Small Molecules in Rare Tumors: Emerging Role of MicroRNAs in GIST
Abstract
1. Introduction
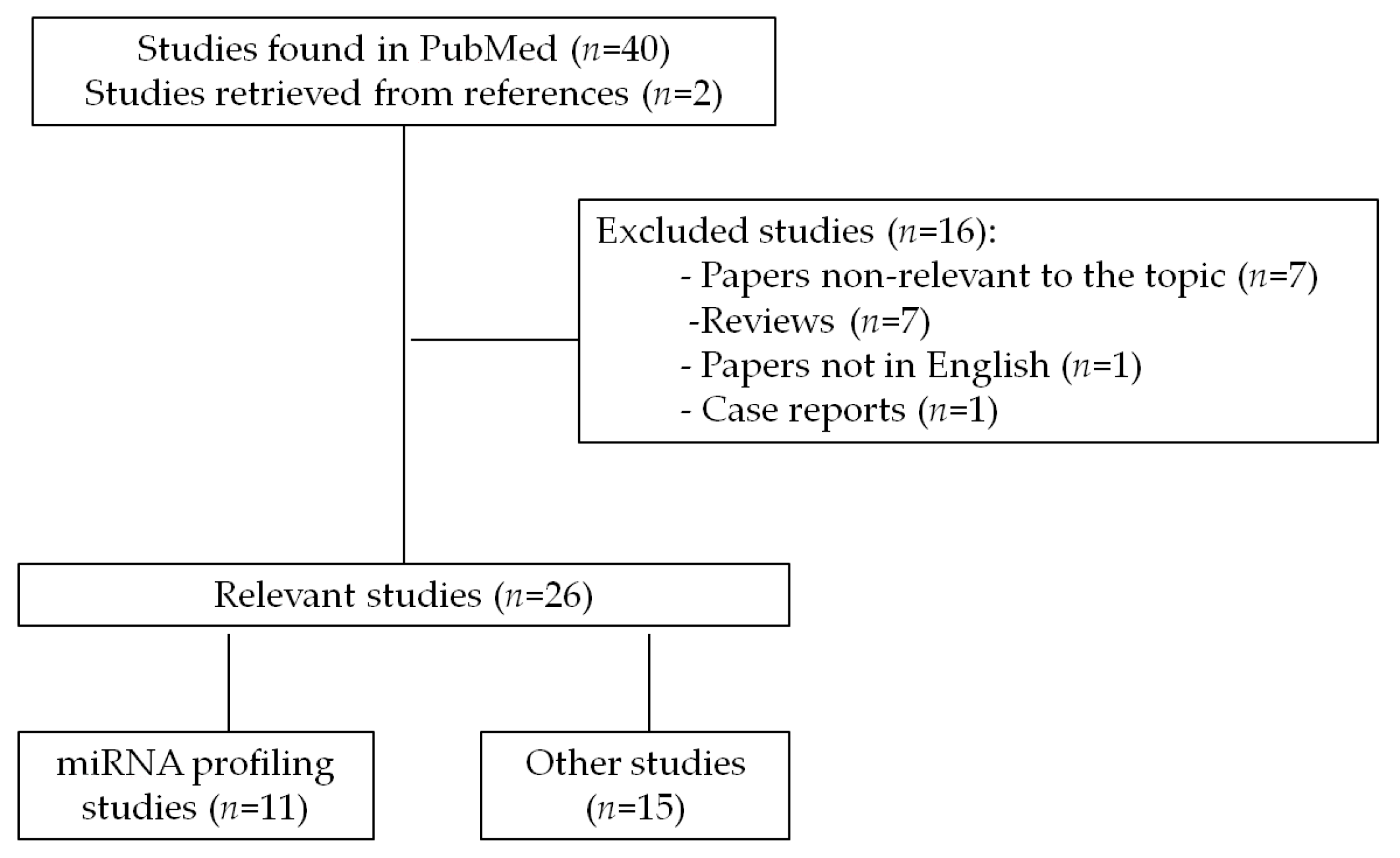
2. Methods
3. miRNA Expression Profiles in GIST
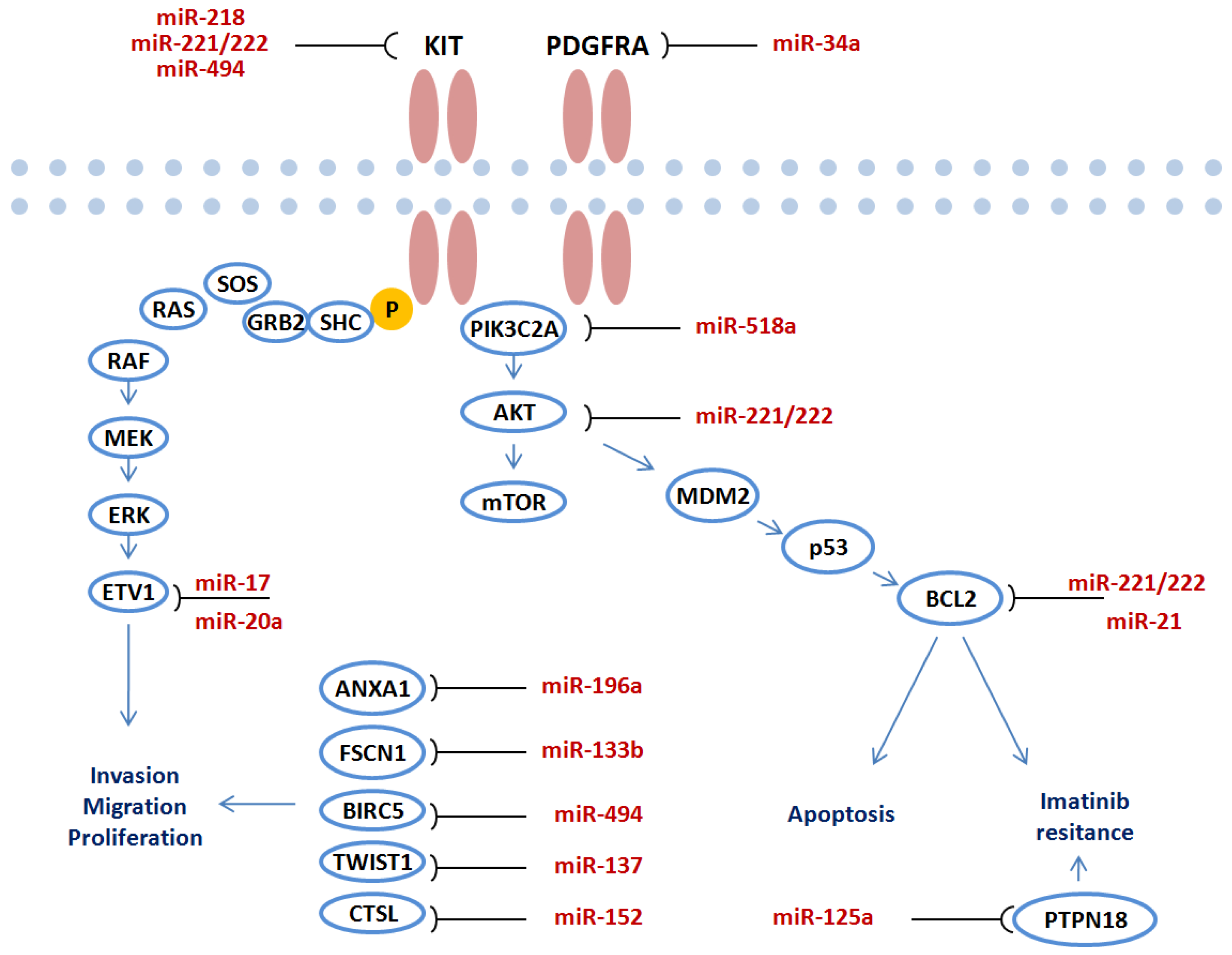
4. Functional Role of miRNAs in GIST
4.1. The Functional Role of miR-221/miR-222
4.2. The Functional Role of miR-494
4.3. The Functional Role of Other miRNAs
4.4. Proposed Functional Role of miRNAs by Bioinformatical Analyses
5. Potential Role of miRNA in GIST Treatment
6. Prognostic Role of miRNAs in GIST
7. Future Perspectives
7.1. Identification of isomiR Profiles and Their Functional Role in GIST
7.2. Circulating miRNAs in Blood of GIST Patients
7.3. Single Cell Profiles of miRNAs in GIST
8. Conclusions
Conflicts of Interest
Abbreviations
| ANXA1 | annexin A1 |
| BCL-2 | B-cell lymphoma 2 |
| BIRC5 | baculoviral IAP repeat-containing protein 5 |
| CTSL | cathepsin L |
| ERK | extracellular signal-regulated kinase |
| ETV1 | ets variant 1 |
| FSCN-1 | fascin actin-bundling protein 1 |
| GIST | gastrointestinal stromal tumor |
| GRB2 | growth factor receptor bound protein 2 |
| isomiRs | isoforms of miRNAs |
| KIT | tyrosine kinase family |
| MDM2 | murine double minute 2 |
| microRNA | miRNA |
| MEK | mitogen-activated protein kinase |
| p53 | tumor protein p53 |
| PDGFRA | platelet derived growth factor |
| PIK3C2A | phosphatidylinositol-4-phosphate 3-kinase C2 domain-containing alpha polypeptide |
| PTPN18 | protein tyrosine phosphatase non-receptor type 18 |
| SDH | succinate dehydrogenase |
| SHC | squalene-hopene cyclase |
| SOS | son of sevenless gene |
| TWIST1 | twist family bHLH transcription factor 1 |
| WT | wild-type |
References
- Hirota, S.; Isozaki, K.; Moriyama, Y.; Hashimoto, K.; Nishida, T.; Ishiguro, S.; Kawano, K.; Hanada, M.; Kurata, A.; Takeda, M.; et al. Gain-of-function mutations of c-kit in human gastrointestinal stromal tumors. Science 1998, 279, 577–580. [Google Scholar] [CrossRef] [PubMed]
- Heinrich, M.C.; Corless, C.L.; Duensing, A.; McGreevey, L.; Chen, C.-J.; Joseph, N.; Singer, S.; Griffith, D.J.; Haley, A.; Town, A.; et al. PDGFRA Activating Mutations in Gastrointestinal Stromal Tumors. Science 2003, 299, 708–710. [Google Scholar] [CrossRef] [PubMed]
- Capelli, L.; Petracci, E.; Quagliuolo, V.; Saragoni, L.; Colombo, P.; Morgagni, P.; Calistri, D.; Tomezzoli, A.; Di Cosmo, M.; Roviello, F.; et al. Italian Gastric Cancer Research Group (GIRCG) Gastric GISTs: Analysis of c-Kit, PDGFRA and BRAF mutations in relation to prognosis and clinical pathological characteristics of patients—A GIRCG study. Eur. J. Surg. Oncol. 2016, 42, 1206–1214. [Google Scholar] [CrossRef] [PubMed]
- Miettinen, M.; Sobin, L.H.; Lasota, J. Gastrointestinal stromal tumors of the stomach: A clinicopathologic, immunohistochemical, and molecular genetic study of 1765 cases with long-term follow-up. Am. J. Surg. Pathol. 2005, 29, 52–68. [Google Scholar] [CrossRef] [PubMed]
- Min, K.W.; Leabu, M. Interstitial cells of Cajal (ICC) and gastrointestinal stromal tumor (GIST): Facts, speculations, and myths. J. Cell. Mol. Med. 2006, 10, 995–1013. [Google Scholar] [CrossRef] [PubMed]
- Yacob, M.; Inian, S.; Sudhakar, C.B. Gastrointestinal Stromal Tumours: Review of 150 Cases from a Single Centre. Indian J. Surg. 2015, 77, 505–510. [Google Scholar] [CrossRef] [PubMed]
- Antonescu, C.R.; Viale, A.; Sarran, L.; Tschernyavsky, S.J.; Gonen, M.; Segal, N.H.; Maki, R.G.; Socci, N.D.; DeMatteo, R.P.; Besmer, P. Gene Expression in Gastrointestinal Stromal Tumors Is Distinguished by KIT Genotype and Anatomic Site. Clin. Cancer Res. 2004, 10, 3282–3290. [Google Scholar] [CrossRef] [PubMed]
- Ravegnini, G.; Nannini, M.; Sammarini, G.; Astolfi, A.; Biasco, G.; Pantaleo, M.; Hrelia, P.; Angelini, S. Personalized Medicine in Gastrointestinal Stromal Tumor (GIST): Clinical Implications of the Somatic and Germline DNA Analysis. Int. J. Mol. Sci. 2015, 16, 15592–15608. [Google Scholar] [CrossRef] [PubMed]
- Shah, K.; Chan, K.K.W.; Ko, Y.J. A systematic review and network meta-analysis of post-imatinib therapy in advanced gastrointestinal stromal tumour. Curr. Oncol. 2017, 24, e531–e539. [Google Scholar] [CrossRef] [PubMed]
- Lujambio, A.; Lowe, S.W. The microcosmos of cancer. Nature 2012, 482, 347–355. [Google Scholar] [CrossRef] [PubMed]
- Steponaitiene, R.; Kupcinskas, J.; Langner, C.; Balaguer, F.; Venclauskas, L.; Pauzas, H.; Tamelis, A.; Skieceviciene, J.; Kupcinskas, L.; Malfertheiner, P.; et al. Epigenetic silencing of miR-137 is a frequent event in gastric carcinogenesis. Mol. Carcinog. 2016, 55, 376–386. [Google Scholar] [CrossRef] [PubMed]
- Link, A.; Kupcinskas, J.; Wex, T.; Malfertheiner, P. Macro-role of MicroRNA in gastric cancer. Dig. Dis. 2012, 30, 255–267. [Google Scholar] [CrossRef] [PubMed]
- Balaguer, F.; Link, A.; Lozano, J.J.; Cuatrecasas, M.; Nagasaka, T.; Boland, C.R.; Goel, A. Epigenetic Silencing of miR-137 Is an Early Event in Colorectal Carcinogenesis. Cancer Res. 2010, 70, 6609–6618. [Google Scholar] [CrossRef] [PubMed]
- Rizzato, C.; Campa, D.; Talar-Wojnarowska, R.; Halloran, C.; Kupcinskas, J.; Butturini, G.; Mohelníková-Duchoňová, B.; Sperti, C.; Tjaden, C.; Ghaneh, P.; et al. Association of genetic polymorphisms with survival of pancreatic ductal adenocarcinoma patients. Carcinogenesis 2016, 37, 957–964. [Google Scholar] [CrossRef] [PubMed]
- Catanzaro, G.; Sabato, C.; Russo, M.; Rosa, A.; Abballe, L.; Besharat, Z.; Po, A.; Miele, E.; Bellavia, D.; Chiacchiarini, M.; et al. Loss of miR-107, miR-181c and miR-29a-3p Promote Activation of Notch2 Signaling in Pediatric High-Grade Gliomas (pHGGs). Int. J. Mol. Sci. 2017, 18, 2742. [Google Scholar] [CrossRef] [PubMed]
- Link, A.; Goel, A. MicroRNA in gastrointestinal cancer: A step closer to reality. Adv. Clin. Chem. 2013, 62, 221–268. [Google Scholar] [PubMed]
- Laneve, P.; Di Marcotullio, L.; Gioia, U.; Fiori, M.E.; Ferretti, E.; Gulino, A.; Bozzoni, I.; Caffarelli, E. The interplay between microRNAs and the neurotrophin receptor tropomyosin-related kinase C controls proliferation of human neuroblastoma cells. Proc. Natl. Acad. Sci. USA 2007, 104, 7957–7962. [Google Scholar] [CrossRef] [PubMed]
- Juzenas, S.; Salteniene, V.; Kupcinskas, J.; Link, A.; Kiudelis, G.; Jonaitis, L.; Jarmalaite, S.; Kupcinskas, L.; Malfertheiner, P.; Skieceviciene, J. Analysis of deregulated micrornas and their target genes in gastric cancer. PLoS ONE 2015, 10, e0135762. [Google Scholar] [CrossRef] [PubMed]
- Yanaihara, N.; Caplen, N.; Bowman, E.; Seike, M.; Kumamoto, K.; Yi, M.; Stephens, R.M.; Okamoto, A.; Yokota, J.; Tanaka, T.; et al. Unique microRNA molecular profiles in lung cancer diagnosis and prognosis. Cancer Cell 2006, 9, 189–198. [Google Scholar] [CrossRef] [PubMed]
- Imig, J.; Motsch, N.; Zhu, J.Y.; Barth, S.; Okoniewski, M.; Reineke, T.; Tinguely, M.; Faggioni, A.; Trivedi, P.; Meister, G.; et al. microRNA profiling in Epstein—Barr virus-associated B-cell lymphoma. Nucleic Acids Res. 2011, 39, 1880–1893. [Google Scholar] [CrossRef] [PubMed]
- Nannini, M.; Ravegnini, G.; Angelini, S.; Astolfi, A.; Biasco, G.; Pantaleo, M.A. miRNA profiling in gastrointestinal stromal tumors: Implication as diagnostic and prognostic markers. Epigenomics 2015, 7, 1033–1049. [Google Scholar] [CrossRef] [PubMed]
- Kupcinskas, J.; Wex, T.; Link, A.; Leja, M.; Bruzaite, I.; Steponaitiene, R.; Juzenas, S.; Gyvyte, U.; Ivanauskas, A.; Ancans, G.; et al. Gene Polymorphisms of Micrornas in Helicobacter pylori-Induced High Risk Atrophic Gastritis and Gastric Cancer. PLoS ONE 2014, 9, e87467. [Google Scholar] [CrossRef] [PubMed]
- Kupcinskas, J.; Bruzaite, I.; Juzenas, S.; Gyvyte, U.; Jonaitis, L.; Kiudelis, G.; Skieceviciene, J.; Leja, M.; Pauzas, H.; Tamelis, A.; et al. Lack of association between miR-27a, miR-146a, miR-196a-2, miR-492 and miR-608 gene polymorphisms and colorectal cancer. Sci. Rep. 2014, 4, 5993. [Google Scholar] [CrossRef] [PubMed]
- Petkevicius, V.; Salteniene, V.; Juzenas, S.; Wex, T.; Link, A.; Leja, M.; Steponaitiene, R.; Skieceviciene, J.; Kupcinskas, L.; Jonaitis, L.; et al. Polymorphisms of microRNA target genes IL12B, INSR, CCND1 and IL10 in gastric cancer. World J. Gastroenterol. 2017, 23, 3556–3564. [Google Scholar] [CrossRef] [PubMed]
- Ding, L.; Xu, Y.; Zhang, W.; Deng, Y.; Si, M.; Du, Y.; Yao, H.; Liu, X.; Ke, Y.; Si, J.; et al. MiR-375 frequently downregulated in gastric cancer inhibits cell proliferation by targeting JAK2. Cell Res. 2010, 20, 784–793. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Hou, J.; Jin, W.; Li, J.; Yue, Y.; Jin, H.; Wang, X. Increased Circulating MicroRNA-155 as a Potential Biomarker for Breast Cancer Screening: A Meta-Analysis. Molecules 2014, 19, 6282–6293. [Google Scholar] [CrossRef] [PubMed]
- Yamakuchi, M.; Lotterman, C.D.; Bao, C.; Hruban, R.H.; Karim, B.; Mendell, J.T.; Huso, D.; Lowenstein, C.J. P53-induced microRNA-107 inhibits HIF-1 and tumor angiogenesis. Proc. Natl. Acad. Sci. USA 2010, 107, 6334–6339. [Google Scholar] [CrossRef] [PubMed]
- Simonson, B.; Das, S. MicroRNA Therapeutics: The Next Magic Bullet? Mini Rev. Med. Chem. 2015, 15, 467–474. [Google Scholar] [CrossRef] [PubMed]
- Ji, W.; Sun, B.; Su, C. Targeting MicroRNAs in Cancer Gene Therapy. Genes 2017, 8, 21. [Google Scholar] [CrossRef] [PubMed]
- Subramanian, S.; Lui, W.O.; Lee, C.H.; Espinosa, I.; Nielsen, T.O.; Heinrich, M.C.; Corless, C.L.; Fire, A.Z.; van de Rijn, M. MicroRNA expression signature of human sarcomas. Oncogene 2008, 27, 2015–2026. [Google Scholar] [CrossRef] [PubMed]
- Choi, H.-J.; Lee, H.; Kim, H.; Kwon, J.E.; Kang, H.J.; You, K.T.; Rhee, H.; Noh, S.H.; Paik, Y.-K.; Hyung, W.J.; et al. MicroRNA expression profile of gastrointestinal stromal tumors is distinguished by 14q loss and anatomic site. Int. J. Cancer 2010, 126, 1640–1650. [Google Scholar] [CrossRef] [PubMed]
- Haller, F.; Zhang, D.J.; Löbke, C.; von Heydebreck, A.; Korf, U.; Füzesi, L.; Sahin, Ö. Multilayer-Analyse der Signaltransduktion und Zellzykluskontrolle in GIST. Pathologe 2010, 31, 134–137. [Google Scholar] [CrossRef] [PubMed]
- Kelly, L.; Bryan, K.; Kim, S.Y.; Janeway, K.A.; Killian, J.K.; Schildhaus, H.-U.; Miettinen, M.; Helman, L.; Meltzer, P.S.; van de Rijn, M.; et al. NIH Pediatric and Wild-Type GIST Clinic Post-transcriptional dysregulation by miRNAs is implicated in the pathogenesis of gastrointestinal stromal tumor [GIST]. PLoS ONE 2013, 8, e64102. [Google Scholar] [CrossRef] [PubMed]
- Niinuma, T.; Suzuki, H.; Nojima, M.; Nosho, K.; Yamamoto, H.; Takamaru, H.; Yamamoto, E.; Maruyama, R.; Nobuoka, T.; Miyazaki, Y.; et al. Upregulation of miR-196a and HOTAIR drive malignant character in gastrointestinal stromal tumors. Cancer Res. 2012, 72, 1126–1136. [Google Scholar] [CrossRef] [PubMed]
- Akçakaya, P.; Caramuta, S.; Åhlen, J.; Ghaderi, M.; Berglund, E.; Östman, A.; Bränström, R.; Larsson, C.; Lui, W.-O. microRNA expression signatures of gastrointestinal stromal tumours: Associations with imatinib resistance and patient outcome. Br. J. Cancer 2014, 111, 2091–2102. [Google Scholar] [CrossRef] [PubMed]
- Tong, H.-X.; Zhou, Y.-H.; Hou, Y.-Y.; Zhang, Y.; Huang, Y.; Xie, B.; Wang, J.-Y.; Jiang, Q.; He, J.-Y.; Shao, Y.-B.; et al. Expression profile of microRNAs in gastrointestinal stromal tumors revealed by high throughput quantitative RT-PCR microarray. World J. Gastroenterol. 2015, 21, 5843–5855. [Google Scholar] [CrossRef] [PubMed]
- Isosaka, M.; Niinuma, T.; Nojima, M.; Kai, M.; Yamamoto, E.; Maruyama, R.; Nobuoka, T.; Nishida, T.; Kanda, T.; Taguchi, T.; et al. A Screen for Epigenetically Silenced microRNA Genes in Gastrointestinal Stromal Tumors. PLoS ONE 2015, 10, e0133754. [Google Scholar] [CrossRef] [PubMed]
- Fan, R.; Zhong, J.; Zheng, S.; Wang, Z.; Xu, Y.; Li, S.; Zhou, J.; Yuan, F. MicroRNA-218 inhibits gastrointestinal stromal tumor cell and invasion by targeting KIT. Tumour Biol. 2014, 35, 4209–4217. [Google Scholar] [CrossRef] [PubMed]
- Ihle, M.A.; Trautmann, M.; Kuenstlinger, H.; Huss, S.; Heydt, C.; Fassunke, J.; Wardelmann, E.; Bauer, S.; Schildhaus, H.-U.; Buettner, R.; et al. miRNA-221 and miRNA-222 induce apoptosis via the KIT/AKT signalling pathway in gastrointestinal stromal tumours. Mol. Oncol. 2015, 9, 1421–1433. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, H.; Kohashi, K.; Fujita, A.; Oda, Y. Fascin-1 overexpression and miR-133b downregulation in the progression of gastrointestinal stromal tumor. Mod. Pathol. 2013, 26, 563–571. [Google Scholar] [CrossRef] [PubMed]
- Niinuma, T.; Kai, M.; Kitajima, H.; Yamamoto, E.; Harada, T.; Maruyama, R.; Nobuoka, T.; Nishida, T.; Kanda, T.; Hasegawa, T.; et al. Downregulation of miR-186 is associated with metastatic recurrence of gastrointestinal stromal tumors. Oncol. Lett. 2017, 14, 5703–5710. [Google Scholar] [CrossRef] [PubMed]
- Fan, R.; Zhong, J.; Zheng, S.; Wang, Z.; Xu, Y.; Li, S.; Zhou, J.; Yuan, F. microRNA-218 increase the sensitivity of gastrointestinal stromal tumor to imatinib through PI3K/AKT pathway. Clin. Exp. Med. 2015, 15, 137–144. [Google Scholar] [CrossRef] [PubMed]
- Shi, Y.; Gao, X.; Hu, Q.; Li, X.; Xu, J.; Lu, S.; Liu, Y.; Xu, C.; Jiang, D.; Lin, J.; et al. PIK3C2A is a gene-specific target of microRNA-518a-5p in imatinib mesylate-resistant gastrointestinal stromal tumor. Lab. Investig. 2016, 96, 652–660. [Google Scholar] [CrossRef] [PubMed]
- Durso, M.; Gaglione, M.; Piras, L.; Mercurio, M.E.; Terreri, S.; Olivieri, M.; Marinelli, L.; Novellino, E.; Incoronato, M.; Grieco, P.; et al. Chemical modifications in the seed region of miRNAs 221/222 increase the silencing performances in gastrointestinal stromal tumor cells. Eur. J. Med. Chem. 2016, 111, 15–25. [Google Scholar] [CrossRef] [PubMed]
- Pantaleo, M.A.; Ravegnini, G.; Astolfi, A.; Simeon, V.; Nannini, M.; Saponara, M.; Urbini, M.; Gatto, L.; Indio, V.; Sammarini, G.; et al. Integrating miRNA and gene expression profiling analysis revealed regulatory networks in gastrointestinal stromal tumors. Epigenomics 2016, 8, 1347–1366. [Google Scholar] [CrossRef] [PubMed]
- Bachet, J.-B.; Tabone-Eglinger, S.; Dessaux, S.; Besse, A.; Brahimi-Adouane, S.; Emile, J.-F.; Blay, J.-Y.; Alberti, L. Gene expression patterns of hemizygous and heterozygous KIT mutations suggest distinct oncogenic pathways: A study in NIH3T3 cell lines and GIST samples. PLoS ONE 2013, 8, e61103. [Google Scholar] [CrossRef] [PubMed]
- Gyvyte, U.; Juzenas, S.; Salteniene, V.; Kupcinskas, J.; Poskiene, L.; Kucinskas, L.; Jarmalaite, S.; Stuopelyte, K.; Steponaitiene, R.; Hemmrich-Stanisak, G.; et al. MiRNA profiling of gastrointestinal stromal tumors by next-generation sequencing. Oncotarget 2017, 8, 37225–37238. [Google Scholar] [CrossRef] [PubMed]
- Gits, C.M.M.; van Kuijk, P.F.; Jonkers, M.B.E.; Boersma, A.W.M.; van Ijcken, W.F.; Wozniak, A.; Sciot, R.; Rutkowski, P.; Schöffski, P.; Taguchi, T.; et al. MiR-17-92 and miR-221/222 cluster members target KIT and ETV1 in human gastrointestinal stromal tumours. Br. J. Cancer 2013, 109, 1625–1635. [Google Scholar] [CrossRef] [PubMed]
- Bornschein, J.; Leja, M.; Kupcinskas, J.; Link, A.; Weaver, J.; Rugge, M.; Malfertheiner, P. Molecular diagnostics in gastric cancer. Front. Biosci. 2014, 19, 312–338. [Google Scholar] [CrossRef]
- Cao, C.-L.; Niu, H.-J.; Kang, S.-P.; Cong, C.-L.; Kang, S.-R. miRNA-21 sensitizes gastrointestinal stromal tumors (GISTs) cells to Imatinib via targeting B-cell lymphoma 2 (Bcl-2). Eur. Rev. Med. Pharmacol. Sci. 2016, 20, 3574–3581. [Google Scholar] [PubMed]
- Liu, S.; Cui, J.; Liao, G.; Zhang, Y.; Ye, K.; Lu, T.; Qi, J.; Wan, G. MiR-137 regulates epithelial-mesenchymal transition in gastrointestinal stromal tumor. Tumour Biol. 2014, 35, 9131–9138. [Google Scholar] [CrossRef] [PubMed]
- Lu, H.-J.; Yan, J.; Jin, P.-Y.; Zheng, G.-H.; Qin, S.-M.; Wu, D.-M.; Lu, J.; Zheng, Y.-L. MicroRNA-152 inhibits tumor cell growth while inducing apoptosis via the transcriptional repression of cathepsin L in gastrointestinal stromal tumor. Cancer Biomark. 2017, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Koelz, M.; Lense, J.; Wrba, F.; Scheffler, M.; Dienes, H.P.; Odenthal, M. Down-regulation of miR-221 and miR-222 correlates with pronounced Kit expression in gastrointestinal stromal tumors. Int. J. Oncol. 2011, 38, 503–511. [Google Scholar] [CrossRef] [PubMed]
- Yun, S.; Kim, W.K.; Kwon, Y.; Jang, M.; Bauer, S.; Kim, H. Survivin Is a Novel Transcription Regulator of KIT and Is Downregulated by miRNA-494 in Gastrointestinal Stromal Tumors. Int. J. Cancer 2017. [Google Scholar] [CrossRef] [PubMed]
- Kim, W.K.; Park, M.; Kim, Y.-K.; Tae, Y.K.; Yang, H.-K.; Lee, J.M.; Kim, H. MicroRNA-494 downregulates KIT and inhibits gastrointestinal stromal tumor cell proliferation. Clin. Cancer Res. 2011, 17, 7584–7594. [Google Scholar] [CrossRef] [PubMed]
- Garofalo, M.; Quintavalle, C.; Romano, G.; Croce, C.M.; Condorelli, G. miR221/222 in cancer: Their role in tumor progression and response to therapy. Curr. Mol. Med. 2012, 12, 27–33. [Google Scholar] [CrossRef] [PubMed]
- Liu, K.; Liu, S.; Zhang, W.; Jia, B.; Tan, L.; Jin, Z.; Liu, Y. miR-494 promotes cell proliferation, migration and invasion, and increased sorafenib resistance in hepatocellular carcinoma by targeting PTEN. Oncol. Rep. 2015, 34, 1003–1010. [Google Scholar] [CrossRef] [PubMed]
- Li, N.; Zhao, X.; Wang, L.; Zhang, S.; Cui, M.; He, J. miR-494 suppresses tumor growth of epithelial ovarian carcinoma by targeting IGF1R. Tumor Biol. 2016, 37, 7767–7776. [Google Scholar] [CrossRef] [PubMed]
- Romano, G.; Acunzo, M.; Garofalo, M.; Di Leva, G.; Cascione, L.; Zanca, C.; Bolon, B.; Condorelli, G.; Croce, C.M. MiR-494 is regulated by ERK1/2 and modulates TRAIL-induced apoptosis in non-small-cell lung cancer through BIM down-regulation. Proc. Natl. Acad. Sci. USA 2012, 109, 16570–16575. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Li, J.; Qin, F.; Dai, S. miR-152 as a tumor suppressor microRNA: Target recognition and regulation in cancer (Review). Oncol. Lett. 2016, 11, 3911–3916. [Google Scholar] [CrossRef] [PubMed]
- Lacombe, J.; Zenhausern, F. Emergence of miR-34a in radiation therapy. Crit. Rev. Oncol. Hematol. 2017, 109, 69–78. [Google Scholar] [CrossRef] [PubMed]
- Zhang, F.; Wang, D. The Pattern of microRNA Binding Site Distribution. Genes 2017, 8, 296. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Chen, Z.; Chen, H.; Hou, Y.; Lu, W.; He, J.; Tong, H.; Zhou, Y.; Cai, W. Genetic Polymorphisms Contribute to the Individual Variations of Imatinib Mesylate Plasma Levels and Adverse Reactions in Chinese GIST Patients. Int. J. Mol. Sci. 2017, 18, 603. [Google Scholar] [CrossRef] [PubMed]
- Anastasiadou, E.; Garg, N.; Bigi, R.; Yadav, S.; Campese, A.F.; Lapenta, C.; Spada, M.; Cuomo, L.; Botta, A.; Belardelli, F.; et al. Epstein-Barr virus infection induces miR-21 in terminally differentiated malignant B cells. Int. J. Cancer 2015, 137, 1491–1497. [Google Scholar] [CrossRef] [PubMed]
- Tu, L.; Wang, M.; Zhao, W.-Y.; Zhang, Z.-Z.; Tang, D.-F.; Zhang, Y.-Q.; Cao, H.; Zhang, Z.-G. miRNA-218-loaded carboxymethyl chitosan-Tocopherol nanoparticle to suppress the proliferation of gastrointestinal stromal tumor growth. Mater. Sci. Eng. C Mater. Biol. Appl. 2017, 72, 177–184. [Google Scholar] [CrossRef] [PubMed]
- Bradshaw, G.; Sutherland, H.G.; Haupt, L.M.; Griffiths, L.R. Dysregulated MicroRNA Expression Profiles and Potential Cellular, Circulating and Polymorphic Biomarkers in Non-Hodgkin Lymphoma. Genes 2016, 7, 130. [Google Scholar] [CrossRef] [PubMed]
- D’Anzeo, M.; Faloppi, L.; Scartozzi, M.; Giampieri, R.; Bianconi, M.; Del Prete, M.; Silvestris, N.; Cascinu, S. The role of Micro-RNAs in Hepatocellular Carcinoma: From Molecular Biology to Treatment. Molecules 2014, 19, 6393–6406. [Google Scholar] [CrossRef] [PubMed]
- Verschoor, A.J.; Bovée, J.V.M.G.; Overbeek, L.I.H.; Hogendoorn, P.C.W.; Gelderblom, H.; Gelderblom, H. The incidence, mutational status, risk classification and referral pattern of gastro-intestinal stromal tumours in the Netherlands: A nationwide pathology registry (PALGA) study. Virchows Arch. 2018. [Google Scholar] [CrossRef] [PubMed]
- Cloonan, N.; Wani, S.; Xu, Q.; Gu, J.; Lea, K.; Heater, S.; Barbacioru, C.; Steptoe, A.L.; Martin, H.C.; Nourbakhsh, E.; et al. MicroRNAs and their isomiRs function cooperatively to target common biological pathways. Genome Biol. 2011, 12, R126. [Google Scholar] [CrossRef] [PubMed]
- Ameres, S.L.; Horwich, M.D.; Hung, J.-H.; Xu, J.; Ghildiyal, M.; Weng, Z.; Zamore, P.D. Target RNA-Directed Trimming and Tailing of Small Silencing RNAs. Science 2010, 328, 1534–1539. [Google Scholar] [CrossRef] [PubMed]
- Telonis, A.G.; Rigoutsos, I. Race disparities in the contribution of miRNA isoforms and tRNA-derived fragments to triple-negative breast cancer. Cancer Res. 2017. [Google Scholar] [CrossRef] [PubMed]
- Li, S.-C.; Liao, Y.-L.; Ho, M.-R.; Tsai, K.-W.; Lai, C.-H.; Lin, W. miRNA arm selection and isomiR distribution in gastric cancer. BMC Genom. 2012, 13, S13. [Google Scholar] [CrossRef] [PubMed]
- Babapoor, S.; Fleming, E.; Wu, R.; Dadras, S.S. A novel miR-451a isomiR, associated with amelanotypic phenotype, acts as a tumor suppressor in melanoma by retarding cell migration and invasion. PLoS ONE 2014, 9, e107502. [Google Scholar] [CrossRef] [PubMed]
- Yu, F.; Pillman, K.A.; Neilsen, C.T.; Toubia, J.; Lawrence, D.M.; Tsykin, A.; Gantier, M.P.; Callen, D.F.; Goodall, G.J.; Bracken, C.P. Naturally existing isoforms of miR-222 have distinct functions. Nucleic Acids Res. 2017, 45, 11371–11385. [Google Scholar] [CrossRef] [PubMed]
- Mitchell, P.S.; Parkin, R.K.; Kroh, E.M.; Fritz, B.R.; Wyman, S.K.; Pogosova-Agadjanyan, E.L.; Peterson, A.; Noteboom, J.; O’Briant, K.C.; Allen, A.; et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc. Natl. Acad. Sci. USA 2008, 105, 10513–10518. [Google Scholar] [CrossRef] [PubMed]
- ElSharawy, A.; Röder, C.; Becker, T.; Habermann, J.K.; Schreiber, S.; Rosenstiel, P.; Kalthoff, H. Concentration of circulating miRNA-containing particles in serum enhances miRNA detection and reflects CRC tissue-related deregulations. Oncotarget 2016, 7, 75353–75365. [Google Scholar] [CrossRef] [PubMed]
- Alemar, B.; Izetti, P.; Gregório, C.; Macedo, G.S.; Castro, M.A.A.; Osvaldt, A.B.; Matte, U.; Ashton-Prolla, P. miRNA-21 and miRNA-34a Are Potential Minimally Invasive Biomarkers for the Diagnosis of Pancreatic Ductal Adenocarcinoma. Pancreas 2016, 45, 84–92. [Google Scholar] [CrossRef] [PubMed]
- Jiang, X.; Wang, W.; Yang, Y.; Du, L.; Yang, X.; Wang, L.; Zheng, G.; Duan, W.; Wang, R.; Zhang, X.; et al. Identification of circulating microRNA signatures as potential noninvasive biomarkers for prediction and prognosis of lymph node metastasis in gastric cancer. Oncotarget 2017, 8, 65132–65142. [Google Scholar] [CrossRef] [PubMed]
- Uotani, K.; Fujiwara, T.; Yoshida, A.; Iwata, S.; Morita, T.; Kiyono, M.; Yokoo, S.; Kunisada, T.; Takeda, K.; Hasei, J.; et al. Circulating MicroRNA-92b-3p as a Novel Biomarker for Monitoring of Synovial Sarcoma. Sci. Rep. 2017, 7, 14634. [Google Scholar] [CrossRef] [PubMed]
- Porichis, F.; Hart, M.G.; Griesbeck, M.; Everett, H.L.; Hassan, M.; Baxter, A.E.; Lindqvist, M.; Miller, S.M.; Soghoian, D.Z.; Kavanagh, D.G.; et al. High-throughput detection of miRNAs and gene-specific mRNA at the single-cell level by flow cytometry. Nat. Commun. 2014, 5, 5641. [Google Scholar] [CrossRef] [PubMed]
- Wu, M.; Piccini, M.E.; Singh, A.K. miRNA Detection at Single-Cell Resolution Using Microfluidic LNA Flow-FISH. In Methods in Molecular Biology; Springer: Clifton, NJ, USA, 2014; Volume 1211, pp. 245–260. [Google Scholar]
- Juzenas, S.; Venkatesh, G.; Hübenthal, M.; Hoeppner, M.P.; Du, Z.G.; Paulsen, M.; Rosenstiel, P.; Senger, P.; Hofmann-Apitius, M.; Keller, A.; et al. A comprehensive, cell specific microRNA catalogue of human peripheral blood. Nucleic Acids Res. 2017, 45, 9290–9301. [Google Scholar] [CrossRef] [PubMed]
| Study | miRNA Profiling Method | Number of Investigated miRNAs | Compared Groups (n) | Number of Upregulated miRNAs | Number of Down-Regulated miRNAs |
|---|---|---|---|---|---|
| Subramanian et al., 2008 [30] | microarray | 328 | GIST (n, 8) vs. sarcomas (n, 25) | 15 | 9 |
| Choi et al., 2010 [31] | microarray | 470 | Gastric (n, 15) vs. intestinal (n, 5) | 20 | 4 |
| Haller et al., 2010 [32] | microarray | 734 | Gastric vs. intestinal (total n, 12) | 4 | 7 |
| Gits et al., 2013 [48] | microarray | 725 | GIST (n, 50) vs. leiomyosarcomas (n, 10) | 7 | 14 |
| Yamamoto et al., 2013 [40] | microarray | 904 | High risk (n, 10) vs. low risk (n, 4) | - | 24 |
| Kelly et al., 2013 [33] | RT-PCR | 667 | pediatric (n, 18) vs. adult GIST (n, 55) | 30 | 10 |
| Bachet et al., 2013 [46] | microarray | 384 | Cell lines (WT, D6, D54, WT/D6 and WT/D54); GIST samples WT (n, 3), PDGFRA mutations (n, 6) or KIT mutations (n, 11) | Different clustering of miRNAs based on mutation profile | |
| Akçakaya et al., 2014 [35] | microaaray | 903 | metastatic (n, 20) vs. non-metastatic (n, 10) | 19 | 25 |
| Tong et al., 2015 [36] | microarray | 849 | benign (n, 9) vs. malignant (n, 30) | 3 | 1 |
| Pantaleo et al., 2016 [45] | microarray | 723 | KIT/PDGFRA mutant (n, 4) vs. KIT/PDGFRA WT-SDH deficient GIST (n, 9) | 16 | 40 |
| Gyvyte et al., 2017 [47] | NGS | 1672 | GIST (n, 15) vs. GISTadjacent tissue (n, 15) | 34 | 66 |
| miRNA | Deregulation Pattern | Target Genes | Biological Effect | Reference |
|---|---|---|---|---|
| miR-17 | downregulated | ETV1 | inhibited cell proliferation, induced apoptosis | Gits et al., 2013 [48] |
| miR-20a | downregulated | ETV1 | inhibited cell proliferation, induced apoptosis | Gits et al., 2013 [48] |
| miR-21 | downregulated | BCL2 | Aggravate the Imatinib-mediated growth inhibition and apoptosis | Cao et al., 2016 [50] |
| miR-34a | downregulated | PDGFRA | Suppressed cell proliferation | Isosaka et al., 2015 [37] |
| miR-125a | upregulated | PTPN18 | Imatinib resistance | Akcakaya et al., 2016 [35] |
| miR-133b | downregulated | FSCN1 | Enhanced proliferation | Yamamoto et al., 2013 [40] |
| miR-137 | downregulated | TWIST1 | inhibits cell migration | Liu et al., 2014 [51] |
| miR-152 | downregulated | CTSL | inhibit proliferation, migration, invasion | Lu et al., 2017 [52] |
| miR-196a | upregulated | ANXA1 | Invasion | Niinuma et al., 2012 [34] |
| miR-218 | downregulated | KIT | inhibit proliferation and invasion | Fan et al., 2014 [38] |
| miR-221/222 | downregulated | KIT, AKT, BCL2 | inhibited cell proliferation, induced apoptosis | Ihle MA et al., 2015 [39] Koeltz M et al., 2011 [39] Gits et al., 2013 [53] |
| miR-494 | downregulated | BIRC5, KIT | Suppressed proliferation; promote apoptosis and inhibite cell growth | Yun S et al., 2017 [54] Kim WK et al., 2011 [55] |
| miR-518a | downregulated | PIK3C2A | reduce proliferation and promote apoptosis | Shi Y et al., 2016 [43] |
© 2018 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kupcinskas, J. Small Molecules in Rare Tumors: Emerging Role of MicroRNAs in GIST. Int. J. Mol. Sci. 2018, 19, 397. https://doi.org/10.3390/ijms19020397
Kupcinskas J. Small Molecules in Rare Tumors: Emerging Role of MicroRNAs in GIST. International Journal of Molecular Sciences. 2018; 19(2):397. https://doi.org/10.3390/ijms19020397
Chicago/Turabian StyleKupcinskas, Juozas. 2018. "Small Molecules in Rare Tumors: Emerging Role of MicroRNAs in GIST" International Journal of Molecular Sciences 19, no. 2: 397. https://doi.org/10.3390/ijms19020397
APA StyleKupcinskas, J. (2018). Small Molecules in Rare Tumors: Emerging Role of MicroRNAs in GIST. International Journal of Molecular Sciences, 19(2), 397. https://doi.org/10.3390/ijms19020397





