The Potential Role of a Soluble γ-Chain Cytokine Receptor as a Regulator of IL-7-Induced Lymphoproliferative Disorders
Abstract
1. Introduction
2. Results and Discussion
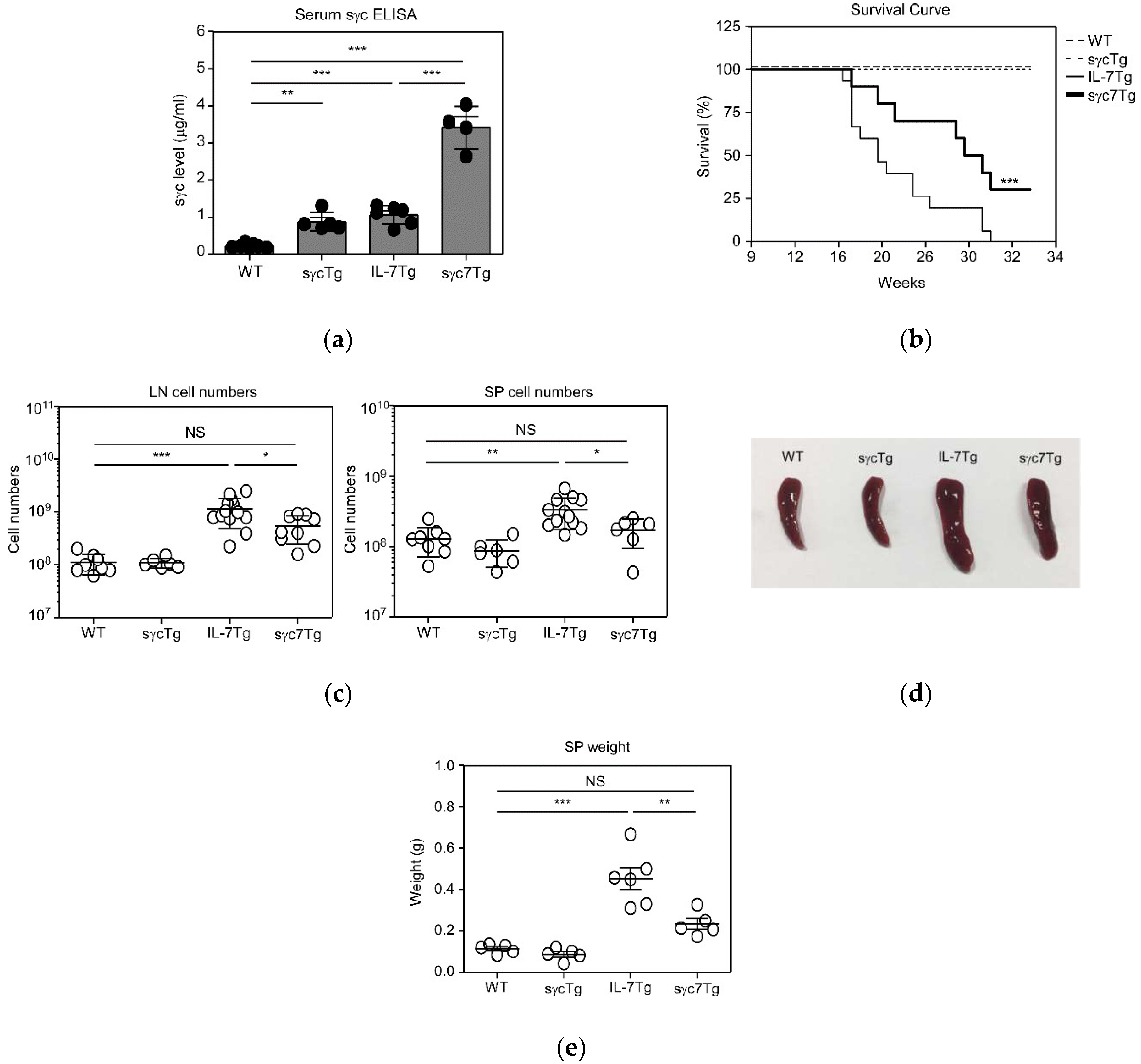
2.1. IL-7-Mediated LPDs Are Regulated by sγc Expression Level
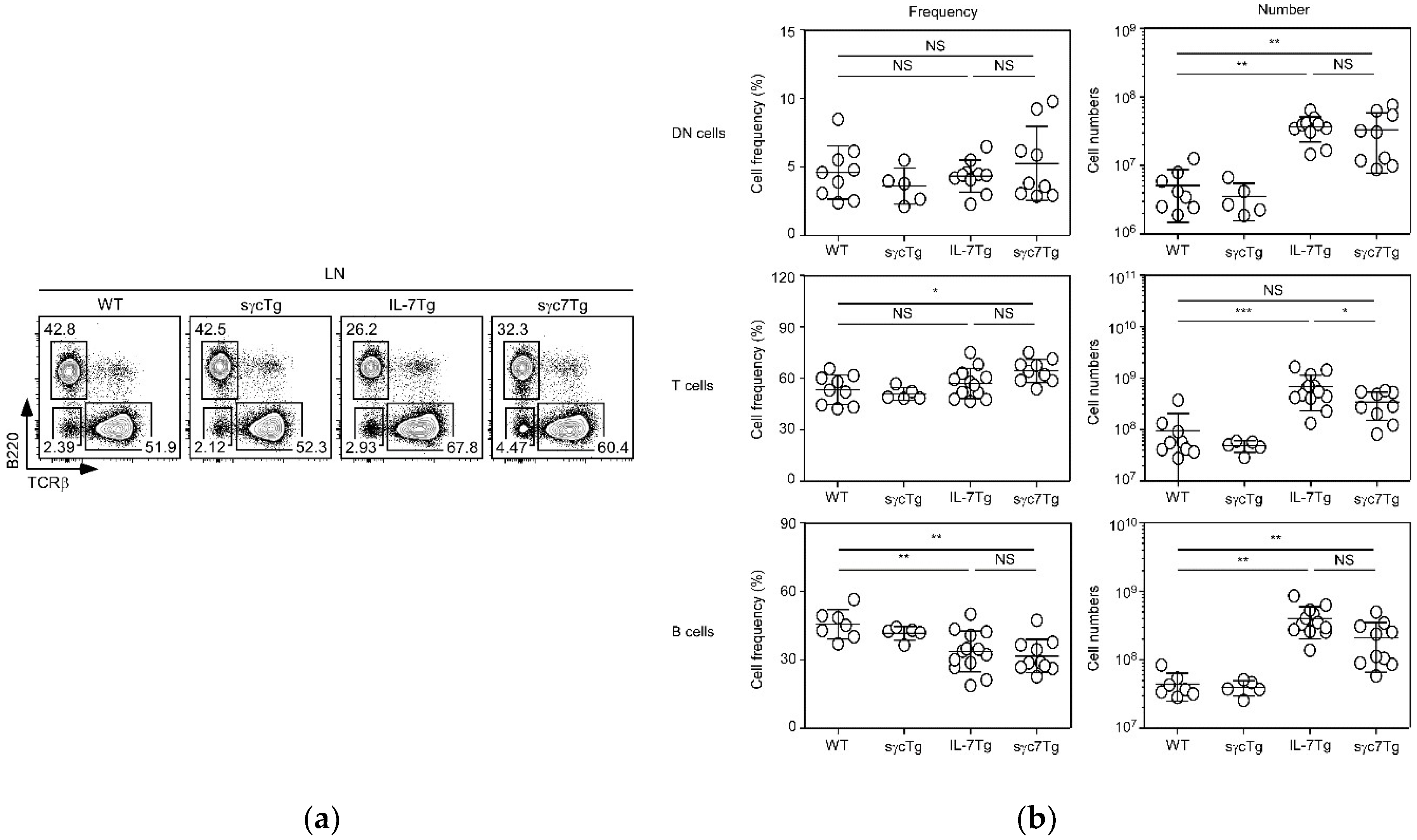
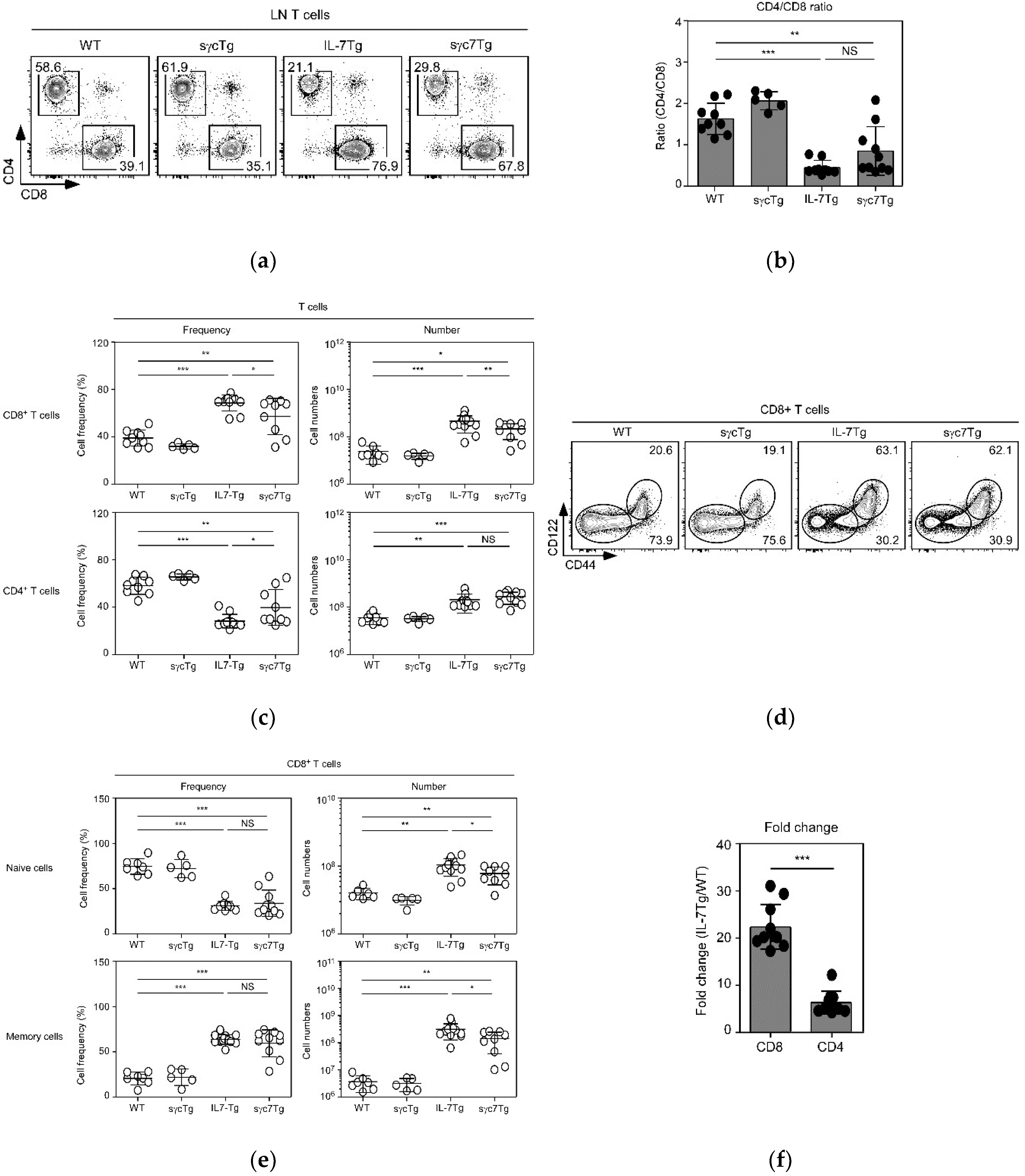
2.2. sγc Suppresses IL-7-Related Expansion of T Cells
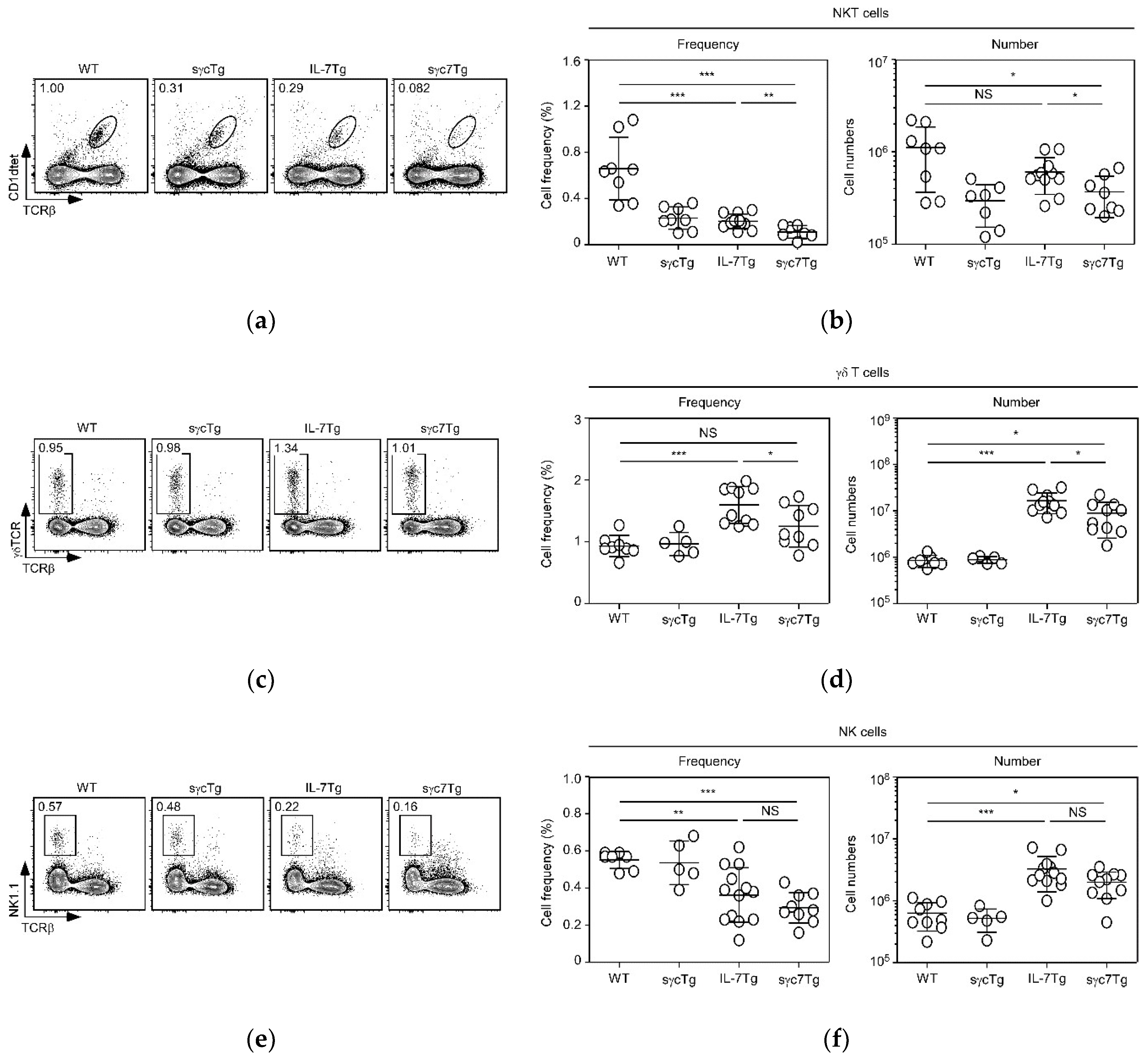
2.3. sγc Inhibits IL-7-Mediated Expansion of γδT and NKT Cells, but Not NK Cells
3. Materials and Methods
3.1. Animal
3.2. Flow Cytometry Analysis
3.3. Detection of sγc Levels
3.4. Statistical Analysis
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| IL | Interleukin |
| TCR | T-cell receptor |
| NK | Natural killer |
| SCID | Severe combined immunodeficiency |
References
- Hong, C.; Luckey, M.A.; Park, J.H. Intrathymic IL-7: The where, when, and why of IL-7 signaling during T cell development. Semin. Immunol. 2012, 24, 151–158. [Google Scholar] [CrossRef] [PubMed]
- Foss, H.D.; Hummel, M.; Gottstein, S.; Ziemann, K.; Falini, B.; Herbst, H.; Stein, H. Frequent expression of IL-7 gene transcripts in tumor cells of classical Hodgkin’s disease. Am. J. Pathol. 1995, 146, 33–39. [Google Scholar] [PubMed]
- Touw, I.; Pouwels, K.; van Agthoven, T.; van Gurp, R.; Budel, L.; Hoogerbrugge, H.; Delwel, R.; Goodwin, R.; Namen, A.; Lowenberg, B. Interleukin-7 is a growth factor of precursor B and T acute lymphoblastic leukemia. Blood 1990, 75, 2097–2101. [Google Scholar] [PubMed]
- Rochman, Y.; Spolski, R.; Leonard, W.J. New insights into the regulation of T cells by gamma(c) family cytokines. Nat. Rev. Immunol. 2009, 9, 480–490. [Google Scholar] [CrossRef] [PubMed]
- Abraham, N.; Ma, M.C.; Snow, J.W.; Miners, M.J.; Herndier, B.G.; Goldsmith, M.A. Haploinsufficiency identifies STAT5 as a modifier of IL-7-induced lymphomas. Oncogene 2005, 24, 5252–5257. [Google Scholar] [CrossRef] [PubMed]
- Rich, B.E.; Campos-Torres, J.; Tepper, R.I.; Moreadith, R.W.; Leder, P. Cutaneous lymphoproliferation and lymphomas in interleukin 7 transgenic mice. J. Exp. Med. 1993, 177, 305–316. [Google Scholar] [CrossRef] [PubMed]
- Lin, J.X.; Migone, T.S.; Tsang, M.; Friedmann, M.; Weatherbee, J.A.; Zhou, L.; Yamauchi, A.; Bloom, E.T.; Mietz, J.; John, S.; et al. The role of shared receptor motifs and common Stat proteins in the generation of cytokine pleiotropy and redundancy by IL-2, IL-4, IL-7, IL-13, and IL-15. Immunity 1995, 2, 331–339. [Google Scholar] [CrossRef]
- Pallard, C.; Stegmann, A.P.; van Kleffens, T.; Smart, F.; Venkitaraman, A.; Spits, H. Distinct roles of the phosphatidylinositol 3-kinase and STAT5 pathways in IL-7-mediated development of human thymocyte precursors. Immunity 1999, 10, 525–535. [Google Scholar] [CrossRef]
- Barata, J.T.; Silva, A.; Brandao, J.G.; Nadler, L.M.; Cardoso, A.A.; Boussiotis, V.A. Activation of PI3K is indispensable for interleukin 7-mediated viability, proliferation, glucose use, and growth of T cell acute lymphoblastic leukemia cells. J. Exp. Med. 2004, 200, 659–669. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Q.; Li, W.Q.; Aiello, F.B.; Mazzucchelli, R.; Asefa, B.; Khaled, A.R.; Durum, S.K. Cell biology of IL-7, a key lymphotrophin. Cytokine Growth Factor Rev. 2005, 16, 513–533. [Google Scholar] [CrossRef] [PubMed]
- Kelly, J.A.; Spolski, R.; Kovanen, P.E.; Suzuki, T.; Bollenbacher, J.; Pise-Masison, C.A.; Radonovich, M.F.; Lee, S.; Jenkins, N.A.; Copeland, N.G.; et al. Stat5 synergizes with T cell receptor/antigen stimulation in the development of lymphoblastic lymphoma. J. Exp. Med. 2003, 198, 79–89. [Google Scholar] [CrossRef] [PubMed]
- Bolotin, E.; Annett, G.; Parkman, R.; Weinberg, K. Serum levels of IL-7 in bone marrow transplant recipients: Relationship to clinical characteristics and lymphocyte count. Bone Marrow Transplant. 1999, 23, 783–788. [Google Scholar] [CrossRef] [PubMed]
- Hong, C.; Luckey, M.A.; Ligons, D.L.; Waickman, A.T.; Park, J.Y.; Kim, G.Y.; Keller, H.R.; Etzensperger, R.; Tai, X.; Lazarevic, V.; et al. Activated T cells secrete an alternatively spliced form of common gamma-chain that inhibits cytokine signaling and exacerbates inflammation. Immunity 2014, 40, 910–923. [Google Scholar] [CrossRef] [PubMed]
- Park, J.Y.; Jo, Y.; Ko, E.; Luckey, M.A.; Park, Y.K.; Park, S.H.; Park, J.H.; Hong, C. Soluble gammac cytokine receptor suppresses IL-15 signaling and impairs iNKT cell development in the thymus. Sci. Rep. 2016, 6, 36962. [Google Scholar] [CrossRef] [PubMed]
- Goh, T.S.; Hong, C. New insights of common gamma chain in hematological malignancies. Cytokine 2017, 89, 179–184. [Google Scholar] [CrossRef] [PubMed]
- Kim, G.; Hwang, H.; Jo, Y.; Lee, B.; Lee, Y.H.; Kim, C.H.; Hong, C. Soluble gammac receptor attenuates anti-tumor responses of CD8(+) T cells in T cell immunotherapy. Int. J. Cancer 2018, 143, 1212–1223. [Google Scholar] [CrossRef] [PubMed]
- Geiselhart, L.A.; Humphries, C.A.; Gregorio, T.A.; Mou, S.; Subleski, J.; Komschlies, K.L. IL-7 administration alters the CD4:CD8 ratio, increases T cell numbers, and increases T cell function in the absence of activation. J. Immunol. 2001, 166, 3019–3027. [Google Scholar] [CrossRef] [PubMed]
- Kieper, W.C.; Tan, J.T.; Bondi-Boyd, B.; Gapin, L.; Sprent, J.; Ceredig, R.; Surh, C.D. Overexpression of interleukin (IL)-7 leads to IL-15-independent generation of memory phenotype CD8+ T cells. J. Exp. Med. 2002, 195, 1533–1539. [Google Scholar] [CrossRef] [PubMed]
- Tan, J.T.; Dudl, E.; LeRoy, E.; Murray, R.; Sprent, J.; Weinberg, K.I.; Surh, C.D. IL-7 is critical for homeostatic proliferation and survival of naive T cells. Proc. Natl. Acad. Sci. USA 2001, 98, 8732–8737. [Google Scholar] [CrossRef] [PubMed]
- Tan, J.T.; Ernst, B.; Kieper, W.C.; LeRoy, E.; Sprent, J.; Surh, C.D. Interleukin (IL)-15 and IL-7 jointly regulate homeostatic proliferation of memory phenotype CD8+ cells but are not required for memory phenotype CD4+ cells. J. Exp. Med. 2002, 195, 1523–1532. [Google Scholar] [CrossRef] [PubMed]
- Ye, S.K.; Maki, K.; Lee, H.C.; Ito, A.; Kawai, K.; Suzuki, H.; Mak, T.W.; Chien, Y.; Honjo, T.; Ikuta, K. Differential roles of cytokine receptors in the development of epidermal gamma delta T cells. J. Immunol. 2001, 167, 1929–1934. [Google Scholar] [CrossRef] [PubMed]
- He, Y.W.; Malek, T.R. Interleukin-7 receptor alpha is essential for the development of gamma delta+ T cells, but not natural killer cells. J. Exp. Med. 1996, 184, 289–293. [Google Scholar] [CrossRef] [PubMed]
- Lodolce, J.P.; Boone, D.L.; Chai, S.; Swain, R.E.; Dassopoulos, T.; Trettin, S.; Ma, A. IL-15 receptor maintains lymphoid homeostasis by supporting lymphocyte homing and proliferation. Immunity 1998, 9, 669–676. [Google Scholar] [CrossRef]
- Webster, K.E.; Kim, H.O.; Kyparissoudis, K.; Corpuz, T.M.; Pinget, G.V.; Uldrich, A.P.; Brink, R.; Belz, G.T.; Cho, J.H.; Godfrey, D.I.; et al. IL-17-producing NKT cells depend exclusively on IL-7 for homeostasis and survival. Mucosal Immunol. 2014, 7, 1058–1067. [Google Scholar] [CrossRef] [PubMed]
- Gordy, L.E.; Bezbradica, J.S.; Flyak, A.I.; Spencer, C.T.; Dunkle, A.; Sun, J.; Stanic, A.K.; Boothby, M.R.; He, Y.W.; Zhao, Z.; et al. IL-15 regulates homeostasis and terminal maturation of NKT cells. J. Immunol. 2011, 187, 6335–6345. [Google Scholar] [CrossRef] [PubMed]
- Matsuda, J.L.; Gapin, L.; Sidobre, S.; Kieper, W.C.; Tan, J.T.; Ceredig, R.; Surh, C.D.; Kronenberg, M. Homeostasis of Vα 14i NKT cells. Nat. Immunol. 2002, 3, 966–974. [Google Scholar] [CrossRef] [PubMed]
- Prlic, M.; Blazar, B.R.; Farrar, M.A.; Jameson, S.C. In vivo survival and homeostatic proliferation of natural killer cells. J. Exp. Med. 2003, 197, 967–976. [Google Scholar] [CrossRef] [PubMed]
- Waldmann, T.A.; Longo, D.L.; Leonard, W.J.; Depper, J.M.; Thompson, C.B.; Kronke, M.; Goldman, C.K.; Sharrow, S.; Bongiovanni, K.; Greene, W.C. Interleukin 2 receptor (Tac antigen) expression in HTLV-I-associated adult T-cell leukemia. Cancer Res. 1985, 45, 4559s–4562s. [Google Scholar] [PubMed]
- Li, Z.; Chen, L.; Qin, Z. Paradoxical roles of IL-4 in tumor immunity. Cell. Mol. Immunol. 2009, 6, 415–422. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Jiang, J.; Wang, Z.; Zhang, J.; Xiao, M.; Wang, C.; Lu, Y.; Qin, Z. Endogenous interleukin-4 promotes tumor development by increasing tumor cell resistance to apoptosis. Cancer Res. 2008, 68, 8687–8694. [Google Scholar] [CrossRef] [PubMed]
- Lange, K.; Uckert, W.; Blankenstein, T.; Nadrowitz, R.; Bittner, C.; Renauld, J.C.; van Snick, J.; Feller, A.C.; Merz, H. Overexpression of NPM-ALK induces different types of malignant lymphomas in IL-9 transgenic mice. Oncogene 2003, 22, 517–527. [Google Scholar] [CrossRef] [PubMed]
- Fehniger, T.A.; Suzuki, K.; Ponnappan, A.; VanDeusen, J.B.; Cooper, M.A.; Florea, S.M.; Freud, A.G.; Robinson, M.L.; Durbin, J.; Caligiuri, M.A. Fatal leukemia in interleukin 15 transgenic mice follows early expansions in natural killer and memory phenotype CD8+ T cells. J. Exp. Med. 2001, 193, 219–231. [Google Scholar] [CrossRef] [PubMed]
- Fehniger, T.A.; Caligiuri, M.A. Interleukin 15: Biology and relevance to human disease. Blood 2001, 97, 14–32. [Google Scholar] [CrossRef] [PubMed]
- Brenne, A.T.; Ro, T.B.; Waage, A.; Sundan, A.; Borset, M.; Hjorth-Hansen, H. Interleukin-21 is a growth and survival factor for human myeloma cells. Blood 2002, 99, 3756–3762. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kim, G.; Jo, Y.; Lee, B.; Ali, L.A.; Lee, B.; Hong, C. The Potential Role of a Soluble γ-Chain Cytokine Receptor as a Regulator of IL-7-Induced Lymphoproliferative Disorders. Int. J. Mol. Sci. 2018, 19, 3375. https://doi.org/10.3390/ijms19113375
Kim G, Jo Y, Lee B, Ali LA, Lee B, Hong C. The Potential Role of a Soluble γ-Chain Cytokine Receptor as a Regulator of IL-7-Induced Lymphoproliferative Disorders. International Journal of Molecular Sciences. 2018; 19(11):3375. https://doi.org/10.3390/ijms19113375
Chicago/Turabian StyleKim, Geona, Yuna Jo, Byunghyuk Lee, Laraib Amir Ali, Boae Lee, and Changwan Hong. 2018. "The Potential Role of a Soluble γ-Chain Cytokine Receptor as a Regulator of IL-7-Induced Lymphoproliferative Disorders" International Journal of Molecular Sciences 19, no. 11: 3375. https://doi.org/10.3390/ijms19113375
APA StyleKim, G., Jo, Y., Lee, B., Ali, L. A., Lee, B., & Hong, C. (2018). The Potential Role of a Soluble γ-Chain Cytokine Receptor as a Regulator of IL-7-Induced Lymphoproliferative Disorders. International Journal of Molecular Sciences, 19(11), 3375. https://doi.org/10.3390/ijms19113375





