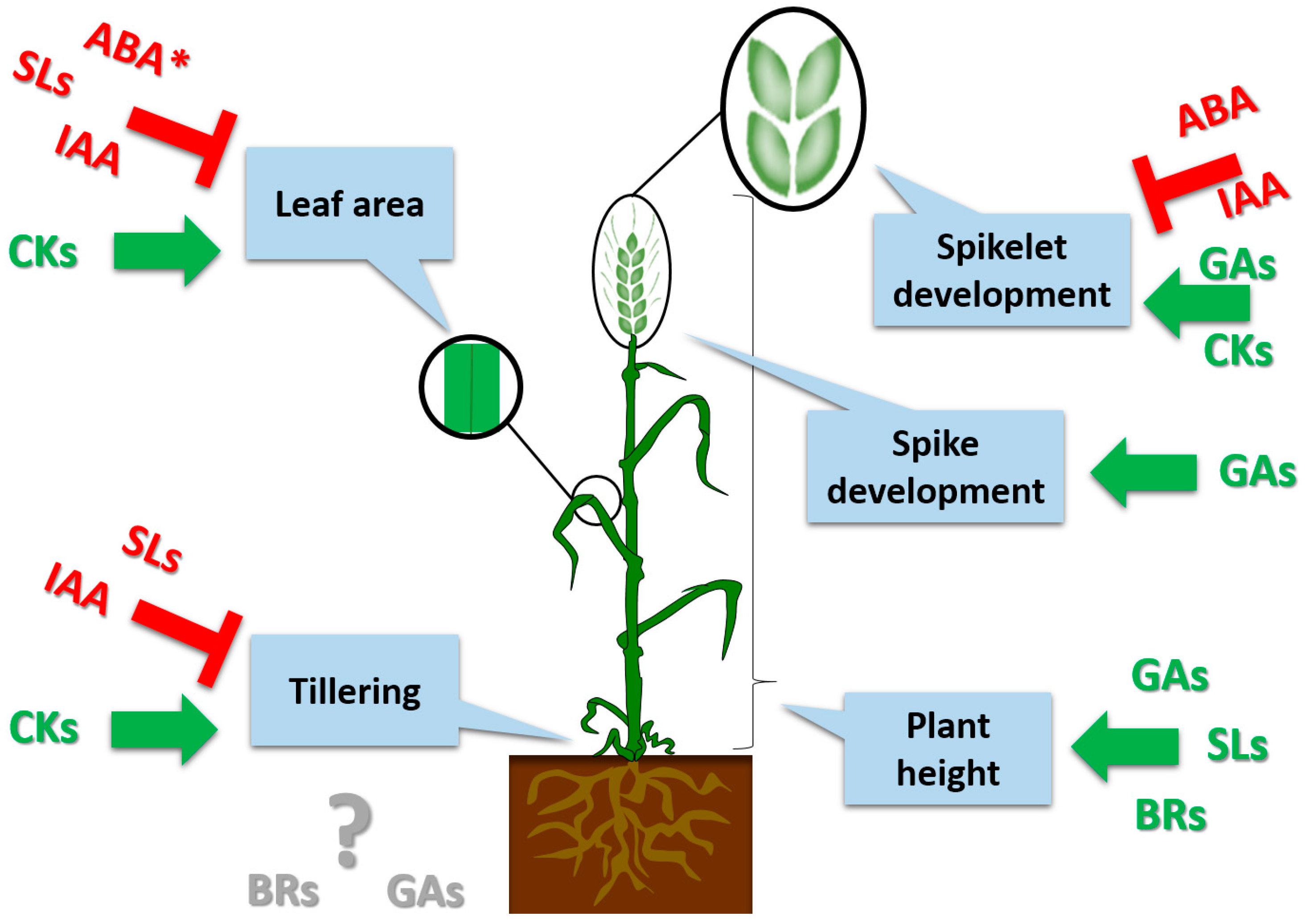
2.1. Roles of Phytohormones in Plant Height
2.2. The Role of Phytohormones in Tillering
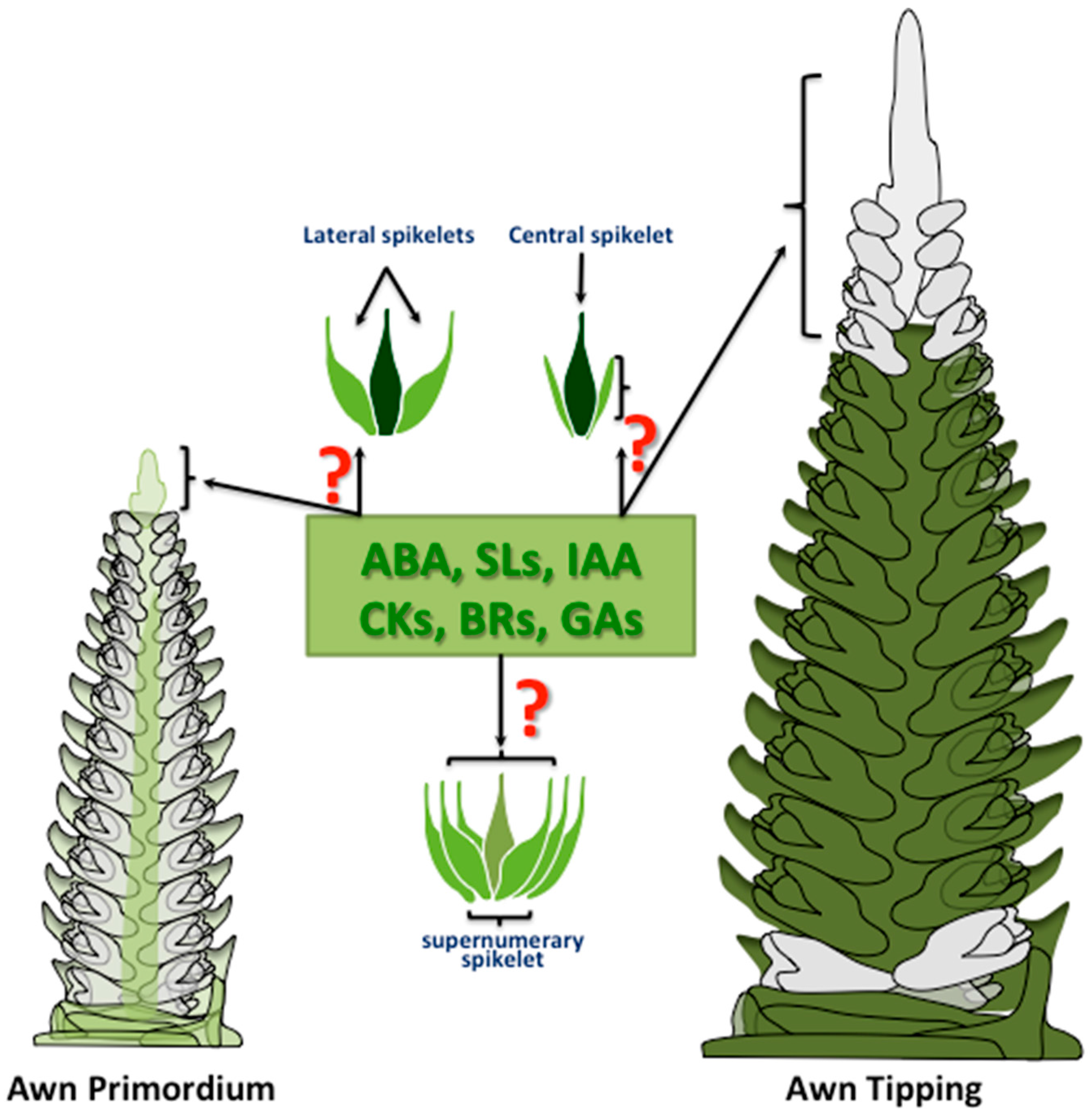
2.3. Phytohormone Regulation of Spike and Spikelet Development and Fertility
2.4. Phytohormone Regulation of the Leaf Area
3. Conclusions and Remarks
Acknowledgments
Author Contributions
Funding
Conflicts of Interest
References
- Ray, D.K.; Gerber, J.S.; MacDonald, G.K.; West, P.C. Climate variation explains a third of global crop yield variability. Nat. Commun. 2015, 6, 5989. [Google Scholar] [CrossRef] [PubMed]
- Donald, C.M.T. The breeding of crop ideotypes. Euphytica 1968, 17, 385–403. [Google Scholar] [CrossRef]
- Tao, F.; Rötter, R.P.; Palosuo, T.; Díaz-Ambron, C.G.H.; Mínguez, M.I.; Semenov, M.A.; Kersebaum, K.C.; Nendel, C.; Cammarano, D.; Hoffmann, H.; et al. Designing future barley ideotypes using a crop model ensemble. Eur. J. Agron. 2017, 82, 144–162. [Google Scholar] [CrossRef]
- Rasmusson, D.C. A plant breeder’s experience with ideotype breeding. Field Crops Res. 1991, 26, 191–200. [Google Scholar] [CrossRef]
- Tilman, D.; Balzer, C.; Hill, J.; Befort, B.L. Global food demand and the sustainable intensification of agriculture. Proc. Natl. Acad. Sci. USA 2011, 108, 20260–20264. [Google Scholar] [CrossRef] [PubMed]
- Rietveld, C.A.; Medland, S.E.; Derringer, J.; Yang, J.; Esko, T.; Martin, N.W.; Westra, H.J.; Shakhbazov, K.; Abdellaoui, A.; Agrawal, A.; et al. GWAS of 126,559 individuals identifies genetic variants associated with educational attainment. Science 2013, 340, 1467–1471. [Google Scholar] [CrossRef] [PubMed]
- Rogers, E.D.; Benfey, P.N. Regulation of plant root system architecture: Implications for crop advancement. Curr. Opin. Biotechnol. 2015, 32, 93–98. [Google Scholar] [CrossRef] [PubMed]
- Peleg, Z.; Blumwald, E. Hormone balance and abiotic stress tolerance in crop plants. Curr. Opin. Plant Biol. 2011, 14, 290–295. [Google Scholar] [CrossRef] [PubMed]
- Youssef, H.M.; Eggert, K.; Koppolu, R.; Alqudah, A.M.; Poursarebani, N.; Fazeli, A.; Sakuma, S.; Tagiri, A.; Rutten, T.; Govind, G.; et al. VRS2 regulates hormone-mediated inflorescence patterning in barley. Nat. Genet. 2017, 49, 157–161. [Google Scholar] [CrossRef] [PubMed]
- Alqudah, A.M.; Koppolu, R.; Wolde, G.M.; Graner, A.; Schnurbusch, T. The Genetic Architecture of Barley Plant Stature. Front. Genet. 2016, 7, 117. [Google Scholar] [CrossRef] [PubMed]
- Alqudah, A.M.; Youssef, H.M.; Graner, A.; Schnurbusch, T. Natural variation and genetic make-up of leaf blade area in spring barley. Theor. Appl. Genet. 2018. [Google Scholar] [CrossRef] [PubMed]
- Reinert, S.; Kortz, A.; Leon, J.; Naz, A.A. Genome-Wide Association Mapping in the Global Diversity Set Reveals New QTL Controlling Root System and Related Shoot Variation in Barley. Front. Plant Sci. 2016, 7, 1061. [Google Scholar] [CrossRef] [PubMed]
- Digel, B.; Tavakol, E.; Verderio, G.; Tondelli, A.; Xu, X.; Cattivelli, L.; Rossini, L.; von Korff, M. Photoperiod-H1 (Ppd-H1) Controls Leaf Size. Plant Physiol. 2016, 172, 405–415. [Google Scholar] [CrossRef] [PubMed]
- Thirulogachandar, V.; Alqudah, A.M.; Koppolu, R.; Rutten, T.; Graner, A.; Hensel, G.; Kumlehn, J.; Bräutigam, A.; Sreenivasulu, N.; Schnurbusch, T.; Kuhlmann, M. Leaf primordium size specifies leaf width and vein number among row-type classes in barley. Plant J. 2017, 91, 601–602. [Google Scholar] [CrossRef] [PubMed]
- Ramsay, L.; Comadran, J.; Druka, A.; Marshall, D.F.; Thomas, W.T.; Macaulay, M.; MacKenzie, K.; Simpson, C.; Fuller, J.; Bonar, N.; et al. INTERMEDIUM-C, a modifier of lateral spikelet fertility in barley, is an ortholog of the maize domestication gene TEOSINTE BRANCHED 1. Nat. Genet. 2011, 43, 169–172. [Google Scholar] [CrossRef] [PubMed]
- Sukumaran, S.; Dreisigacker, S.; Lopes, M.; Chavez, P.; Reynolds, M.P. Genome-wide association study for grain yield and related traits in an elite spring wheat population grown in temperate irrigated environments. Theor. Appl. Genet. 2015, 128, 353–363. [Google Scholar] [CrossRef] [PubMed]
- Hedden, P. The genes of the Green Revolution. Trends Genet. 2003, 19, 5–9. [Google Scholar] [CrossRef]
- Marzec, M.; Gruszka, D.; Tylec, P.; Szarejko, I. Identification and functional analysis of the HvD14 gene involved in strigolactone signaling in Hordeum vulgare. Physiol. Plant 2016, 158, 341–355. [Google Scholar] [CrossRef] [PubMed]
- Dockter, C.; Gruszka, D.; Braumann, I.; Druka, A.; Druka, I.; Franckowiak, J.; Gough, S.P.; Janeczko, A.; Kurowska, M.; Lundqvist, J.; et al. Induced variations in brassinosteroid genes define barley height and sturdiness, and expand the green revolution genetic toolkit. Plant Physiol. 2014, 166, 1912–1927. [Google Scholar] [CrossRef] [PubMed]
- Gupta, R.; Chakrabarty, S.K. Gibberellic acid in plant. Plant Signal. Behav. 2013, 8, e25504. [Google Scholar] [CrossRef] [PubMed]
- Tong, H.; Xiao, Y.; Liu, D.; Gao, S.; Liu, L.; Yin, Y.; Jin, Y.; Qian, Q.; Chu, C. Brassinosteroid Regulates Cell Elongation by Modulating Gibberellin Metabolism in Rice. Plant Cell 2014, 26, 4376–4393. [Google Scholar] [CrossRef] [PubMed]
- De Saint Germain, A.; Ligerot, Y.; Dun, E.A.; Pillot, J.-P.; Ross, J.J.; Beveridge, C.A.; Rameau, C. Strigolactones Stimulate Internode Elongation Independently of Gibberellins. Plant Physiol. 2013, 163, 1012–1025. [Google Scholar] [CrossRef] [PubMed]
- Daoura, B.G.; Chen, L.; Du, Y.; Hu, Y.G. Genetic effects of dwarfing gene Rht-5 on agronomic traits in common wheat (Triticum aestivum L.) and QTL analysis on its linked traits. Field Crops Res. 2014, 156, 22–29. [Google Scholar] [CrossRef]
- Nagano, H.; Onishi, K.; Ogasawara, M.; Horiuchi, Y.; Sano, Y. Genealogy of the “Green Revolution” gene in rice. Genes Genet. Syst. 2005, 80, 351–356. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Hellewell, K.B.; Rasmusson, D.C.; Gallo-Meagher, M. Enhancing yield in semidwarf barley. Crop Sci. 2000, 40, 352–358. [Google Scholar] [CrossRef]
- Al-Babili, S.; Bouwmeester, H.J. Strigolactones, a novel carotenoid-derived plant hormone. Annu. Rev. Plant Biol. 2015, 66, 161–186. [Google Scholar] [CrossRef] [PubMed]
- Kebrom, T.H.; Spielmeyer, W.; Finnegan, E.J. Grasses provide new insights into regulation of shoot branching. Trends Plant Sci. 2013, 18, 41–48. [Google Scholar] [CrossRef] [PubMed]
- Leopold, A.C. The control of tillering in grasses by auxin. Am. J. Bot. 1949, 36, 437–440. [Google Scholar] [CrossRef] [PubMed]
- Sharif, R.; Dale, J.E. Growth-regulating substances and the growth of tiller buds in barley; effects of cytokinins. J. Exp. Bot. 1980, 31, 921–930. [Google Scholar] [CrossRef]
- Liu, Y.; Gu, D.; Ding, Y.; Wang, Q.; Li, G.; Wang, S. The relationship between nitrogen, auxin and cytokinin in the growth regulation of rice (Oryza sativa L.) tiller buds. Aust. J.Crop Sci. 2011, 5, 1019–1026. [Google Scholar]
- Sakamoto, T.; Sakakibara, H.; Kojima, M.; Yamamoto, Y.; Nagasaki, H.; Inukai, Y.; Sato, Y.; Matsuoka, M. Ectopic expression of KNOTTED1-like homeobox protein induces expression of cytokinin biosynthesis genes in rice. Plant Physiol. 2006, 142, 54–62. [Google Scholar] [CrossRef] [PubMed]
- Umehara, M.; Hanada, A.; Yoshida, S.; Akiyama, K.; Arite, T.; Takeda-Kamiya, N.; Magome, H.; Kamiya, Y.; Shirasu, K.; Yoneyama, K.; et al. Inhibition of shoot branching by new terpenoid plant hormones. Nature 2008, 455, 195–200. [Google Scholar] [CrossRef] [PubMed]
- Marzec, M.; Muszynska, A. In Silico Analysis of the Genes Encoding Proteins that Are Involved in the Biosynthesis of the RMS/MAX/D Pathway Revealed New Roles of Strigolactones in Plants. Int. J. Mol. Sci. 2015, 16, 6757–6782. [Google Scholar] [CrossRef] [PubMed]
- Lo, S.F.; Yang, S.Y.; Chen, K.T.; Hsing, Y.I.; Zeevaart, J.A.; Chen, L.J.; Yu, S.M. A novel class of gibberellin 2-oxidases control semidwarfism, tillering, and root development in rice. Plant Cell 2008, 20, 2603–2618. [Google Scholar] [CrossRef] [PubMed]
- Jia, Q.; Zhang, X.Q.; Westcott, S.; Broughton, S.; Cakir, M.; Yang, J.; Lance, R.; Li, C. Expression level of a gibberellin 20-oxidase gene is associated with multiple agronomic and quality traits in barley. Theor. Appl. Genet. 2011, 122, 1451–1460. [Google Scholar] [CrossRef] [PubMed]
- Ito, S.; Yamagami, D.; Umehara, M.; Hanada, A.; Yoshida, S.; Sasaki, Y.; Yajima, S.; Kyozuka, J.; Ueguchi-Tanaka, M.; Matsuoka, M.; et al. Regulation of Strigolactone Biosynthesis by Gibberellin Signaling. Plant Physiol. 2017, 174, 1250–1259. [Google Scholar] [CrossRef] [PubMed]
- Marzec, M. Strigolactones and Gibberellins: A New Couple in the Phytohormone World? Trends Plant Sci. 2017, 22, 813–815. [Google Scholar] [CrossRef] [PubMed]
- Tong, H.; Jin, Y.; Liu, W.; Li, F.; Fang, J.; Yin, Y.; Qian, Q.; Zhu, L.; Chu, C. DWARF AND LOW-TILLERING, a new member of the GRAS family, plays positive roles in brassinosteroid signaling in rice. Plant J. 2009, 58, 803–816. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.Y.; Trieu, A.; Radhakrishnan, P.; Kwok, S.F.; Harris, S.; Zhang, K.; Wang, J.; Wan, J.; Zhai, H.; Takatsuto, S.; et al. Brassinosteroids regulate grain filling in rice. Plant Cell 2008, 20, 2130–2145. [Google Scholar] [CrossRef] [PubMed]
- Chono, M.; Honda, I.; Zeniya, H.; Yoneyama, K.; Saisho, D.; Takeda, K.; Takatsuto, S.; Hoshino, T.; Watanabe, Y. A Semidwarf Phenotype of Barley uzu Results from a Nucleotide Substitution in the Gene Encoding a Putative Brassinosteroid Receptor. Plant Physiol. 2003, 133, 1209–1219. [Google Scholar] [CrossRef] [PubMed]
- Gruszka, D.; Szarejko, I.; Maluszynski, M. New allele of HvBRI1 gene encoding brassinosteroid receptor in barley. J. Appl. Genet. 2011, 52, 257–268. [Google Scholar] [CrossRef] [PubMed]
- Pearce, S.; Vanzetti, L.S.; Dubcovsky, J. Exogenous gibberellins induce wheat spike development under short days only in the presence of VERNALIZATION1. Plant Physiol. 2013, 163, 1433–1445. [Google Scholar] [CrossRef] [PubMed]
- Pearce, S.; Huttly, A.K.; Prosser, I.M.; Li, Y.D.; Vaughan, S.P.; Gallova, B.; Patil, A.; Coghill, J.A.; Dubcovsky, J.; Hedden, P.; et al. Heterologous expression and transcript analysis of gibberellin biosynthetic genes of grasses reveals novel functionality in the GA3ox family. BMC Plant Biol. 2015, 15, 130. [Google Scholar] [CrossRef] [PubMed]
- Flintham, J.E.; Börner, A.; Worland, A.J.; Gale, M.D. Optimizing wheat grain yield: Effects of Rht (gibberellin-insensitive) dwarfing genes. J. Agric. Sci. 1997, 128, 11–25. [Google Scholar] [CrossRef]
- Boden, S.A.; Weiss, D.; Ross, J.J.; Davies, N.W.; Trevaskis, B.; Chandler, P.M.; Swain, S.M. EARLY FLOWERING3 Regulates Flowering in Spring Barley by Mediating Gibberellin Production and FLOWERING LOCUS T Expression. Plant Cell 2014, 26, 1557–1569. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Cao, W.; Dai, T.; Zhou, Q. Effects of exogenous hormones on floret development and grain set in wheat. Plant Growth Regul. 2001, 35, 225–231. [Google Scholar] [CrossRef]
- Zheng, C.; Zhu, Y.; Zhu, H.; Kang, G.; Guo, T.; Wang, C. Floret development and grain setting characteristics in winter wheat in response to pre-anthesis applications of 6-benzylaminopurine and boron. Field Crops Res. 2014, 169, 70–76. [Google Scholar] [CrossRef]
- Patel, R.; Mohapatra, P.K. Regulation of Spikelet Development in Rice by Hormones. J. Exp. Bot. 1992, 43, 257–262. [Google Scholar] [CrossRef]
- Cao, W.X.; Wang, Z.; Dai, T.B. Changes in levels of endogenous plant hormones during floret development in wheat genotypes of different spike sizes. J. Integr. Plant Biol. 2000, 42, 1026–1032. [Google Scholar]
- Wang, R.; Yu, Z.; Pan, Q.; Xu, Y. Changes of endogenous plant hormone contents during grain development in wheat. Zuo Wu Xue Bao 1999, 25, 227–231. [Google Scholar]
- Cai, Q.; Yuan, Z.; Chen, M.J.; Yin, C.S.; Luo, Z.J.; Zhao, X.X.; Liang, W.Q.; Hu, J.P.; Zhang, D.B. Jasmonic acid regulates spikelet development in rice. Nat. Commun. 2014, 5, 13. [Google Scholar] [CrossRef] [PubMed]
- Alqudah, A.M.; Schnurbusch, T. Awn primordium to tipping is the most decisive developmental phase for spikelet survival in barley. Funct. Plant Biol. 2014, 41, 424–436. [Google Scholar] [CrossRef]
- Alqudah, A.M.; Schnurbusch, T. Heading Date Is Not Flowering Time in Spring Barley. Front. Plant Sci. 2017, 8, 896. [Google Scholar] [CrossRef] [PubMed]
- Chhun, T.; Aya, K.; Asano, K.; Yamamoto, E.; Morinaka, Y.; Watanabe, M.; Kitano, H.; Ashikari, M.; Matsuoka, M.; Ueguchi-Tanaka, M. Gibberellin Regulates Pollen Viability and Pollen Tube Growth in Rice. Plant Cell 2007, 19, 3876–3888. [Google Scholar] [CrossRef] [PubMed]
- Ikeda, A.; Ueguchi-Tanaka, M.; Sonoda, Y.; Kitano, H.; Koshioka, M.; Futsuhara, Y.; Matsuoka, M.; Yamaguchi, J. slender Rice, a Constitutive Gibberellin Response Mutant, Is Caused by a Null Mutation of the SLR1 Gene, an Ortholog of the Height-Regulating Gene GAI/RGA/RHT/D8. Plant Cell 2001, 13, 999–1010. [Google Scholar] [CrossRef] [PubMed]
- Yuo, T.; Yamashita, Y.; Kanamori, H.; Matsumoto, T.; Lundqvist, U.; Sato, K.; Ichii, M.; Jobling, S.A.; Taketa, S. A SHORT INTERNODES (SHI) family transcription factor gene regulates awn elongation and pistil morphology in barley. J. Exp. Bot. 2012, 63, 5223–5232. [Google Scholar] [CrossRef] [PubMed]
- Eklund, D.M.; Ståldal, V.; Valsecchi, I.; Cierlik, I.; Eriksson, C.; Hiratsu, K.; Ohme-Takagi, M.; Sundström, J.F.; Thelander, M.; Ezcurra, I.; Sundberg, E. The Arabidopsis thaliana STYLISH1 Protein Acts as a Transcriptional Activator Regulating Auxin Biosynthesis. Plant Cell 2010, 22, 349–363. [Google Scholar] [CrossRef] [PubMed]
- Fridborg, I.; Kuusk, S.; Robertson, M.; Sundberg, E. The Arabidopsis Protein SHI Represses Gibberellin Responses in Arabidopsis and Barley. Plant Physiol. 2001, 127, 937–948. [Google Scholar] [CrossRef] [PubMed]
- Aya, K.; Ueguchi-Tanaka, M.; Kondo, M.; Hamada, K.; Yano, K.; Nishimura, M.; Matsuoka, M. Gibberellin Modulates Anther Development in Rice via the Transcriptional Regulation of GAMYB. Plant Cell 2009, 21, 1453–1472. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.M.; Cihlar, J. Plant canopy gap-size analysis theory for improving optical measurements of leaf-area index. Appl. Opt. 1995, 34, 6211–6222. [Google Scholar] [CrossRef] [PubMed]
- Jiang, D.; Fang, J.; Lou, L.; Zhao, J.; Yuan, S.; Yin, L.; Sun, W.; Peng, L.; Guo, B.; Li, X. Characterization of a null allelic mutant of the rice NAL1 gene reveals its role in regulating cell division. PLoS ONE 2015, 10, e0118169. [Google Scholar] [CrossRef] [PubMed]
- Driever, S.M.; Lawson, T.; Andralojc, P.; Raines, C.A.; Parry, M. Natural variation in photosynthetic capacity, growth, and yield in 64 field-grown wheat genotypes. J. Exp. Bot. 2014, 65, 4959–4973. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.M.; Huang, J.F.; Lou, Z.H. A comparison of three methods for estimating leaf area index of paddy rice from optimal hyperspectral bands. Precis. Agric. 2011, 12, 439–447. [Google Scholar] [CrossRef]
| No. | Chr. | cM (POP SEQ) | Gene | Barley High Conf. Gene | Contig Identifier | Trait | Hormone |
|---|---|---|---|---|---|---|---|
| 1 | 1H | 5.38 | BRASSINOSTEROID-6-OXIDASE (HvBRD1) | AK372445 | morex_contig_244330 CAJW010244330 | Tillering | BRs |
| 2 | 1H | 55.52 | GIBBERELLIN INSENSITIVE DWARF1 (HvGID1) | AK356665 | morex_contig_137029 CAJW010137029 | Tillering | GAs |
| 3 | 1H | 94.75 | GIBBERELLIN 20 OXIDASE 4 (HvGA2ox4) | MLOC_13981.1 | morex_contig_1566970 CAJW011566970 | Tillering and leaf area | GAs |
| 4 | 2H | 58.78 | GIBBERELLIN-INSENSITIVE DWARF 2 (HvGID2) | MLOC_61457.1 | morex_contig_41142 | Tillering | GAs |
| 5 | 2H | 59.91 | DWARF 11 (HvD11), CYTOCHROME P450 724B1 | AK371371 | morex_contig_45000 CAJW010045000 | Tillering | BRs |
| 6 | 3H | 44.26 | DWARF 2 (HvD2), CYTOCHROME P450 90D2 | MLOC_62829.1 | morex_contig_47012 CAJW010047012 | Tillering | BRs |
| 7 | 3H | 46.03 | GIBBERELLIN 3 OXIDASE 2 (HvGA3ox2) | MLOC_12855.1 | morex_contig_51542 CAJW010051542 | Tillering | GAs |
| 8 | 3H | 46.03 | DWARF 18 (HvD18) | MLOC_12855.1 | morex_contig_51542 CAJW010051542 | Tillering | GAs |
| 9 | 3H | 51.34 | BRASSINOSTEROID INSENSITIVE 1 /SEMIBRACHYTIC/Dwarf61 (HvBRI1/ uzu1 HvD61/) | MLOC_5176.2 | morex_contig_58772 CAJW010058772 | Tillering and leaf area | BRs |
| 10 | 3H | 62.93 | MORE AXILLARY BRANCHES 4/ CAROTENOID CLEAVAGE DIOXYGENASE 8 /DWARF 10 (HvMAX4/CHvCD8/ HvD10) | MLOC_66551.1 | morex_contig_51744 CAJW010051744 | Tillering and leaf area | SLs |
| 11 | 3H | 64.16 | GIBBERELLIN 20 OXIDASE 1 (HvGA2ox1) | AK364775 | morex_contig_2550522 CAJW012550522 | Tillering and leaf area | GAs |
| 12 | 3H | 106.02 | GIBBERELLIN 20 OXIDASE 3 (HvGA20ox3) | MLOC_66389.1 | morex_contig_51490 CAJW010051490 | Tillering | GAs |
| 13 | 4H | 59.63 | DWARF 4 (HvD4) | AK355174 | morex_contig_61948 CAJW010061948 | Plant height | BRs |
| 14 | 5H | 44.02 | BRASSINOSTEROID C-23 HYDROXYLASE (HvCPD) | MLOC_10658.1 | morex_contig_1559549 CAJW011559549 | Tillering, plant height and leaf area | BRs |
| 15 | 5H | 46.59 | DWARF 53 (HvD53) | AK372211 | morex_contig_244827 CAJW010244827 | Leaf area | SLs |
| 16 | 5H | 47.22 | BRITTLE CULM12/ GIBBERELLIN-DEFICIENT DWARF 1 (HvBC12/GGD1) | AK373790 | morex_contig_45441 CAJW010045441 | Tillering | GAs |
| 17 | 5H | 80.8 | DWARF RICE WITH OVEREXPRESSION OF GIBBERELLIN-INDUCED GENE (HvDOG)s | AK359310 | morex_contig_1575121 CAJW011575121 | Tillering | GAs |
| 18 | 7H | 29.95 | MORE AXILLARY BRANCHES 2 (HvMAX2) | MLOC_4044.5 | morex_contig_134615 CAJW01013461 | Leaf area | SLs |
| 19 | 7H | 77.4 | DWARF 35 (HvD35), CYTOCHROME P450 701A6 | AK369327 | morex_contig_1575857 CAJW011575857 | Tillering and leaf area | GAs |
| 20 | 7H | 140.65 | BRASSINOSTEROID DEFICIENT DWARF 2/ DIMINUTO, DWARF1 (HvBRD2/HvDIM/HvDWF1) | MLOC_52405.2 | morex_contig_37512 CAJW010037512 | Tillering and leaf area | BRs |
| Chr | Physical pos. | Gene | Barley HC Gene | Annotation | Accession | GO Term | Function | Hormone |
|---|---|---|---|---|---|---|---|---|
| 3H | 627490751-627493520 | EXTRA GLUME 1 EG1 | HORVU3Hr1G089140 | Phospholipase A1-II 1 | 24647160 | GO:0006629 | Regulation of spikelet development [51] | JA [51] |
| 2H | 725648938–725655525 | EXTRA GLUME 2 EG2 | HORVU2Hr1G112360 | jasmonate-zim-domain protein 12 | AK069326 | none | Regulation of spikelet development [51] | JA [51] |
| 4H | 16668801–16671743 | SLENDER 1 SL1 | HORVU4Hr1G006930 | DELLA protein | AB262980 | none | Regulation of floral development and spikelet fertility | GAs |
| 1H | 441473716–441477633 | GIBBERELLIN INSENSITIVE DWARF1 GID1 | HORVU1Hr1G060810 | Gibberellin receptor GID1 | AK074026 | GO:0008152, GO:0016787 | GAs | |
| 5H | 564406197–564410417 | Six-rowed spike 2 VRS2 | HORVU5Hr1G081450.1 | SHI-related sequence 5 | KX601696.1 | none | Regulation of spikelet development and fertility [9], awn elongation and pistil shape [56] | IAA [57], GAs [58] |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Marzec, M.; Alqudah, A.M. Key Hormonal Components Regulate Agronomically Important Traits in Barley. Int. J. Mol. Sci. 2018, 19, 795. https://doi.org/10.3390/ijms19030795
Marzec M, Alqudah AM. Key Hormonal Components Regulate Agronomically Important Traits in Barley. International Journal of Molecular Sciences. 2018; 19(3):795. https://doi.org/10.3390/ijms19030795
Chicago/Turabian StyleMarzec, Marek, and Ahmad M. Alqudah. 2018. "Key Hormonal Components Regulate Agronomically Important Traits in Barley" International Journal of Molecular Sciences 19, no. 3: 795. https://doi.org/10.3390/ijms19030795
APA StyleMarzec, M., & Alqudah, A. M. (2018). Key Hormonal Components Regulate Agronomically Important Traits in Barley. International Journal of Molecular Sciences, 19(3), 795. https://doi.org/10.3390/ijms19030795