Genome-Wide Analysis of the NF-YB Gene Family in Gossypium hirsutum L. and Characterization of the Role of GhDNF-YB22 in Embryogenesis
Abstract
1. Introduction
2. Results
2.1. Identification of NF-YB Genes in Cotton
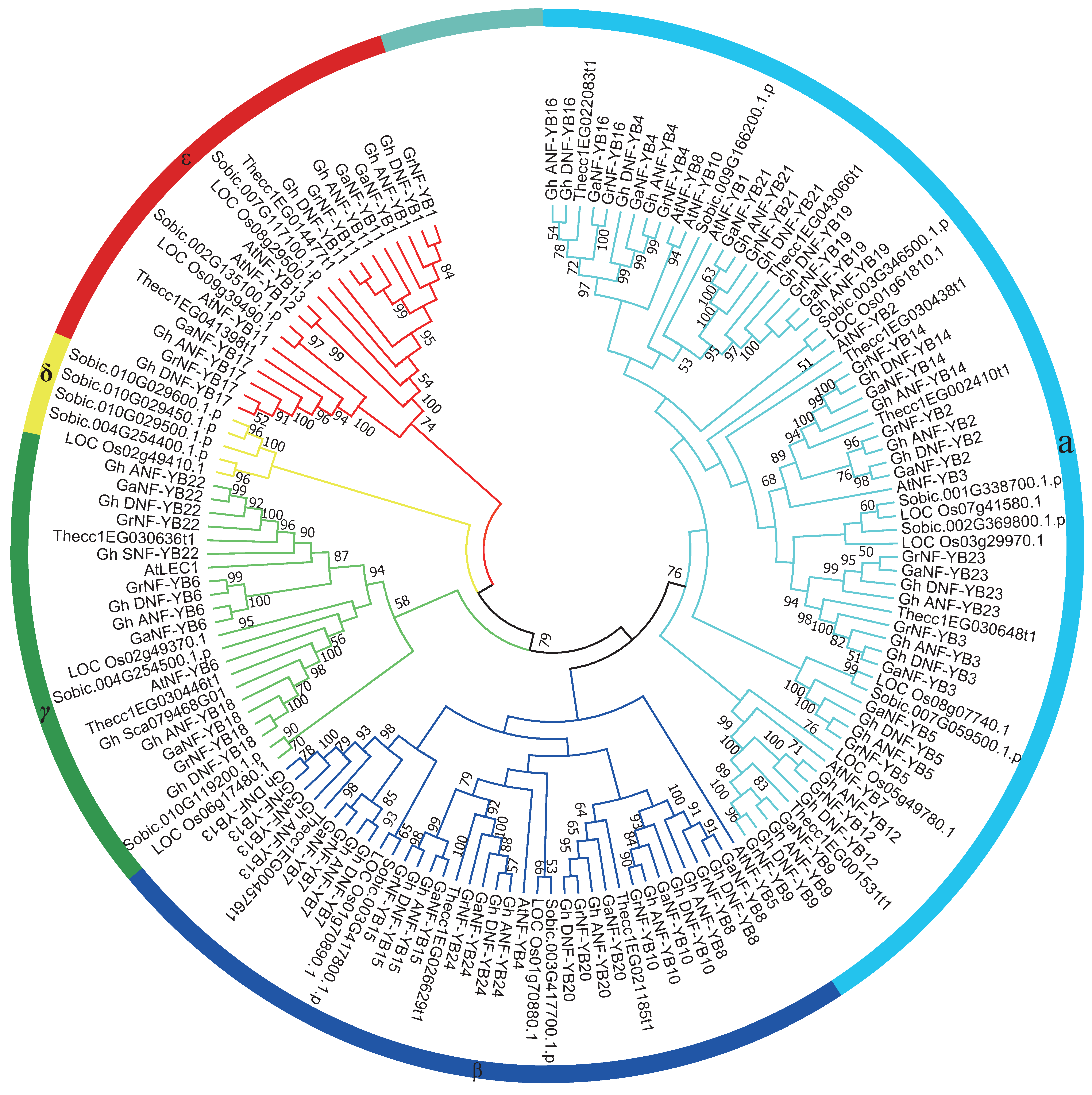
2.2. Phylogenetic Analysis of the NF-YB Gene Family
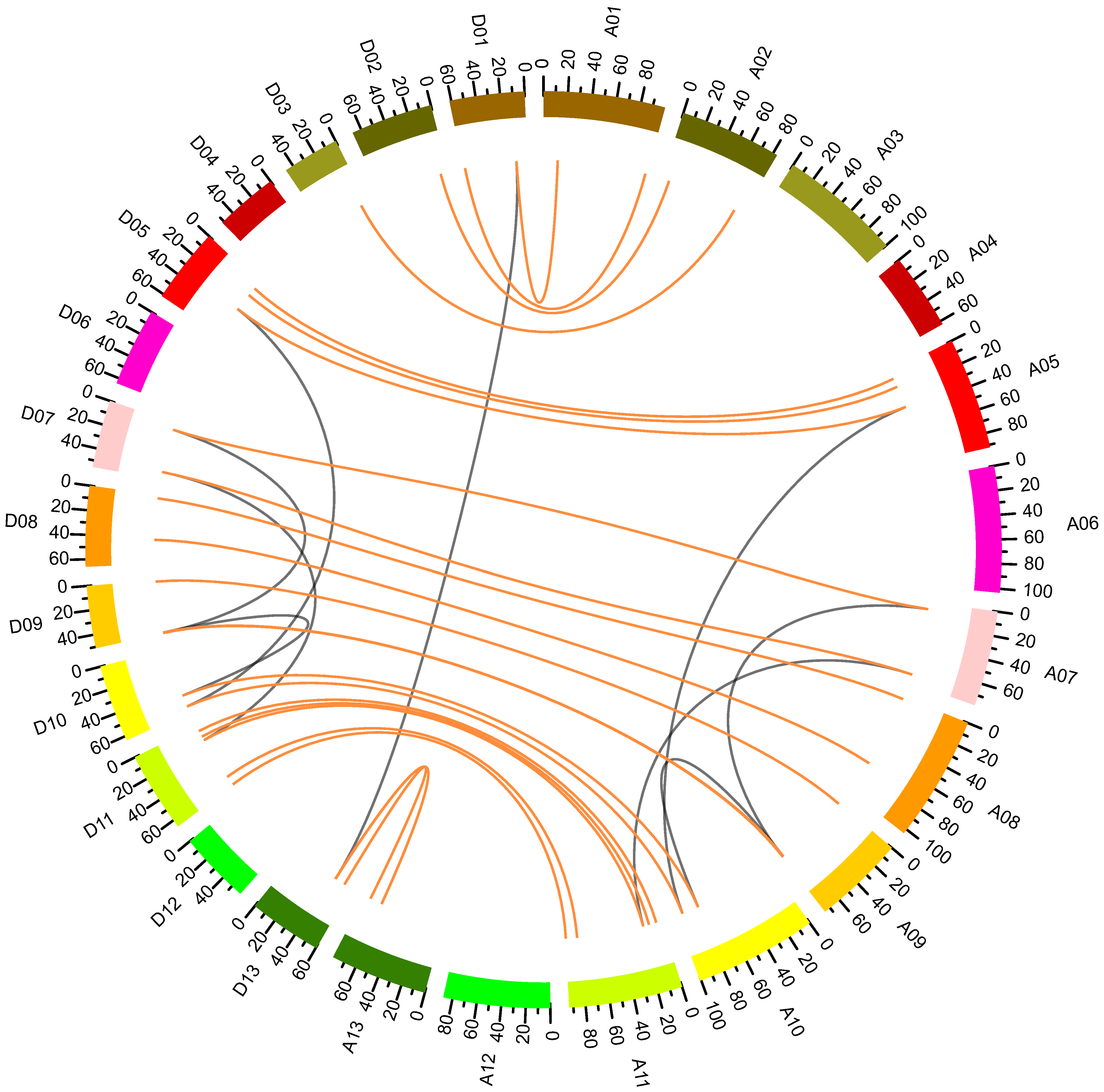
2.3. Chromosomal Distribution and Synteny Analysis of GhNF-YB Genes
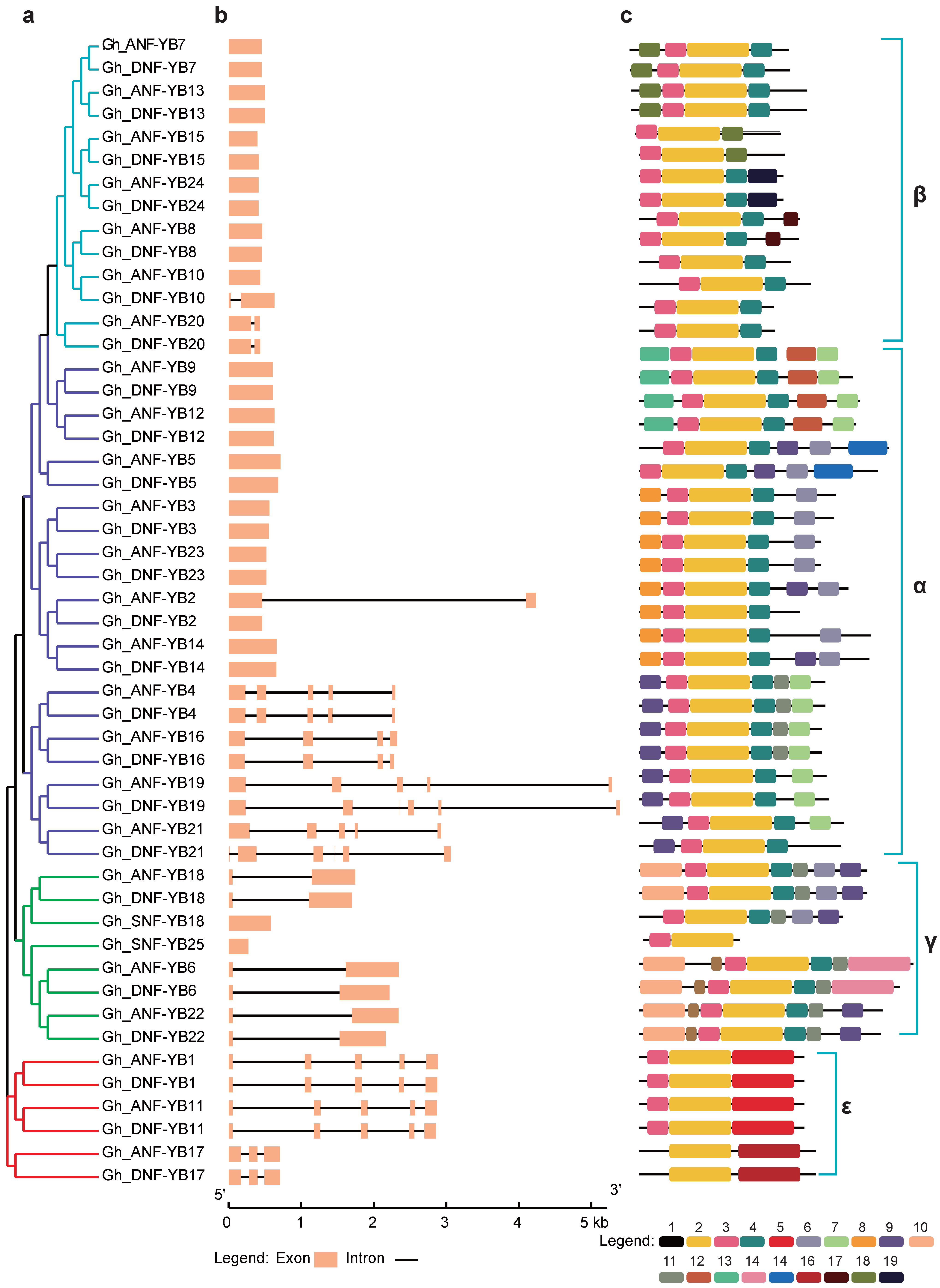
2.4. Gene Structure and Analysis of Conserved Motifs
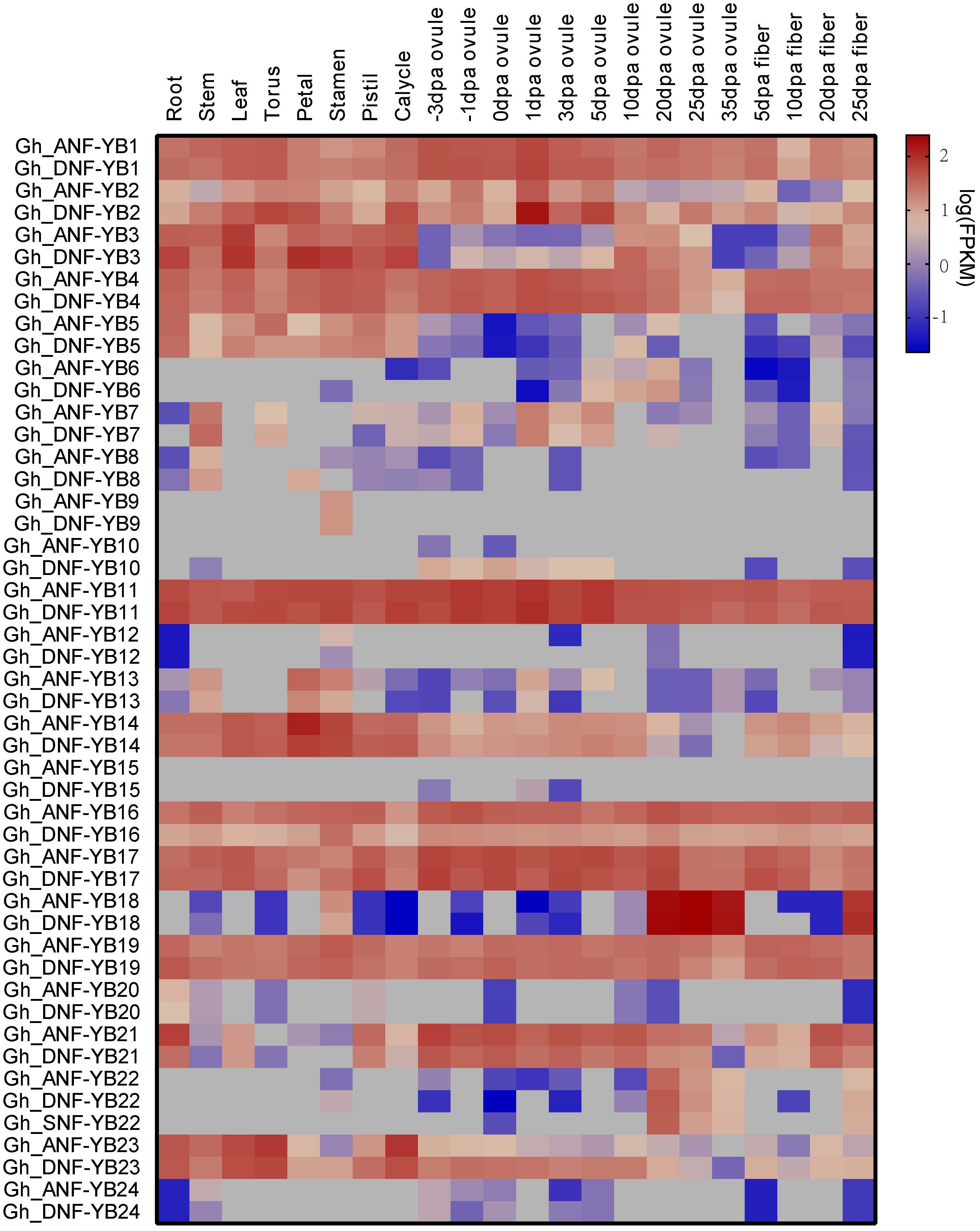
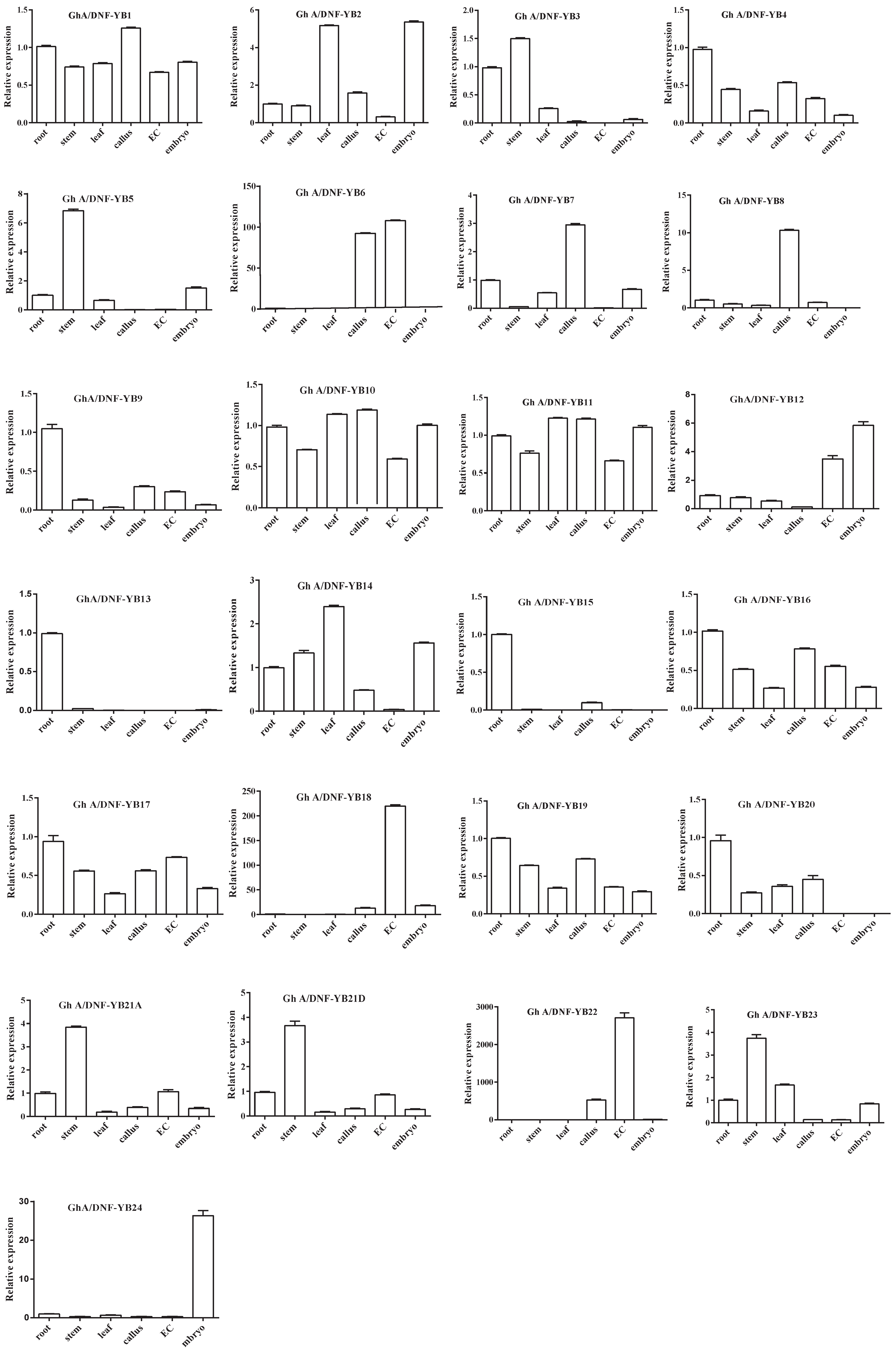
2.5. Analyses of Tissue-Specific Expression Patterns of 50 G. hirsutum NF-YB Genes
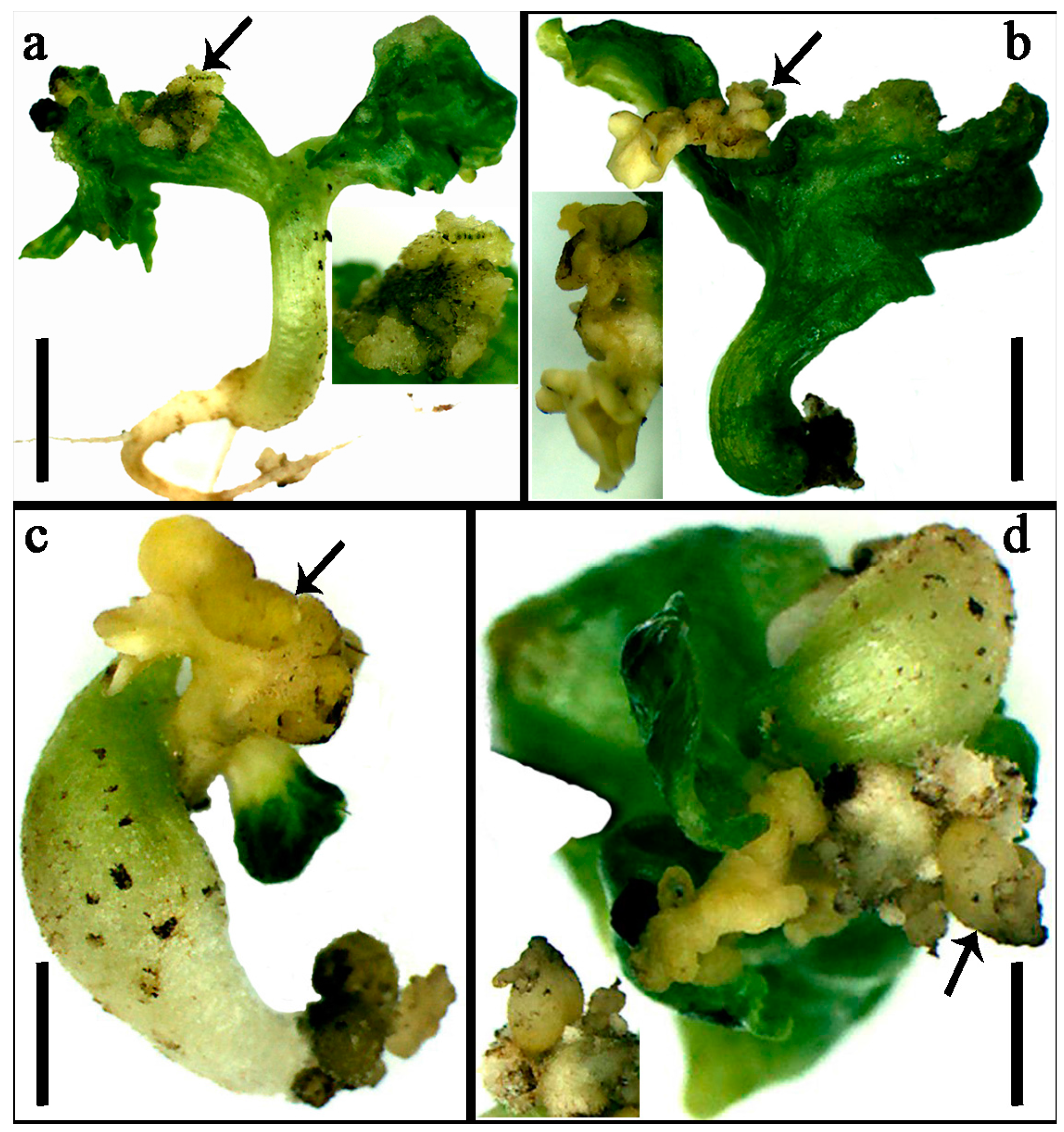
2.6. Overexpression of GhDNF-YB22 in Cotton Affects Embryogenesis
3. Discussion
3.1. Variation in the NF-YB Gene Family in G. hirsutum
3.2. Expression Patterns of NF-YB Genes in G. hirsutum
3.3. Role of GhDNF-YB22 in Embryogenesis
4. Materials and Methods
4.1. Identification of the NF-YB Gene Family
4.2. Phylogenetic Analyses
4.3. Chromosome Locations and Collinearity Analyses
4.4. Estimating Ka/Ks Rates
4.5. Analysis of Transposable Elements
4.6. Gene Structure and Conserved Motifs Analysis
4.7. Gene Expression Heat Map
4.8. RNA Isolation and qRT-PCR Verification
4.9. Gene Cloning and Transformation into Cotton
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Dorn, A.; Bollekens, J.; Staub, A.; Benoist, C.; Mathis, D. A multiplicity of CCAAT box-binding proteins. Cell 1987, 50, 863–872. [Google Scholar] [CrossRef]
- Romier, C.; Cocchiarella, F.; Mantovani, R.; Moras, D. The NF-YB/NF-YC structure gives insight into DNA binding and transcription regulation by CCAAT factor NF-Y. J. Biol. Chem. 2003, 278, 1336–1345. [Google Scholar] [CrossRef] [PubMed]
- Gusmaroli, G.; Tonelli, C.; Mantovani, R. Regulation of the CCAAT-Binding NF-Y subunits in Arabidopsis thaliana. Gene 2001, 264, 173–185. [Google Scholar] [CrossRef]
- Yang, S.; Wang, Y.; Yin, H.; Guo, H.; Gao, M.; Zhu, H.; Chen, Y. Identification and characterization of NF-YB family genes in tung tree. Mol. Genet. Genom. 2015, 290, 2187–2198. [Google Scholar] [CrossRef] [PubMed]
- Sinha, S.; Kim, I.S.; Sohn, K.Y.; DeCrombrugghe, B.; Maity, S.N. Three classes of mutations in the A subunit of the CCAAT-Binding factor CBF delineate functional domains involved in the three-step assembly of the CBF-DNA complex. Mol. Cell. Biol. 1996, 16, 328–337. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Lee, H.S.; Fischer, R.L.; Goldberg, R.B.; Harada, J.J. Arabidopsis LEAFY COTYLEDON1 represents a functionally specialized subunit of the CCAAT binding transcription factor. Proc. Natl. Acad. Sci. USA 2003, 100, 2152–2156. [Google Scholar] [CrossRef] [PubMed]
- Kwong, R.W.; Bui, A.Q.; Lee, H.; Kwong, L.W.; Fischer, R.L.; Goldberg, R.B.; Harada, J.J. LEAFY COTYLEDON1-like defines a class of regulators essential for embryo development. Plant Cell 2003, 15, 5–18. [Google Scholar] [CrossRef] [PubMed]
- Edwards, D.; Murray, J.A.H.; Smith, A.G. Multiple genes encoding the conserved CCAAT-box transcription factor complex are expressed in Arabidopsis. Plant Physiol. 1998, 117, 1015–1022. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Siefers, N.; Dang, K.K.; Kumimoto, R.W.; Bynum, W.E.; Tayrose, G.; Holt, B.F., III. Tissue-specific expression patterns of arabidopsis NF-Y transcription factors suggest potential for extensive combinatorial complexity. Plant Physiol. 2009, 149, 625–641. [Google Scholar] [CrossRef] [PubMed]
- Thirumurugan, T.; Ito, Y.; Kubo, T.; Serizawa, A.; Kurata, N. Identification, characterization and interaction of hap family genes in rice. Mol. Genet. Genom. 2008, 279, 279–289. [Google Scholar] [CrossRef] [PubMed]
- Stephenson, T.J.; McIntyre, C.L.; Collet, C.; Xue, G.-P. Genome-wide identification and expression analysis of the NF-Y family of transcription factors in triticum aestivum. Plant Mol. Biol. 2007, 65, 77–92. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Li, K.; Ju, Z.; Cao, D.Y.; Fu, D.Q.; Zhu, H.L.; Zhu, B.Z.; Luo, Y.B. Genome-wide analysis of tomato NF-Y factors and their role in fruit ripening. BMC Genom. 2016, 17, 36. [Google Scholar] [CrossRef] [PubMed]
- Liang, M.; Yin, X.; Lin, Z.; Zheng, Q.; Liu, G.; Zhao, G. Identification and characterization of NF-Y transcription factor families in canola (Brassica napus L.). Planta 2014, 239, 107–126. [Google Scholar] [CrossRef] [PubMed]
- Quach, T.; Nguyen, H.; Valliyodan, B.; Joshi, T.; Xu, D.; Nguyen, H. Genome-wide expression analysis of soybean NF-Y genes reveals potential function in development and drought response. Mol. Genet. Genom. 2015, 290, 1095–1115. [Google Scholar] [CrossRef] [PubMed]
- Ren, C.; Zhang, Z.; Wang, Y.; Li, S.H.; Liang, Z.C. Genome-wide identification and characterization of the NF-Y gene family in grape (Vitis vinifera L.). BMC Genom. 2016, 17, 605. [Google Scholar] [CrossRef] [PubMed]
- Mu, J.Y.; Tan, H.L.; Zheng, Q.; Fu, F.Y.; Liang, Y.; Zhang, J.A.; Yang, X.H.; Wang, T.; Chong, K.; Wang, X.J.; et al. LEAFY COTYLEDON1 is a key regulator of fatty acid biosynthesis in arabidopsis. Plant Physiol. 2008, 148, 1042–1054. [Google Scholar] [CrossRef] [PubMed]
- Rider, S.D.; Henderson, J.T.; Jerome, R.E.; Edenberg, H.J.; Romero-Severson, J.; Ogas, J. Coordinate repression of regulators of embryonic identity by pickle during germination in arabidopsis. Plant J. 2003, 35, 33–43. [Google Scholar] [CrossRef]
- Nelson, D.E.; Repetti, P.P.; Adams, T.R.; Creelman, R.A.; Wu, J.; Warner, D.C.; Anstrom, D.C.; Bensen, R.J.; Castiglioni, P.P.; Donnarummo, M.G.; et al. Plant nuclear factor Y (NF-Y) B subunits confer drought tolerance and lead to improved corn yields on water-limited acres. Proc. Natl. Acad. Sci. USA 2007, 104, 16450–16455. [Google Scholar] [CrossRef] [PubMed]
- Warpeha, K.M.; Upadhyay, S.; Yeh, J.; Adamiak, J.; Hawkins, S.I.; Lapik, Y.R.; Anderson, M.B.; Kaufman, L.S. The GCR1, GPA1, PRN1, Nf-Y signal chain mediates both blue light and abscisic acid responses in arabidopsis. Plant Physiol. 2007, 143, 1590–1600. [Google Scholar] [CrossRef] [PubMed]
- Kumimoto, R.W.; Adam, L.; Hymus, G.J.; Repetti, P.P.; Reuber, T.L.; Marion, C.M.; Hempel, F.D.; Ratcliffe, O.J. The nuclear factor Y subunits NF-YB2 and NF-YB3 play additive roles in the promotion of flowering by inductive long-day photoperiods in arabidopsis. Planta 2008, 228, 709–723. [Google Scholar] [CrossRef] [PubMed]
- Ballif, J.; Endo, S.; Kotani, M.; MacAdam, J.; Wu, Y. Over-expression of HAP3b enhances primary root elongation in Arabidopsis. Plant Physiol. Biochem. 2011, 49, 579–583. [Google Scholar] [CrossRef] [PubMed]
- Shen, B.; Allen, W.B.; Zheng, P.; Li, C.; Glassman, K.; Ranch, J.; Nubel, D.; Tarczynski, M.C. Expression of ZmLEC1 and ZmWRI1 increases seed oil production in maize. Plant Physiol. 2010, 153, 980–987. [Google Scholar] [CrossRef] [PubMed]
- Tan, H.; Yang, X.; Zhang, F.; Zheng, X.; Qu, C.; Mu, J.; Fu, F.; Li, J.; Guan, R.; Zhang, H.; et al. Enhanced seed oil production in canola by conditional expression of Brassica napus LEAFY COTYLEDON1 and LEC1-LIKE in developing seeds. Plant Physiol. 2011, 156, 1577–1588. [Google Scholar] [CrossRef] [PubMed]
- Miyoshi, K.; Ito, Y.; Serizawa, A.; Kurata, N. OsHAP3 genes regulate chloroplast biogenesis in rice. Plant J. 2003, 36, 532–540. [Google Scholar] [CrossRef] [PubMed]
- Yadav, D.; Shavrukov, Y.; Bazanova, N.; Chirkova, L.; Borisjuk, N.; Kovalchuk, N.; Ismagul, A.; Parent, B.; Langridge, P.; Hrmova, M.; et al. Constitutive overexpression of the TANF-YB4 gene in transgenic wheat significantly improves grain yield. J. Exp. Bot. 2015, 66, 6635–6650. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; Wang, Z.; Li, F.; Ye, W.; Wang, J.; Song, G.; Yue, Z.; Cong, L.; Shang, H.; Zhu, S.; et al. The draft genome of a diploid cotton Gossypium raimondii. Nat. Genet. 2012, 44, 1098. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Fan, G.; Wang, K.; Sun, F.; Yuan, Y.; Song, G.; Li, Q.; Ma, Z.; Lu, C.; Zou, C.; et al. Genome sequence of the cultivated cotton Gossypium arboreum. Nat. Genet. 2014, 46, 567–572. [Google Scholar] [CrossRef] [PubMed]
- Zhang, T.; Hu, Y.; Jiang, W.; Fang, L.; Guan, X.; Chen, J.; Zhang, J.; Saski, C.A.; Scheffler, B.E.; Stelly, D.M.; et al. Sequencing of allotetraploid cotton (Gossypium hirsutum L. acc. TM-1) provides a resource for fiber improvement. Nat. Biotechnol. 2015, 33, 531. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Fan, G.; Lu, C.; Xiao, G.; Zou, C.; Kohel, R.J.; Ma, Z.; Shang, H.; Ma, X.; Wu, J.; et al. Genome sequence of cultivated upland cotton (Gossypium hirsutum TM-1) provides insights into genome evolution. Nat. Biotechnol. 2015, 33, 524. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.; Gong, Q.; Qin, W.; Yang, Z.; Cheng, Y.; Lu, L.; Ge, X.; Zhang, C.; Wu, Z.; Li, F. Genome-wide analysis of wox genes in upland cotton and their expression pattern under different stresses. BMC Plant Biol. 2017, 17, 113. [Google Scholar] [CrossRef] [PubMed]
- Xu, G.; Guo, C.; Shan, H.; Kong, H. Divergence of duplicate genes in exon-intron structure. Proc. Natl. Acad. Sci. USA 2012, 109, 1187–1192. [Google Scholar] [CrossRef] [PubMed]
- Lynch, M.; Conery, J.S. The evolutionary fate and consequences of duplicate genes. Science 2000, 290, 1151–1155. [Google Scholar] [CrossRef] [PubMed]
- Parisod, C.; Alix, K.; Just, J.; Petit, M.; Sarilar, V.; Mhiri, C.; Ainouche, M.; Chalhoub, B.; Grandbastien, M.-A. Impact of transposable elements on the organization and function of allopolyploid genomes. New Phytol. 2010, 186, 37–45. [Google Scholar] [CrossRef] [PubMed]
- Grandbastien, M.; Audeon, C.; Bonnivard, E.; Casacuberta, J.M.; Chalhoub, B.; Costa, A.P.P.; Le, Q.H.; Melayah, D.; Petit, M.; Poncet, C.; et al. Stress activation and genomic impact of Tnt1 retrotransposons in solanaceae. Cytogenet. Genome Res. 2005, 110, 229–241. [Google Scholar] [CrossRef] [PubMed]
- Trapnell, C.; Roberts, A.; Goff, L.; Pertea, G.; Kim, D.; Kelley, D.R.; Pimentel, H.; Salzberg, S.L.; Rinn, J.L.; Pachter, L. Differential gene and transcript expression analysis of RNA-seq experiments with tophat and cufflinks. Nat. Protoc. 2012, 7, 562. [Google Scholar] [CrossRef] [PubMed]
- Lotan, T.; Ohto, M.; Yee, K.M.; West, M.A.L.; Lo, R.; Kwong, R.W.; Yamagishi, K.; Fischer, R.L.; Goldberg, R.B.; Harada, J.J. Arabidopsis LEAFY COTYLEDON1 is sufficient to induce embryo development in Vegetative cells. Cell 1998, 93, 1195–1205. [Google Scholar] [CrossRef]
- Paterson, A.H.; Bowers, J.E.; Chapman, B.A. Ancient polyploidization predating divergence of the cereals, and its consequences for comparative genomics. Proc. Natl. Acad. Sci. USA 2004, 101, 9903–9908. [Google Scholar] [CrossRef] [PubMed]
- Van de Peer, Y.; Fawcett, J.A.; Proost, S.; Sterck, L.; Vandepoele, K. The flowering world: A tale of duplications. Trends Plant Sci. 2009, 14, 680–688. [Google Scholar] [CrossRef] [PubMed]
- Jiao, Y.; Wickett, N.J.; Ayyampalayam, S.; Chanderbali, A.S.; Landherr, L.; Ralph, P.E.; Tomsho, L.P.; Hu, Y.; Liang, H.; Soltis, P.S.; et al. Ancestral polyploidy in seed plants and angiosperms. Nature 2011, 473, 97–100. [Google Scholar] [CrossRef] [PubMed]
- Scarpino, S.V.; Levin, D.A.; Meyers, L.A. Polyploid formation shapes flowering plant diversity. Am. Nat. 2014, 184, 456–465. [Google Scholar] [CrossRef] [PubMed]
- Soltis, D.E.; Burleigh, J.G. Surviving the K-T mass extinction: New perspectives of polyploidization in angiosperms. Proc. Natl. Acad. Sci. USA 2009, 106, 5455–5456. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Guo, H.; Wang, J.; Lei, T.; Liu, T.; Wang, Z.; Li, Y.; Lee, T.-H.; Li, J.; Tang, H.; et al. Comparative genomic de-convolution of the cotton genome revealed a decaploid ancestor and widespread chromosomal fractionation. New Phytol. 2016, 209, 1252–1263. [Google Scholar] [CrossRef] [PubMed]
- Xie, Z.Y.; Li, X.; Glover, B.J.; Bai, S.N.; Rao, G.Y.; Luo, J.C.; Yang, J. Duplication and functional diversification of HAP3 genes leading to the origin of the seed-developmental regulatory gene, LEAFY COTYLEDON1 (LEC1), in nonseed plant genomes. Mol. Biol. Evol. 2008, 25, 1581–1592. [Google Scholar] [CrossRef] [PubMed]
- Rogozin, I.B.; Wolf, Y.I.; Sorokin, A.V.; Mirkin, B.G.; Koonin, E.V. Remarkable interkingdom conservation of intron positions and massive, lineage-specific intron loss and gain in eukaryotic evolution. Curr. Biol. 2003, 13, 1512–1517. [Google Scholar] [CrossRef]
- Roy, S.W.; Penny, D. Patterns of intron loss and gain in plants: Intron loss-dominated evolution and genome-wide comparison of O-sativa and A-thaliana. Mol. Biol. Evol. 2007, 24, 171–181. [Google Scholar] [CrossRef] [PubMed]
- Zhao, H.; Wu, D.; Kong, F.; Lin, K.; Zhang, H.; Li, G. The arabidopsis thaliana nuclear factor Y transcription factors. Front. Plant Sci. 2017, 8, 2045. [Google Scholar] [CrossRef] [PubMed]
- Zimmermann, P.; Hirsch-Hoffmann, M.; Hennig, L.; Gruissem, W. GENEVESTIGATOR. Arabidopsis microarray database and analysis toolbox. Plant Physiol. 2004, 136, 2621–2632. [Google Scholar] [CrossRef] [PubMed]
- Cao, S.; Kumimoto, R.W.; Gnesutta, N.; Calogero, A.M.; Mantovani, R.; Holt, B.F. A distal ccaat nuclear factor Y complex promotes chromatin looping at the flowering locus T promoter and regulates the timing of flowering in arabidopsis. Plant Cell 2014, 26, 1009–1017. [Google Scholar] [CrossRef] [PubMed]
- Liang, M.; Hole, D.; Wu, J.; Blake, T.; Wu, Y. Expression and functional analysis of nuclear factor-Y, subunit B genes in barley. Planta 2012, 235, 779–791. [Google Scholar] [CrossRef] [PubMed]
- Huang, M.; Hu, Y.; Liu, X.; Li, Y.; Hou, X. Arabidopsis LEAFY COTYLEDON1 controls cell fate determination during post-embryonic development. Front. Plant Sci. 2015, 6, 955. [Google Scholar] [CrossRef] [PubMed]
- Meinke, D.W. A homoeotic mutant of arabidopsis thaliana with LEAFY COTYLEDONS. Science 1992, 258, 1647–1650. [Google Scholar] [CrossRef] [PubMed]
- Pelletier, J.M.; Kwong, R.W.; Park, S.; Le, B.H.; Baden, R.; Cagliari, A.; Hashimoto, M.; Munoz, M.D.; Fischer, R.L.; Goldberg, R.B.; et al. LEC1 sequentially regulates the transcription of genes involved in diverse developmental processes during seed development. Proc. Natl. Acad. Sci. USA 2017, 114, E6710–E6719. [Google Scholar] [CrossRef] [PubMed]
- Braybrook, S.A.; Harada, J.J. Lecs go crazy in embryo development. Trends Plant Sci. 2008, 13, 624–630. [Google Scholar] [CrossRef] [PubMed]
- Kakar, K.U.; Nawaz, Z.; Kakar, K.; Ali, E.; Almoneafy, A.A.; Ullah, R.; Ren, X.-L.; Shu, Q.-Y. Comprehensive genomic analysis of the CNGC gene family in brassica oleracea: Novel insights into synteny, structures, and transcript profiles. BMC Genom. 2017, 18, 869. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. Mega7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- McGinnis, S.; Madden, T.L. Blast: At the core of a powerful and diverse set of sequence analysis tools. Nucleic Acids Res. 2004, 32, W20–W25. [Google Scholar] [CrossRef] [PubMed]
- Krzywinski, M.; Schein, J.; Birol, İ.; Connors, J.; Gascoyne, R.; Horsman, D.; Jones, S.J.; Marra, M.A. Circos: An information aesthetic for comparative genomics. Genome Res. 2009, 19, 1639–1645. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z. Paml 4: Phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 2007, 24, 1586–1591. [Google Scholar] [CrossRef] [PubMed]
- Xu, Z.; Wang, H. LTR_FINDER: An efficient tool for the prediction of full-length LTR retrotransposons. Nucleic Acids Res. 2007, 35, W265–W268. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C.; Myers, E.W. PILER: Identification and classification of genomic repeats. Bioinformatics 2005, 21, i152–i158. [Google Scholar] [CrossRef] [PubMed]
- Sturn, A.; Quackenbush, J.; Trajanoski, Z. Genesis: Cluster analysis of microarray data. Bioinformatics 2002, 18, 207–208. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Yu, S.; Fan, S.; Zhang, J.; Li, F. Inheritance of somatic embryogenesis using leaf petioles as explants in upland cotton. Euphytica 2011, 181, 55–63. [Google Scholar] [CrossRef]
- Yang, Z.; Zhang, C.; Yang, X.; Liu, K.; Wu, Z.; Zhang, X.; Zheng, W.; Xun, Q.; Liu, C.; Lu, L.; et al. PAG1, a cotton brassinosteroid catabolism gene, modulates fiber elongation. New Phytol. 2014, 203, 437–448. [Google Scholar] [CrossRef] [PubMed]
- Ren, Z.; Yu, D.; Yang, Z.; Li, C.; Qanmber, G.; Li, Y.; Li, J.; Liu, Z.; Lu, L.; Wang, L.; et al. Genome-wide identification of the MIKC-type MADS-box gene family in Gossypium hirsutum L. Unravels their roles in flowering. Front. Plant Sci. 2017, 8, 384. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Ge, X.; Yang, Z.; Zhang, C.; Zhao, G.; Chen, E.; Liu, J.; Zhang, X.; Li, F. Genome-wide identification and characterization of SnRK2 gene family in cotton (Gossypium hirsutum L.). BMC Genet. 2017, 18, 54. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCt method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.; Li, C.; Wang, Y.; Zhang, C.; Wu, Z.; Zhang, X.; Liu, C.; Li, F. Ghagl15s, preferentially expressed during somatic embryogenesis, promote embryogenic callus formation in cotton (Gossypium hirsutum L.). Mol. Genet. Genom. 2014, 289, 873–883. [Google Scholar] [CrossRef] [PubMed]
- Shang, H.-H.; Liu, C.-L.; Zhang, C.-J.; Li, F.-L.; Hong, W.-D.; Li, F.-G. Histological and ultrastructural observation reveals significant cellular differences between agrobacterium transformed embryogenic and non-embryogenic calli of cotton. J. Integr. Plant Biol. 2009, 51, 456–465. [Google Scholar] [CrossRef] [PubMed]

| Homologous Pairs | Ka | Ks | Ka/Ks | |
|---|---|---|---|---|
| Gh_ANF-YB21 | Gh_ANF-YB19 | 0.071 | 0.376 | 0.189 |
| Gh_ANF-YB11 | Gh_ANF-YB1 | 0.025 | 0.563 | 0.044 |
| Gh_ANF-YB14 | Gh_ANF-YB2 | 0.061 | 0.621 | 0.098 |
| Gh_ANF-YB20 | Gh_ANF-YB10 | 0.203 | 1.113 | 0.182 |
| Gh_DNF-YB23 | Gh_DNF-YB3 | 0.067 | 0.486 | 0.139 |
| Gh_DNF-YB21 | Gh_DNF-YB19 | 0.183 | 0.474 | 0.386 |
| Gh_DNF-YB11 | Gh_DNF-YB1 | 0.025 | 0.481 | 0.052 |
| Gh_DNF-YB14 | Gh_DNF-YB2 | 0.083 | 0.500 | 0.165 |
| Gh_DNF-YB20 | Gh_DNF-YB10 | 0.245 | 0.846 | 0.289 |
| Type | Elements | Number of Elements | Length Occupied (bp) | Percentage of Sequence (%) | Number of Elements | Length Occupied (bp) | Percentage of Sequence (%) |
|---|---|---|---|---|---|---|---|
| 10,000 bp region | 2000 bp region | ||||||
| DNA transposons | 1 | 91 | 0.10 | 0 | 0 | 0 | |
| CMC-EnSpm | 0 | 0 | 0 | 0 | 0 | 0 | |
| MULE-MuDR | 0 | 0 | 0 | 0 | 0 | 0 | |
| PIF-Harbinger | 0 | 0 | 0 | 0 | 0 | 0 | |
| TcMar-Pogo | 0 | 0 | 0 | 0 | 0 | 0 | |
| hAT | 0 | 0 | 0 | 0 | 0 | 0 | |
| hAT-Ac | 1 | 91 | 0.10 | 0 | 0 | 0 | |
| hAT-Charlie | 0 | 0 | 0 | 0 | 0 | 0 | |
| hAT-Tag1 | 0 | 0 | 0 | 0 | 0 | 0 | |
| hAT-Tip100 | 0 | 0 | 0 | 0 | 0 | 0 | |
| Retroelements | 53 | 17,673 | 18.90 | 3 | 1038 | 7.43 | |
| LINE: | 10 | 2923 | 3.13 | 1 | 91 | 0.65 | |
| L1 | 10 | 2923 | 3.13 | 1 | 91 | 0.65 | |
| LTR: | 43 | 14,750 | 15.78 | 2 | 947 | 6.78 | |
| Caulimovirus | 0 | 0 | 0 | 0 | 0 | 0 | |
| Copia | 33 | 12,359 | 13.22 | 2 | 947 | 6.7 | |
| Gypsy | 10 | 2391 | 2.56 | 0 | 0 | 0 | |
| RC: | 0 | 0 | 0 | 0 | 0 | 0 | |
| Helitron | 0 | 0 | 0 | 0 | 0 | 0 | |
| DNA | 1 | 72 | 0.08 | 0 | 0 | 0 | |
| Low_complexity | 166 | 9514 | 10.18 | 47 | 2479 | 17.74 | |
| Simple_repeat | 586 | 25176 | 26.93 | 221 | 8121 | 58.12 | |
| Unspecified | 151 | 49452 | 52.90 | 16 | 3681 | 26.35 | |
| tRNA | 1 | 30 | 0.03 | 0 | 0 | 0 | |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chen, Y.; Yang, Z.; Xiao, Y.; Wang, P.; Wang, Y.; Ge, X.; Zhang, C.; Zhang, X.; Li, F. Genome-Wide Analysis of the NF-YB Gene Family in Gossypium hirsutum L. and Characterization of the Role of GhDNF-YB22 in Embryogenesis. Int. J. Mol. Sci. 2018, 19, 483. https://doi.org/10.3390/ijms19020483
Chen Y, Yang Z, Xiao Y, Wang P, Wang Y, Ge X, Zhang C, Zhang X, Li F. Genome-Wide Analysis of the NF-YB Gene Family in Gossypium hirsutum L. and Characterization of the Role of GhDNF-YB22 in Embryogenesis. International Journal of Molecular Sciences. 2018; 19(2):483. https://doi.org/10.3390/ijms19020483
Chicago/Turabian StyleChen, Yanli, Zhaoen Yang, Yanqing Xiao, Peng Wang, Ye Wang, Xiaoyang Ge, Chaojun Zhang, Xianlong Zhang, and Fuguang Li. 2018. "Genome-Wide Analysis of the NF-YB Gene Family in Gossypium hirsutum L. and Characterization of the Role of GhDNF-YB22 in Embryogenesis" International Journal of Molecular Sciences 19, no. 2: 483. https://doi.org/10.3390/ijms19020483
APA StyleChen, Y., Yang, Z., Xiao, Y., Wang, P., Wang, Y., Ge, X., Zhang, C., Zhang, X., & Li, F. (2018). Genome-Wide Analysis of the NF-YB Gene Family in Gossypium hirsutum L. and Characterization of the Role of GhDNF-YB22 in Embryogenesis. International Journal of Molecular Sciences, 19(2), 483. https://doi.org/10.3390/ijms19020483




