Key Players of Cisplatin Resistance: Towards a Systems Pharmacology Approach
Abstract
1. Introduction
2. Results
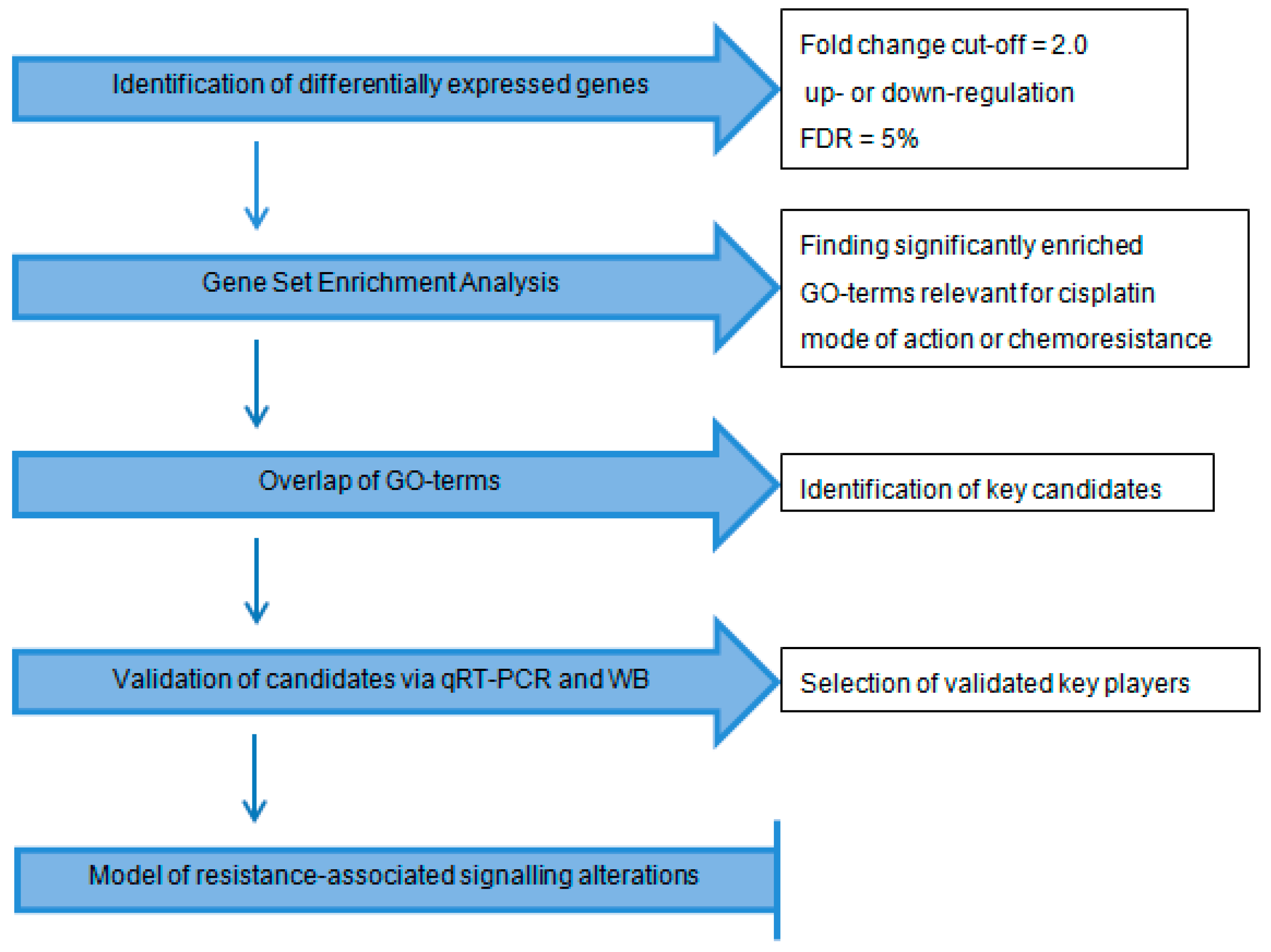
2.1. Workflow
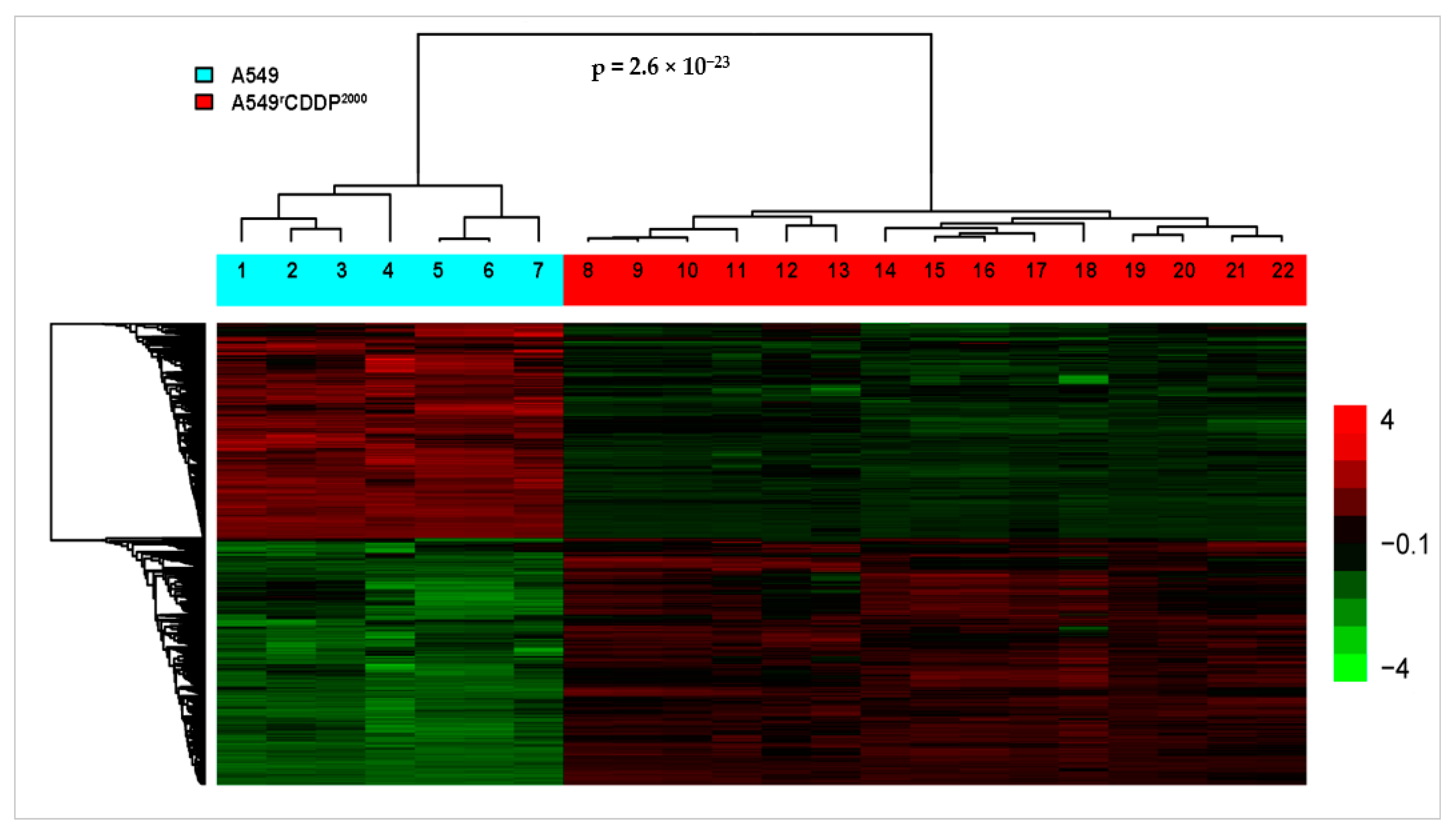
2.2. Microarray Analysis
2.2.1. Differentially Expressed Genes
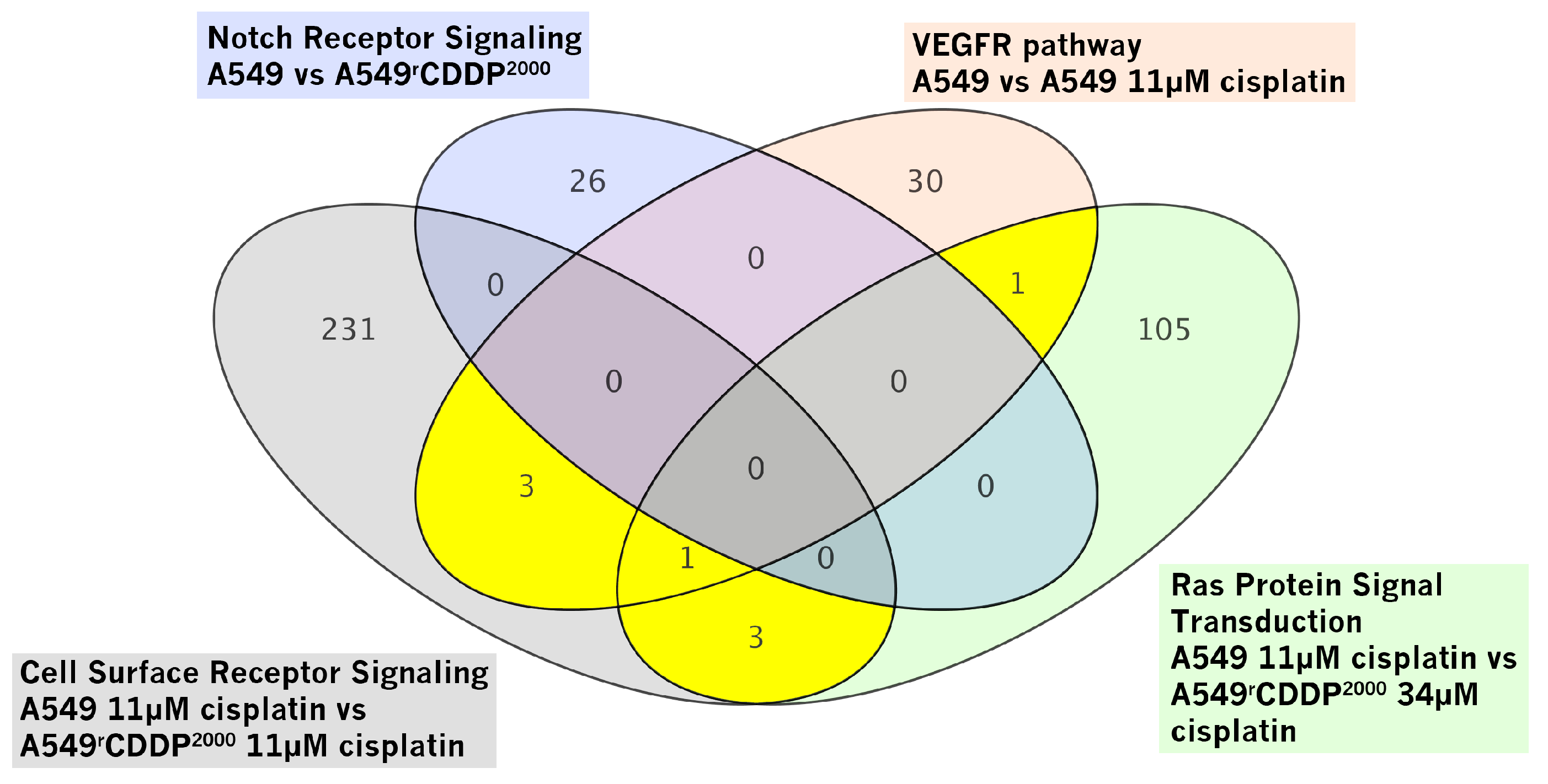
2.2.2. Gene Set Enrichment Analysis
2.3. Evaluation of the Identified Candidates
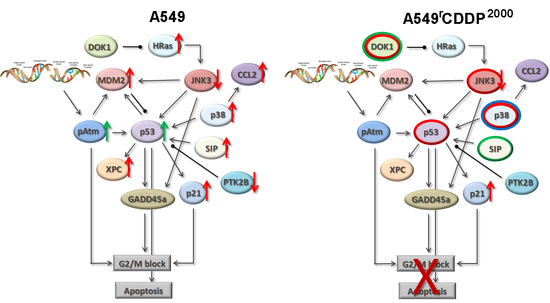
2.4. Extended Model of Resistance-Associated Signalling Alterations
3. Discussion
3.1. Systems Pharmacology Approach
3.2. Role of the Identified Key Players
3.3. Model of Resistance-Associated Signalling Alterations
4. Materials and Methods
4.1. Drugs
4.2. Cell Lines
4.3. Microarray
4.4. RNA Isolation, cDNA Synthesis and qRT-PCR
4.5. SDS-PAGE and Western Blot
4.6. Statistical Analysis
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| NSCLC | non-small cell lung cancer |
| CDDP, DDP | cis-diamminedichloroplatinum (II) (cisplatin) |
| GO | Gene Ontology |
| GSEA | Gene Set Enrichment Analysis |
| FDR | false discovery rate |
| VEGFR | vascular endothelial growth factor receptor |
| TBS | Tris-buffered saline |
| MAPK | mitogen-activated protein kinase |
| ERK | Extracellular-signal Regulated Kinase |
| DOK1 | Docking protein 1 |
| PTK2B, Pyk2 | Protein tyrosine kinase 2 beta |
| CCL2 | C-C motif chemokine ligand 2 |
| MCP-1 | monocyte chemotactic protein 1 |
| MAPKAPK2 | MAP kinase-activated protein kinase 2 |
| JNK | c-Jun N-terminal kinase |
| MDM2 | mouse double minute 2 homolog |
| GAPDH | glyceraldehyde 3-phosphate dehydrogenase |
References
- Besse, B.; Adjei, A.; Baas, P.; Meldgaard, P.; Nicolson, M.; Paz-Ares, L.; Reck, M.; Smit, E.F.; Syrigos, K.; Stahel, R.; et al. 2nd ESMO Consensus Conference on Lung Cancer: Non-small-cell lung cancer first-line/second and further lines of treatment in advanced disease. Ann. Oncol. 2014, 25, 1475–1484. [Google Scholar] [CrossRef] [PubMed]
- Shtivelman, E.; Hensing, T.; Simon, G.R.; Dennis, P.A.; Otterson, G.A.; Bueno, R.; Salgia, R. Molecular pathways and therapeutic targets in lung cancer. Oncotarget 2014, 5, 1392–1433. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Liu, Z.M.; Li, X.C.; Yao, Y.T.; Yin, Z.X. Activation of ERK1/2 and Akt is associated with cisplatin resistance in human lung cancer cells. J. Chemother. 2013, 25, 162–169. [Google Scholar] [CrossRef] [PubMed]
- Jordan, P.; Carmo-Fonseca, M. Molecular mechanisms involved in cisplatin cytotoxicity. Cell. Mol. Life Sci. 2000, 57, 1229–1235. [Google Scholar] [CrossRef] [PubMed]
- Fajac, A.; Da Silva, J.; Ahomadegbe, J.C.; Rateau, J.G.; Bernaudin, J.F.; Riou, G.; Bénard, J. Cisplatin-induced apoptosis and p53 gene status in a cisplatin-resistant human ovarian carcinoma cell line. Int. J. Cancer 1996, 68, 67–74. [Google Scholar] [CrossRef] [PubMed]
- Sarin, N.; Engel, F.; Kalayda, G.V.; Mannewitz, M.; Cinatl, J.; Rothweiler, F.; Michaelis, M.; Saafan, H.; Ritter, C.A.; Jaehde, U.; et al. Cisplatin resistance in non-small cell lung cancer cells is associated with an abrogation of cisplatin-induced G2/M cell cycle arrest. PLoS ONE 2017, 12, e0181081. [Google Scholar] [CrossRef] [PubMed]
- Galluzzi, L.; Vitale, I.; Michels, J.; Brenner, C.; Szabadkai, G.; Harel-Bellan, A.; Castedo, M.; Kroemer, G. Systems biology of cisplatin resistance: Past, present and future. Cell Death Dis. 2014, 5, e1257. [Google Scholar] [CrossRef] [PubMed]
- Galluzzi, L.; Vitale, I.; Senovilla, L.; Eisenberg, T.; Carmona-Gutierrez, D.; Vacchelli, E.; Robert, T.; Ripoche, H.; Jägemann, N.; Paccard, C.; et al. Independent transcriptional reprogramming and apoptosis induction by cisplatin. Cell Cycle 2012, 11, 3472–3480. [Google Scholar] [CrossRef] [PubMed]
- Toshimitsu, H.; Hashimoto, K.; Tangoku, A.; Iizuka, N.; Yamamoto, K.; Kawauchi, S.; Oga, A.; Furuya, T.; Oka, M.; Sasaki, K. Molecular signature linked to acquired resistance to cisplatin in oesophageal cancer cells. Cancer Lett. 2004, 211, 69–78. [Google Scholar] [CrossRef] [PubMed]
- Gatti, L.; Chen, D.; Beretta, G.L.; Rustici, G.; Carenini, N.; Corna, E.; Colangelo, D.; Zunino, F.; Bähler, J.; Perego, P. Global gene expression of fission yeast in response to cisplatin. Cell. Mol. Life Sci. 2004, 61, 2253–2263. [Google Scholar] [CrossRef] [PubMed]
- Cheng, T.C.; Manorek, G.; Samimi, G.; Lin, X.; Berry, C.C.; Howell, S.B. Identification of genes whose expression is associated with cisplatin resistance in human ovarian carcinopma cells. Cancer Chemother. Pharmacol. 2006, 58, 384–395. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Li, H.; Hou, S.; Hu, B.; Liu, J.; Wang, J. Differences in gene expression profiles and carcinogenesis pathways involved in cisplatin resistance of four types of cancer. Oncol. Rep. 2013, 30, 596–614. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Li, H.; Hou, S.; Hu, B.; Liu, J.; Wang, J. The noncoding RNA expression profile and the effect of lncRNA AK126698 on cisplatin resistance in non-small-cell lung cancer cell. PLoS ONE 2013, 8, e65309. [Google Scholar] [CrossRef] [PubMed]
- Hu, L.; Chen, J.; Zhang, F.; Wang, J.; Pan, J.; Chen, J.; Wang, Y. Aberrant Long Noncoding RNAs Expression Profiles Affect Cisplatin Resistance in Lung Adenocarcinoma. BioMed Res. Int. 2017, 2017, 7498151. [Google Scholar] [CrossRef] [PubMed]
- Fang, Y.; Zhang, C.; Wu, T.; Wang, Q.; Liu, J.; Dai, P. Transcriptome Sequencing Reveals Key Pathways and Genes Associated with Cisplatin Resistance in Lung Adenocarcinoma A549 Cells. PLoS ONE 2017, 12, e0170609. [Google Scholar] [CrossRef] [PubMed]
- Stewart, J.J.; White, J.T.; Yan, X.; Collins, S.; Drescher, C.W.; Urban, N.D.; Hood, L.; Lin, B. Proteins associated with Cisplatin resistance in ovarian cancer cells identified by quantitative proteomic technology and integrated with mRNA expression levels. Mol. Cell. Proteom. 2006, 5, 433–443. [Google Scholar] [CrossRef] [PubMed]
- Zeng, H.-Z.; Qu, Y.-Q.; Zhang, W.-J.; Xiu, B.; Deng, A.-M.; Liang, A.-B. Proteomic analysis identified DJ-1 as a cisplatin resistant marker in non-small cell lung cancer. Int. J. Mol. Sci. 2011, 12, 3489–3499. [Google Scholar] [CrossRef] [PubMed]
- Stark, A.L.; Madian, A.G.; Williams, S.W.; Chen, V.; Wing, C.; Hause, R.J.; To, L.A.; Gill, A.L.; Myers, J.L.; Gorsic, L.K.; et al. Identification of Novel Protein Expression Changes Following Cisplatin Treatment and Application to Combination Therapy. J. Proteome Res. 2017, 16, 4227–4236. [Google Scholar] [CrossRef] [PubMed]
- Wist, A.D.; Berger, S.I.; Iyengar, R. Systems pharmacology and genome medicine: A future perspective. Genome Med. 2009, 1, 11. [Google Scholar] [CrossRef] [PubMed]
- Subramanian, A.; Tamayo, P.; Mootha, V.K.; Mukherjee, S.; Ebert, B.L.; Gillette, M.A.; Paulovich, A.; Pomeroy, S.L.; Golub, T.R.; Lander, E.S.; et al. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 2005, 102, 15545–15550. [Google Scholar] [CrossRef] [PubMed]
- Goeman, J.J.; van de Geer, S.A.; de Kort, F.; van Houwelingen, H.C. A global test for groups of genes: Testing association with a clinical outcome. Bioinformatics 2004, 20, 93–99. [Google Scholar] [CrossRef] [PubMed]
- Harris, M.A.; Clark, J.; Ireland, A.; Lomax, J.; Ashburner, M.; Foulger, R.; Eilbeck, K.; Lewis, S.; Marshall, B.; Mungall, C.; et al. The Gene Ontology (GO) database and informatics resource. Nucleic Acids Res. 2004, 32, D258–D261. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Terfve, C.; Rose, J.C.; Markowetz, F. HTSanalyzeR: An R/Bioconductor package for integrated network analysis of high-throughput screens. Bioinformatics 2011, 27, 879–880. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Mao, Z.; Huang, J.; Xie, S.; Liu, T.; Mao, Z. Blocking the NOTCH pathway can inhibit the growth of CD133-positive A549 cells and sensitize to chemotherapy. Biochem. Biophys. Res. Commun. 2014, 444, 670–675. [Google Scholar] [CrossRef] [PubMed]
- McAuliffe, S.M.; Morgan, S.L.; Wyant, G.A.; Tran, L.T.; Muto, K.W.; Chen, Y.S.; Chin, K.T.; Partridge, J.C.; Poole, B.B.; Cheng, K.-H.; et al. Targeting Notch, a key pathway for ovarian cancer stem cells, sensitizes tumors to platinum therapy. Proc. Natl. Acad. Sci. USA 2012, 109, E2939–E2948. [Google Scholar] [CrossRef] [PubMed]
- Daenen, L.G.M.; Roodhart, J.M.L.; van Amersfoort, M.; Dehnad, M.; Roessingh, W.; Ulfman, L.H.; Derksen, P.W.B.; Voest, E.E. Chemotherapy enhances metastasis formation via VEGFR-1-expressing endothelial cells. Cancer Res. 2011, 71, 6976–6985. [Google Scholar] [CrossRef] [PubMed]
- Sini, P.; Samarzija, I.; Baffert, F.; Littlewood-Evans, A.; Schnell, C.; Theuer, A.; Christian, S.; Boos, A.; Hess-Stumpp, H.; Foekens, J.A.; et al. Inhibition of multiple vascular endothelial growth factor receptors (VEGFR) blocks lymph node metastases but inhibition of VEGFR-2 is sufficient to sensitize tumor cells to platinum-based chemotherapeutics. Cancer Res. 2008, 68, 1581–1592. [Google Scholar] [CrossRef] [PubMed]
- Granados, M.L.; Hudson, L.G.; Samudio-Ruiz, S.L. Contributions of the Epidermal Growth Factor Receptor to Acquisition of Platinum Resistance in Ovarian Cancer Cells. PLoS ONE 2015, 10, e0136893. [Google Scholar] [CrossRef] [PubMed]
- Juliachs, M.; Muñoz, C.; Moutinho, C.A.; Vidal, A.; Condom, E.; Esteller, M.; Graupera, M.; Casanovas, O.; Germà, J.R.; Villanueva, A.; et al. The PDGFRβ-AKT pathway contributes to CDDP-acquired resistance in testicular germ cell tumors. Clin. Cancer Res. 2014, 20, 658–667. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, T.; Tsigelny, I.F.; Götz, A.W.; Howell, S.B. Cisplatin inhibits MEK1/2. Oncotarget 2015, 6, 23510–23522. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Zhou, Y.; Li, Y.; Zhou, J.; Wu, Y.; Cui, Y.; Yang, G.; Hong, Y. Mutations of p53 and KRAS activate NF-κB to promote chemoresistance and tumorigenesis via dysregulation of cell cycle and suppression of apoptosis in lung cancer cells. Cancer Lett. 2015, 357, 520–526. [Google Scholar] [CrossRef] [PubMed]
- Cuenda, A.; Rousseau, S. p38 MAP-kinases pathway regulation, function and role in human diseases. Biochim. Biophys. Acta 2007, 1773, 1358–1375. [Google Scholar] [CrossRef] [PubMed]
- Rane, M.J.; Coxon, P.Y.; Powell, D.W.; Webster, R.; Klein, J.B.; Pierce, W.; Ping, P.; McLeish, K.R. p38 Kinase-dependent MAPKAPK-2 activation functions as 3-phosphoinositide-dependent kinase-2 for Akt in human neutrophils. J. Biol. Chem. 2001, 276, 3517–3523. [Google Scholar] [CrossRef] [PubMed]
- Cho, J.; Gruol, D.L. The chemokine CCL2 activates p38 mitogen-activated protein kinase pathway in cultured rat hippocampal cells. J. Neuroimmunol. 2008, 199, 94–103. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Warner, G.M.; Yin, P.; Knudsen, B.E.; Cheng, J.; Butters, K.A.; Lien, K.R.; Gray, C.E.; Garovic, V.D.; Lerman, L.O.; et al. Inhibition of p38 MAPK attenuates renal atrophy and fibrosis in a murine renal artery stenosis model. Am. J. Physiol. Ren. Physiol. 2013, 304, F938–F947. [Google Scholar] [CrossRef] [PubMed]
- Mercier, P.-L.; Bachvarova, M.; Plante, M.; Gregoire, J.; Renaud, M.-C.; Ghani, K.; Têtu, B.; Bairati, I.; Bachvarov, D. Characterization of DOK1, a candidate tumor suppressor gene, in epithelial ovarian cancer. Mol. Oncol. 2011, 5, 438–453. [Google Scholar] [CrossRef] [PubMed]
- Geng, W.; Ng, K.T.P.; Sun, C.K.W.; Yau, W.L.; Liu, X.B.; Cheng, Q.; Poon, R.T.P.; Lo, C.M.; Man, K.; Fan, S.T. The role of proline rich tyrosine kinase 2 (Pyk2) on cisplatin resistance in hepatocellular carcinoma. PLoS ONE 2011, 6, e27362. [Google Scholar] [CrossRef] [PubMed]
- Lim, S.-T.; Miller, N.L.G.; Nam, J.-O.; Chen, X.L.; Lim, Y.; Schlaepfer, D.D. Pyk2 inhibition of p53 as an adaptive and intrinsic mechanism facilitating cell proliferation and survival. J. Biol. Chem. 2010, 285, 1743–1753. [Google Scholar] [CrossRef] [PubMed]
- Mashima, R.; Hishida, Y.; Tezuka, T.; Yamanashi, Y. The roles of Dok family adapters in immunoreceptor signaling. Immunol. Rev. 2009, 232, 273–285. [Google Scholar] [CrossRef] [PubMed]
- Binétruy, B.; Smeal, T.; Karin, M. Ha-Ras augments c-Jun activity and stimulates phosphorylation of its activation domain. Nature 1991, 351, 122–127. [Google Scholar] [CrossRef] [PubMed]
- Smeal, T.; Binetruy, B.; Mercola, D.A.; Birrer, M.; Karin, M. Oncogenic and transcriptional cooperation with Ha-Ras requires phosphorylation of c-Jun on serines 63 and 73. Nature 1991, 354, 494–496. [Google Scholar] [CrossRef] [PubMed]
- Fuchs, S.Y.; Adler, V.; Pincus, M.R.; Ronai, Z. MEKK1/JNK signaling stabilizes and activates p53. Proc. Natl. Acad. Sci. USA 1998, 95, 10541–10546. [Google Scholar] [CrossRef] [PubMed]
- Wu, G.S. The functional interactions between the p53 and MAPK signaling pathways. Cancer Biol. Ther. 2004, 3, 156–161. [Google Scholar] [CrossRef] [PubMed]
- Hildesheim, J.; Fornace, A.J. Gadd45a: An elusive yet attractive candidate gene in pancreatic cancer. Clin. Cancer Res. 2002, 8, 2475–2479. [Google Scholar] [CrossRef] [PubMed]
- Sanchez-Prieto, R.; Rojas, J.M.; Taya, Y.; Gutkind, J.S. A role for the p38 mitogen-acitvated protein kinase pathway in the transcriptional activation of p53 on genotoxic stress by chemotherapeutic agents. Cancer Res. 2000, 60, 2464–2472. [Google Scholar] [PubMed]
- Wang, X.; Martindale, J.L.; Holbrook, N.J. Requirement for ERK activation in cisplatin-induced apoptosis. J. Biol. Chem. 2000, 275, 39435–39443. [Google Scholar] [CrossRef] [PubMed]
- Galluzzi, L.; Vitale, I.; Senovilla, L.; Olaussen, K.A.; Pinna, G.; Eisenberg, T.; Goubar, A.; Martins, I.; Michels, J.; Kratassiouk, G.; et al. Prognostic impact of vitamin B6 metabolism in lung cancer. Cell Rep. 2012, 2, 257–269. [Google Scholar] [CrossRef] [PubMed]
- Maier, T.; Guell, M.; Serrano, L. Correlation of mRNA and protein in complex biological samples. FEBS Lett. 2009, 583, 3966–3973. [Google Scholar] [CrossRef] [PubMed]
- Arbiser, J.L.; Moses, M.A.; Fernandez, C.A.; Ghiso, N.; Cao, Y.; Klauber, N.; Frank, D.; Brownlee, M.; Flynn, E.; Parangi, S.; et al. Oncogenic H-ras stimulates tumor angiogenesis by two distinct pathways. Proc. Natl. Acad. Sci. USA 1997, 94, 861–866. [Google Scholar] [CrossRef] [PubMed]
- To, M.D.; Wong, C.E.; Karnezis, A.N.; Del Rosario, R.; Di Lauro, R.; Balmain, A. Kras regulatory elements and exon 4A determine mutation specificity in lung cancer. Nat. Genet. 2008, 40, 1240–1244. [Google Scholar] [CrossRef] [PubMed]
- Sklar, M.D. Increased resistance to cis-diamminedichloroplatinum(II) in NIH 3T3 cells transformed by ras oncogenes. Cancer Res. 1988, 48, 793–797. [Google Scholar] [PubMed]
- Wagner, E.F.; Nebreda, A.R. Signal integration by JNK and p38 MAPK pathways in cancer development. Nat. Rev. Cancer 2009, 9, 537–549. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Atayar, C.; Rosati, S.; Bosga-Bouwer, A.; Kluin, P.; Visser, L. JNK is constitutively active in mantle cell lymphoma: Cell cycle deregulation and polyploidy by JNK inhibitor SP600125. J. Pathol. 2009, 218, 95–103. [Google Scholar] [CrossRef] [PubMed]
- Niedner, H.; Christen, R.; Lin, X.; Kondo, A.; Howell, S.B. Identification of genes that mediate sensitivity to cisplatin. Mol. Pharmacol. 2001, 60, 1153–1160. [Google Scholar] [CrossRef] [PubMed]
- Shih, C.-M.; Chen, K.; Wang, Y.-C.; Lee, P.-J.; Wang, Y.-C. Elevated p53 and p21waf1 mRNA expression in blood lymphocytes from lung cancer patients with chemoresistance. Cancer Detect. Prev. 2007, 31, 366–370. [Google Scholar] [CrossRef] [PubMed]
- Vega, G.G.; Avilés-Salas, A.; Chalapud, J.R.; Martinez-Paniagua, M.; Pelayo, R.; Mayani, H.; Hernandez-Pando, R.; Martinez-Maza, O.; Huerta-Yepez, S.; Bonavida, B.; et al. P38 MAPK expression and activation predicts failure of response to CHOP in patients with Diffuse Large B-Cell Lymphoma. BMC Cancer 2015, 15, 722. [Google Scholar] [CrossRef] [PubMed]
- Pereira, L.; Igea, A.; Canovas, B.; Dolado, I.; Nebreda, A.R. Inhibition of p38 MAPK sensitizes tumour cells to cisplatin-induced apoptosis mediated by reactive oxygen species and JNK. EMBO Mol. Med. 2013, 5, 1759–1774. [Google Scholar] [CrossRef] [PubMed]
- Tsai, M.-S.; Weng, S.-H.; Chen, H.-J.; Chiu, Y.-F.; Huang, Y.-C.; Tseng, S.-C.; Kuo, Y.-H.; Lin, Y.-W. Inhibition of p38 MAPK-dependent excision repair cross-complementing 1 expression decreases the DNA repair capacity to sensitize lung cancer cells to etoposide. Mol. Cancer Ther. 2012, 11, 561–571. [Google Scholar] [CrossRef] [PubMed]
- Fader, A.N.; Rasool, N.; Vaziri, S.A.J.; Kozuki, T.; Faber, P.W.; Elson, P.; Biscotti, C.V.; Michener, C.M.; Rose, P.G.; Rojas-Espaillat, L.; et al. CCL2 expression in primary ovarian carcinoma is correlated with chemotherapy response and survival outcomes. Anticancer Res. 2010, 30, 4791–4798. [Google Scholar] [PubMed]
- Levina, V.; Su, Y.; Nolen, B.; Liu, X.; Gordin, Y.; Lee, M.; Lokshin, A.; Gorelik, E. Chemotherapeutic drugs and human tumor cells cytokine network. Int. J. Cancer 2008, 123, 2031–2040. [Google Scholar] [CrossRef] [PubMed]
- Michaelis, M.; Rothweiler, F.; Barth, S.; Cinatl, J.; van Rikxoort, M.; Löschmann, N.; Voges, Y.; Breitling, R.; von Deimling, A.; Rödel, F.; et al. Adaptation of cancer cells from different entities to the MDM2 inhibitor nutlin-3 results in the emergence of p53-mutated multi-drug-resistant cancer cells. Cell Death Dis. 2011, 2, e243. [Google Scholar] [CrossRef] [PubMed]
- Ritchie, M.E.; Silver, J.; Oshlack, A.; Holmes, M.; Diyagama, D.; Holloway, A.; Smyth, G.K. A comparison of background correction methods for two-colour microarrays. Bioinformatics 2007, 23, 2700–2707. [Google Scholar] [CrossRef] [PubMed]
- Bolstad, B.M.; Irizarry, R.A.; Astrand, M.; Speed, T.P. A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics 2003, 19, 185–193. [Google Scholar] [CrossRef] [PubMed]
- Smyth, G.K. Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat. Appl. Genet. Mol. Biol. 2004, 3. [Google Scholar] [CrossRef] [PubMed]

 ) and A549rCDDP2000 (
) and A549rCDDP2000 ( ) before (ctrl) and after treatment with 11 µM cisplatin (11) or 34 µM cisplatin (34) presented as mean ± SEM; as well as representative Western blots. * p < 0.05; ** p < 0.01; *** p < 0.01.
) before (ctrl) and after treatment with 11 µM cisplatin (11) or 34 µM cisplatin (34) presented as mean ± SEM; as well as representative Western blots. * p < 0.05; ** p < 0.01; *** p < 0.01.
 ) and A549rCDDP2000 (
) and A549rCDDP2000 ( ) before (ctrl) and after treatment with 11 µM cisplatin (11) or 34 µM cisplatin (34) presented as mean ± SEM; as well as representative Western blots. * p < 0.05; ** p < 0.01; *** p < 0.01.
) before (ctrl) and after treatment with 11 µM cisplatin (11) or 34 µM cisplatin (34) presented as mean ± SEM; as well as representative Western blots. * p < 0.05; ** p < 0.01; *** p < 0.01.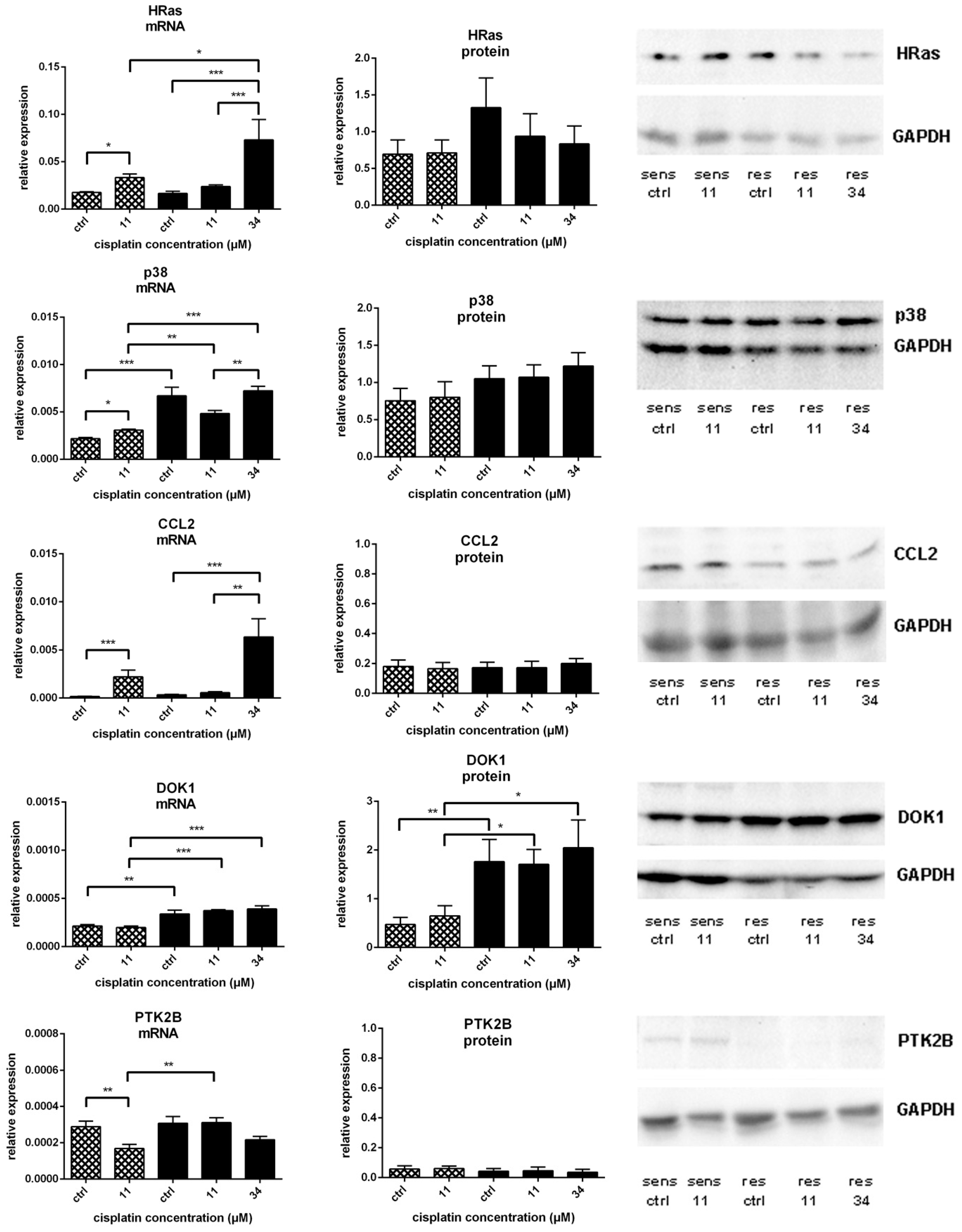
 ) and A549rCDDP2000 (
) and A549rCDDP2000 ( ) after treatment with 11 µM cisplatin (11) or 34 µM cisplatin (34) expressed as mean ± SEM, as well as representative Western blots.* p < 0.05; ** p < 0.01; *** p < 0.01.
) after treatment with 11 µM cisplatin (11) or 34 µM cisplatin (34) expressed as mean ± SEM, as well as representative Western blots.* p < 0.05; ** p < 0.01; *** p < 0.01.
 ) and A549rCDDP2000 (
) and A549rCDDP2000 ( ) after treatment with 11 µM cisplatin (11) or 34 µM cisplatin (34) expressed as mean ± SEM, as well as representative Western blots.* p < 0.05; ** p < 0.01; *** p < 0.01.
) after treatment with 11 µM cisplatin (11) or 34 µM cisplatin (34) expressed as mean ± SEM, as well as representative Western blots.* p < 0.05; ** p < 0.01; *** p < 0.01.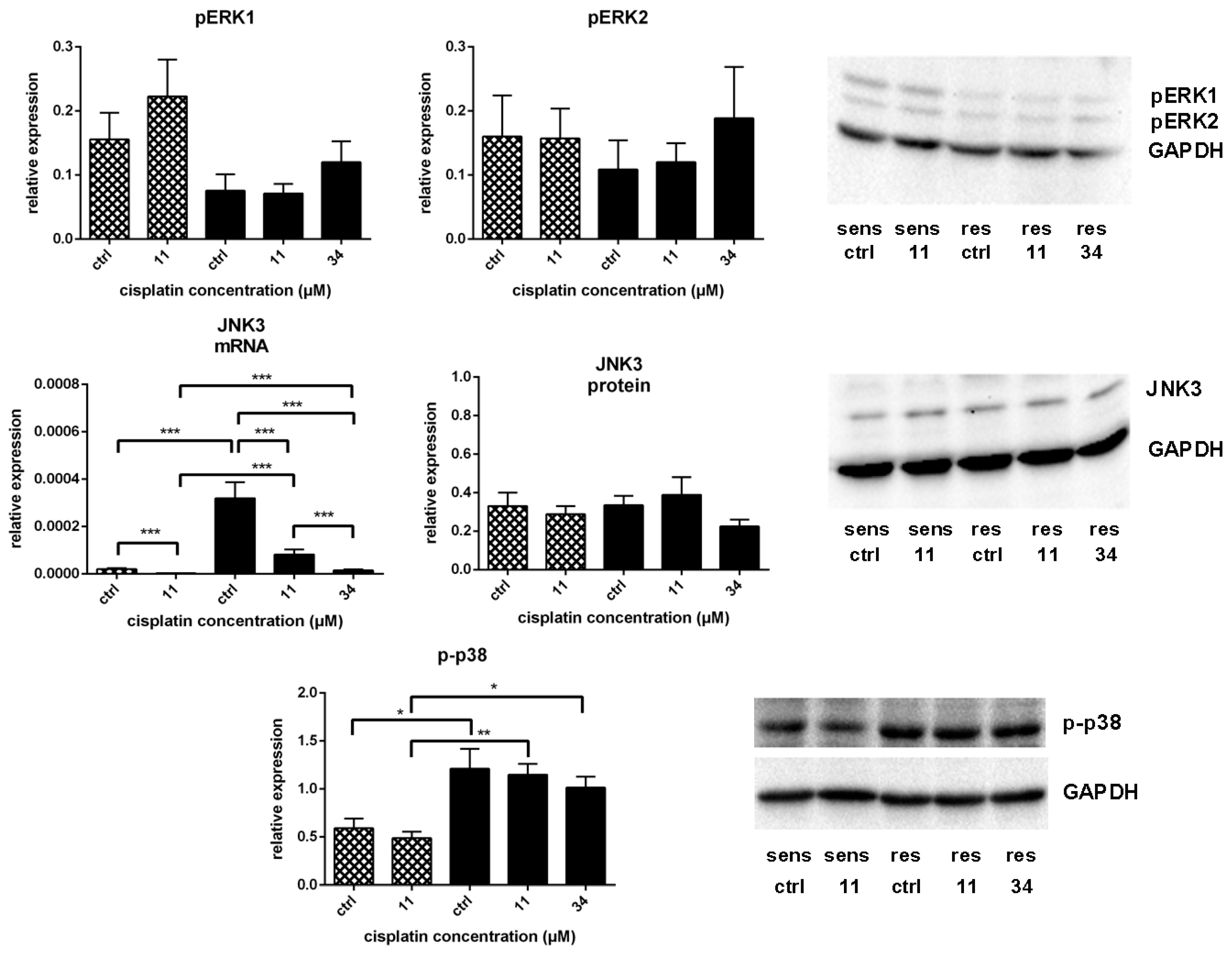

| Treatment Condition 1 | Treatment Condition 2 | Number of Differentially Expressed Genes |
|---|---|---|
| A549, untreated | A549rCDDP2000, untreated | 3697 |
| A549, 11 µM cisplatin | A549rCDDP2000, 11 µM cisplatin | 4394 |
| A549rCDDP2000, untreated | A549rCDDP2000, 11 µM cisplatin | 27 |
| A549rCDDP2000, untreated | A549rCDDP2000, 34 µM cisplatin | 708 |
| A549, untreated | A549, 11 µM cisplatin | 1191 |
| A549, 11 µM cisplatin | A549rCDDP2000, 34 µM cisplatin | 3670 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sarin, N.; Engel, F.; Rothweiler, F.; Cinatl, J.; Michaelis, M.; Frötschl, R.; Fröhlich, H.; Kalayda, G.V. Key Players of Cisplatin Resistance: Towards a Systems Pharmacology Approach. Int. J. Mol. Sci. 2018, 19, 767. https://doi.org/10.3390/ijms19030767
Sarin N, Engel F, Rothweiler F, Cinatl J, Michaelis M, Frötschl R, Fröhlich H, Kalayda GV. Key Players of Cisplatin Resistance: Towards a Systems Pharmacology Approach. International Journal of Molecular Sciences. 2018; 19(3):767. https://doi.org/10.3390/ijms19030767
Chicago/Turabian StyleSarin, Navin, Florian Engel, Florian Rothweiler, Jindrich Cinatl, Martin Michaelis, Roland Frötschl, Holger Fröhlich, and Ganna V. Kalayda. 2018. "Key Players of Cisplatin Resistance: Towards a Systems Pharmacology Approach" International Journal of Molecular Sciences 19, no. 3: 767. https://doi.org/10.3390/ijms19030767
APA StyleSarin, N., Engel, F., Rothweiler, F., Cinatl, J., Michaelis, M., Frötschl, R., Fröhlich, H., & Kalayda, G. V. (2018). Key Players of Cisplatin Resistance: Towards a Systems Pharmacology Approach. International Journal of Molecular Sciences, 19(3), 767. https://doi.org/10.3390/ijms19030767