Causal Relationship between Diet-Induced Gut Microbiota Changes and Diabetes: A Novel Strategy to Transplant Faecalibacterium prausnitzii in Preventing Diabetes
Abstract
1. Introduction
2. Interactions between the Gut Microbiota and Dietary Nutrients
3. Role of Gut Microbiota in Diabetes
4. Multi-Skilled Commensal bacterium F. prausnitzii: A Diagnostic and Therapeutic Biomarker for Gut-Associated Diseases
5. Dietary Interventions Modulate F. prausnitzii
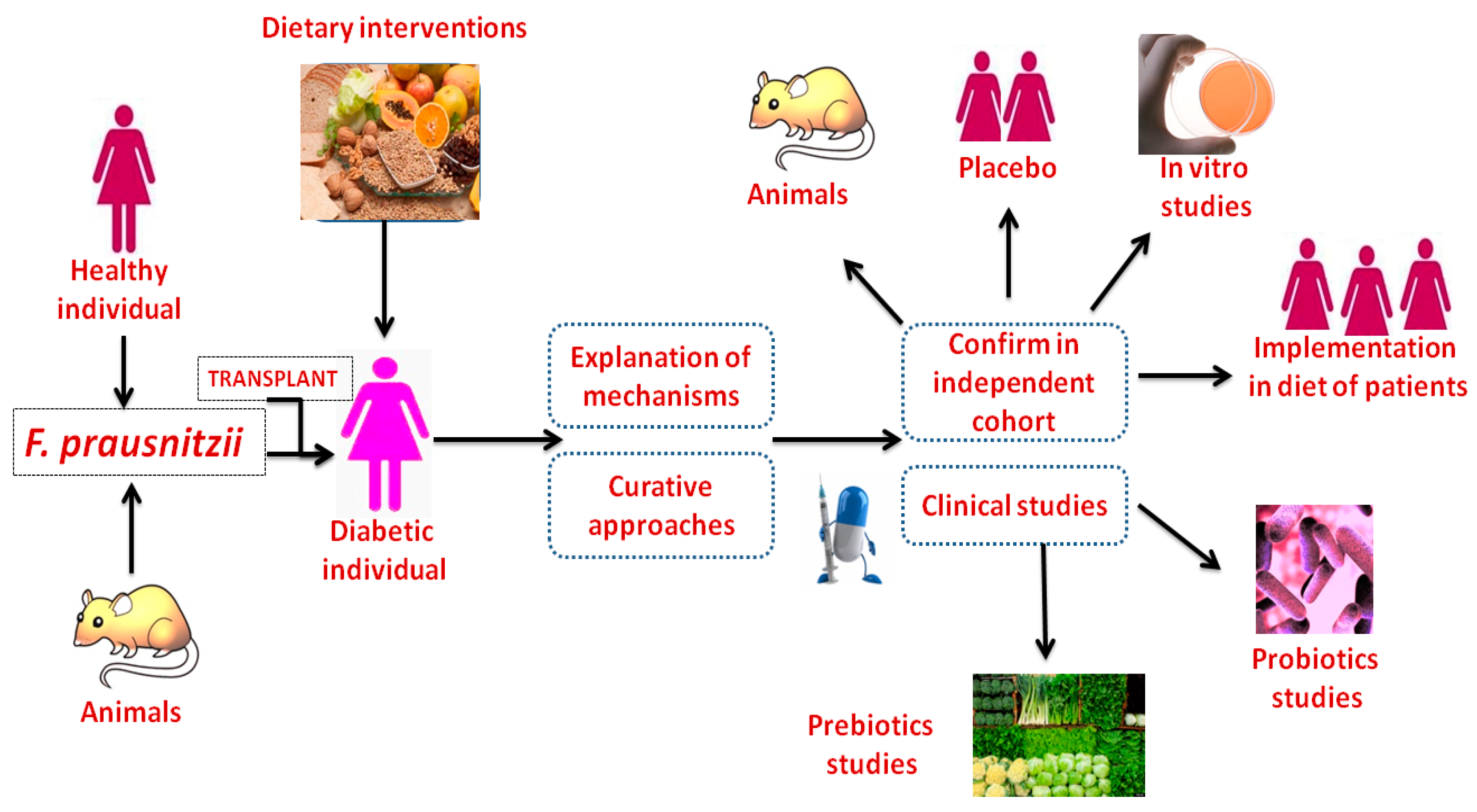
6. F. prausnitzii Transplantation Improves Diabetes
7. Conclusions
Author Contributions
Acknowledgments
Conflicts of Interest
Abbreviation
| AP1 | Activator protein 1 |
| ATP | Adenosine monophosphate |
| BUT | Butyryl-CoA:acetate CoA-transferase |
| cAMP | Cyclic adenosine monophosphate |
| CD14 | Cluster of differentiation 14 |
| CREB | Cyclic adenosine monophosphate responsive element-binding protein |
| DPP-IV | Dipeptidyl peptidase IV |
| Epac1 and Epac2 | Exchange protein directly activated by cyclic adenosine monophosphate 1 and 2 |
| ES | Embryonic stem cells |
| EX 4 | Exendin 4 |
| GI tract | Gastrointestinal tract |
| GIP | Gastric inhibitory peptide |
| GLP-1 | Glucagon-like peptide-1 |
| GPCR | G-protein coupled receptors |
| GPR41 and GPR43 | G-protein receptors 41 and 43 |
| IEC | Intestinal epithelial cell |
| IFN-γ | Interferon gamma |
| IL | Interleukin |
| IRAKs | Interleukin 1receptor associated kinases |
| IRF | Interferon regulatory factors |
| LBP | Lipopolysaccharide binding protein |
| LFHc | Low-fat, high-complex carbohydrate diet |
| LPS | Lipopolysaccharide |
| MAL | Myeloid differentiation primary response protein 88-adaptor-like protein |
| MAPK | Mitogen-activated protein kinases |
| MLDS | multiple low-dose streptozotocin |
| MYD88 | Myeloid differentiation primary response protein 88 |
| NF-κB | Nuclear factor kappa B |
| PCNA | Proliferating cell nuclear antigen |
| PDX-1 | Pancreatic and duodenal homeobox 1 |
| PKA | Protein kinase A |
| PYY | Peptide YY |
| SCFA | Short chain fatty acid |
| SUR1 | Sulfonylurea receptor 1 |
| TGFβ | Tumour growth factor beta |
| TLR4 | Toll-like receptor 4 |
| TNF | Tumour necrosis factor |
| TRAF | Tumour necrosis factor receptor-associated factors |
| Tregs | Regulatory T cells |
References
- Tang, A.T.; Choi, J.P.; Kotzin, J.J.; Yang, Y.; Hong, C.C.; Hobson, N.; Girard, R.; Zeineddine, H.A.; Lightle, R.; Moore, T.; et al. Endothelial TLR4 and the microbiome drive cerebral cavernous malformations. Nature 2017, 545, 305–310. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Firrman, J.; Zhang, L.; Arango-Argoty, G.; Tomasula, P.; Liu, L.; Xiao, W.; Yam, K. Apigenin impacts the growth of the gut microbiota and alters the gene expression of Enterococcus. Molecules 2017, 22, 1292. [Google Scholar] [CrossRef] [PubMed]
- Gomes, A.C.; Hoffmann, C.; Mota, J.F. The human gut microbiota: Metabolism and perspective in obesity. Gut Microbes 2018, 9, 308–325. [Google Scholar] [CrossRef] [PubMed]
- Ruiz, L.; Delgado, S.; Ruas-Madiedo, P.; Sánchez, B.; Margolles, A. Bifidobacteria and their molecular communication with the immune system. Front. Microbiol. 2017, 8, 2345. [Google Scholar] [CrossRef] [PubMed]
- Graf, D.; Di Cagno, R.; Fåk, F.; Flint, H.J.; Nyman, M.; Saarela, M.; Watzl, B. Contribution of diet to the composition of the human gut microbiota. Microb. Ecol. Health Dis. 2015, 26, 26164. [Google Scholar] [CrossRef] [PubMed]
- Soverini, M.; Turroni, S.; Biagi, E.; Quercia, S.; Brigidi, P.; Candela, M.; Rampelli, S. Variation of carbohydrate-active enzyme patterns in the gut microbiota of Italian healthy subjects and Type 2 Diabetes Patients. Front. Microbiol. 2017, 8, 2079. [Google Scholar] [CrossRef] [PubMed]
- Reichardt, N.; Duncan, S.H.; Young, P.; Belenguer, A.; McWilliam, L.C.; Scott, K.P.; Flint, H.J.; Louis, P. Phylogenetic distribution of three pathways for propionate production within the human gut microbiota. ISME J. 2014, 8, 1323–1335. [Google Scholar] [CrossRef] [PubMed]
- Moreno-Indias, I.; Cardona, F.; Tinahones, F.J.; Queipo-Ortuño, M.I. Impact of the gut microbiota on the development of obesity and type 2 diabetes mellitus. Front. Microbiol. 2014, 5, 190. [Google Scholar] [CrossRef] [PubMed]
- Manfredo Vieira, S.; Hiltensperger, M.; Kumar, V.; Zegarra-Ruiz, D.; Dehner, C.; Khan, N.; Costa, F.R.C.; Tiniakou, E.; Greiling, T.; Ruff, W.; et al. Translocation of a gut pathobiont drives autoimmunity in mice and humans. Science 2018, 359, 1156–1161. [Google Scholar] [CrossRef] [PubMed]
- Kobaek-Larsen, M.; Nielsen, D.S.; Kot, W.; Krych, Ł.; Christensen, L.P.; Baatrup, G. Effect of the dietary polyacetylenes falcarinol and falcarindiol on the gut microbiota composition in a rat model of colorectal cancer. BMC Res. Notes 2018, 11, 411. [Google Scholar] [CrossRef] [PubMed]
- Cani, P.D.; Jordan, B.F. Gut microbiota-mediated inflammation in obesity: A link with gastrointestinal cancer. Nat. Rev. Gastroenterol. Hepatol. 2018. [Google Scholar] [CrossRef] [PubMed]
- Rea, D.; Coppola, G.; Palma, G.; Barbieri, A.; Luciano, A.; Del Prete, P.; Rossetti, S.; Berretta, M.; Facchini, G.; Perdonà, S.; et al. Microbiota effects on cancer: From risks to therapies. Oncotarget 2018, 9, 17915–17927. [Google Scholar] [CrossRef] [PubMed]
- Bellocchi, C.; Volkmann, E.R. Update on the gastrointestinal microbiome in systemic sclerosis. Curr. Rheumatol. Rep. 2018, 20, 49. [Google Scholar] [CrossRef] [PubMed]
- Chu, F.; Shi, M.; Lang, Y.; Shen, D.; Jin, T.; Zhu, J.; Cui, L. Gut microbiota in multiple sclerosis and experimental autoimmune encephalomyelitis: Current applications and future perspectives. Mediat. Inflamm. 2018, 2018, 8168717. [Google Scholar] [CrossRef] [PubMed]
- Endesfelder, D.; zu Castell, W.; Ardissone, A.; Davis-Richardson, A.G.; Achenbach, P.; Hagen, M.; Pflueger, M.; Gano, K.A.; Fagen, J.R.; Drew, J.C.; et al. Compromised gut microbiota networks in children with anti-islet cell autoimmunity. Diabetes 2014, 63, 2006–2014. [Google Scholar] [CrossRef] [PubMed]
- Zhong, X.S.; Winston, J.H.; Luo, X.; Kline, K.T.; Nayeem, S.Z.; Cong, Y.; Savidge, T.C.; Dashwood, R.H.; Powell, D.W.; Li, Q. Neonatal colonic inflammation epigenetically aggravates epithelial inflammatory responses to injury in adult life. Cell. Mol. Gastroenterol. Hepatol. 2018, 6, 65–78. [Google Scholar] [CrossRef] [PubMed]
- Xun, Z.; Zhang, Q.; Xu, T.; Chen, N.; Chen, F. Dysbiosis and Ecotypes of the Salivary microbiome associated with inflammatory bowel diseases and the assistance in diagnosis of diseases using oral bacterial profiles. Front. Microbiol. 2018, 9, 1136. [Google Scholar] [CrossRef] [PubMed]
- Saroli Palumbo, C.; Restellini, S.; Chao, C.Y.; Aruljothy, A.; Lemieux, C.; Wild, G.; Afif, W.; Lakatos, P.L.; Bitton, A.; Cocciolillo, S.; et al. Screening for nonalcoholic fatty liver disease in inflammatory bowel diseases: A cohort study using Transient Elastography. Inflamm. Bowel Dis. 2018. [Google Scholar] [CrossRef] [PubMed]
- Alkanani, A.K.; Hara, N.; Gottlieb, P.A.; Ir, D.; Robertson, C.E.; Wagner, B.D.; Frank, D.N.; Zipris, D. Alterations in intestinal microbiota correlate with susceptibility to Type 1 Diabetes. Diabetes 2015, 64, 3510–3520. [Google Scholar] [CrossRef] [PubMed]
- Jena, P.K.; Prajapati, B.; Mishra, P.K.; Seshadri, S. Influence of gut microbiota on inflammation and pathogenesis of sugar rich diet-induced diabetes. Immun. Res. 2016, 12, 109. [Google Scholar]
- Aydin, Ö.; Nieuwdorp, M.; Gerdes, V. The gut microbiome as a target for the treatment of Type 2 Diabetes. Curr. Diab. Rep. 2018, 18, 55. [Google Scholar] [CrossRef] [PubMed]
- Zhu, G.; Ma, F.; Wang, G.; Wang, Y.; Zhao, J.; Zhang, H.; Chen, W. Bifidobacteria attenuate the development of metabolic disorders, with inter- and intra-species differences. Food Funct. 2018, 9, 3509–3522. [Google Scholar] [CrossRef] [PubMed]
- Kim, T.T.; Parajuli, N.; Sung, M.M.; Bairwa, S.C.; Levasseur, J.; Soltys, C.M.; Wishart, D.S.; Madsen, K.; Schertzer, J.D.; Dyck, J.R.B. Fecal transplant from resveratrol-fed donors improves glycaemia and cardiovascular features of the metabolic syndrome in mice. Am. J. Physiol. Endocrinol. Metab. 2018. [Google Scholar] [CrossRef] [PubMed]
- Fontané, L.; Benaiges, D.; Goday, A.; Llauradó, G.; Pedro-Botet, J. Influence of the microbiota and probiotics in obesity. Clin. Investig. Arterioscler. 2018, 30, 271–279. [Google Scholar]
- Bouter, K.; Bakker, G.J.; Levin, E.; Hartstra, A.V.; Kootte, R.S.; Udayappan, S.D.; Katiraei, S.; Bahler, L.; Gilijamse, P.W.; Tremaroli, V.; et al. Differential metabolic effects of oral butyrate treatment in lean versus metabolic syndrome subjects. Clin. Transl. Gastroenterol. 2018, 9, 155. [Google Scholar] [CrossRef] [PubMed]
- Yang, T.; Santisteban, M.M.; Rodriguez, V.; Li, E.; Ahmari, N.; Carvajal, J.M.; Zadeh, M.; Gong, M.; Qi, Y.; Zubcevic, J.; et al. Gut dysbiosis is linked to hypertension. Hypertension 2015, 65, 1331–1340. [Google Scholar] [CrossRef] [PubMed]
- Bidu, C.; Escoula, Q.; Bellenger, S.; Spor, A.; Galan, M.; Geissler, A.; Bouchot, A.; Dardevet, D.; Morio-Liondor, B.; Cani, P.D.; et al. The Transplantation of ω3 PUFA-altered gut microbiota of fat-1 mice to wild-type littermates prevents obesity and associated metabolic disorders. Diabetes 2018. [Google Scholar] [CrossRef] [PubMed]
- Gomes, A.C.; Bueno, A.A.; de Souza, R.G.; Mota, J.F. Gut microbiota, probiotics and diabetes. Nutr. J. 2014, 13, 60. [Google Scholar] [CrossRef] [PubMed]
- Sochocka, M.; Donskow-Łysoniewska, K.; Diniz, B.S.; Kurpas, D.; Brzozowska, E.; Leszek, J. The gut microbiome alterations and inflammation-driven pathogenesis of Alzheimer’s Disease—A critical review. Mol. Neurobiol. 2018. [Google Scholar] [CrossRef] [PubMed]
- Dinan, T.G.; Stilling, R.M.; Stanton, C.; Cryan, J.F. Collective unconscious: How gut microbes shape human behavior. J. Psychiatr. Res. 2015, 63, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Strandwitz, P. Neurotransmitter modulation by the gut microbiota. Brain Res. 2018, 1693, 128–133. [Google Scholar] [CrossRef] [PubMed]
- Naseribafrouei, A.; Hestad, K.; Avershina, E.; Sekelja, M.; Linløkken, A.; Wilson, R.; Rudi, K. Correlation between the human fecal microbiota and depression. Neurogastroenterol. Motil. 2014, 26, 1155–1162. [Google Scholar] [CrossRef] [PubMed]
- Scheperjans, F.; Aho, V.; Pereira, P.A.; Koskinen, K.; Paulin, L.; Pekkonen, E.; Haapaniemi, E.; Kaakkola, S.; Eerola-Rautio, J.; Pohja, M.; et al. Gut microbiota are related to Parkinson’s disease and clinical phenotype. Mov. Disord. 2015, 30, 350–358. [Google Scholar] [CrossRef] [PubMed]
- Patterson, E.; Ryan, P.M.; Cryan, J.F.; Dinan, T.G.; Ross, R.P.; Fitzgerald, G.F.; Stanton, C. Gut microbiota, obesity and diabetes. Postgrad. Med. J. 2016, 92, 286–300. [Google Scholar] [CrossRef] [PubMed]
- Zietek, T.; Rath, E. Inflammation meets metabolic disease: Gut feeling mediated by GLP-1. Front. Immunol. 2016, 7, 154. [Google Scholar] [CrossRef] [PubMed]
- Meyer, K.A.; Bennett, B.J. Diet and gut microbial function in metabolic and cardiovascular disease risk. Curr. Diab. Rep. 2016, 16, 93. [Google Scholar] [CrossRef] [PubMed]
- Wong, J.M. Gut microbiota and cardiometabolic outcomes: Influence of dietary patterns and their associated components. Am. J. Clin. Nutr. 2014, 100, 369S–377S. [Google Scholar] [CrossRef] [PubMed]
- Xu, Z.; Knight, R. Dietary effects on human gut microbiome diversity. Br. J. Nutr. 2015, 113, S1–S5. [Google Scholar] [CrossRef] [PubMed]
- Singh, S.P.; Jadaun, J.S.; Narnoliya, L.K.; Pandey, A. Prebiotic oligosaccharides: Special focus on fructooligosaccharides, its biosynthesis and bioactivity. Appl. Biochem. Biotechnol. 2017, 183, 613–615. [Google Scholar] [CrossRef] [PubMed]
- Rea, K.; O’Mahony, S.M.; Dinan, T.G.; Cryan, J.F. The role of the gastrointestinal microbiota in visceral pain. Handb. Exp. Pharmacol. 2017, 239, 269–287. [Google Scholar] [PubMed]
- Burke, D.G.; Fouhy, F.; Harrison, M.J.; Rea, M.C.; Cotter, P.D.; O’Sullivan, O.; Stanton, C.; Hill, C.; Shanahan, F.; Plant, B.J.; et al. The altered gut microbiota in adults with cystic fibrosis. BMC Microbiol. 2017, 17, 58. [Google Scholar]
- Yamaguchi, Y.; Adachi, K.; Sugiyama, T.; Shimozato, A.; Ebi, M.; Ogasawara, N.; Funaki, Y.; Goto, C.; Sasaki, M.; Kasugai, K. Association of intestinal microbiota with metabolic markers and dietary habits in patients with type 2 Diabetes. Digestion 2016, 94, 66–72. [Google Scholar] [CrossRef] [PubMed]
- Sanz, Y.; Olivares, M.; Moya-Pérez, Á.; Agostoni, C. Understanding the role of gut microbiome in metabolic disease risk. Pediatr. Res. 2015, 77, 236–244. [Google Scholar] [CrossRef] [PubMed]
- Salonen, A.; de Vos, W.M. Impact of diet on human intestinal microbiota and health. Annu. Rev. Food Sci. Technol. 2014, 5, 239–262. [Google Scholar] [CrossRef] [PubMed]
- Musilova, S.; Modrackova, N.; Hermanova, P.; Hudcovic, T.; Svejstil, R.; Rada, V.; Tejnecky, V.; Bunesova, V. Assessment of the synbiotic properites of human milk oligosaccharides and Bifidobacterium longum subsp. infantis in vitro and in humanised mice. Benef. Microbes 2017, 8, 281–289. [Google Scholar] [CrossRef] [PubMed]
- Blatchford, P.; Stoklosinski, H.; Eady, S.; Wallace, A.; Butts, C.; Gearry, R.; Gibson, G.; Ansell, J. Consumption of kiwifruit capsules increases Faecalibacterium prausnitzii abundance in functionally constipated individuals: A randomised controlled human trial. J. Nutr. Sci. 2017, 6, e52. [Google Scholar] [CrossRef] [PubMed]
- Sonnenburg, J.L.; Bäckhed, F. Diet-microbiota interactions as moderators of human metabolism. Nature 2016, 535, 56–64. [Google Scholar] [CrossRef] [PubMed]
- Zoetendal, E.G.; de Vos, W.M. Effect of diet on the intestinal microbiota and its activity. Curr. Opin. Gastroenterol. 2014, 30, 189–195. [Google Scholar] [CrossRef] [PubMed]
- Shen, W.; Gaskins, H.R.; McIntosh, M.K. Influence of dietary fat on intestinal microbes, inflammation, barrier function and metabolic outcomes. J. Nutr. Biochem. 2014, 25, 270–280. [Google Scholar] [CrossRef] [PubMed]
- Kashyap, P.C.; Marcobal, A.; Ursell, L.K.; Smits, S.A.; Sonnenburg, E.D.; Costello, E.K.; Higginbottom, S.K.; Domino, S.E.; Holmes, S.P.; Relman, D.A.; et al. Genetically dictated change in host mucus carbohydrate landscape exerts a diet-dependent effect on the gut microbiota. Proc. Natl. Acad. Sci. USA 2013, 110, 17059–17064. [Google Scholar] [CrossRef] [PubMed]
- Zimmer, J.; Lange, B.; Frick, J.S.; Sauer, H.; Zimmermann, K.; Schwiertz, A.; Rusch, K.; Klosterhalfen, S.; Enck, P. A vegan or vegetarian diet substantially alters the human colonic faecal microbiota. Eur. J. Clin. Nutr. 2012, 66, 53–60. [Google Scholar] [CrossRef] [PubMed]
- Biragyn, A.; Ferrucci, L. Gut dysbiosis: A potential link between increased cancer risk in ageing and inflammaging. Lancet Oncol. 2018, 19, e295–e304. [Google Scholar] [CrossRef]
- Savin, Z.; Kivity, S.; Yonath, H.; Yehuda, S. Smoking and the intestinal microbiome. Arch. Microbiol. 2018, 200, 677–684. [Google Scholar] [CrossRef] [PubMed]
- Martin, G.; Kolida, S.; Marchesi, J.R.; Want, E.; Sidaway, J.E.; Swann, J.R. In Vitro Modeling of Bile Acid Processing by the Human Fecal Microbiota. Front. Microbiol. 2018, 9, 1153. [Google Scholar] [CrossRef] [PubMed]
- Allaire, J.M.; Morampudi, V.; Crowley, S.M.; Stahl, M.; Yu, H.; Bhullar, K.; Knodler, L.A.; Bressler, B.; Jacobson, K.; Vallance, B.A. Frontline defenders: Goblet cell mediators dictate host-microbe interactions in the intestinal tract during health and disease. Am. J. Physiol. Gastrointest. Liver Physiol. 2018, 314, G360–G377. [Google Scholar] [CrossRef] [PubMed]
- Conlon, M.A.; Bird, A.R. The impact of diet and lifestyle on gut microbiota and human health. Nutrients 2014, 7, 17–44. [Google Scholar] [CrossRef] [PubMed]
- Pannaraj, P.S.; Ly, M.; Cerini, C.; Saavedra, M.; Aldrovandi, G.M.; Saboory, A.A.; Johnson, K.M.; Pride, D.T. Shared and distinct features of human milk and infant stool viromes. Front. Microbiol. 2018, 9, 1162. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.-W.; Suda, W.; Kim, S.; Oshima, K.; Fukuda, S.; Ohno, H.; Morita, H.; Hattori, M. Robustness of gut microbiota of healthy adults in response to probiotic intervention revealed by high-throughput pyrosequencing. DNA Res. 2013, 20, 241–253. [Google Scholar] [CrossRef] [PubMed]
- Luo, J.; Han, L.; Liu, L.; Gao, L.; Xue, B.; Wang, Y.; Ou, S.; Miller, M.; Peng, X. Catechin supplemented in a FOS diet induces weight loss by altering cecal microbiota and gene expression of colonic epithelial cells. Food Funct. 2018, 9, 2962–2969. [Google Scholar] [CrossRef] [PubMed]
- Mu, C.; Yang, Y.; Luo, Z.; Guan, L.; Zhu, W. The colonic microbiome and epithelial transcriptome are altered in rats fed a high-protein diet compared with a normal-protein diet. J. Nutr. 2016, 146, 474–483. [Google Scholar] [CrossRef] [PubMed]
- Wu, G.D.; Chen, J.; Hoffmann, C.; Bittinger, K.; Chen, Y.Y.; Keilbaugh, S.A.; Bewtra, M.; Knights, D.; Walters, W.A.; Knight, R.; et al. Linking long-term dietary patterns with gut microbial enterotypes. Science 2011, 334, 105–108. [Google Scholar] [CrossRef] [PubMed]
- Winglee, K.; Howard, A.G.; Sha, W.; Gharaibeh, R.Z.; Liu, J.; Jin, D.; Fodor, A.A.; Gordon-Larsen, P. Recent urbanization in China is correlated with a Westernized microbiome encoding increased virulence and antibiotic resistance genes. Microbiome 2017, 5, 121. [Google Scholar] [CrossRef] [PubMed]
- Martinez-Medina, M.; Denizot, J.; Dreux, N.; Robin, F.; Billard, E.; Bonnet, R.; Darfeuille-Michaud, A.; Barnich, N. Western diet induces dysbiosis with increased E coli in CEABAC10 mice, alters host barrier function favouring AIEC colonisation. Gut 2014, 63, 116–124. [Google Scholar] [CrossRef] [PubMed]
- Chaves, D.F.S.; Carvalho, P.C.; Brasili, E.; Rogero, M.M.; Hassimotto, N.A.; Diedrich, J.K.; Moresco, J.J.; Yates, J.R., 3rd; Lajolo, F.M. Proteomic analysis of peripheral blood mononuclear cells after a high-fat, high-carbohydrate meal with orange juice. J. Proteome Res. 2017, 16, 4086–4092. [Google Scholar] [CrossRef] [PubMed]
- Emerson, S.R.; Kurti, S.P.; Harms, C.A.; Haub, M.D.; Melgarejo, T.; Logan, C.; Rosenkranz, S.K. Magnitude and timing of the postprandial inflammatory response to a high-fat meal in healthy adults: A Systematic Review. Adv. Nutr. 2017, 8, 213–225. [Google Scholar] [CrossRef] [PubMed]
- Herieka, M.; Erridge, C. High-fat meal induced postprandial inflammation. Mol. Nutr. Food Res. 2014, 58, 136–146. [Google Scholar] [CrossRef] [PubMed]
- Bibbò, S.; Dore, M.P.; Pes, G.M.; Delitala, G.; Delitala, A.P. Is there a role for gut microbiota in type 1 diabetes pathogenesis? Ann. Med. 2017, 49, 11–22. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Chen, Y.; Sun, Y.; Wang, R.; Zhang, J.; Jia, Z. Plateau hypoxia attenuates the metabolic activity of intestinal flora to enhance the bioavailability of nifedipine. Drug Deliv. 2018, 25, 1175–1181. [Google Scholar] [CrossRef] [PubMed]
- German, J.B.; Freeman, S.L.; Lebrilla, C.B.; Mills, D.A. Human milk oligosaccharides: Evolution, structures and bioselectivity as substrates for intestinal bacteria. Nestle Nutr. Workshop Ser. Pediatr. Program. 2008, 62, 205–218. [Google Scholar] [PubMed]
- Marcobal, A.; Barboza, M.; Sonnenburg, E.D.; Pudlo, N.; Martens, E.C.; Desai, P.; Lebrilla, C.B.; Weimer, B.C.; Mills, D.A.; German, J.B.; et al. Bacteroides in the infant gut consume milk oligosaccharides via mucus-utilization pathways. Cell Host Microbe 2011, 10, 507–514. [Google Scholar] [CrossRef] [PubMed]
- Marcobal, A.; Sonnenburg, J.L. Human milk oligosaccharide consumption by intestinal microbiota. Clin. Microbiol. Infect. 2012, 18, 12–15. [Google Scholar] [CrossRef] [PubMed]
- Martinez, I.; Lattimer, J.M.; Hubach, K.L.; Case, J.A.; Yang, J.; Weber, C.G.; Louk, J.A.; Rose, D.J.; Kyureghian, G.; Peterson, D.A.; et al. Resistant starches types 2 and 4 have differential effects on the composition of the fecal microbiota in human subjects. PLoS ONE 2010, 5, e15046. [Google Scholar] [CrossRef] [PubMed]
- Linetzky, W.D.; Alves Pereira, C.C.; Logullo, L.; Manzoni, J.T.; Almeida, D.; Teixeira da Silva, M.L.; Matos de Miranda Torrinhas, R.S. Microbiota benefits after inulin and partially hydrolized guar gum supplementation: A randomized clinical trial in constipated women. Nutr. Hosp. 2012, 27, 123–129. [Google Scholar]
- Baer, D.J.; Stote, K.S.; Henderson, T.; Paul, D.R.; Okuma, K.; Tagami, H.; Kanahori, S.; Gordon, D.T.; Rumpler, W.V.; Ukhanova, M.; et al. The metabolizable energy of dietary resistant maltodextrin is variable and alters fecal microbiota composition in adult men. J. Nutr. 2014, 144, 1023–1029. [Google Scholar] [CrossRef] [PubMed]
- Ramirez-Farias, C.; Slezak, K.; Fuller, Z.; Duncan, A.; Holtrop, G.; Louis, P. Effect of inulin on the human gut microbiota: Stimulation of Bifidobacterium adolescentis and Faecalibacterium prausnitzii. Br. J. Nutr. 2009, 101, 541–550. [Google Scholar] [CrossRef] [PubMed]
- Louis, P.; Young, P.; Holtrop, G.; Flint, H.J. Diversity of human colonic butyrate-producing bacteria revealed by analysis of the butyryl-CoA: Acetate CoA transferase gene. Environ. Microbiol. 2010, 12, 304–314. [Google Scholar] [CrossRef] [PubMed]
- Ramnani, P.; Gaudier, E.; Bingham, M.; van Bruggen, P.; Tuohy, K.M.; Gibson, G.R. Prebiotic effect of fruit and vegetable shots containing Jerusalem artichoke inulin: A human intervention study. Br. J. Nutr. 2010, 104, 233–240. [Google Scholar] [CrossRef] [PubMed]
- Costabile, A.; Kolida, S.; Klinder, A.; Gietl, E.; Bauerlein, M.; Frohberg, C.; Landschütze, V.; Gibson, G.R. A double-blind, placebo-controlled, cross-over study to establish the bifidogenic effect of a very-long-chain inulin extracted from globe artichoke (Cynara scolymus) in healthy human subjects. Br. J. Nutr. 2010, 104, 1007–1017. [Google Scholar] [CrossRef] [PubMed]
- Lecerf, J.M.; Depeint, F.; Clerc, E.; Dugenet, Y.; Niamba, C.N.; Rhazi, L.; Cayzeele, A.; Abdelnour, G.; Jaruga, A.; Younes, H.; et al. Xylo-oligosaccharide (XOS) in combination with inulin modulates both the intestinal environment and immune status in healthy subjects, while XOS alone only shows prebiotic properties. Br. J. Nutr. 2012, 108, 1847–1858. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Peris, P.; Velasco, C.; Lozano, M.A.; Moreno, Y.; Paron, L.; de la Cuerda, C.; Bretón, I.; Camblor, M.; García-Hernández, J.; Guarner, F.; et al. Effect of a mixture of inulin and fructo-oligosaccharide on Lactobacillus and Bifidobacterium intestinal microbiota of patients receiving radiotherapy: A randomized, double-blind, placebo-controlled trial. Nutr. Hosp. 2012, 27, 1908–1915. [Google Scholar] [PubMed]
- Dewulf, E.M.; Cani, P.D.; Claus, S.P.; Fuentes, S.; Puylaert, P.G.; Neyrinck, A.M.; Bindels, L.B.; de Vos, W.M.; Gibson, G.R.; Thissen, J.P.; et al. Insight into the prebiotic concept: Lessons from an exploratory, double blind intervention study with inulin-type fructans in obese women. Gut 2013, 62, 1112–1121. [Google Scholar] [CrossRef] [PubMed]
- Vivatvakin, B.; Mahayosnond, A.; Theamboonlers, A.; Steenhout, P.G.; Conus, N.J. Effect of a whey-predominant starter formula containing LCPUFAs and oligosaccharides (FOS/GOS) on gastrointestinal comfort in infants. Asia Pac. J. Clin. Nutr. 2010, 19, 473–480. [Google Scholar] [PubMed]
- Vulevic, J.; Juric, A.; Tzortzis, G.; Gibson, G.R. A mixture of trans-galactooligosaccharides reduces markers of metabolic syndrome and modulates the fecal microbiota and immune function of overweight adults. J. Nutr. 2013, 143, 324–331. [Google Scholar] [CrossRef] [PubMed]
- Hooda, S.; Boler, B.M.; Serao, M.C.; Brulc, J.M.; Staeger, M.A.; Boileau, T.W.; Dowd, S.E.; Fahey, G.C., Jr.; Swanson, K.S. 454 pyrosequencing reveals a shift in fecal microbiota of healthy adult men consuming polydextrose or soluble corn fiber. J. Nutr. 2012, 142, 1259–1265. [Google Scholar] [CrossRef] [PubMed]
- Costabile, A.; Fava, F.; Roytio, H.; Forssten, S.D.; Olli, K.; Klievink, J.; Rowland, I.R.; Ouwehand, A.C.; Rastall, R.A.; Gibson, G.R.; et al. Impact of polydextrose on the faecal microbiota: A double-blind, crossover, placebo-controlled feeding study in healthy human subjects. Br. J. Nutr. 2012, 108, 471–481. [Google Scholar] [CrossRef] [PubMed]
- Cloetens, L.; Broekaert, W.F.; Delaedt, Y.; Ollevier, F.; Courtin, C.M.; Delcour, J.A.; Rutgeerts, P.; Verbeke, K. Tolerance of arabinoxylan-oligosaccharides and their prebiotic activity in healthy subjects: A randomised, placebo-controlled cross-over study. Br. J. Nutr. 2010, 103, 703–713. [Google Scholar] [CrossRef] [PubMed]
- Walton, G.E.; Lu, C.; Trogh, I.; Arnaut, F.; Gibson, G.R. A randomised, double-blind, placebo controlled cross-over study to determine the gastrointestinal effects of consumption of arabinoxylan-oligosaccharides enriched bread in healthy volunteers. Nutr. J. 2012, 11, 36. [Google Scholar] [CrossRef] [PubMed]
- Shinohara, K.; Ohashi, Y.; Kawasumi, K.; Terada, A.; Fujisawa, T. Effect of apple intake on fecal microbiota and metabolites in humans. Anaerobe 2010, 16, 510–515. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.K.; Low, K.Y.; Siah, K.; Drummond, L.M.; Kok An Gwee, K.A. Kiwifruit (Actinidia deliciosa) changes intestinal microbial profile. Microb. Ecol. Health Dis. 2012, 23, 18572–18576. [Google Scholar] [CrossRef] [PubMed]
- Mitsou, E.K.; Kougia, E.; Nomikos, T.; Yannakoulia, M.; Mountzouris, K.C.; Kyriacou, A. Effect of banana consumption on faecal microbiota: A randomized, controlled trial. Anaerobe 2011, 17, 384–387. [Google Scholar] [CrossRef] [PubMed]
- Louis, P.; Scott, K.P.; Duncan, S.H.; Flint, H.J. Understanding the effects of diet on bacterial metabolism in the large intestine. J. Appl. Microbiol. 2007, 102, 1197–1208. [Google Scholar] [CrossRef] [PubMed]
- Topping, D. Cereal complex carbohydrates and their contribution to human health. J. Cereal Sci. 2007, 46, 220–229. [Google Scholar] [CrossRef]
- Pieper, R.; Bindelle, J.; Rossnagel, B.; Van Kessel, A.; Pascal Leterme, P. Effect of carbohydrate composition in barley and oat cultivars on microbial ecophysiology and proliferation of Salmonella enteric in an in vitro model of the porcine gastrointestinal tract. Appl. Environ. Microbiol. 2009, 75, 7006–7016. [Google Scholar] [CrossRef] [PubMed]
- Bindelle, J.; Pieper, R.; Montoya, C.A.; Van Kessel, A.G.; Leterme, P. Nonstarch polysaccharide degrading enzymes alter the microbial community and the fermentation patterns of barley cultivars and wheat products in an in vitro model of the porcine gastrointestinal tract. FEMS Microbiol. Ecol. 2011, 76, 553–563. [Google Scholar] [CrossRef] [PubMed]
- Walker, A.W.; Ince, J.; Duncan, S.H.; Webster, L.M.; Holtrop, G.; Ze, X.; Brown, D.; Stares, M.D.; Scott, P.; Bergerat, A.; et al. Dominant and diet-responsive groups of bacteria within the human colonic microbiota. ISME J. 2011, 5, 220–230. [Google Scholar] [CrossRef] [PubMed]
- Duncan, S.H.; Belenguer, A.; Holtrop, G.; Johnstone, A.M.; Flint, H.J.; Lobley, G.E. Reduced dietary intake of carbohydrates by obese subjects results in decreased concentrations of butyrate and butyrate-producing bacteria in feces. Appl. Environ. Microbiol. 2007, 73, 1073–1078. [Google Scholar] [CrossRef] [PubMed]
- Leitch, E.C.M.; Walker, A.W.; Duncan, S.H.; Holtrop, G.; Flint, H.J. Selective colonization of insoluble substrates by human colonic bacteria. Environ. Microbiol. 2007, 72, 667–669. [Google Scholar] [CrossRef] [PubMed]
- Abell, G.C.J.; Cooke, C.M.; Bennett, C.N.; Conlon, M.A.; McOrist, A.L. Phylotypes related to Ruminococcus bromii are abundant in the large bowel of humans and increase in response to a diet high in resistant starch. FEMS Microbiol. Ecol. 2008, 66, 505–515. [Google Scholar] [CrossRef] [PubMed]
- Benno, Y.; Endo, K.; Miyoshi, H.; Okuda, T.; Koishi, H.; Mitsuoka, T. Effect of rice fiber on human fecal microflora. Microbiol. Immunol. 1989, 33, 435–440. [Google Scholar] [CrossRef] [PubMed]
- De Filippo, C.; Cavalieri, D.; Di Paola, M.; Ramazzotti, M.; Poullet, J.B.; Massart, S.; Collini, S.; Pieraccini, G.; Lionetti, P. Impact of diet in shaping gut microbiota revealed by a comparative study in children from Europe and rural Africa. Proc. Natl. Acad. Sci. USA 2010, 107, 14691–14696. [Google Scholar] [CrossRef] [PubMed]
- Costabile, A.; Klinder, A.; Fava, F.; Napolitano, A.; Fogliano, V.; Leonard, C.; Gibson, G.R.; Tuohy, K.M. Whole-grain wheat breakfast cereal has a prebiotic effect on the human gut microbiota: A double-blind, placebo-controlled, crossover study. Br. J. Nutr. 2008, 99, 110–120. [Google Scholar] [CrossRef] [PubMed]
- Neyrinck, A.M.; Sam Possemiers, S.; Druart, C.; van de Wiele, T.; De Backer, F.; Cani, P.D.; Larondelle, Y.; Delzenne, N.M. Prebiotic effects of wheat arabinoxylan related to the increase in Bifidobacteria, Roseburia and Bacteroides/Prevotella in diet-induced obese mice. PLoS ONE 2011, 6, e20944. [Google Scholar] [CrossRef] [PubMed]
- Carvalho-Wells, A.L.; Helmolz, K.; Nodet, C.; Molzer, C.; Leonard, C.; McKevith, B.; Thielecke, F.; Jackson, K.G.; Tuohy, K.M. Determination of the in vivo prebiotic potential of a maize-based whole grain breakfast cereal: A human feeding study. Br. J. Nutr. 2010, 104, 1353–1356. [Google Scholar] [CrossRef] [PubMed]
- Martinez, I.; Lattimer, J.M.; Hubach, K.L.; Case, J.A.; Yang, J.; Weber, C.G.; Louk, J.A.; Rose, D.J.; Kyureghian, G.; Peterson, D.A.; et al. Gut microbiome composition is linked to whole grain-induced immunological improvements. ISME J. 2013, 7, 269–280. [Google Scholar] [CrossRef] [PubMed]
- Lappi, J.; Salojarvi, J.; Kolehmainen, M.; Mykkanen, H.; Poutanen, K.; de Vos, W.M.; Salonen, A. Intake of whole-grain and fiber-rich rye bread versus refined wheat bread does not differentiate intestinal microbiota composition in Finnish adults with metabolic syndrome. J. Nutr. 2013, 143, 648–655. [Google Scholar] [CrossRef] [PubMed]
- Vendrame, S.; Guglielmetti, S.; Riso, P.; Arioli, S.; Klimis-Zacas, D.; Porrini, M. Six-week consumption of a wild blueberry powder drink increases Bifidobacteria in the human gut. J. Agric. Food Chem. 2011, 59, 12815–12820. [Google Scholar] [CrossRef] [PubMed]
- Guglielmetti, S.; Fracassetti, D.; Taverniti, V.; Del Bo, C.; Vendrame, S.; Klimis-Zacas, D.; Arioli, S.; Riso, P.; Porrini, M. Differential modulation of human intestinal bifidobacterium populations after consumption of a wild blueberry (Vaccinium angustifolium) drink. J. Agric. Food Chem. 2013, 61, 8134–8140. [Google Scholar] [CrossRef] [PubMed]
- Queipo-Ortuno, M.I.; Boto-Ordonez, M.; Murri, M.; Gomez-Zumaquero, J.M.; Clemente-Postigo, M.; Estruch, R.; Cardona Diaz, F.; Andrés-Lacueva, C.; Tinahones, F.J. Influence of red wine polyphenols and ethanol on the gut microbiota ecology and biochemical biomarkers. Am. J. Clin. Nutr. 2012, 95, 1323–1334. [Google Scholar] [CrossRef] [PubMed]
- Ukhanova, M.; Wang, X.; Baer, D.J.; Novotny, J.A.; Fredborg, M.; Mai, V. Effects of almond and pistachio consumption on gut microbiota composition in a randomized cross-over human feeding study. Br. J. Nutr. 2014, 111, 2146–2152. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Lin, X.; Huang, G.; Zhang, W.; Rao, P.; Ni, L. Prebiotic effects of almonds and almond skins on intestinal microbiota in healthy adult humans. Anaerobe 2014, 26, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Cani, P.D.; Amar, J.; Iglesias, M.A.; Poggi, M.; Knauf, C.; Bastelica, D.; Neyrinck, A.M.; Fava, F.; Tuohy, K.M.; Chabo, C.; et al. Metabolic endotoxemia initiates obesity and insulin resistance. Diabetes 2007, 56, 1761–1772. [Google Scholar] [CrossRef] [PubMed]
- Cani, P.D.; Delzenne, N.M. Benefits of bariatric surgery: An issue of microbial-host metabolism interactions? Gut 2011, 60, 1166–1167. [Google Scholar] [CrossRef] [PubMed]
- Weisberg, S.P.; McCann, D.; Desai, M.; Rosenbaum, M.; Leibel, R.L.; Ferrante, A.W., Jr. Obesity is associated with macrophage accumulation in adipose tissue. J. Clin. Investig. 2003, 112, 1796–1808. [Google Scholar] [CrossRef] [PubMed]
- Kabeerdoss, J.; Devi, R.S.; Mary, R.R.; Ramakrishna, B.S. Faecal microbiota composition in vegetarians: Comparison with omnivores in a cohort of young women in southern India. Br. J. Nutr. 2012, 108, 953–957. [Google Scholar] [CrossRef] [PubMed]
- McAllan, L.; Skuse, P.; Cotter, P.D.; O’Connor, P.; Cryan, J.F.; Ross, R.P.; Fitzgerald, G.; Roche, H.M.; Nilaweera, K.N. Protein quality and the protein to carbohydrate ratio within a high fat diet influences energy balance and the gut microbiota in C57BL/6J mice. PLoS ONE 2014, 9, e88904. [Google Scholar] [CrossRef] [PubMed]
- Fernando, W.M.; Hill, J.E.; Zello, G.A.; Tyler, R.T.; Dahl, W.J.; Van Kessel, A.G. Diets supplemented with chickpea or its main oligosaccharide component raffinose modify faecal microbial composition in healthy adults. Benef. Microbes 2010, 1, 197–207. [Google Scholar] [CrossRef] [PubMed]
- Fernandez-Raudales, D.; Hoeflinger, J.L.; Bringe, N.A.; Cox, S.B.; Dowd, S.E.; Miller, M.J.; Gonzalez de Mejia, E. Consumption of different soymilk formulations differentially affects the gut microbiomes of overweight and obese men. Gut Microbes 2012, 3, 490–500. [Google Scholar] [CrossRef] [PubMed]
- Cichewicz, R.H.; Thorpe, P.A. The antimicrobial properties of chile peppers (Capsicum species) and their uses in Mayan medicine. J. Ethnopharmacol. 1996, 52, 61–70. [Google Scholar] [CrossRef]
- Sivam, G.P.; Lampe, J.W.; Ulness, B.; Swanzy, S.R.; Potter, J.D. Helicobacter pylori–in vitro susceptibility to garlic (Allium sativum) extract. Nutr. Cancer 1997, 27, 118–121. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.C.; Jenner, A.M.; Low, C.S.; Lee, Y.K. Effect of tea phenolics and their aromatic fecal bacterial metabolites on intestinal microbiota. Res. Microbiol. 2006, 157, 876–884. [Google Scholar] [CrossRef] [PubMed]
- Molan, A.L.; Lila, A.A.; Mawson, J.; De, S. In vitro and in vivo evaluation of the prebiotic activity of water-soluble blueberry extracts. World J. Microbiol. Biotechnol. 2009, 25, 1243–1249. [Google Scholar] [CrossRef]
- Tzounis, X.; Vulevic, J.; Kuhnle, G.G.; George, T.; Leonczak, J.; Gibson, G.R.; Kwik-Uribe, C.; Spencer, J.P. Flavanol monomer-induced changes to the human faecal microflora. Br. J. Nutr. 2008, 99, 782–792. [Google Scholar] [CrossRef] [PubMed]
- Medina, E.; Garcia, A.; Romero, C.; de Castro, A.; Brenes, M. Study of the anti-lactic acid bacteria compounds in table olives. Int. J. Food Sci. Technol. 2009, 7, 1286–1291. [Google Scholar] [CrossRef]
- Wang, W.B.; Lai, H.C.; Hsueh, P.R.; Chiou, R.Y.Y.; Lin, S.B.; Liaw, S.J. Inhibition of swarming and virulence factor expression in Proteus mirabilis by resveratrol. J. Med. Microbiol. 2006, 55, 1313–1321. [Google Scholar] [CrossRef] [PubMed]
- Larrosa, M.; Yanéz-Gascón, M.J.; Selma, M.V.; González-Sarrías, A.; Toti, S.; Cerón, J.J.; Tomás-Barberán, F.; Dolara, P.; Espín, J.C. Effect of a low dose of dietary resveratrol on colon microbiota, inflammation and tissue damage in a DSS-induced colitis rat model. J. Agric. Food Chem. 2009, 57, 2211–2220. [Google Scholar] [CrossRef] [PubMed]
- Puupponen-Pimiä, R.; Nohynek, L.; Hartmann-Schmidlin, S.; Kähkönen, M.; Heinonen, M.; Määttä-Riihinen, K.; Oksman-Caldentey, K.M. Berry phenolics selectively inhibit the growth of intestinal pathogens. J. Appl. Microbiol. 2005, 98, 991–1000. [Google Scholar] [CrossRef] [PubMed]
- Nohynek, L.J.; Alakomi, H.L.; Kahkonen, M.P.; Heinonen, M.; Helander, I.M.; Oksman-Caldentey, K.M.; Puupponen-Pimiä, R.H. Berry phenolics: Antimicrobial properties and mechanisms of action against severe human pathogens. Nutr. Cancer 2006, 54, 18–32. [Google Scholar] [CrossRef] [PubMed]
- Salonen, A.; Lahti, L.; Salojärvi, J.; Holtrop, G.; Korpela, K.; Duncan, S.H.; Date, P.; Farquharson, F.; Johnstone, A.M.; Lobley, G.E.; et al. Impact of diet and individual variation on intestinal microbiota composition and fermentation products in obese men. ISME J. 2014, 8, 2218–2230. [Google Scholar] [CrossRef] [PubMed]
- Yin, D.T.; Fu, Y.; Zhao, X.H. In vitro activities of inulin fermentation products to HCT-116 cells enhanced by the cooperation between exogenous strains and adult faecal microbiota. Int. J. Food Sci. Nutr. 2018, 69, 814–823. [Google Scholar] [CrossRef] [PubMed]
- Chung, W.S.; Walker, A.W.; Louis, P.; Parkhill, J.; Vermeiren, J.; Bosscher, D.; Duncan, S.H.; Flint, H.J. Modulation of the human gut microbiota by dietary fibres occurs at the species level. BMC Biol. 2016, 14, 3. [Google Scholar] [CrossRef] [PubMed]
- Ang, Z.; Ding, J.L. GPR41 and GPR43 in obesity and inflammation—Protective or causative? Front. Immunol. 2016, 7, 28. [Google Scholar] [CrossRef] [PubMed]
- Buenaventura, T.; Kanda, N.; Douzenis, P.C.; Jones, B.; Bloom, S.R.; Chabosseau, P.; Corrêa, I.R., Jr.; Bosco, D.; Piemonti, L.; Marchetti, P.; et al. A targeted rnai screen identifies endocytic trafficking factors that control GLP-1 receptor signaling in pancreatic β-Cells. Diabetes 2018, 67, 385–399. [Google Scholar] [CrossRef] [PubMed]
- Kai, A.K.; Lam, A.K.; Chen, Y.; Tai, A.C.; Zhang, X.; Lai, A.K.; Yeung, P.K.; Tam, S.; Wang, J.; Lam, K.S.; et al. Exchange protein activated by cAMP 1 (Epac1)-deficient mice develop β-cell dysfunction and metabolic syndrome. FASEB J. 2013, 27, 4122–4135. [Google Scholar] [CrossRef] [PubMed]
- Hwang, M.; Go, Y.; Park, J.H.; Shin, S.K.; Song, S.E.; Oh, B.C.; Im, S.S.; Hwang, I.; Jeon, Y.H.; Lee, I.K.; et al. Epac2a-null mice exhibit obesity-prone nature more susceptible to leptin resistance. Int. J. Obes. 2017, 41, 279–288. [Google Scholar] [CrossRef] [PubMed]
- Sfairopoulos, D.; Liatis, S.; Tigas, S.; Liberopoulos, E. Clinical pharmacology of glucagon-like peptide-1 receptor agonists. Hormones 2018. [Google Scholar] [CrossRef] [PubMed]
- Guja, C.; Frías, J.P.; Somogyi, A.; Jabbour, S.; Wang, H.; Hardy, E.; Rosenstock, J. Effect of exenatide QW or placebo, both added to titrated insulin glargine, in uncontrolled type 2 diabetes: The DURATION-7 randomized study. Diabetes Obes. Metab. 2018, 20, 1602–1614. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.S.; Jun, H.S. Glucagon-like peptide-1 receptor agonist and glucagon increase glucose-stimulated insulin secretion in beta cells via distinct adenylyl cyclases. Int. J. Med. Sci. 2018, 15, 603. [Google Scholar] [CrossRef] [PubMed]
- Nygaard, G.; Herfindal, L.; Asrud, K.S.; Bjørnstad, R.; Kopperud, R.K.; Oveland, E.; Berven, F.S.; Myhren, L.; Hoivik, E.A.; Lunde, T.H.F.; et al. Epac1-deficient mice have bleeding phenotype and thrombocytes with decreased GPIbβ expression. Sci. Rep. 2017, 7, 8725. [Google Scholar] [CrossRef] [PubMed]
- Nannipieri, M.; Baldi, S.; Mari, A.; Colligiani, D.; Guarino, D.; Camastra, S.; Barsotti, E.; Berta, R.; Moriconi, D.; Bellini, R.; et al. Roux-en-Y gastric bypass and sleeve gastrectomy: Mechanisms of diabetes remission and role of gut hormones. J. Clin. Endocrinol. Metab. 2013, 98, 4391–4399. [Google Scholar] [CrossRef] [PubMed]
- Li, A.Q.; Zhao, L.; Zhou, T.F.; Zhang, M.Q.; Qin, X.M. Exendin-4 promotes endothelial barrier enhancement via PKA- and Epac1-dependent Rac1 activation. Am. J. Physiol. Cell Physiol. 2015, 308, C164–C175. [Google Scholar] [CrossRef] [PubMed]
- Tremaroli, V.; Backhed, F. Functional interactions between the gut microbiota and host metabolism. Nature 2012, 489, 242–249. [Google Scholar] [CrossRef] [PubMed]
- Everard, A.; Cani, P.D. Diabetes, obesity and gut microbiota. Best Pract. Res. Clin. Gastroenterol. 2013, 27, 73–83. [Google Scholar] [CrossRef] [PubMed]
- Viggiano, D.; Ianiro, G.; Vanella, G.; Bibbo, S.; Bruno, G.; Simeone, G.; Mele, G. Gut barrier in health and disease: Focus on childhood. Eur. Rev. Med. Pharmacol. Sci. 2015, 19, 1077–1085. [Google Scholar] [PubMed]
- Gulden, E.; Wong, F.S.; Wen, L. The gut microbiota and type 1 diabetes. Clin. Immunol. 2015, 159, 143–153. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Atkinson, M.A. The role for gut permeability in the pathogenesis of type 1 diabetes—A solid or leaky concept? Pediat. Diabetes 2015, 16, 485–492. [Google Scholar] [CrossRef] [PubMed]
- Bodin, J.; Bølling, A.K.; Becher, R.; Kuper, F.; Løvik, M.; Nygaard, U.C. Transmaternal bisphenol A exposure accelerates diabetes type 1 development in NOD mice. Toxicol. Sci. 2014, 137, 311–323. [Google Scholar] [CrossRef] [PubMed]
- Trepanowski, J.F.; Mey, J.; Varady, K.A. Fetuin-A: A novel link between obesity and related complications. Int. J. Obes. 2015, 39, 734–741. [Google Scholar] [CrossRef] [PubMed]
- Lee, A.S.; Gibson, D.L.; Zhang, Y.; Sham, H.P.; Vallance, B.A.; Dutz, J.P. Gut barrier disruption by an enteric bacterial pathogen accelerates insulitis in NOD mice. Diabetologia 2010, 53, 741–748. [Google Scholar] [CrossRef] [PubMed]
- Sabatino, A.; Regolisti, G.; Cosola, C.; Gesualdo, L.; Fiaccadori, E. Intestinal microbiota in type 2 diabetes and chronic kidney disease. Curr. Diabetes Rep. 2017, 17, 16. [Google Scholar] [CrossRef] [PubMed]
- Mejia-Leon, M.E.; Barca, A.M. Diet, microbiota and immune system in type 1 diabetes development and evolution. Nutrients 2015, 7, 9171–9184. [Google Scholar] [CrossRef] [PubMed]
- Knip, M.; Siljander, H. The role of the intestinal microbiota in type 1 diabetes mellitus. Nat. Rev. Endocrinol. 2016, 12, 154–167. [Google Scholar] [CrossRef] [PubMed]
- Guilloteau, P.; Martin, L.; Eeckhaut, V.; Ducatelle, R.; Zabielski, R.; Van Immerseel, F. From the gut to the peripheral tissues: The multiple effects of butyrate. Nutr. Res. Rev. 2010, 23, 366–384. [Google Scholar] [CrossRef] [PubMed]
- Zhou, L.; Zhang, M.; Wang, Y.; Dorfman, R.G.; Liu, H.; Yu, T.; Chen, X.; Tang, D.; Xu, L.; Yin, Y.; et al. Faecalibacterium prausnitzii produces butyrate to maintain th17/treg balance and to ameliorate colorectal colitis by inhibiting histone deacetylase 1. Inflamm. Bowel Dis. 2018. [Google Scholar] [CrossRef] [PubMed]
- De Andrés, J.; Manzano, S.; García, C.; Rodríguez, J.M.; Espinosa-Martos, I.; Jiménez, E. Modulatory effect of three probiotic strains on infants’ gut microbial composition and immunological parameters on a placebo-controlled, double-blind, randomised study. Benef. Microbes 2018, 9, 573–584. [Google Scholar]
- Van den Abbeele, P.; Belzer, C.; Goossens, M.; Kleerebezem, M.; De Vos, W.M.; Thas, O.; De Weirdt, R.; Kerckhof, F.M.; Van de Wiele, T. Butyrate-producing Clostridium cluster XIVa species specifically colonize mucins in an in vitro gut model. ISME J. 2013, 7, 949–961. [Google Scholar] [CrossRef] [PubMed]
- Qin, J.; Li, Y.; Cai, Z.; Li, S.; Zhu, J.; Zhang, F.; Liang, S.; Zhang, W.; Guan, Y.; Shen, D.; et al. A metagenome-wide association study of gut microbiota in type 2 diabetes. Nature 2012, 490, 55–60. [Google Scholar] [CrossRef] [PubMed]
- Brown, L.M. Helicobacter pylori: Epidemiology and routes of transmission. Epidemiol. Rev. 2000, 22, 283–297. [Google Scholar] [CrossRef] [PubMed]
- Devaraj, S.; Hemarajata, P.; Versalovic, J. The human gut microbiome and body metabolism: Implications for obesity and diabetes. Clin. Chem. 2013, 59, 617–628. [Google Scholar] [CrossRef] [PubMed]
- Hippe, B.; Remely, M.; Aumueller, E.; Pointner, A.; Magnet, U.; Haslberger, A.G. Faecalibacterium prausnitzii phylotypes in type two diabetic, obese, and lean control subjects. Benef. Microbes 2016, 7, 511–517. [Google Scholar] [CrossRef] [PubMed]
- Burrows, M.P.; Volchkov, P.; Kobayashi, K.S.; Chervonsky, A.V. Microbiota regulates type 1 diabetes through Toll-like receptors. Proc. Natl. Acad. Sci. USA 2015, 112, 9973–9977. [Google Scholar] [CrossRef] [PubMed]
- Karlsson, F.H.; Tremaroli, V.; Nookaew, I.; Bergstrom, G.; Behre, C.J.; Fagerberg, B.; Nielsen, J.; Backhed, F. Gut metagenome in European women with normal, impaired and diabetic glucose control. Nature 2013, 498, 99–103. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Shen, D.; Fang, Z.; Jie, Z.; Qiu, X.; Zhang, C.; Chen, Y.; Ji, L. Human gut microbiota changes reveal the progression of glucose intolerance. PLoS ONE 2013, 8, e71108. [Google Scholar] [CrossRef] [PubMed]
- Giongo, A.; Gano, K.A.; Crabb, D.B.; Mukherjee, N.; Novelo, L.L.; Casella, G.; Drew, J.C.; Ilonen, J.; Knip, M.; Hyöty, H.; Veijola, R.; et al. Toward defining the autoimmune microbiome for type 1 diabetes. ISME J. 2011, 5, 82–91. [Google Scholar] [CrossRef] [PubMed]
- Murri, M.; Leiva, I.; Gomez-Zumaquero, J.M.; Inahones, F.J.; Cardona, F.; Soriguer, F.; Queipo-Ortuño, M.I. Gut microbiota in children with type-1 diabetes differs from that in healthy children: A case-control study. BMC Med. 2013, 11, 46. [Google Scholar] [CrossRef] [PubMed]
- Soyucen, E.; Gulcan, A.; Aktuglu-Zeybek, A.C.; Onal, H.; Kiykim, E.; Aydin, A. Differences in the gut microbiota of healthy children and those with type-1 diabetes. Pediatr. Int. 2014, 56, 336–343. [Google Scholar] [CrossRef] [PubMed]
- McLean, M.H.; Dieguez, D., Jr.; Miller, L.M.; Young, H.A. Does the microbiota play a role in the pathogenesis of autoimmune diseases? Gut 2015, 64, 332–341. [Google Scholar] [CrossRef] [PubMed]
- Xie, L.; Li, Q.; Dong, R.; Zhao, K.; Feng, Y.; Bao, Z.; Zhou, M. Critical regulation of inflammation via class A scavenger receptor. Int. J. Chron. Obstruct. Pulmon. Dis. 2018, 13, 1145–1155. [Google Scholar] [CrossRef] [PubMed]
- Loeser, K.; Seemann, S.; König, S.; Lenhardt, I.; Abdel-Tawab, M.; Koeberle, A.; Werz, O.; Lupp, A. Protective effect of Casperome®, an orally bioavailable frankincense extract, on lipopolysaccharide-induced systemic inflammation in mice. Front. Pharmacol. 2018, 9, 387. [Google Scholar] [CrossRef] [PubMed]
- Geng, S.; Wang, S.; Zhu, W.; Xie, C.; Li, X.; Wu, J.; Zhu, J.; Jiang, Y.; Yang, X.; Li, Y.; et al. Curcumin attenuates BPA-induced insulin resistance in HepG2 cells through suppression of JNK/p38 pathways. Toxicol. Lett. 2017, 272, 75–83. [Google Scholar] [CrossRef] [PubMed]
- Cani, P.D.; Delzenne, N.M. The gut microbiome as a therapeutic target. Pharmacol. Ther. 2011, 130, 202–212. [Google Scholar] [CrossRef] [PubMed]
- Hochdorfer, T.; Tiedje, C.; Stumpo, D.J.; Blackshear, P.J.; Gaestel, M.; Huber, M. LPS-induced production of TNF-a and IL-6 in mast cells is dependent on p38 but independent of TTP. Cell Signal. 2013, 25, 1339–1347. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Liu, Y.X. LncRNA HOTTIP improves diabetic retinopathy by regulating the p38-MAPK pathway. Eur. Rev. Med. Pharmacol. Sci. 2018, 22, 2941–2948. [Google Scholar] [PubMed]
- Shen, K.P.; Lin, H.L.; Yen, H.W.; Hsieh, S.L.; An, L.M.; Wu, B.N. Eugenosedin-A improves glucose metabolism and inhibits MAPKs expression in streptozotocin/nicotinamide-induced diabetic rats. Kaohsiung J. Med. Sci. 2018, 34, 142–149. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Zhao, L.G.; Wu, Q.J.; Ma, X.; Xiang, Y.B. Association between dietary fiber and lower risk of all-cause mortality: A meta-analysis of cohort studies. Am. J. Epidemiol. 2015, 181, 83–91. [Google Scholar] [CrossRef] [PubMed]
- Amyot, J.; Semache, M.; Ferdaoussi, M.; Fontés, G.; Poitout, V. Lipopolysaccharides impair insulin gene expression in isolated islets of Langerhans via Toll-Like Receptor-4 and NF-κB signalling. PLoS ONE 2012, 7, e36200. [Google Scholar] [CrossRef] [PubMed]
- Haenen, D.; Zhang, J.; Souza da Silva, C.; Bosch, G.; van der Meer, I.M.; van Arkel, J.; van den Borne, J.J.; Pérez Gutiérrez, O.; Smidt, H.; Kemp, B.; et al. A diet high in resistant starch modulates microbiota composition, SCFA concentrations, and gene expression in pig intestine. J. Nutr. 2013, 143, 274–283. [Google Scholar] [CrossRef] [PubMed]
- Castillo, M.; Skene, G.; Roca, M.; Anguita, M.; Badiola, I.; Duncan, S.H.; Flint, H.J.; Martín-Orúe, S.M. Application of 16S rRNA gene-targetted fluorescence in situ hybridization and restriction fragment length polymorphism to study porcine microbiota along the gastrointestinal tract in response to different sources of dietary fibre. FEMS Microbiol. Ecol. 2007, 59, 138–146. [Google Scholar] [CrossRef] [PubMed]
- Meimandipour, A.; Shuhaimi, M.; Soleimani, A.F.; Azhar, K.; Hair-Bejo, M.; Kabeir, B.M.; Javanmard, A.; Muhammad Anas, O.; Yazid, A.M. Selected microbial groups and short-chain fatty acids profile in a simulated chicken cecum supplemented with two strains of Lactobacillus. Poult. Sci. 2010, 89, 470–476. [Google Scholar] [CrossRef] [PubMed]
- Oikonomou, G.; Teixeira, A.G.; Foditsch, C.; Bicalho, M.L.; Machado, V.S.; Bicalho, R.C. Associations of Faecalibacterium species with health and growth fecal microbial diversity in pre-weaned dairy calves as described by pyrosequencing of metagenomic 16S rDNA. PLoS ONE 2013, 8, e63157. [Google Scholar] [CrossRef] [PubMed]
- Nava, G.M.; Stappenbeck, T.S. Diversity of the autochthonous colonic microbiota. Gut Microbes 2011, 2, 99–104. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Siles, M.; Enrich-Capó, N.; Aldeguer, X.; Sabat-Mir, M.; Duncan, S.H.; Garcia-Gil, L.J.; Martinez-Medina, M. Alterations in the abundance and co-occurrence of Akkermansia muciniphila and Faecalibacterium prausnitzii in the colonic mucosa of inflammatory bowel disease subjects. Front. Cell Infect. Microbiol. 2018, 8, 281. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Siles, M.; Martinez-Medina, M.; Busquets, D.; Sabat-Mir, M.; Duncan, S.H.; Flint, H.J.; Aldeguer, X.; Garcia-Gil, L.J. Mucosa-associated Faecalibacterium prausnitzii and Escherichia coli co-abundance can distinguish irritable bowel syndrome and inflammatory bowel disease phenotypes. Int. J. Med. Microbiol. 2014, 304, 464–475. [Google Scholar] [CrossRef] [PubMed]
- Eppinga, H.; Sperna Weiland, C.J.; Thio, H.B.; van der Woude, C.J.; Nijsten, T.E.; Peppelenbosch, M.P.; Konstantinov, S.R. Similar depletion of protective Faecalibacterium prausnitzii in psoriasis and inflammatory bowel disease, but not in Hidradenitis Suppurativa. J. Crohns Colitis 2016, 10, 1067–1075. [Google Scholar] [CrossRef] [PubMed]
- Barkas, F.; Liberopoulos, E.; Kei, A.; Elisaf, M. Electrolyte and acid-base disorders in inflammatory bowel disease. Ann. Gastroenterol. 2013, 26, 23–28. [Google Scholar] [PubMed]
- Foditsch, C.; Santos, T.M.; Teixeira, A.G.; Pereira, R.V.; Dias, J.M.; Gaeta, N.; Bicalho, R.C. Isolation and characterization of Faecalibacterium prausnitzii from calves and piglets. PLoS ONE 2014, 9, e116465. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Siles, M.; Martinez-Medina, M.; Surís-Valls, R.; Aldeguer, X.; Sabat-Mir, M.; Duncan, S.H.; Flint, H.J.; Garcia-Gil, L.J. Changes in the abundance of Faecalibacterium prausnitzii phylogroups I and II in the intestinal mucosa of inflammatory bowel disease and patients with colorectal cancer. Inflamm. Bowel Dis. 2016, 22, 28–41. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Siles, M.; Martinez-Medina, M.; Abellà, C.; Busquets, D.; Sabat-Mir, M.; Duncan, S.H.; Aldeguer, X.; Flint, H.J.; Garcia-Gil, L.J. Mucosa-associated Faecalibacterium prausnitzii phylotype richness is reduced in patients with inflammatory bowel disease. Appl. Environ. Microbiol. 2015, 81, 7582–7592. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Siles, M.; Khan, T.M.; Duncan, S.H.; Harmsen, H.J.; Garcia-Gil, L.J.; Flint, H.J. Cultured representatives of two major phylogroups of human colonic Faecalibacterium prausnitzii can utilize pectin, uronic acids, and host-derived substrates for growth. Appl. Environ. Microbiol. 2012, 78, 420–428. [Google Scholar] [CrossRef] [PubMed]
- Martín, R.; Bermúdez-Humarán, L.G.; Langella, P. Searching for the bacterial effector: The example of the multi-skilled commensal bacterium Faecalibacterium prausnitzii. Front. Microbiol. 2018, 9, 346. [Google Scholar] [CrossRef] [PubMed]
- Martín, R.; Miquel, S.; Chain, F.; Natividad, J.M.; Jury, J.; Lu, J.; Sokol, H.; Theodorou, V.; Bercik, P.; Verdu, E.F.; et al. Faecalibacterium prausnitzii prevents physiological damages in a chronic low-grade inflammation murine model. BMC Microbiol. 2015, 15, 67. [Google Scholar] [CrossRef] [PubMed]
- Zhang, M.; Qiu, X.; Zhang, H.; Yang, X.; Hong, N.; Yang, Y.; Chen, H.; Yu, C. Faecalibacterium prausnitzii inhibits interleukin-17 to ameliorate colorectal colitis in rats. PLoS ONE 2014, 9, e109146. [Google Scholar] [CrossRef] [PubMed]
- Quévrain, E.; Maubert, M.A.; Michon, C.; Chain, F.; Marquant, R.; Tailhades, J.; Miquel, S.; Carlier, L.; Bermúdez-Humarán, L.G.; Pigneur, B.; et al. Identification of an anti-inflammatory protein from Faecalibacterium prausnitzii, a commensal bacterium deficient in Crohn’s disease. Gut 2016, 65, 415–425. [Google Scholar] [CrossRef] [PubMed]
- Sokol, H.; Pigneur, B.; Watterlot, L.; Lakhdari, O.; Bermúdez-Humarán, L.G.; Gratadoux, J.J.; Blugeon, S.; Bridonneau, C.; Furet, J.P.; Corthier, G.; et al. Faecalibacterium prausnitzii is an anti-inflammatory commensal bacterium identified by gut microbiota analysis of Crohn disease patients. Proc. Natl. Acad. Sci. USA 2008, 105, 16731–16736. [Google Scholar] [CrossRef] [PubMed]
- Qiu, X.; Zhang, M.; Yang, X.; Hong, N.; Yu, C. Faecalibacterium prausnitzii upregulates regulatory T cells and anti-inflammatory cytokines in treating TNBS-induced colitis. J. Crohns Colitis 2013, 7, e558–e568. [Google Scholar] [CrossRef] [PubMed]
- Breyner, N.M.; Michon, C.; de Sousa, C.S.; Vilas Boas, P.B.; Chain, F.; Azevedo, V.A.; Langella, P.; Chatel, J.M. Microbial anti-inflammatory molecule (MAM) from Faecalibacterium prausnitzii shows a protective effect on DNBS and DSS-induced colitis model in mice through inhibition of NF-κB Pathway. Front. Microbiol. 2017, 8, 114. [Google Scholar] [CrossRef] [PubMed]
- Martín, R.; Miquel, S.; Benevides, L.; Bridonneau, C.; Robert, V.; Hudault, S.; Chain, F.; Berteau, O.; Azevedo, V.; Chatel, J.M.; et al. Functional Characterization of Novel Faecalibacterium prausnitzii Strains Isolated from Healthy Volunteers: A Step Forward in the Use of F. prausnitzii as a Next-Generation Probiotic. Front. Microbiol. 2017, 8, 1226. [Google Scholar] [CrossRef] [PubMed]
- Heinken, A.; Khan, M.T.; Paglia, G.; Rodionov, D.A.; Harmsen, H.J.; Thiele, I. Functional metabolic map of Faecalibacterium prausnitzii, a beneficial human gut microbe. J. Bacteriol. 2014, 196, 3289–3302. [Google Scholar] [CrossRef] [PubMed]
- McGill, C.R.; Fulgoni, V.L.; Devareddy, L. Ten-year trends in fiber and whole grain intakes and food sources for the United States population: National Health and Nutrition Examination Survey 2001–2010. Nutrients 2015, 7, 1119–1130. [Google Scholar] [CrossRef] [PubMed]
- Furusawa, Y.; Obata, Y.; Fukuda, S.; Endo, T.A.; Nakato, G.; Takahashi, D.; Nakanishi, Y.; Uetake, C.; Kato, K.; Kato, T.; et al. Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 2013, 504, 446–450. [Google Scholar] [CrossRef] [PubMed]
- Haro, C.; Garcia-Carpintero, S.; Alcala-Diaz, J.F.; Gomez-Delgado, F.; Delgado-Lista, J.; Perez-Martinez, P.; Rangel Zuniga, O.A.; Quintana-Navarro, G.M.; Landa, B.B.; Clemente, J.C.; et al. The gut microbial community in metabolic syndrome patients is modified by diet. J. Nutr. Biochem. 2016, 27, 27–31. [Google Scholar] [CrossRef] [PubMed]
- Sokol, H.; Seksik, P.; Furet, J.P.; Firmesse, O.; Nion-Larmurier, I.; Beaugerie, L.; Cosnes, J.; Corthier, G.; Marteau, P.; Doré, J. Low counts of Faecalibacterium prausnitzii in colitis microbiota. Inflamm. Bowel Dis. 2009, 15, 1183–1189. [Google Scholar] [CrossRef] [PubMed]
- Scott, K.P.; Gratz, S.W.; Sheridan, P.O.; Flint, H.J.; Duncan, S.H. The influence of diet on the gut microbiota. Pharmacol. Res. 2013, 69, 52–60. [Google Scholar] [CrossRef] [PubMed]
- Miquel, S.; Martín, R.; Rossi, O.; Bermúdez-Humarán, L.G.; Chatel, J.M.; Sokol, H.; Thomas, M.; Wells, J.M.; Langella, P. Faecalibacterium prausnitzii and human intestinal health. Curr. Opin. Microbiol. 2013, 16, 255–261. [Google Scholar] [CrossRef] [PubMed]
- Thorburn, A.N.; Macia, L.; Mackay, C.R. Diet, metabolites, and “western-lifestyle” inflammatory diseases. Immunity 2014, 40, 833–842. [Google Scholar] [CrossRef] [PubMed]
- Lu, Y.; Fan, C.; Liang, A.; Fan, X.; Wang, R.; Li, P.; Qi, K. Effects of SCFA on the DNA methylation pattern of adiponectin and resistin in high-fat-diet-induced obese male mice. Br. J. Nutr. 2018, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Mathis, D.; Benoist, C. The influence of the microbiota on type-1 diabetes: On the threshold of a leap forward in our understanding. Immunol. Rev. 2012, 245, 239–249. [Google Scholar] [CrossRef] [PubMed]
- Roesch, L.F.; Lorca, G.L.; Casella, G.; Giongo, A.; Naranjo, A.; Pionzio, A.M.; Li, N.; Mai, V.; Wasserfall, C.H.; Schatz, D.; et al. Culture-independent identification of gut bacteria correlated with the onset of diabetes in a rat model. ISME J. 2009, 3, 536–548. [Google Scholar] [CrossRef] [PubMed]
- Markle, J.G.; Frank, D.N.; Mortin-Toth, S.; Robertson, C.E.; Feazel, L.M.; Rolle-Kampczyk, U.; von Bergen, M.; McCoy, K.D.; Macpherson, A.J.; Danska, J.S. Sex differences in the gut microbiome drive hormonedependent regulation of autoimmunity. Science 2013, 339, 1084–1088. [Google Scholar] [CrossRef] [PubMed]
- Yurkovetskiy, L.; Burrows, M.; Khan, A.A.; Graham, L.; Volchkov, P.; Becker, L.; Antonopoulos, D.; Umesaki, Y.; Chervonsky, A.V. Gender bias in autoimmunity is influenced by microbiota. Immunity 2013, 39, 400–412. [Google Scholar] [CrossRef] [PubMed]
- Furet, J.; Kong, L.; Tap, J.; Poitou, C.; Basdevant, A.; Bouillot, J.L.; Mariat, D.; Corthier, G.; Doré, J.; Henegar, C.; et al. Differential adaption of human gut microbiota to bariatric surgery induced weight loss: Links with metabolic and low-grade inflammation markers. Diabetes 2010, 59, 3049–3057. [Google Scholar] [CrossRef] [PubMed]
- Gerritsen, J.; Smidt, H.; Rijkers, G.T.; De Vos, W.M. Intestinal microbiota in human health and disease: The impact of probiotics. Genes Nutr. 2011, 6, 209–240. [Google Scholar] [CrossRef] [PubMed]
- Fujimoto, T.; Imaeda, H.; Takahashi, K.; Kasumi, E.; Bamba, S.; Fujiyama, Y.; Andoh, A. Decreased abundance of Faecalibacterium prausnitzii in the gut microbiota of Crohn’s disease. J. Gastroenterol. Hepatol. 2013, 28, 613–619. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Lian, F.; Zhao, L.; Chen, X.; Zhang, X.; Guo, Y.; Zhang, C.; Zhou, Q.; Xue, Z.; Pang, X.; et al. Structural modulation of gut microbiota during alleviation of type 2 diabetes with a Chinese herbal formula. ISME J. 2015, 9, 552–562. [Google Scholar] [CrossRef] [PubMed]
- Balamurugan, R.; George, G.; Kabeerdoss, J.; Hepsiba, J.; Chandragunasekaran, A.M.; Ramakrishna, B.S. Quantitative differences in intestinal Faecalibacterium prausnitzii in obese Indian children. Br. J. Nutr. 2010, 103, 335–338. [Google Scholar] [CrossRef] [PubMed]
- Hong, J.; Jia, Y.; Pan, S.; Jia, L.; Li, H.; Han, Z.; Cai, D.; Zhao, R. Butyrate alleviates high fat diet-induced obesity through activation of adiponectin-mediated pathway and stimulation of mitochondrial function in the skeletal muscle of mice. Oncotarget 2016, 7, 56071–56082. [Google Scholar] [CrossRef] [PubMed]
- Lin, H.V.; Frassetto, A.; Kowalik, E.J., Jr.; Nawrocki, A.R.; Lu, M.M.; Kosinski, J.R.; Hubert, J.A.; Szeto, D.; Yao, X.; Forrest, G.; et al. Butyrate and propionate protect against diet-induced obesity and regulate gut hormones via free fatty acid receptor 3-independent mechanisms. PLoS ONE 2012, 7, e35240. [Google Scholar] [CrossRef] [PubMed]
- Vrieze, A.; Van Nood, E.; Holleman, F.; Salojärvi, J.; Kootte, R.S.; Bartelsman, J.F.; Dallinga-Thie, G.M.; Ackermans, M.T.; Serlie, M.J.; Oozeer, R.; et al. Transfer of intestinal microbiota from lean donors increases insulin sensitivity in individuals with metabolic syndrome. Gastroenterology 2012, 143, 913–917. [Google Scholar] [CrossRef] [PubMed]
- Gao, Z.; Yin, J.; Zhang, J.; Ward, R.E.; Martin, R.J.; Lefevre, M.; Cefalu, W.T.; Ye, J. Butyrate improves insulin sensitivity and increases energy expenditure in mice. Diabetes 2009, 58, 1509–1517. [Google Scholar] [CrossRef] [PubMed]
- Van Nood, E.; Vrieze, A.; Nieuwdorp, M.; Fuentes, S.; Zoetendal, E.G.; de Vos, W.M.; Visser, C.E.; Kuijper, E.J.; Bartelsman, J.F.; Tijssen, J.G.; et al. Duodenal infusion of donor feces for recurrent Clostridium difficile. N. Engl. J. Med. 2013, 368, 407–415. [Google Scholar] [CrossRef] [PubMed]
| Diet Components/Sources | Consumption of Dietary Sources | Changes in the Gut Bacteria |
|---|---|---|
| Carbohydrates: Indigestible complex oligosaccharides | Human milk glycans [69,70,71] | ↑Bifidobacterium infantis, Bacteroides |
| Resistant starch (type 2,3,4) [72,73] | RS2: ↑Ruminococcus bromii, ↑Eubacterium rectale RS3: ↑Ruminococcus bromii, ↑Ocillobactor, ↑Eubacterium rectale RS4: ↑Bifidobacterium adoloscentis, ↑Parabacteroides distasonis | |
| Resistant maltodextrin [74] | ↑Ruminococcus, ↑Eubacterium, ↑Lachnospiraceae, ↑Bacteriodes, ↑Holdemania, ↑Faecalibacterium | |
| Jerusalem artichoke inulin [75,76,77] | ↑Bifidobacterium, ↑Lactobacillus, ↑Enterococcus, ↑Faecalibacterium prausnitzii, ↑Clostridial cluster XIVa | |
| Inulin and partially hydrolysed guar gum, fructo-oligosaccharides, Long chain inulin, Xylo- oligosaccharides [78,79,80,81] | ↑Lactobacillus/Enterococcus, ↑Bifidobacterium, ↓Clostridium, ↓Bacteroides, ↓Prevotell | |
| Galacto-oligosaccharides, fructo-oligosaccharides [82,83] | ↑Bifidobacterium | |
| Polydextrose and soluble corn fibre [84,85] | ↑Clostridiaceae, ↑Faecalibacterium prausnitzii, ↑Phasolarciobacterium, ↑Dialister, ↑Lactobacillus, ↑Ruminococcus intestinalis, ↓Eubacteiaceae, Actinobacteria | |
| Simple sugars Digestible carbohydrates | Arabinoxylans [86,87] | ↑Bifidobacteria, ↑Eubacteriumrectale, ↑Roseburia/Eubacterium, ↑Faecalibacterium prausnitzii, ↑Bacteroides |
| Sugars in food [61] | ↑Prevotella | |
| Insoluble complex carbohydrates | Apple [88] | ↑Bifidobacteria, ↓Clostridia perfringens |
| Kiwifruit [89] | ↑Bifidobacteria, ↑Lactobacillus, ↑Clostridia | |
| Banana [90] | ↑Bifidobacterium | |
| Insoluble non-starch polysaccharides | Cereal cellulose [91,92,93,94] | ↑Ruminococcus flavefaciens, ↑Clostridium xylanolyticum |
| Cereal amylose [91,92,93,94] | ↑Clostridium butyricum | |
| Amylopectin and Starch [91,92,93,94,95,96,97,98] | ↑Clostridium ramosum, ↑Clostridium cluster XIVa, ↑Bacteroides | |
| ↑Ruminococcus bromii, ↑Eubacterium rectale, ↑Roseburia | ||
| ↑E. rectale, ↑Roseburia | ||
| Dietary fiber [99] | ↑Clostridium, ↑Bacteroides, ↑Bacillus subtilis, ↑Bifidobacterium,↑Fusobacterium | |
| Soybean, radishes, cabbage, fish, seaweed and green tea [41] Western diet (high in meat) [41] | ↓Bacteroides fragilis; ↑Lactobacillus, ↑E. coli, ↑Proteus, ↑Klebsiella, ↑Staphylococcus, ↑Streptococci, ↑Clostridium, ↑Eubacterium, ↑Ruminococcus | |
| Cereal (millet, grain, sorghum), Legumes (black-eyed peas) and Vegetables [100] | ↑Bacteroidetes, ↑Prevotella, ↑Xylanibacter, ↓Firmicutes | |
| Whole grain wheat [101,102] | ↑Bifidobacteria animalis, ↑Roseburia, ↑Bacteroides, ↑Prevotella, ↑Clostridium ↑Lactobacillus/Enterococci | |
| Maize-based whole grains and cereals [103] | ↑Bifidobacterium spp., ↑Atobobium cluster spp. | |
| Whole grain barley, brown rice or mix [104] | ↑Firmicutes, ↑Blautia, ↑Roseburia, ↑Bifidobacterium, ↓Bacteroides, ↓odoribacter | |
| Rye bread [105] | ↓Bryantella formatexiagans | |
| Wild blueberry drink [106,107] | ↑Bifidobacterium spp., ↑Lactobacillus acidophilus, ↑B. longum sub sp. infantis | |
| Red wine, dealcoholized red wine, gin [108] | ↑Bacteroidetes, ↑Bacteroides, ↑B. uniformis, ↑Firmicutes, ↑E. rectale group, ↑Prevotella, ↑Fusobacteria, ↑Proteobacteria, ↑Bifidobacterium, ↑Eggerthellalenta, ↑Enterococcus | |
| Almonds and pistachios [109,110] | ↑Bifidobacterium, ↑Lactobacillus spp., ↓Lactic acid bacteria, ↓Clostridumperfringens | |
| Fat and fatty acids | High-fat diet [111,112] | ↑E. rectal, ↑C. coccoides, ↓Bifidobacterium, ↓Bacteroides, |
| Protein | Meat [113,114] | ↑Bacteroides, ↑Bifidobacterium, ↑Peptococcus, ↑Lactobacillus, ↑Clostridium cluster XIVa, ↑Clostridium coccoides, ↑Roseburia, ↑E. rectal |
| A variety of amino acids and saturated fats [61,100] | ↑Bacteroides, ↑Clostridium | |
| Whey protein isolate [115] | ↑Lactobacillaceae, ↓Clostridiaceae | |
| Chickpea or raffinose [116] | ↓Clostridium cluster I, II XI | |
| Soymilk, low glycinin soymilk, bovine milk [117] | ↑Bacteroides, ↑prevotella, ↑Lactobacillus, ↓Bifidobacterium | |
| Non-nutrients (Phytochemicals) | Red pepper (Capsicum annuum) and Garlic (Allium sativum) [118,119] | ↓Bacillus cereus, ↑B. subtilis, ↑C. tetani, ↑Helicobacter pylori |
| Tea polyphenols [120] | ↓Bacteroides, ↓Clostridium perfringens, ↓C. difficile, ↓E. coli, ↓Salmonella typhimurium | |
| Wild blueberries (Vaccinium angustifolium) [106,121] | ↑Bifidobacterium, ↑Lactobacillus acidophilus, ↑Bacteroides, ↑Prevotella, ↑Enterococcus, ↑C. coccoides | |
| Coffee (catechin and epicatechin) [122] | ↑C. coccoides, ↑E. rectale group, ↑E. coli, ↓C. histolyticum | |
| Dietary polyphenol [123] | ↑Bifidobacterium, ↑Lactobacillus | |
| Wine (resveratrol) [124,125] | ↑Bifidobacterium, ↑Lactobacillus, ↓Proteus mirabilis | |
| Berries (anthocyanins) [126,127] | ↓Staphylococcus, ↓Salmonella, ↓H. pylori, ↓B. cereus |
| Name of the Prevalence Bacteria | Model | References |
|---|---|---|
| ↑Akkermansia muciniphila, ↑Bacteroides intestinalis, ↑Bacteroides sp. ↑Clostridium bolteae, ↑Clostridium ramosum, ↑Clostridium sp. HGF2, ↑Clostridium symbiosum, ↑Colstridium hathewayi, ↑Desulfovibrio sp., ↑Eggerthellalenta, ↑Escherichia coli | Human | [156] |
| ↑Bacteroides, ↑Prevotella, ↑Clostridia, ↑Betaproteobacteria, ↑Bacteroidetes/Firmicutes ratio, ↓Firmicutes, ↓Clostridia, ↓Eubacterium rectale | Human | [157] |
| ↑ Bacteroidetes thetaiotaomicron, ↑Akkermansia muciniphila,↑E. coli | Human | [142,158] |
| ↓Faecalibacterium prausnitzii phylotypes | Human | [159] |
| ↑Betaproteobacteria, ↓Firmicutes (Clostridia) | Mice | [27] |
| ↓Bifidobacterium, ↓Bacteroides vulgatus | Human | [61] |
| ↓Bacteroidescaccae, ↓Eubacteriumrectale, ↓Faecalibacterium prausnitzii, ↓Roseburia intestinalis, ↓Roseburiainulinivorans, ↑Clostridium hathewayi, ↑Clostridium ramosum, ↑Clostridium symbiosum, ↑Eggerthellalenta, ↑Escherichia coli, ↑Akkermansia muciniphila, ↑Desulfovibrio, ↑Clostridiales sp. SS3/4, | Mice | [160] |
| ↑Lactobacillus spp., ↑Clostridium clostridioforme, ↓Roseburia, ↓Clostridium spp. | Human | [161] |
| ↑Akkermansia muciniphila, ↑Faecalibacterium prausnitzii, ↓Bacteroides | Human | [162] |
| ↑Bifidobacterium, ↑Bacteroides, ↑Lactobacillus, ↑Lactococcus, ↑Streptococcus, ↑Veillonella, ↑Alistipes, ↓Prevotella, ↓Akkermansia, ↓Eubacterium, ↓Fusobacterium, ↓Anaerostipes, ↓Roseburia, ↓Subdoligranulum, ↓Faecalibacterium | Human | [13] |
| ↑Bacteroides, ↑Bacteroidesovatus, ↑Eubacterium, ↓Faecalibacterium, ↓Bacteroides vulgatus, ↓Bacteroidesfragilis | Human | [163] |
| ↑Bacteroides, ↑Veillonella, ↑Clostridium, ↑Prevotella, ↓Lactobacillus, ↓Bifidobacterium, ↓Blautiacoccoides, ↓Eubacteriumrectale | Human | [164] |
| ↑Cantidaalbicans, ↑Enterobacteriaceae, ↑Echerichia coli, ↓Bifidobacterium | Human | [165] |
| Gut-Associated Diseases | Findings | Implications | References |
|---|---|---|---|
| Diagnostic implications ofF. prausnitzii | |||
| Inflammatory bowel diseases | ↑ F. prausnitzii counts in feces | F. prausnitzii assay might play a potentially useful adjunct role in non-invasive screening and diagnosis of inflammatory bowel diseases | [181] |
| Inflammatory bowel diseases associated with skin disorders | ↓ F. prausnitzii and ↑ E. coli | F. prausnitzii assay aids to identify IBD-associated skin disorders | [182,183] |
| Crohn’s disease | ↑ F. prausnitzii counts with acidic stool | F. prausnitzii assay gives a promising diagnostic biomarker for early Crohn’s disease | [184] |
| Crohn’s disease | ↑ bilirubin concentrations along with F. prausnitzii counts with acidic stool | F. prausnitzii analysis contributes a promising diagnostic biomarker for Crohn’s disease | [185] |
| Colorectal cancer | ↓ F. prausnitzii counts in feces | F. prausnitzii assay holds great promising as a diagnostic biomarker for early colon cancer detection and monitoring and has considerable potential for developing an anticancer therapy | [186] |
| Ulcerative colitis | ↓ F. prausnitzii counts in feces | F. prausnitzii analysis contributes a promising diagnostic biomarker for Ulcerative colitis | [186] |
| Irritable bowel syndrome | ↓ F. prausnitzii counts in feces | F. prausnitzii phylotypes quantified as a putative biomarker and depicting the significance of the loss of these subtypes in Irritable bowel syndrome pathogenesis. | [187] |
| Crohn’s disease, ulcerative colitis, and colorectal cancer | ↓ F. prausnitzii phylogroup I was found in subjects with Crohn’s disease, ulcerative colitis, and colorectal cancer, whereas phylogroup II was specifically reduced in with Crohn’s disease. | Quantification of F. prausnitzii phylogroups and E. coli may help to identify gut disorders and to classify inflammatory bowel disease location. | [188] |
| Therapeutic implications of F. prausnitzii in gut-associated diseases | |||
| Gut-associated diseases | Treatment with F. prausnitzii as probiotics can inhibit gut-associated diseases, including malignancy | F. prausnitzii as next-generation probiotics might be useful in the treatment of various cancers with gut-associated diseases | [189] |
| Low-grade inflammation | Treatment with F. prausnitzii as probiotics exhibited intestinal permeability, tissue cytokines, and serotonin levels | F. prausnitzii might be beneficial effects on intestinal epithelial barrier impairment in a chronic low-grade inflammation model. | [190] |
| Inflammatory bowel diseases | Treatment with F. prausnitzii as probiotics showed plasma anti-Th17 cytokines (IL-10 and IL-12) and reduced IL-17 levels in both plasma and colonic mucosa, with ameliorated colonic colitis lesions | F. prausnitzii protected the colon mucosa against the development of Inflammatory bowel diseases and suggesting a promising therapy for Inflammatory bowel diseases. | [191] |
| Crohn’s disease | Seven peptides were identified in the F. prausnitzii culture, known as anti-inflammatory molecules. These molecules reduce the activation of the NF-kB pathway with a dose-dependent effect in the dinitrobenzene sulfonic acid induced-colitis model | F. prausnitzii protected the colon mucosa against the development of Inflammation and suggesting a promising treatment for Crohn’s disease | [192] |
| Ileal Crohn’s disease | Oral administration of F. prausnitzii as probiotics showed as anti-inflammatory properties. They reduce IL-1beta-induced NF-κB pathway in dinitrobenzene sulfonic acid induced-colitis model | F. prausnitzii as a probiotic is a promising strategy in Crohn’s disease | [193,194] |
| Ulcerative Colitis | Oral administration of F. prausnitzii reduced Th1, Th2, and Th17 immune response and increased TGFβ production. | F. prausnitzii as a probiotic is a promising strategy in Colitis | [195] |
| Crohn’s disease | Oral administration of F. prausnitzii as probiotics showed as anti-inflammatory properties. They induced IL-10, an anti-inflammatory cytokine, in peripheral blood mononuclear cells | F. prausnitzii strains could represent good candidates as next-generation probiotic. | [196] |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ganesan, K.; Chung, S.K.; Vanamala, J.; Xu, B. Causal Relationship between Diet-Induced Gut Microbiota Changes and Diabetes: A Novel Strategy to Transplant Faecalibacterium prausnitzii in Preventing Diabetes. Int. J. Mol. Sci. 2018, 19, 3720. https://doi.org/10.3390/ijms19123720
Ganesan K, Chung SK, Vanamala J, Xu B. Causal Relationship between Diet-Induced Gut Microbiota Changes and Diabetes: A Novel Strategy to Transplant Faecalibacterium prausnitzii in Preventing Diabetes. International Journal of Molecular Sciences. 2018; 19(12):3720. https://doi.org/10.3390/ijms19123720
Chicago/Turabian StyleGanesan, Kumar, Sookja Kim Chung, Jairam Vanamala, and Baojun Xu. 2018. "Causal Relationship between Diet-Induced Gut Microbiota Changes and Diabetes: A Novel Strategy to Transplant Faecalibacterium prausnitzii in Preventing Diabetes" International Journal of Molecular Sciences 19, no. 12: 3720. https://doi.org/10.3390/ijms19123720
APA StyleGanesan, K., Chung, S. K., Vanamala, J., & Xu, B. (2018). Causal Relationship between Diet-Induced Gut Microbiota Changes and Diabetes: A Novel Strategy to Transplant Faecalibacterium prausnitzii in Preventing Diabetes. International Journal of Molecular Sciences, 19(12), 3720. https://doi.org/10.3390/ijms19123720






