AMPK: An Epigenetic Landscape Modulator
Abstract
1. Introduction
2. Histone Modification
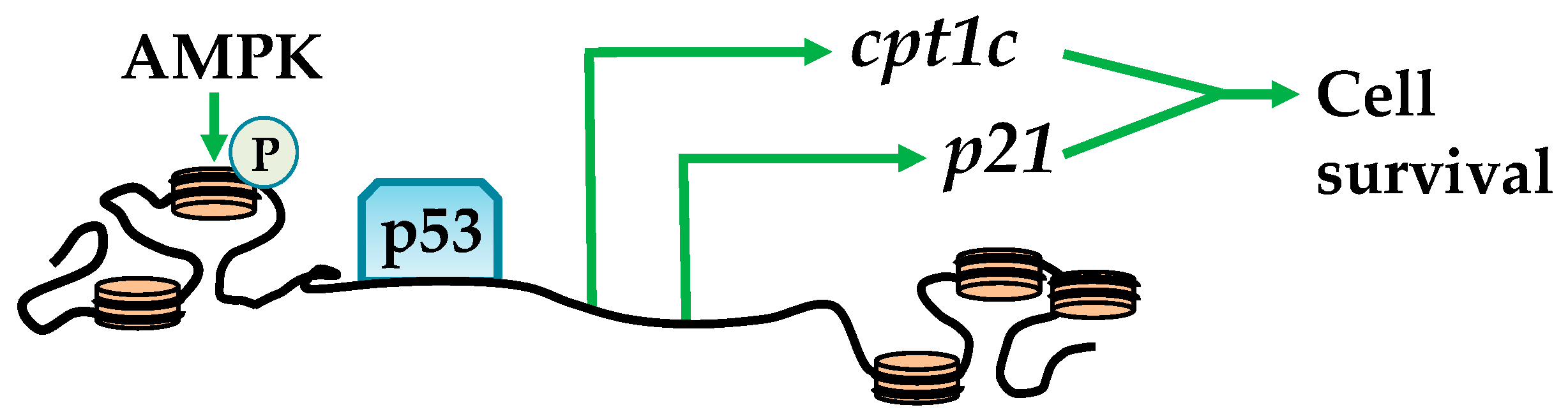
2.1. Histone Phosphorylation
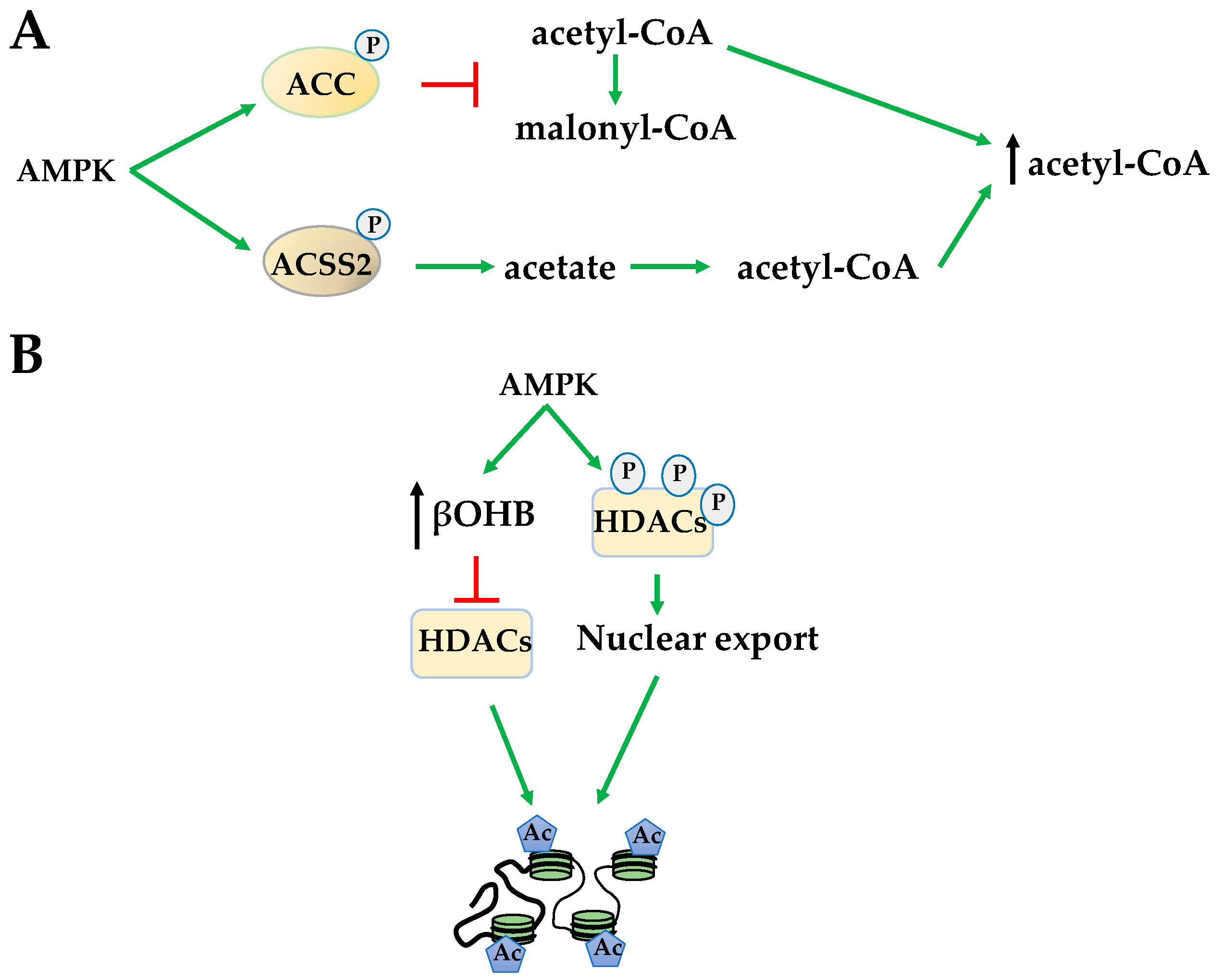
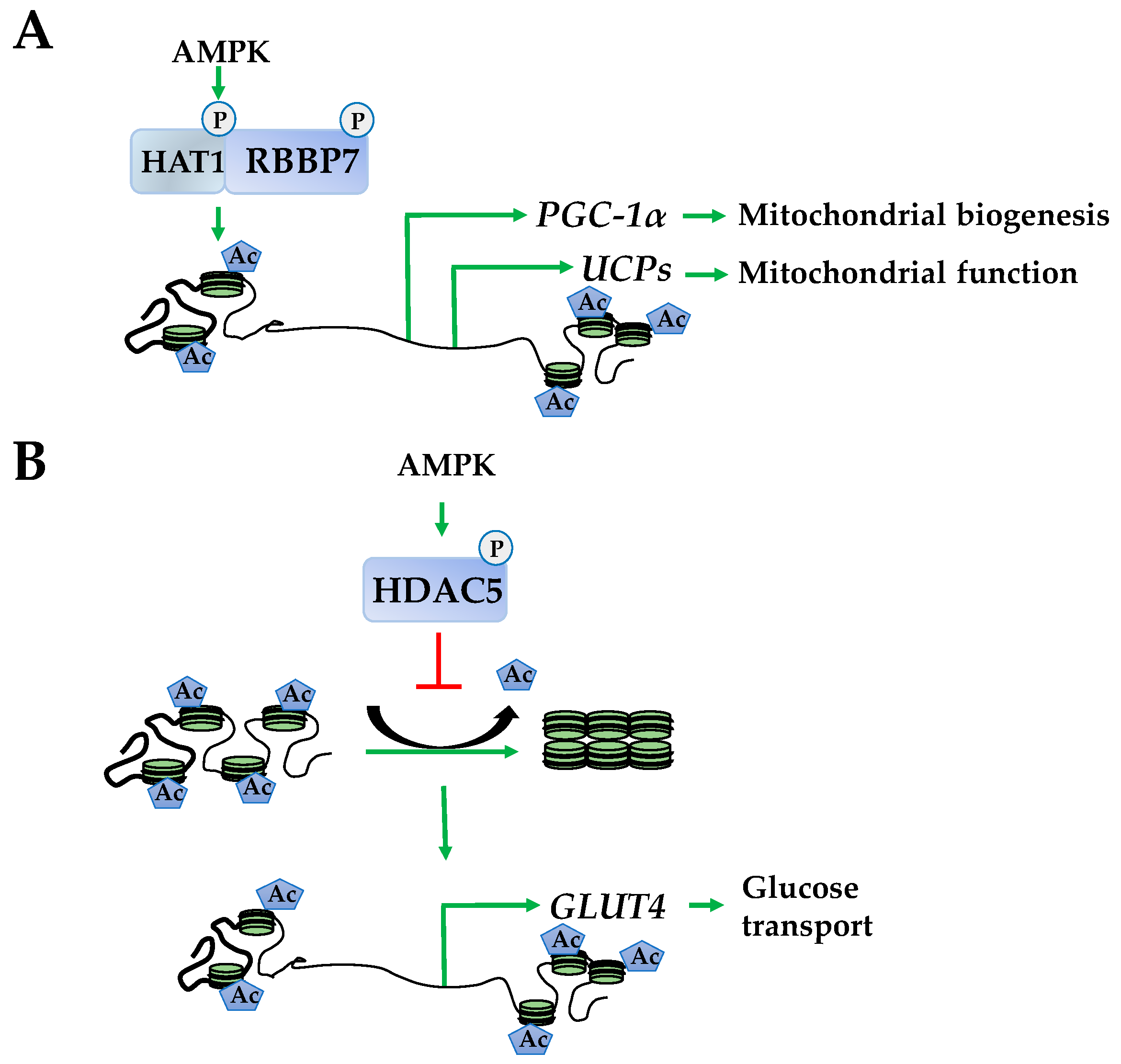
2.2. Histone Acetylation
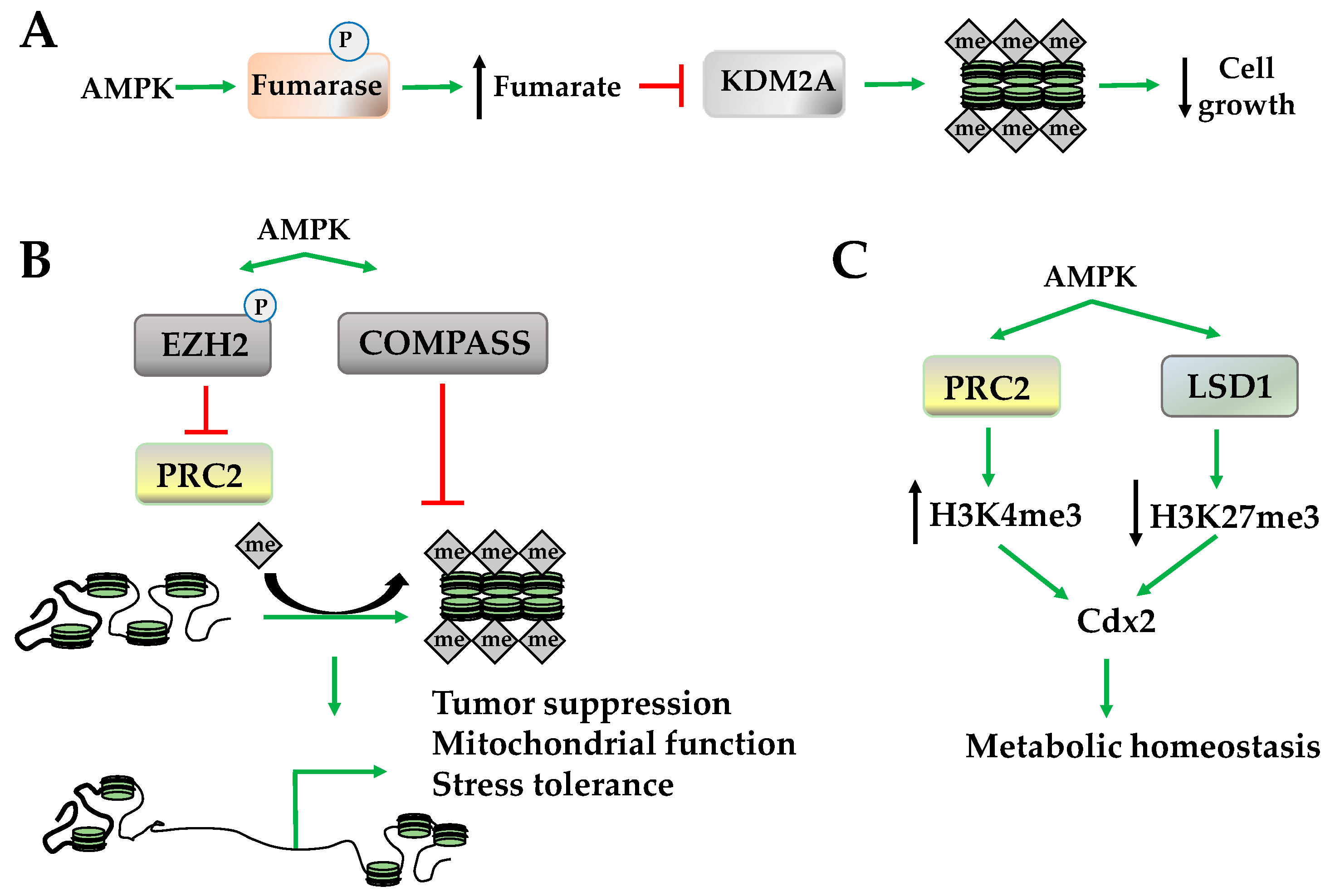
2.3. Histone Methylation
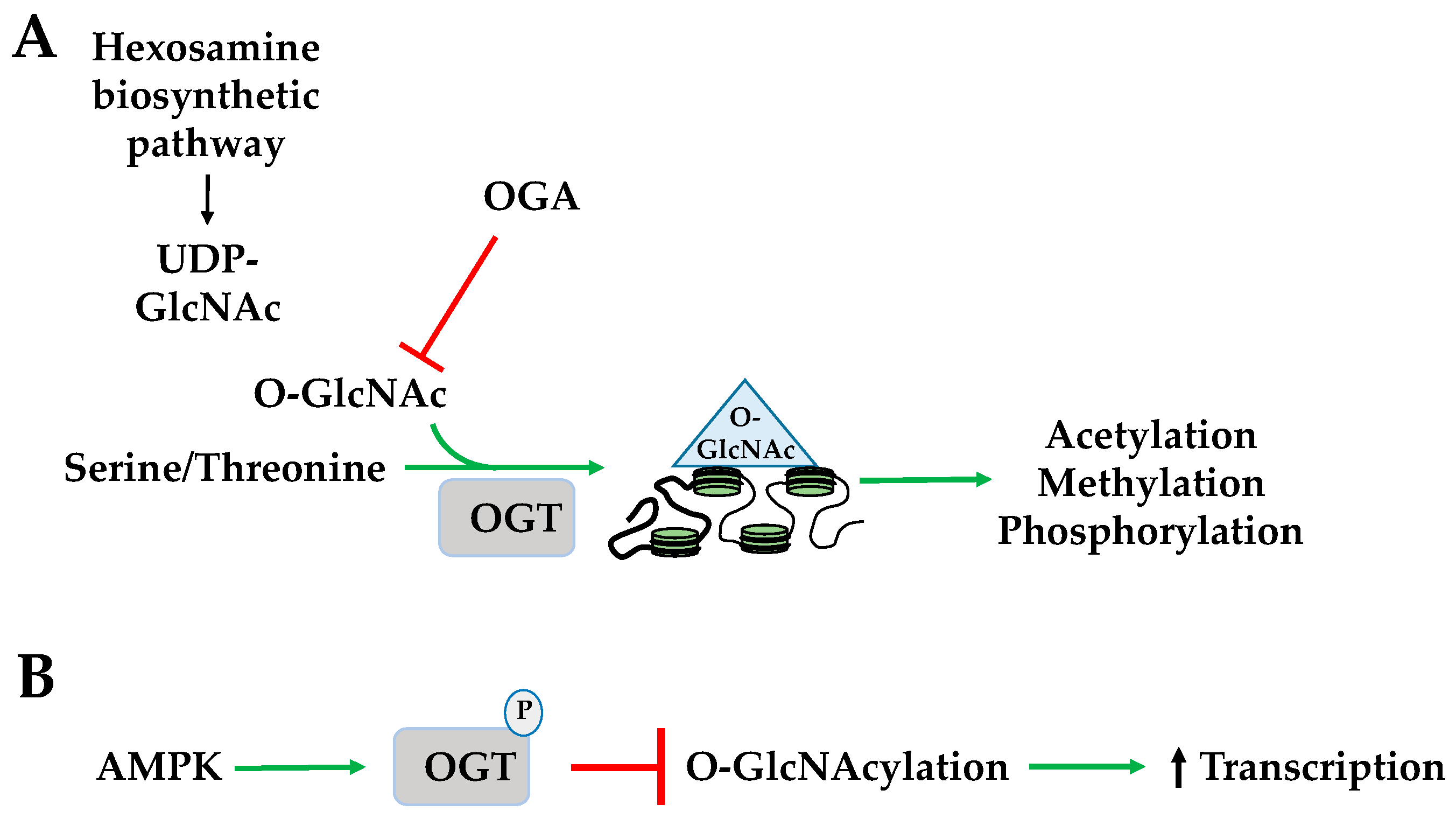
2.4. Histone O-GlcNAcylation
2.5. Histone Ribosylation
3. DNA Modification
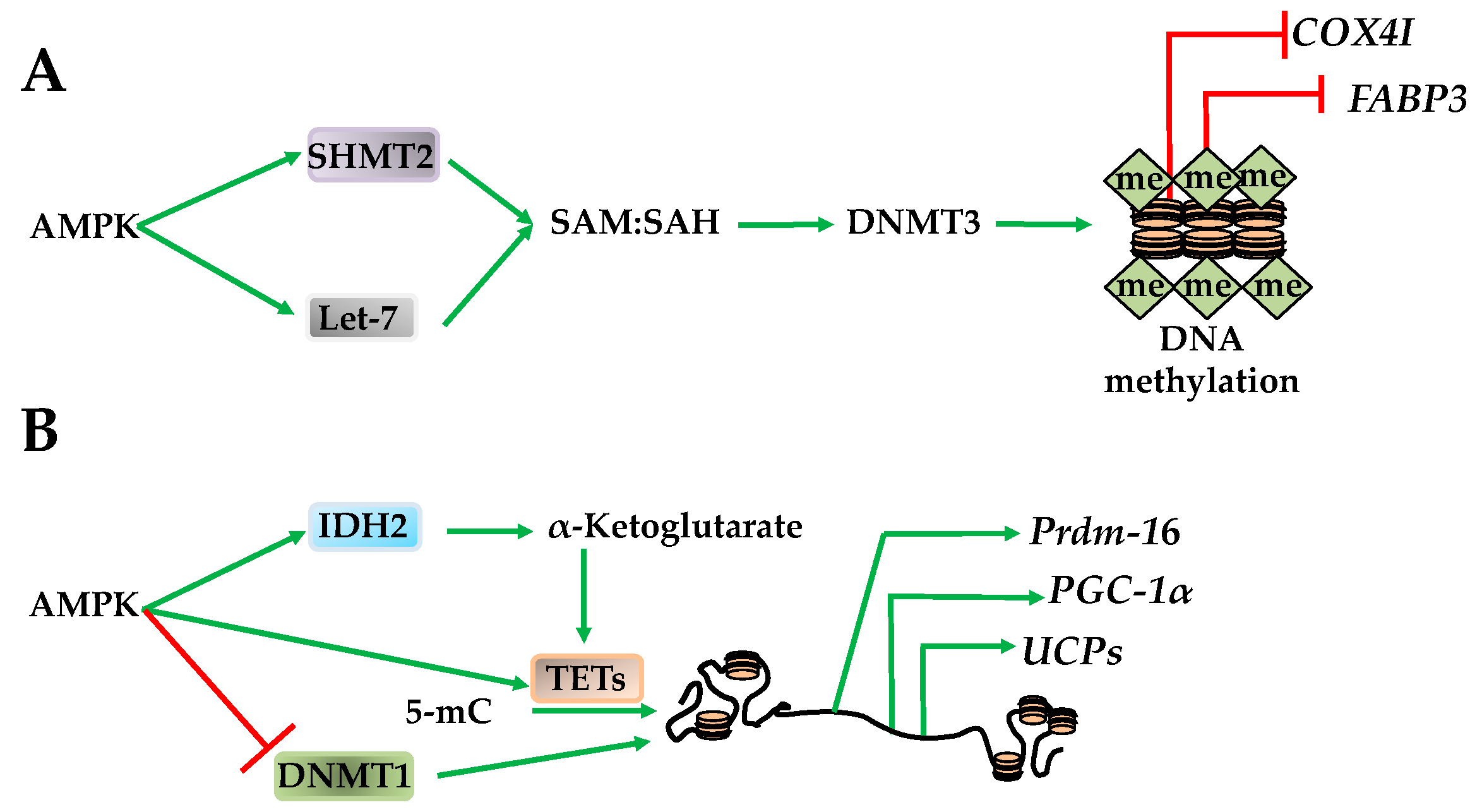
DNA Methylation
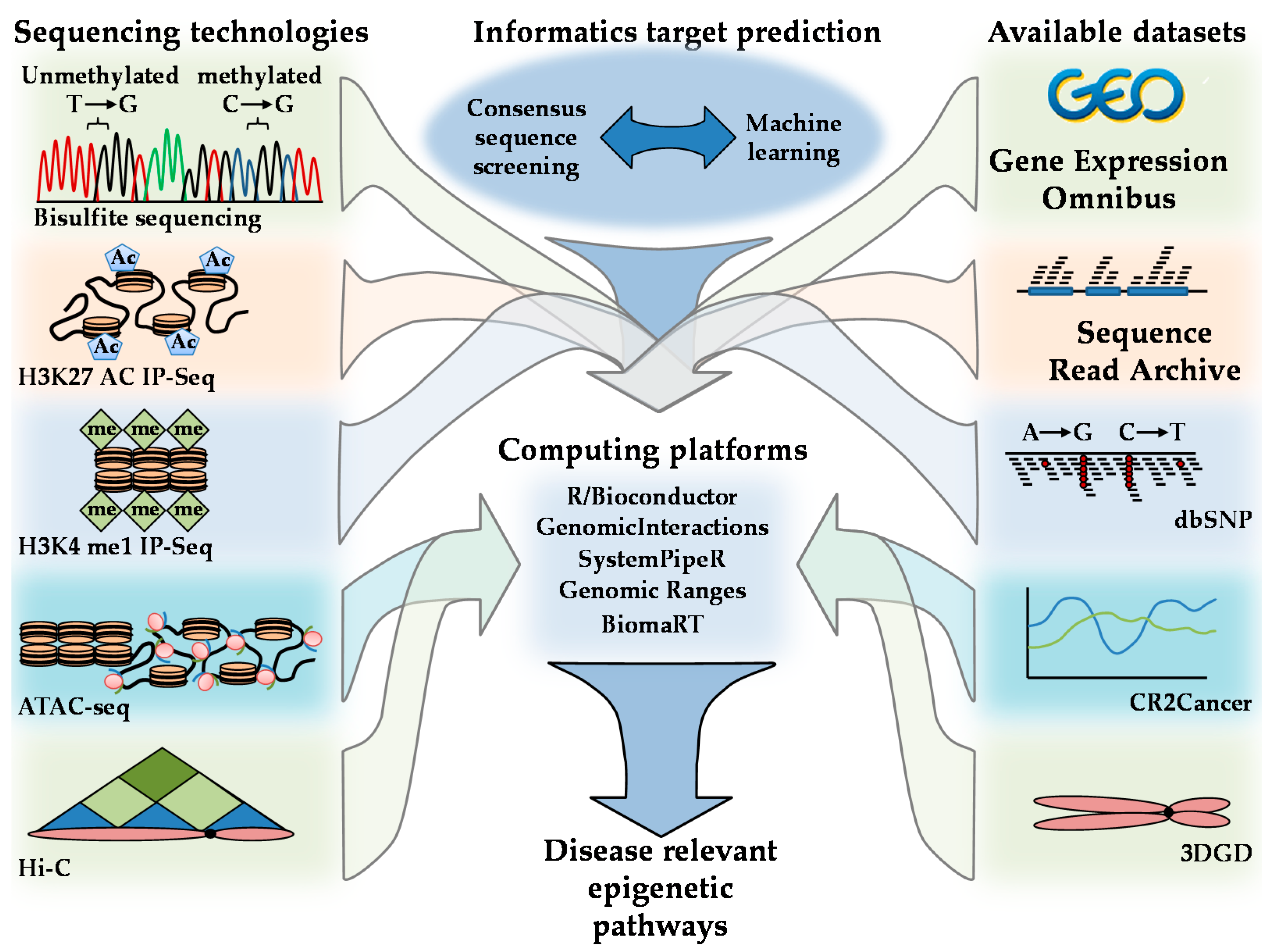
4. Approaches to Elucidating the AMPK-Modulated Epigenetic Landscape
5. Conclusions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| AMPK | AMP-activated protein kinase |
| CPT1C | carnitine palmitoyltransferase 1C |
| p21 | cyclin-dependent kinase inhibitor |
| p53 | TP53 or tumor protein |
| HAT | histone deacetylases |
| acetyl-CoA | acetyl coenzyme-A |
| HDAC | histone deacetylases |
| AICAR | 5-Aminoimidazole-4-carboxamide ribonucleotide |
| ACC | acetyl-CoA carboxylase |
| ACSS2 | acetyl-CoA synthetase short-chain family member 2 |
| TFEB | transcription factor EB |
| SIRT | sirtuin |
| NAD+ | nicotinamide adenine dinucleotide |
| βOHB | β-hydroxybutyrate |
| RBBP7 | retinoblastoma binding protein 7 |
| PGC-1α | peroxisome proliferator-activated receptor gamma coactivator-1α |
| Tfam | transcription factor A |
| NRF | nuclear respiratory factors |
| UCP | uncoupling proteins |
| SAM | S-adenosyl methionine |
| GLUT-4 | glucose transporter type 4 |
| KDM2A | lysine-specific demethylase 2A |
| KDM5 | demethyltransferase lysine demethylase 5 |
| LSD1 | lysine-specific histone demethylase-1 |
| PRC2 | polycomb repressive complex 2 |
| COMPASS | complex proteins associated with Set1 |
| EZH2 | enhancer of zeste homolog 2 |
| Cdx2 | caudal type homeobox2 |
| SLC5A8 | solute carrier family 5 member 8 |
| O-GlcNAc | O-linked N-acetylglucosamine |
| OGT | O-GlcNAc transferase |
| UDP-GlcNAc | uridine diphosphate N-acetylglucosamine |
| HBP | hexosamine biosynthesis pathway |
| ART | mono-ADP ribosyltransferase |
| PTM | Post translational modification |
| PARP | poly-(ADP-ribose) polymerase |
| Bcl-6 | B-cell Lymphoma protein 6 |
| CpG | cytosine-guanine dinucleotides |
| DNMT | DNA methyltransferase |
| MBD | methyl-CpG binding domain proteins |
| 5-mC | 5-methycytosine |
| 5-hmC | 5-hydroxymethylcytosine |
| 5-fC | 5-formylcytosine |
| 5-caC | 5-carboxylcytosine |
| TETs | ten-eleven translocation hydroxylases |
| SAH | S-adenosylhomocysteine |
| SHMT2 | serine hydroxymethyltransferase 2 |
| SAHH | S-adenosylhomocysteine hydrolase |
| COX4I | cytochrome C oxidase subunit 4I1 |
| FABP3 | fatty acid binding protein 3 |
| PPARδ | peroxisome proliferator-activated receptor δ |
| TCA | tricarboxylic acid |
| IDH2 | isocitrase dehydrogenase 2 |
| IP-Seq | immunoprecipitation sequencing |
| ATAC-seq | assay for transposase-accessible chromatin using sequencing |
| GEO | gene expression omnibus |
| SRA | sequence read archive |
| dbSNP | single nucleotide polymorphism database |
| 3DGD | 3D genome database |
References
- Trerotola, M.; Relli, V.; Simeone, P.; Alberti, S. Epigenetic inheritance and the missing heritability. Hum. Genom. 2015, 28, 9–17. [Google Scholar] [CrossRef] [PubMed]
- Bunkar, N.; Pathak, N.; Lohiya, N.K.; Mishra, P.K. Epigenetics: A key paradigm in reproductive health. Clin. Exp. Reprod. Med. 2016, 43, 59–81. [Google Scholar] [CrossRef] [PubMed]
- Boland, M.J.; Nazor, K.L.; Loring, J.F. Epigenetic regulation of pluripotency and differentiation. Circ. Res. 2014, 115, 311–324. [Google Scholar] [CrossRef] [PubMed]
- Barrès, R.; Osler, M.E.; Yan, J.; Rune, A.; Fritz, T.; Caidahl, K.; Krook, A.; Zierath, J.R. Non-CpG methylation of the PGC-1alpha promoter through DNMT3B controls mitochondrial density. Cell Metab. 2009, 10, 189–198. [Google Scholar] [CrossRef] [PubMed]
- Barrès, R.; Yan, J.; Egan, B.; Treebak, J.T.; Rasmussen, M.; Fritz, T.; Caidahl, K.; Krook, A.; O’Gorman, D.J.; Zierath, J.R. Acute exercise remodels promoter methylation in human skeletal muscle. Cell Metab. 2012, 15, 405–411. [Google Scholar] [CrossRef] [PubMed]
- Nitert, M.D.; Dayeh, T.; Volkov, P.; Elgzyri, T.; Hall, E.; Nilsson, E.; Yang, B.T.; Lang, S.; Parikh, H.; Wessman, Y.; et al. Impact of an exercise intervention on DNA methylation in skeletal muscle from first-degree relatives of patients with type 2 diabetes. Diabetes 2012, 61, 3322–3332. [Google Scholar] [CrossRef] [PubMed]
- Ferraro, E.; Giammarioli, A.M.; Chiandotto, S.; Spoletini, I.; Rosano, G. Exercise-induced skeletal muscle remodeling and metabolic adaptation: Redox signaling and role of autophagy. Antioxid. Redox Signal. 2014, 21, 154–176. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Yang, G.; Kim, Y.; Ha, J. AMPK activators: Mechanisms of action and physiological activities. Exp. Mol. Med. 2016, 48, e224. [Google Scholar] [CrossRef] [PubMed]
- Rabinovitch, R.C.; Samborska, B.; Faubert, B.; Ma, E.H.; Gravel, S.P.; Andrzejewski, S.; Raissi, T.C.; Pause, A.; St-Pierre, J.; Jones, R.G. AMPK Maintains Cellular Metabolic Homeostasis through Regulation of Mitochondrial Reactive Oxygen Species. Cell Rep. 2017, 21, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Ross, F.A.; Jensen, T.E.; Hardie, D.G. Differential regulation by AMP and ADP of AMPK complexes containing different gamma subunit isoforms. Biochem. J. 2016, 473, 189–199. [Google Scholar] [CrossRef] [PubMed]
- Vara-Ciruelos, D.; Dandapani, M.; Gray, A.; Egbani, E.O.; Evans, A.M.; Hardie, D.G. Genotoxic Damage Activates the AMPK-α1 Isoform in the Nucleus viaCa2+/CaMKK2 Signaling to Enhance Tumor Cell Survival. Mol. Cancer Res. 2018, 16, 345–357. [Google Scholar] [CrossRef] [PubMed]
- Langendorf, C.G.; Ngoei, K.R.; Scott, J.W.; Ling, N.X.; Issa, S.M.; Gorman, M.A.; Parker, M.W.; Sakamoto, K.; Oakhill, J.S.; Kemp, B.E. Structural basis of allosteric and synergistic activation of AMPK by furan-2-phosphonic derivative C2 binding. Nat. Commun. 2016, 7, 10912. [Google Scholar] [CrossRef] [PubMed]
- Marin, T.L.; Gongol, B.; Zhang, F.; Martin, M.; Johnson, D.A.; Xiao, H.; Wang, Y.; Subramaniam, S.; Chien, S.; Shyy, J.Y. AMPK promotes mitochondrial biogenesis and function by phosphorylating the epigenetic factors DNMT1, RBBP7, and HAT1. Sci. Signal. 2017, 10, eaaf7478. [Google Scholar] [CrossRef] [PubMed]
- Ratman, D.; Mylka, V.; Bougarne, N.; Pawlak, M.; Caron, S.; Hennuyer, N.; Paumelle, R.; De Cauwer, L.; Thommis, J.; Rider, M.H.; et al. Chromatin recruitment of activated AMPK drives fasting response genes co-controlled by GR and PPARα. Nucleic Acids Res. 2016, 44, 10539–10553. [Google Scholar] [CrossRef] [PubMed]
- Satake, M.; Kawata, M.; McLysaght, A.; Makino, T. Evolution of vertebrate tissues driven by differential modes of gene duplication. DNA Res. 2012, 19, 305–316. [Google Scholar] [CrossRef] [PubMed]
- Ross, F.A.; MacKintosh, C.; Hardie, D.G. AMP-activated protein kinase: A cellular energy sensor that comes in 12 flavours. FEBS J. 2016, 283, 2987–3001. [Google Scholar] [CrossRef] [PubMed]
- Hardie, D.G. Keeping the home fires burning: AMP-activated protein kinase. J. R. Soc. Interface 2018, 15, 20170774. [Google Scholar] [CrossRef] [PubMed]
- Laker, R.C.; Wlodek, M.E.; Connelly, J.J.; Yan, Z. Epigenetic origins of metabolic disease: The impact of the maternal condition to the offspring epigenome and later health consequences. Food Sci. Hum. Wellness 2013, 2, 1–11. [Google Scholar] [CrossRef]
- Zhu, M.J.; Han, B.; Tong, J.; Ma, C.; Kimzey, J.M.; Underwood, K.R.; Xiao, Y.; Hess, B.W.; Ford, S.P.; Nathanielsz, P.W.; et al. AMP-activated protein kinase signaling pathways are down regulated and skeletal muscle development impaired in fetuses of obese, over-nourished sheep. J. Physiol. 2008, 586, 2651–2664. [Google Scholar] [CrossRef] [PubMed]
- Demoinet, E.; Li, S.; Roy, R. AMPK blocks starvation-inducible transgenerational defects in Caenorhabditis elegans. Proc. Natl. Acad. Sci. USA 2017, 114, E2689–E2698. [Google Scholar] [CrossRef] [PubMed]
- Bönisch, C.; Hake, S.B. Histone H2A variants in nucleosomes and chromatin: More or less stable? Nucleic Acids Res. 2012, 40, 10719–10741. [Google Scholar] [CrossRef] [PubMed]
- Zlatanova, J.S.; Srebreva, L.N.; Banchev, T.B.; Tasheva, B.T.; Tsanev, R.G. Cytoplasmic pool of histone H1 in mammalian cells. J. Cell Sci. 1990, 96, 461–468. [Google Scholar] [PubMed]
- Mariño-Ramírez, L.; Kann, M.G.; Shoemaker, B.A.; Landsman, D. Histone structure and nucleosome stability. Expert Rev. Proteom. 2005, 2, 719–729. [Google Scholar] [CrossRef] [PubMed]
- Sawicka, A.; Seiser, C. Sensing core histone phosphorylation- a matter of perfect timing. Biochim. Biophys. Acta 2014, 1839, 711–718. [Google Scholar] [CrossRef] [PubMed]
- Marin, T.L.; Gongol, B.; Martin, M.; King, S.J.; Smith, L.; Johnson, D.A.; Subramaniam, S.; Chien, S.; Shyy, J.Y. Identification of AMP-activated protein kinase targets by a consensus sequence search of the proteome. BMC Syst. Biol. 2015, 9, 13. [Google Scholar] [CrossRef] [PubMed]
- Jones, R.G.; Plas, D.R.; Kubek, S.; Buzzai, M.; Mu, J.; Xu, Y.; Birnbaum, M.J.; Thompson, C.B. AMP-activated protein kinase induces a p53-dependent metabolic checkpoint. Mol. Cell 2005, 18, 283–293. [Google Scholar] [CrossRef] [PubMed]
- Bungard, D.; Fuerth, B.J.; Zeng, P.Y.; Faubert, B.; Maas, N.L.; Viollet, B.; Carling, D.; Thompson, C.B.; Jones, R.G.; Berger, S.L. Signaling kinase AMPK activates stress-promoted transcription via histone H2B phosphorylation. Science 2010, 329, 1201–1205. [Google Scholar] [CrossRef] [PubMed]
- Roa-Mansergas, X.; Fadó, R.; Atari, M.; Mir, J.F.; Muley, H.; Serra, D.; Casals, N. CPT1C promotes human mesenchymal stem cells survival under glucose deprivation through the modulation of autophagy. Sci. Rep. 2018, 8, 6997. [Google Scholar] [CrossRef] [PubMed]
- Lee, I.H.; Kawai, Y.; Fergusson, M.M.; Rovira, I.I.; Bishop, A.J.; Motoyama, N.; Cao, L.; Finkel, T. Atg7 modulates p53 activity to regulate cell cycle and survival during metabolic stress. Science 2012, 336, 225–228. [Google Scholar] [CrossRef] [PubMed]
- Itahana, Y.; Itahana, K. Emerging Roles of p53 Family Members in Glucose Metabolism. Int. J. Mol. Sci. 2018, 19, 776. [Google Scholar] [CrossRef] [PubMed]
- Stauffer, D.R.; Howard, T.L.; Nyun, T.; Hollenberg, S.M. CHMP1 is a novel nuclear matrix protein affecting chromatin structure and cell-cycle progression. J. Cell Sci. 2001, 114, 2383–2393. [Google Scholar] [PubMed]
- Javaid, N.; Choi, S. Acetylation- and Methylation-Related Epigenetic Proteins in the Context of Their Targets. Genes 2017, 8, 196. [Google Scholar] [CrossRef] [PubMed]
- Marmorstein, R.; Zhou, M.M. Writers and readers of histone acetylation: Structure, mechanism, and inhibition. Cold Spring Harb. Perspect. Biol. 2014, 6, a018762. [Google Scholar] [CrossRef] [PubMed]
- Bulusu, V.; Tumanov, S.; Michalopoulou, E.; van den Broek, N.J.; MacKay, G.; Nixon, C.; Dhayade, S.; Schug, Z.T.; Vande Voorde, J.; Blyth, K.; et al. Acetate recapturing by nuclear acetyl-CoA synthetase 2 prevents loss of histone acetylation during oxygen and serum limitation. Cell Rep. 2017, 18, 647–658. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Yu, W.; Qian, X.; Xia, Y.; Zheng, Y.; Lee, J.H.; Li, W.; Lyu, J.; Rao, G.; Zhang, X.; et al. Nucleus-translocated ACSS2 promotes gene transcription for lysosomal biogenesis and autophagy. Mol. Cell 2017, 66, 684–697. [Google Scholar] [CrossRef] [PubMed]
- Galdieri, L.; Gatla, H.; Vancurova, I.; Vancura, A. Activation of AMP-activated Protein Kinase by Metformin Induces Protein Acetylation in Prostate and Ovarian Cancer Cells. J. Biol. Chem. 2016, 291, 25154–25166. [Google Scholar] [CrossRef] [PubMed]
- Cuyàs, E.; Fernández-Arroyo, S.; Joven, J.; Menendez, J.A. Metformin targets histone acetylation in cancer prone epithelial cells. Cell Cycle. 2016, 15, 3355–3361. [Google Scholar] [CrossRef] [PubMed]
- Ruderman, N.B.; Xu, X.J.; Nelson, L.; Cacicedo, J.M.; Saha, A.K.; Lan, F.; Ido, Y. AMPK and SIRT1: A long-standing partnership? Am. J. Physiol. Endocrinol. Metab. 2010, 298, E751–E760. [Google Scholar] [CrossRef] [PubMed]
- Salminen, A.; Kauppinen, A.; Kaarniranta, K. AMPK/Snf1 signaling regulates histone acetylation: Impact on gene expression and epigenetic functions. Cell. Signal. 2016, 28, 887–895. [Google Scholar] [CrossRef] [PubMed]
- Zhang, T.; Kraus, W.L. SIRT1-dependent regulation of chromatin and transcription: Linking NAD(+) metabolism and signaling to the control of cellular functions. Biochim. Biophys. Acta 2010, 1804, 1666–1675. [Google Scholar] [CrossRef] [PubMed]
- Shimazu, T.; Hirschey, M.D.; Newman, J.; He, W.; Shirakawa, K.; Le Moan, N.; Grueter, C.A.; Lim, H.; Saunders, L.R.; Stevens, R.D.; et al. Suppression of oxidative stress by β-hydroxybutyrate, an endogenous histone deacetylase inhibitor. Science 2013, 339, 211–214. [Google Scholar] [CrossRef] [PubMed]
- Mihaylova, M.M.; Vasquez, D.S.; Ravnskjaer, K.; Denechaud, P.D.; Yu, R.T.; Alvarez, J.G.; Downes, M.; Evans, R.M.; Montminy, M.; Shaw, R.J. Class IIa histone deacetylases are hormone-activated regulators of FOXO and mammalian glucose homeostasis. Cell 2011, 145, 607–621. [Google Scholar] [CrossRef] [PubMed]
- McGee, S.L.; van Denderen, B.J.; Howlett, K.F.; Mollica, J.; Schertzer, J.D.; Kemp, B.E.; Hargreaves, M. AMP-activated protein kinase regulates GLUT4 transcription by phosphorylating histone deacetylase 5. Diabetes 2008, 57, 860–867. [Google Scholar] [CrossRef] [PubMed]
- Ng, S.; Yue, W.; Oppermann, U.; Klose, R. Dynamic protein methylation in chromatin biology. Cell. Mol. Life Sci. 2009, 66, 407–422. [Google Scholar] [CrossRef] [PubMed]
- Feitag, M. Histone Methylation by SET Domain Proteins in Fungi. Annu. Rev. Microbiol. 2017, 71, 413–439. [Google Scholar] [CrossRef] [PubMed]
- Wei, S.; Li, C.; Yin, Z.; Wen, J.; Meng, H.; Xue, L.; Wang, J. Histone methylation in DNA repair and clinical practice: New findings during the past 5-years. J. Cancer 2018, 9, 2072–2081. [Google Scholar] [CrossRef] [PubMed]
- Yang, Q.; Liang, X.; Sun, X.; Zhang, L.; Fu, X.; Rogers, C.; Berim, A.; Zhang, S.; Wang, S.; Wang, B.; et al. AMPK/α-ketoglutarate axis dynamically mediates DNA demethylation in the Prdm16 promoter and brown adipogenesis. Cell Metab. 2016, 24, 542–554. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Yu, Q.; Li, J.; Hu, B.; Zhao, Q.; Ma, C.; Huang, W.; Zhuo, L.; Fang, H.; Liao, L.; et al. O-GlcNAcylation of fumarase maintains tumour growth under glucose deficiency. Nat. Cell Biol. 2017, 19, 833–843. [Google Scholar] [CrossRef] [PubMed]
- Eissenberg, J.C.; Shilatifard, A. Histone H3 lysine 4 (H3K4) methylation in development and differentiation. Dev. Biol. 2010, 339, 240–249. [Google Scholar] [CrossRef] [PubMed]
- Wan, L.; Xu, K.; Wei, Y.; Zhang, J.; Han, T.; Fry, C.; Zhang, Z.; Wang, Y.V.; Huang, L.; Yuan, M.; et al. Phosphorylation of EZH2 by AMPK Suppresses PRC2 Methyltransferase Activity and Oncogenic Function. Mol. Cell 2018, 69, 279–291. [Google Scholar] [CrossRef] [PubMed]
- Tang, G.; Guo, J.; Zhu, Y.; Huang, Z.; Liu, T.; Cai, J.; Yu, L.; Wang, Z. Metformin inhibits ovarian cancer via decreasing H3K27 trimethylation. Int. J. Oncol. 2018, 52, 1899–1911. [Google Scholar] [CrossRef] [PubMed]
- Aloia, L.; Di Stefano, B.; Di Croce, L. Polycomb complexes in stem cells and embryonic development. Development 2013, 140, 2525–2534. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.; Yang, Q.; Rogers, C.J.; Du, M.; Zhu, M.J. AMPK improves gut epithelial differentiation and barrier function via regulating Cdx2 expression. Cell Death Differ. 2017, 24, 819–831. [Google Scholar] [CrossRef] [PubMed]
- Kakizaki, F.; Aoki, K.; Miyoshi, H.; Carrasco, N.; Aoki, M.; Taketo, M.M. CDX transcription factors positively regulate expression of solute carrier family 5, member 8 in the colonic epithelium. Gastroenterology 2010, 138, 627–635. [Google Scholar] [CrossRef] [PubMed]
- Gurav, A.; Sivaprakasam, S.; Bhutia, Y.D.; Boettger, T.; Singh, N.; Ganapathy, V. Slc5a8, a Na+-coupled high-affinity transporter for short-chain fatty acids, is a conditional tumour suppressor in colon that protects against colitis and colon cancer under low-fibre dietary conditions. Biochem. J. 2015, 469, 267–278. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Qian, K. Protein O-GlcNAcylation: Emerging mechanisms and functions. Nat. Rev. Mol. Cell Biol. 2017, 18, 452–465. [Google Scholar] [CrossRef] [PubMed]
- Dehennaut, V.; Leprince, D.; Lefebvre, T. O-GlcNAcylation, an Epigenetic Mark. Focus on the Histone Code, TETFamily Proteins, and Polycomb Group Proteins. Front. Endocrinol. 2014, 5, 155. [Google Scholar] [CrossRef] [PubMed]
- Poirier, G.G.; de Murcia, G.; Jongstra-Bilen, J.; Niedergang, C.; Mandel, P. Poly(ADP-ribosyl)ation of polynucleosomes causes relaxation of chromatin structure. Proc. Natl. Acad. Sci. USA 1982, 79, 3423–3427. [Google Scholar] [CrossRef] [PubMed]
- Ciccarone, F.; Zampieri, M.; Caiafa, P. PARP1 orchestrates epigenetic events setting up chromatin domains. Semin. Cell Dev. Biol. 2017, 63, 123–134. [Google Scholar] [CrossRef] [PubMed]
- Walker, J.W.; Jijon, H.B.; Madsen, K.L. AMP-activated protein kinase is a positive regulator of poly(ADP-ribose) polymerase. Biochem. Biophys. Res. Commun. 2006, 342, 336–341. [Google Scholar] [CrossRef] [PubMed]
- Gongol, B.; Marin, T.; Peng, I.C.; Woo, B.; Martin, M.; King, S.; Sun, W.; Johnson, D.A.; Chien, S.; Shyy, J.Y. AMPKα2 exerts its anti-inflammatory effects through PARP-1 and Bcl-6. Proc. Natl. Acad. Sci. USA 2013, 110, 3161–3166. [Google Scholar] [CrossRef] [PubMed]
- Lodhi, N.; Kossenkov, A.V.; Tulin, A.V. Bookmarking promoters in mitotic chromatin: Poly (ADP-ribose) polymerase-1 as an epigenetic mark. Nucleic Acids Res. 2014, 42, 7028–7038. [Google Scholar] [CrossRef] [PubMed]
- Dunn, J.; Qiu, H.; Kim, S.; Jjingo, D.; Hoffman, R.; Kim, C.W.; Jang, I.; Son, D.J.; Kim, D.; Pan, C.; et al. Flow-dependent epigenetic DNA methylation regulates endothelial gene expression and atherosclerosis. J. Clin. Investig. 2014, 124, 3187–3199. [Google Scholar] [CrossRef] [PubMed]
- Hill, P.W.; Amouroux, R.; Hajkova, P. DNA demethylation, Tet proteins and 5-hydroxymethylcytosine in epigenetic reprogramming: An emerging complex story. Genomics 2014, 104, 324–333. [Google Scholar] [CrossRef] [PubMed]
- Ficz, G.; Branco, M.R.; Seisenberger, S.; Santos, F.; Krueger, F.; Hore, T.A.; Marques, C.J.; Andrews, S.; Reik, W. Dynamic regulation of 5-hydroxymethylcytosine in mouse ES cells and during differentiation. Nature 2011, 473, 398–402. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.X.; Riggs, A.D. DNA methylation and demethylation in mammals. J. Biol. Chem. 2011, 286, 18347–18353. [Google Scholar] [CrossRef] [PubMed]
- Cuyàs, E.; Fernández-Arroyo, S.; Verdura, S.; García, R.Á.; Stursa, J.; Werner, L.; Blanco-González, E.; Montes-Bayón, M.; Joven, J.; Viollet, B.; et al. Metformin regulates global DNA methylation via mitochondrial one-carbon metabolism. Oncogene 2018, 37, 963–970. [Google Scholar] [CrossRef] [PubMed]
- Zhong, T.; Men, Y.; Lu, L.; Geng, T.; Zhou, J.; Mitsuhashi, A.; Shozu, M.; Maihle, N.J.; Carmichael, G.G.; Taylor, H.S.; et al. Metformin alters DNA methylation genome-wide via the H19/SAHH axis. Oncogene 2017, 36, 2345–2354. [Google Scholar] [CrossRef] [PubMed]
- Lane, S.C.; Camera, D.M.; Lassiter, D.G.; Areta, J.L.; Bird, S.R.; Yeo, W.K.; Jeacocke, N.A.; Krook, A.; Zierath, J.R.; Burke, L.M.; et al. Effects of sleeping with reduced carbohydrate availability on acute training responses. J. Appl. Physiol. 2015, 119, 643–655. [Google Scholar] [CrossRef] [PubMed]
- Snyder, C.M.; Chandel, N.S. Mitochondrial regulation of cell survival and death during low-oxygen conditions. Antioxid. Redox Signal. 2009, 11, 2673–2683. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.; Zhang, Y. Reversing DNA methylation: Mechanisms, genomics, and biological functions. Cell 2014, 156, 45–68. [Google Scholar] [CrossRef] [PubMed]
- Jiang, S.; Mortazavi, A. Integrating ChIP-seq with other functional genomics data. Brief. Funct. Genom. 2018, 17, 104–115. [Google Scholar] [CrossRef] [PubMed]
- Xue, L.; Tao, W.A. Current technologies to identify protein kinase substrates in high throughput. Front. Biol. 2013, 8, 216–227. [Google Scholar] [CrossRef] [PubMed]
- Harmston, N.; Ing-Simmons, E.; Perry, M.; Barešić, A.; Lenhard, B. GenomicInteractions: An R/Bioconductor package for manipulating and investigating chromatin interaction data. BMC Genom. 2015, 16, 963. [Google Scholar] [CrossRef] [PubMed]
- Backman, T.W.H.; Girke, T. systemPipeR: NGS workflow and report generation environment. BMC Bioinform. 2016, 20, 388. [Google Scholar] [CrossRef] [PubMed]
- Deininger, P.; Morales, M.E.; White, T.B.; Baddoo, M.; Hedges, D.J.; Servant, G.; Srivastav, S.; Smither, M.E.; Concha, M.; DeHaro, D.L.; et al. A comprehensive approach to expressionof L1 loci. Nucleic Acids Res. 2017, 45, e31. [Google Scholar] [CrossRef] [PubMed]
- Bhagwat, M. Searching NCBI’s dbSNP database. Curr. Protoc. Bioinform. 2010. [Google Scholar] [CrossRef]
- Li, C.; Dong, X.; Fan, H.; Wang, C.; Ding, G.; Li, Y. The 3DGD: A database of genome 3D structure. Bioinformatics 2014, 30, 1640–1642. [Google Scholar] [CrossRef] [PubMed]
- Ru, B.; Sun, J.; Tong, Y.; Wong, C.N.; Chandra, A.; Tang, A.T.S.; Chow, L.K.Y.; Wun, W.L.; Levitskaya, Z.; Zhang, J. CR2Cancer: A database for chromatin regulators in human cancer. Nucleic Acids Res. 2018, 46, D918–D924. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gongol, B.; Sari, I.; Bryant, T.; Rosete, G.; Marin, T. AMPK: An Epigenetic Landscape Modulator. Int. J. Mol. Sci. 2018, 19, 3238. https://doi.org/10.3390/ijms19103238
Gongol B, Sari I, Bryant T, Rosete G, Marin T. AMPK: An Epigenetic Landscape Modulator. International Journal of Molecular Sciences. 2018; 19(10):3238. https://doi.org/10.3390/ijms19103238
Chicago/Turabian StyleGongol, Brendan, Indah Sari, Tiffany Bryant, Geraldine Rosete, and Traci Marin. 2018. "AMPK: An Epigenetic Landscape Modulator" International Journal of Molecular Sciences 19, no. 10: 3238. https://doi.org/10.3390/ijms19103238
APA StyleGongol, B., Sari, I., Bryant, T., Rosete, G., & Marin, T. (2018). AMPK: An Epigenetic Landscape Modulator. International Journal of Molecular Sciences, 19(10), 3238. https://doi.org/10.3390/ijms19103238



