Can Aluminum Tolerant Wheat Cultivar Perform Better under Phosphate Deficient Conditions?
Abstract
1. Introduction
2. Results
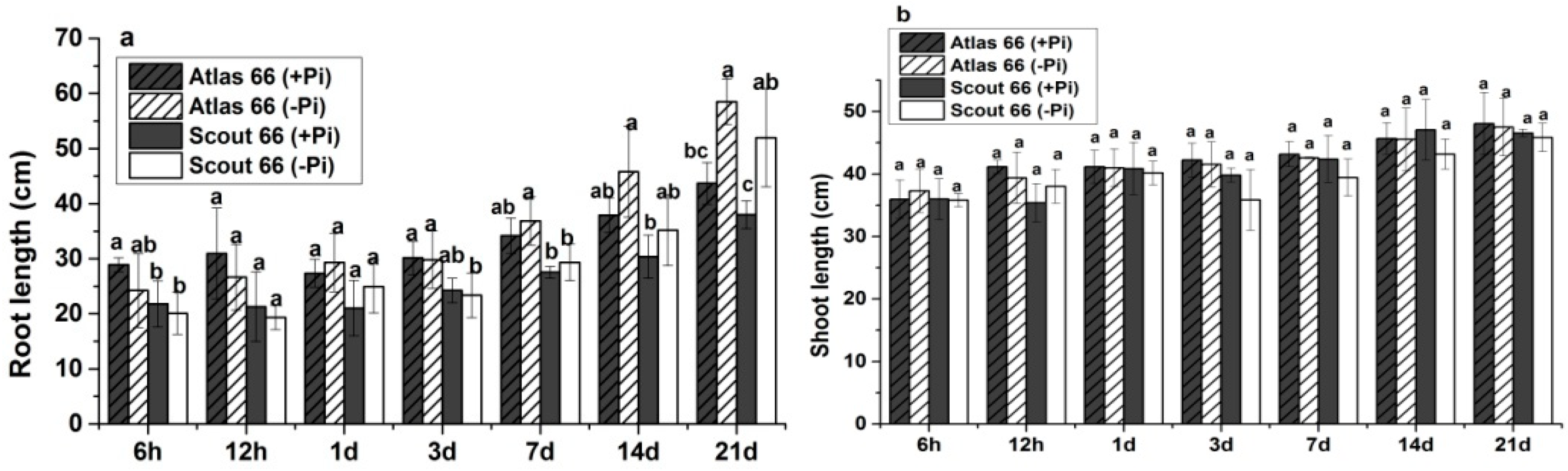
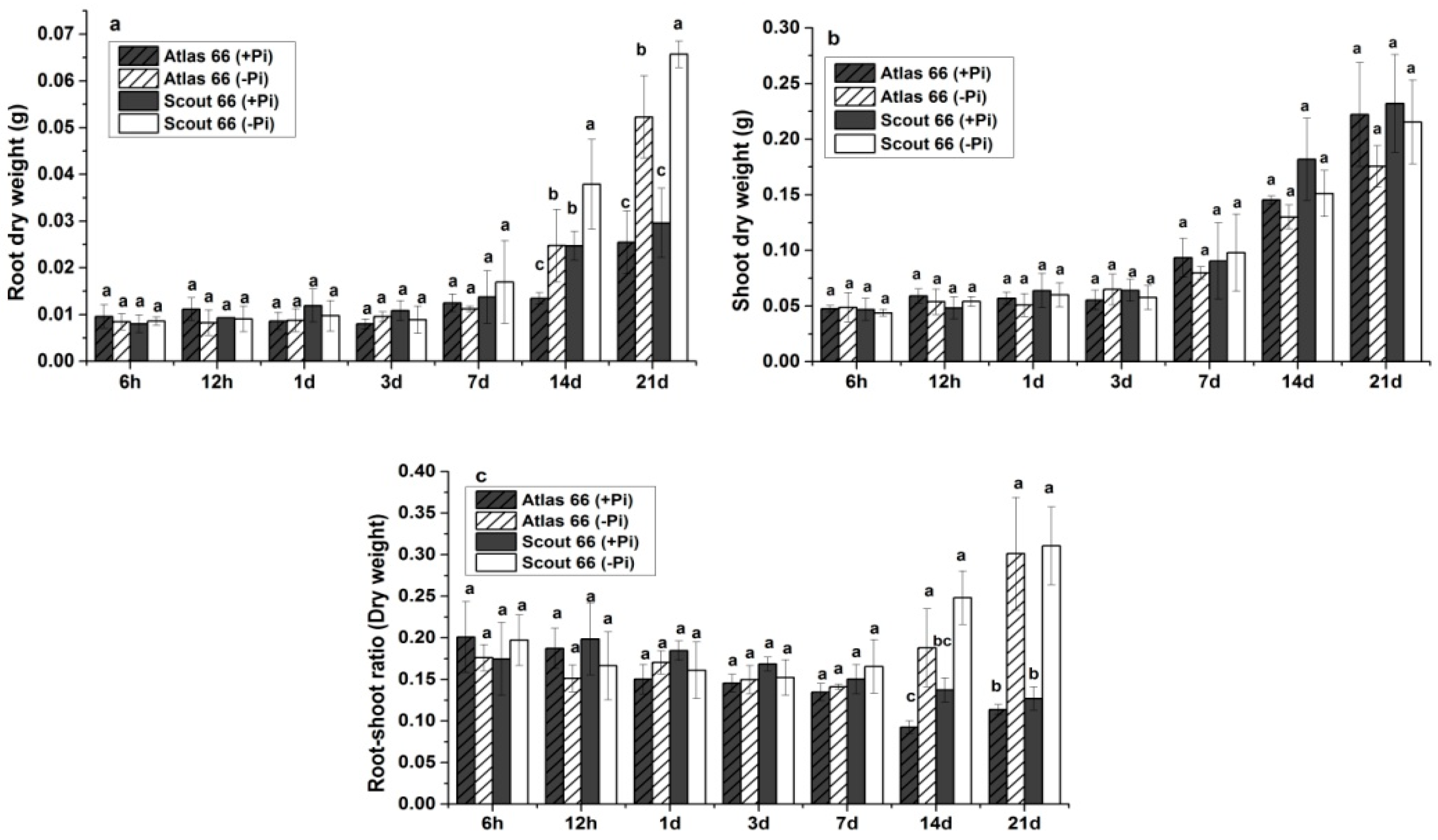
2.1. Scout 66 Showed Higher Root and Shoot Biomass under Phosphate Starvation
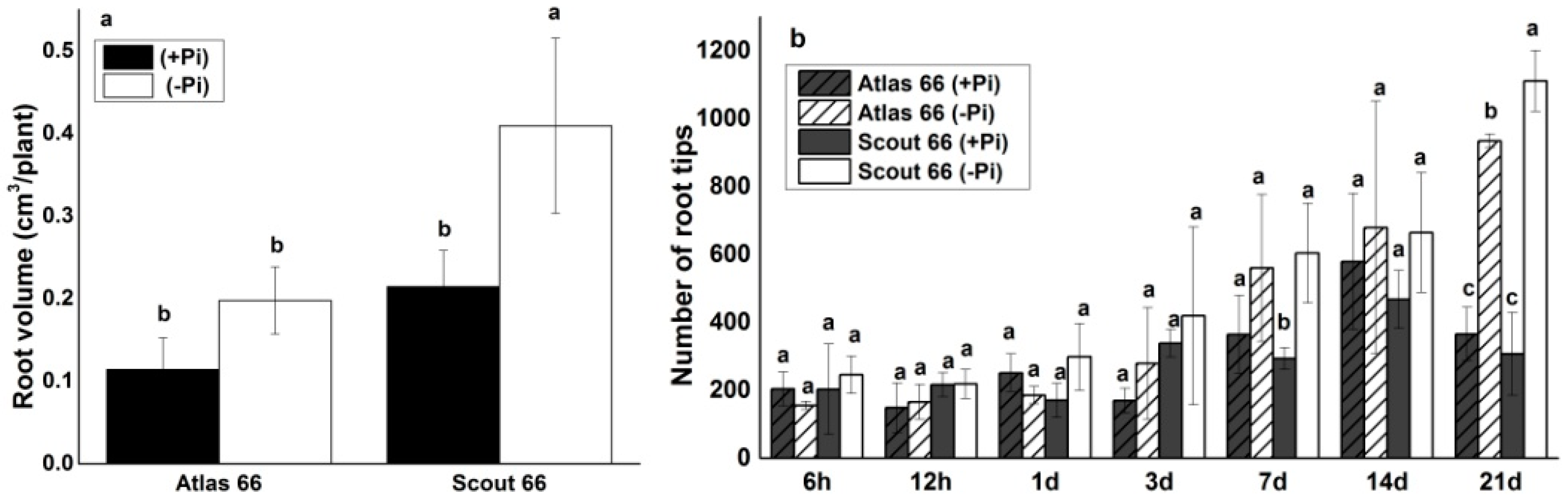
2.2. Scout 66 Produced Higher Root Volume and Had More Root Tip Number under Phosphate Deficiency
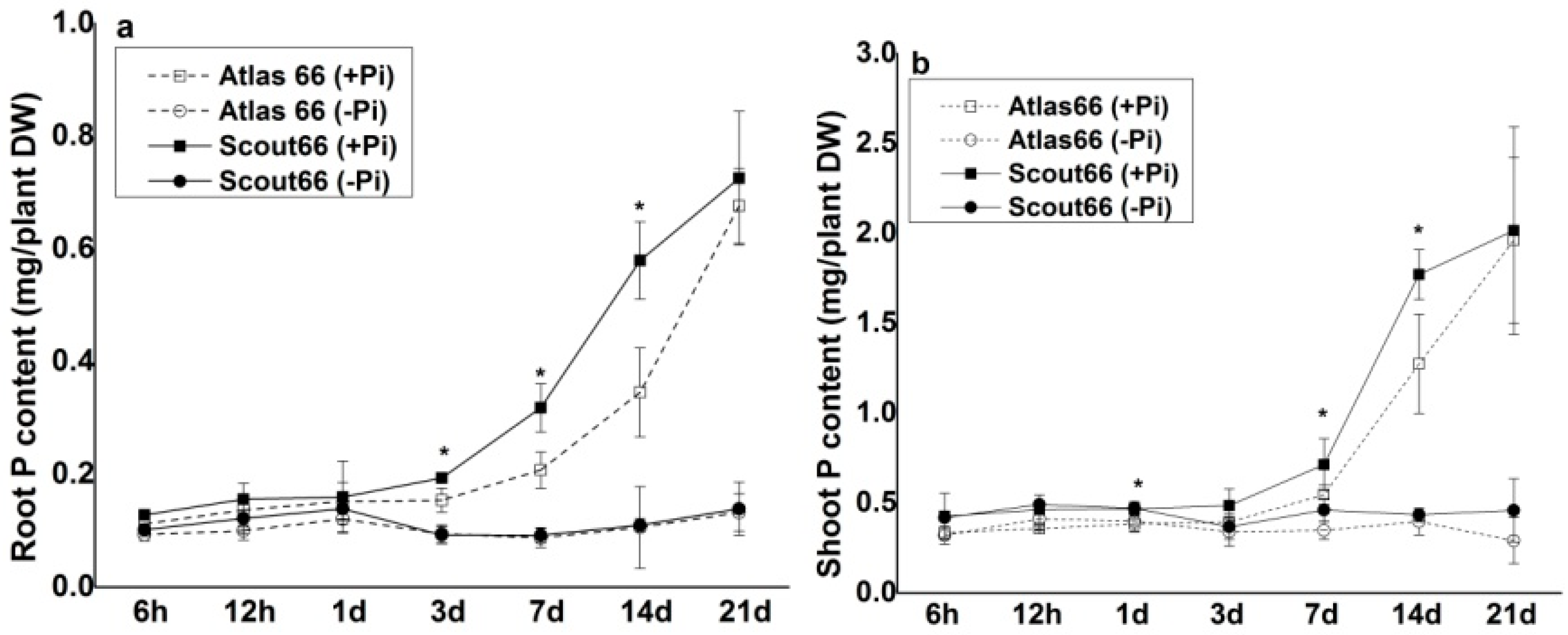
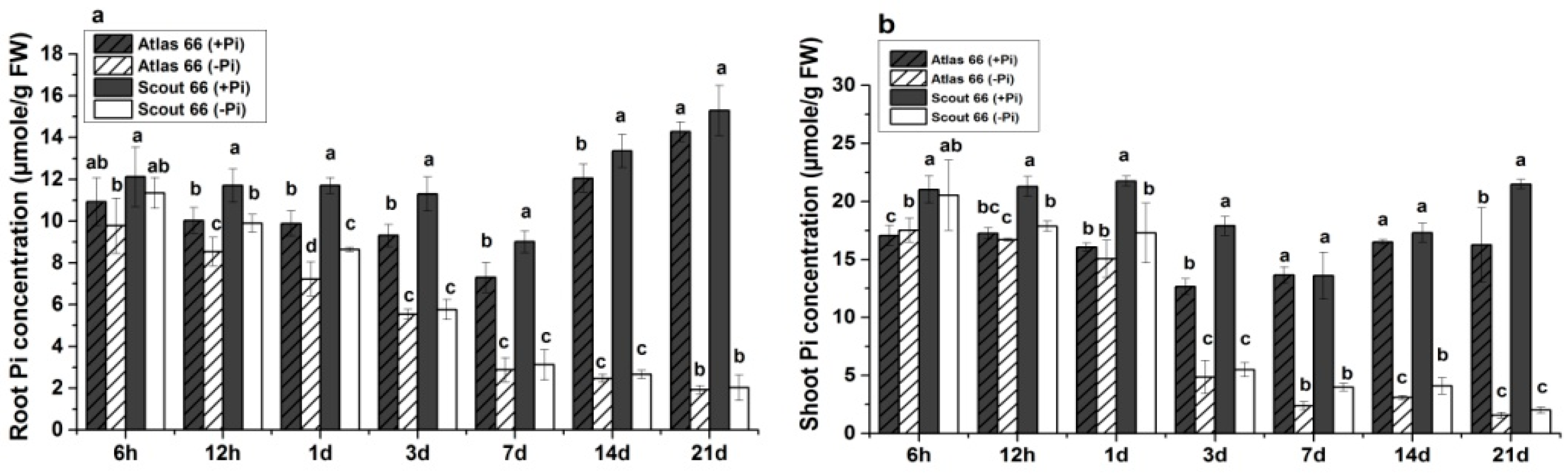
2.3. Higher Total P and Soluble Phosphate Concentrations in Scout 66 under Phosphate Deficiency
2.4. Scout 66 Contains Higher Iron (Fe) and Zinc (Zn) Contents in the Roots under Phosphate Deficiency
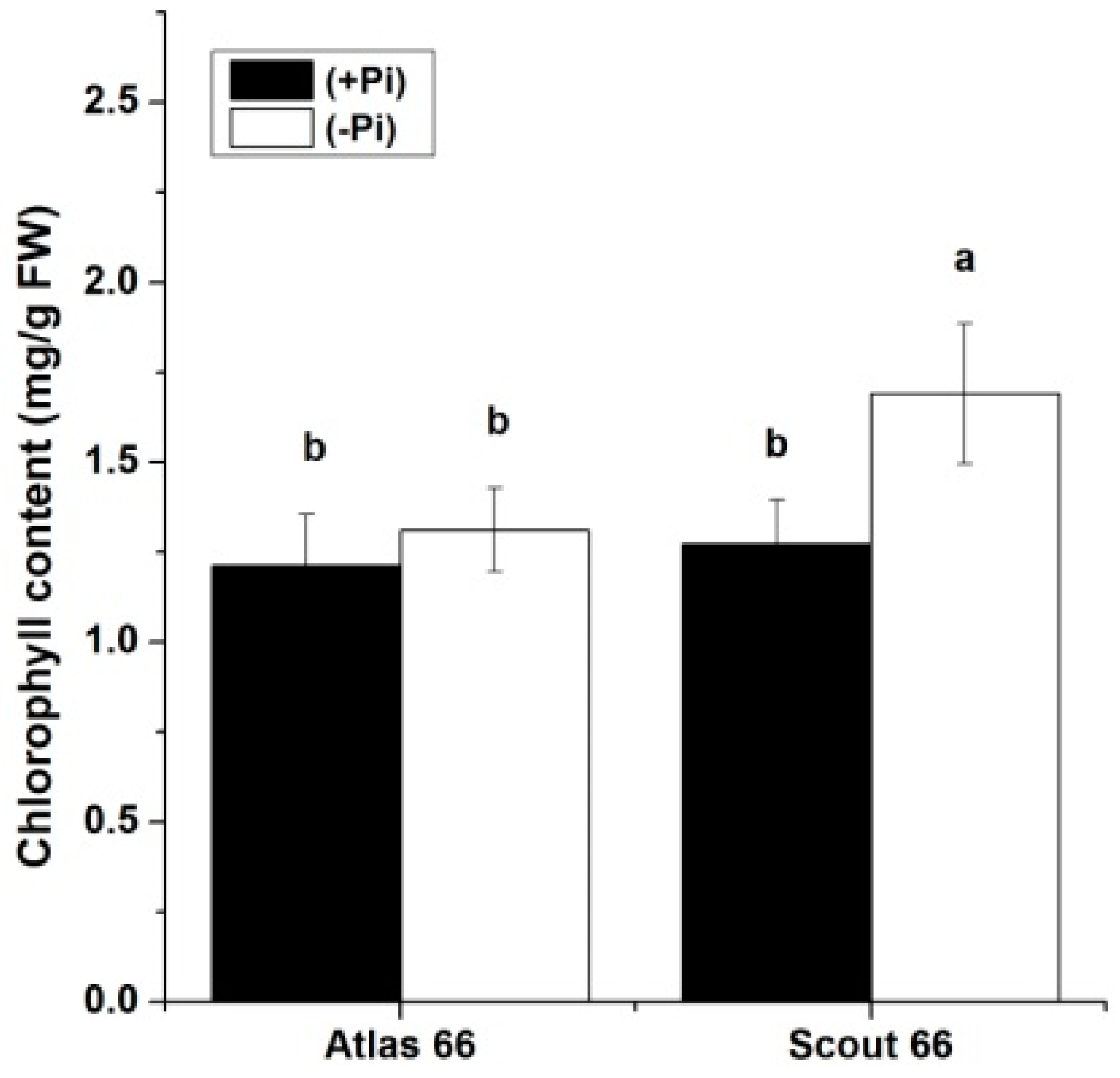
2.5. Scout 66 Can Maintain Higher Chlorophyll Content under Phosphate Deficiency
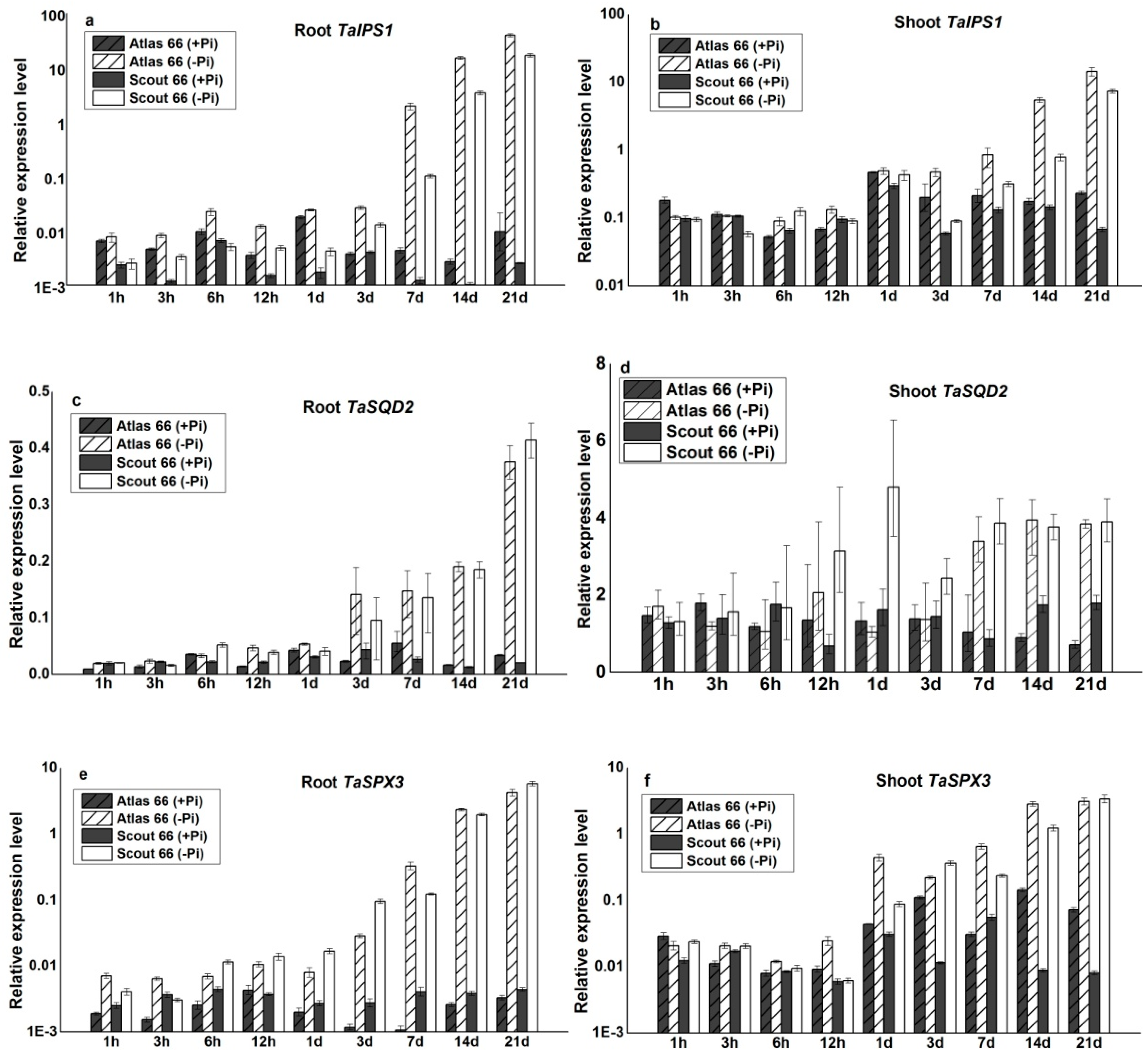
2.6. The Molecular Responses of the Two Cultivars under Phosphate Deficiency
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Growth Conditions
4.2. Root and Shoot Length, Biomass, Root Volume and Root Tip Numbers
4.3. Measurement of Total Phosphorus and Soluble Phosphate Concentration
4.4. Measurement of Iron and Zinc Concentration
4.5. Measurement of Chlorophyll Content
4.6. Phosphate Induced Marker Genes Expression Analysis by qRT-PCR
4.7. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Raghothama, K.G. Phosphate acquisition. Annu. Rev. Plant Physiol. Plant Mol. Biol. 1999, 50, 665–693. [Google Scholar] [CrossRef] [PubMed]
- Chiou, T.J.; Lin, S.I. Signaling network in sensing phosphate availability in plants. Annu. Rev. Plant Biol. 2011, 62, 185–206. [Google Scholar] [CrossRef] [PubMed]
- Bouain, N.; Doumas, P.; Rouached, H. Recent advances in understanding the molecular mechanisms regulating the root system response to phosphate deficiency in arabidopsis. Curr. Genom. 2016, 17, 308–314. [Google Scholar] [CrossRef] [PubMed]
- Rouached, H.; Tran, L.S. Regulation of plant mineral nutrition: Transport, sensing and signaling. Int. J. Mol. Sci. 2015, 16, 29717–29719. [Google Scholar] [CrossRef] [PubMed]
- Lambers, H.; Plaxton, W.C. Phosphorus: Back to the Roots; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2015. [Google Scholar]
- Secco, D.; Bouain, N.; Rouached, A.; Prom, U.T.C.; Hanin, M.; Pandey, A.K.; Rouached, H. Phosphate, phytate and phytases in plants: From fundamental knowledge gained in arabidopsis to potential biotechnological applications in wheat. Crit. Rev. Biotechnol. 2017, 37, 898–910. [Google Scholar] [CrossRef] [PubMed]
- Ham, B.K.; Chen, J.; Yan, Y.; Lucas, W.J. Insights into plant phosphate sensing and signaling. Curr. Opin. Biotechnol. 2018, 49, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Abel, S. Phosphate scouting by root tips. Curr. Opin. Plant Biol. 2017, 39, 168–177. [Google Scholar] [CrossRef] [PubMed]
- Shahzad, Z.; Amtmann, A. Food for thought: How nutrients regulate root system architecture. Curr. Opin. Plant Biol. 2017, 39, 80–87. [Google Scholar] [CrossRef] [PubMed]
- Gutierrez-Alanis, D.; Ojeda-Rivera, J.O.; Yong-Villalobos, L.; Cardenas-Torres, L.; Herrera-Estrella, L. Adaptation to phosphate scarcity: Tips from arabidopsis roots. Trends Plant Sci. 2018, 23, 721–730. [Google Scholar] [CrossRef] [PubMed]
- Lan, P.; Li, W.; Schmidt, W. ‘Omics’ approaches towards understanding plant phosphorus acquisition and use. In Annual Plant Reviews; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2015; Volume 48, pp. 65–97. [Google Scholar]
- Niu, Y.F.; Chai, R.S.; Jin, G.L.; Wang, H.; Tang, C.X.; Zhang, Y.S. Responses of root architecture development to low phosphorus availability: A review. Ann. Bot. 2012, 112, 391–408. [Google Scholar] [CrossRef] [PubMed]
- Vance, C.P. Symbiotic nitrogen fixation and phosphorus acquisition. Plant nutrition in a world of declining renewable resources. Plant Physiol. 2001, 127, 390–397. [Google Scholar] [CrossRef] [PubMed]
- Vance, C.P.; Uhde-Stone, C.; Allan, D.L. Phosphorus acquisition and use: Critical adaptations by plants for securing a nonrenewable resource. New Phytol. 2010, 157, 423–447. [Google Scholar] [CrossRef]
- Shen, J.; Yuan, L.; Zhang, J.; Li, H.; Bai, Z.; Chen, X.; Zhang, W.; Zhang, F. Phosphorus dynamics: From soil to plant. Plant Physiol. 2011, 156, 997–1005. [Google Scholar] [CrossRef] [PubMed]
- Yuan, H.; Liu, D. Signaling components involved in plant responses to phosphate starvation. J. Integr. Plant Biol. 2008, 50, 849–859. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Liu, D. Functions and regulation of phosphate starvation-induced secreted acid phosphatases in higher plants. Plant Sci. Int. J. Exp. Plant Biol. 2018, 271, 108–116. [Google Scholar] [CrossRef] [PubMed]
- Lan, P.; Li, W.; Schmidt, W. Complementary proteome and transcriptome profiling in phosphate-deficient arabidopsis roots reveals multiple levels of gene regulation. Mol. Cell. Proteom. MCP 2012, 11, 1156–1166. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Sun, J.; Miao, J.; Guo, J.; Shi, Z.; He, M.; Chen, Y.; Zhao, X.; Li, B.; Han, F.; et al. A phosphate starvation response regulator ta-phr1 is involved in phosphate signalling and increases grain yield in wheat. Ann. Bot. 2013, 111, 1139–1153. [Google Scholar] [CrossRef] [PubMed]
- Qi, W.; Manfield, I.W.; Muench, S.P.; Baker, A. Atspx1 affects the atphr1-DNA-binding equilibrium by binding monomeric atphr1 in solution. Biochem. J. 2017, 474, 3675–3687. [Google Scholar] [CrossRef] [PubMed]
- Rouached, H.; Secco, D.; Arpat, B.; Poirier, Y. The transcription factor phr1 plays a key role in the regulation of sulfate shoot-to-root flux upon phosphate starvation in arabidopsis. BMC Plant Biol. 2011, 11, 19. [Google Scholar] [CrossRef] [PubMed]
- Jung, J.Y.; Ried, M.K.; Hothorn, M.; Poirier, Y. Control of plant phosphate homeostasis by inositol pyrophosphates and the spx domain. Curr. Opin. Biotechnol. 2018, 49, 156–162. [Google Scholar] [CrossRef] [PubMed]
- Rubio, V.; Linhares, F.; Solano, R.; Martin, A.C.; Iglesias, J.; Leyva, A.; Paz-Ares, J. A conserved myb transcription factor involved in phosphate starvation signaling both in vascular plants and in unicellular algae. Genes Dev. 2001, 15, 2122–2133. [Google Scholar] [CrossRef] [PubMed]
- Xue, Y.B.; Xiao, B.X.; Zhu, S.N.; Mo, X.H.; Liang, C.Y.; Tian, J.; Liao, H.; Miriam, G. Gmphr25, a gmphr member up-regulated by phosphate starvation, controls phosphate homeostasis in soybean. J. Exp. Bot. 2017, 68, 4951–4967. [Google Scholar] [CrossRef] [PubMed]
- Zhong, Y.; Wang, Y.; Guo, J.; Zhu, X.; Shi, J.; He, Q.; Liu, Y.; Wu, Y.; Zhang, L.; Lv, Q.; et al. Rice spx6 negatively regulates the phosphate starvation response through suppression of the transcription factor phr2. New Phytol. 2018, 219, 135–148. [Google Scholar] [CrossRef] [PubMed]
- Nilsson, L.; Lundmark, M.; Jensen, P.E.; Nielsen, T.H. The arabidopsis transcription factor phr1 is essential for adaptation to high light and retaining functional photosynthesis during phosphate starvation. Physiol. Plant. 2012, 144, 35–47. [Google Scholar] [CrossRef] [PubMed]
- Lv, Q.; Zhong, Y.; Wang, Y.; Wang, Z.; Zhang, L.; Shi, J.; Wu, Z.; Liu, Y.; Mao, C.; Yi, K.; et al. Spx4 negatively regulates phosphate signaling and homeostasis through its interaction with phr2 in rice. Plant Cell 2014, 26, 1586–1597. [Google Scholar] [CrossRef] [PubMed]
- Nilsson, L.; Muller, R.; Nielsen, T.H. Increased expression of the myb-related transcription factor, phr1, leads to enhanced phosphate uptake in arabidopsis thaliana. Plant Cell Environ. 2007, 30, 1499–1512. [Google Scholar] [CrossRef] [PubMed]
- Bari, R.; Datt Pant, B.; Stitt, M.; Scheible, W.R. Pho2, microrna399, and phr1 define a phosphate-signaling pathway in plants. Plant Physiol. 2006, 141, 988–999. [Google Scholar] [CrossRef] [PubMed]
- Guo, M.; Ruan, W.; Li, C.; Huang, F.; Zeng, M.; Liu, Y.; Yu, Y.; Ding, X.; Wu, Y.; Wu, Z.; et al. Integrative comparison of the role of the phosphate response1 subfamily in phosphate signaling and homeostasis in rice. Plant Physiol. 2015, 168, 1762–1776. [Google Scholar] [CrossRef] [PubMed]
- Sun, L.; Song, L.; Zhang, Y.; Zheng, Z.; Liu, D. Arabidopsis phl2 and phr1 act redundantly as the key components of the central regulatory system controlling transcriptional responses to phosphate starvation. Plant Physiol. 2016, 170, 499–514. [Google Scholar] [CrossRef] [PubMed]
- Bustos, R.; Castrillo, G.; Linhares, F.; Puga, M.I.; Rubio, V.; Perez-Perez, J.; Solano, R.; Leyva, A.; Paz-Ares, J. A central regulatory system largely controls transcriptional activation and repression responses to phosphate starvation in arabidopsis. PLoS Genet. 2010, 6, e1001102. [Google Scholar] [CrossRef] [PubMed]
- Miura, K.; Rus, A.; Sharkhuu, A.; Yokoi, S.; Karthikeyan, A.S.; Raghothama, K.G.; Baek, D.; Koo, Y.D.; Jin, J.B.; Bressan, R.A.; et al. The arabidopsis sumo e3 ligase siz1 controls phosphate deficiency responses. Proc. Natl. Acad. Sci. USA 2005, 102, 7760–7765. [Google Scholar] [PubMed]
- Puga, M.I.; Mateos, I.; Charukesi, R.; Wang, Z.; Franco-Zorrilla, J.M.; de Lorenzo, L.; Irigoyen, M.L.; Masiero, S.; Bustos, R.; Rodriguez, J.; et al. Spx1 is a phosphate-dependent inhibitor of phosphate starvation response 1 in arabidopsis. Proc. Natl. Acad. Sci. USA 2014, 111, 14947–14952. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Ruan, W.; Shi, J.; Zhang, L.; Xiang, D.; Yang, C.; Li, C.; Wu, Z.; Liu, Y.; Yu, Y.; et al. Rice spx1 and spx2 inhibit phosphate starvation responses through interacting with phr2 in a phosphate-dependent manner. Proc. Natl. Acad. Sci. USA 2014, 111, 14953–14958. [Google Scholar] [CrossRef] [PubMed]
- Wild, R.; Gerasimaite, R.; Jung, J.Y.; Truffault, V.; Pavlovic, I.; Schmidt, A.; Saiardi, A.; Jessen, H.J.; Poirier, Y.; Hothorn, M.; et al. Control of eukaryotic phosphate homeostasis by inositol polyphosphate sensor domains. Science 2016, 352, 986–990. [Google Scholar] [CrossRef] [PubMed]
- Zheng, L.; Lan, P.; Shen, R.F.; Li, W.F. Proteomics of aluminum tolerance in plants. Proteomics 2014, 14, 566–578. [Google Scholar] [CrossRef] [PubMed]
- Du, H.; Yang, C.; Ding, G.; Shi, L.; Xu, F. Genome-wide identification and characterization of spx domain-containing members and their responses to phosphate deficiency in brassica napus. Front. Plant Sci. 2017, 8, 35. [Google Scholar] [CrossRef] [PubMed]
- Ma, J.F.; Zheng, S.J.; Li, X.F.; Takeda, K.; Matsumoto, H. A rapid hydroponic screening for aluminium tolerance in barley. Plant Soil 1997, 191, 133–137. [Google Scholar] [CrossRef]
- Tabuchi, A.; Kikui, S.; Matsumoto, H. Differential effects of aluminium on osmotic potential and sugar accumulation in the root cells of al-resistant and al-sensitive wheat. Physiol. Plant. 2004, 120, 106–112. [Google Scholar] [CrossRef] [PubMed]
- Cocker, K.M.; Evans, D.E.; Hodson, M.J. The amelioration of aluminium toxicity by silicon in wheat (Triticum aestivum L.): Malate exudation as evidence for an in planta mechanism. Planta 1998, 204, 318–323. [Google Scholar] [CrossRef]
- Scheible, W.R.; Rojas-Triana, M. Sensing, signalling, and control of phosphate starvation in plants: Molecular players and applications. In Annual Plant Reviews; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2015; Volume 48, p. 474. [Google Scholar]
- Lambers, H.; Clode, P.L.; Hawkins, H.J.; Laliberté, E.; Oliveira, R.S.; Reddell, P.; Shane, M.W.; Stitt, M.; Weston, P. Metabolic adaptations of the non-mycotrophic proteaceae to soils with low phosphorus availability. In Annual Plant Reviews; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2015; Volume 48, pp. 289–335. [Google Scholar]
- Misson, J.; Raghothama, K.G.; Jain, A.; Jouhet, J.; Block, M.A.; Bligny, R.; Ortet, P.; Creff, A.; Somerville, S.; Rolland, N.; et al. A genome-wide transcriptional analysis using arabidopsis thaliana affymetrix gene chips determined plant responses to phosphate deprivation. Proc. Natl. Acad. Sci. USA 2005, 102, 11934–11939. [Google Scholar] [CrossRef] [PubMed]
- Briat, J.F.; Rouached, H.; Tissot, N.; Gaymard, F.; Dubos, C. Integration of p, s, fe, and zn nutrition signals in arabidopsis thaliana: Potential involvement of phosphate starvation response 1 (phr1). Front. Plant Sci. 2015, 6, 290. [Google Scholar] [CrossRef] [PubMed]
- Chaiwong, N.; Prom, U.T.C.; Bouain, N.; Lacombe, B.; Rouached, H. Individual versus combinatorial effects of silicon, phosphate, and iron deficiency on the growth of lowland and upland rice varieties. Int. J. Mol. Sci. 2018, 19, 899. [Google Scholar] [CrossRef] [PubMed]
- Dong, J.; Pineros, M.A.; Li, X.; Yang, H.; Liu, Y.; Murphy, A.S.; Kochian, L.V.; Liu, D. An arabidopsis abc transporter mediates phosphate deficiency-induced remodeling of root architecture by modulating iron homeostasis in roots. Mol. Plant 2017, 10, 244–259. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.Y.; Shirley, N.; Genc, Y.; Shi, B.; Langridge, P. Phosphate utilization efficiency correlates with expression of low-affinity phosphate transporters and noncoding rna, ips1, in barley. Plant Physiol. 2011, 156, 1217–1229. [Google Scholar] [CrossRef] [PubMed]
- Hammond, J.P.; Broadley, M.R.; White, P.J. Genetic responses to phosphorus deficiency. Ann. Bot. 2004, 94, 323–332. [Google Scholar] [CrossRef] [PubMed]
- Yang, G.; Ding, G.; Shi, L.; Cai, H.; Xu, F. Characterization of phosphorus starvation-induced gene bnspx3 in brassica napus. Plant Soil 2012, 350, 339–351. [Google Scholar] [CrossRef]
- Gamuyao, R.; Chin, J.H.; Pariasca-Tanaka, J.; Pesaresi, P.; Catausan, S.; Dalid, C.; Slamet-Loedin, I.; Tecson-Mendoza, E.M.; Wissuwa, M.; Heuer, S. The protein kinase pstol1 from traditional rice confers tolerance of phosphorus deficiency. Nature 2012, 488, 535–539. [Google Scholar] [CrossRef] [PubMed]
- Chevalier, F.; Rossignol, M. Proteomic analysis of arabidopsis thaliana ecotypes with contrasted root architecture in response to phosphate deficiency. J. Plant Physiol. 2011, 168, 1885–1890. [Google Scholar] [CrossRef] [PubMed]
- Lan, P.; Li, W.; Schmidt, W. Genome-wide co-expression analysis predicts protein kinases as important regulators of phosphate deficiency-induced root hair remodeling in arabidopsis. BMC Genom. 2013, 14, 210. [Google Scholar] [CrossRef] [PubMed]
- Xue, Y.; Xia, H.; Christie, P.; Zhang, Z.; Li, L.; Tang, C. Crop acquisition of phosphorus, iron and zinc from soil in cereal/legume intercropping systems: A critical review. Ann. Bot. 2016, 117, 363–377. [Google Scholar] [CrossRef] [PubMed]
- Bouain, N.; Shahzad, Z.; Rouached, A.; Khan, G.A.; Berthomieu, P.; Abdelly, C.; Poirier, Y.; Rouached, H. Phosphate and zinc transport and signalling in plants: Toward a better understanding of their homeostasis interaction. J. Exp. Bot. 2014, 65, 5725–5741. [Google Scholar] [CrossRef] [PubMed]
- Kisko, M.; Bouain, N.; Safi, A.; Medici, A.; Akkers, R.C.; Secco, D.; Fouret, G.; Krouk, G.; Aarts, M.G.; Busch, W. Lpcat1 controls phosphate homeostasis in a zinc-dependent manner. Elife 2018, 7, e32077. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Lan, P. Genome-wide analysis of overlapping genes regulated by iron deficiency and phosphate starvation reveals new interactions in arabidopsis roots. BMC Res. Notes 2015, 8, 555. [Google Scholar] [CrossRef] [PubMed]
- WenXuan, M.; Tian, X.; Gale, W.J.; Yang, X.; XinChun, L. Tolerance to zn deficiency and p-zn interaction in wheat seedlings cultured in chelator-buffered solutions. J. Arid Land 2011, 3, 206–213. [Google Scholar]
- Shi, J.; Hu, H.; Zhang, K.; Zhang, W.; Yu, Y.; Wu, Z.; Wu, P. The paralogous spx3 and spx5 genes redundantly modulate pi homeostasis in rice. J. Exp. Bot. 2014, 65, 859–870. [Google Scholar] [PubMed]
- Murphy, J.; Riley, J.P. A modified single solution method for the determination of phosphate in natural waters. Anal. Chim. Acta 1962, 27, 31–36. [Google Scholar] [CrossRef]
- Chiou, T.-J.; Aung, K.; Lin, S.-I.; Wu, C.-C.; Chiang, S.-F.; Su, C.-L. Regulation of phosphate homeostasis by microrna in arabidopsis. Plant Cell 2006, 18, 412–421. [Google Scholar] [CrossRef] [PubMed]
- Arnon, D.I. Copper enzymes in isolated chloroplasts. Polyphenoloxidase in beta vulgaris. Plant Physiol. 1949, 24, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative pcr and the 2−δδct method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
| Cultivars | Phosphate Treatment | Fe (mg/g DW) | Zn (µg/g DW) | ||
|---|---|---|---|---|---|
| Root | Shoot | Root | Shoot | ||
| Atlas 66 | (+Pi) | 7.08 ± 2.64 b | 0.13 ± 0.004 a | 128.5 ± 5.0 b | 65.6 ± 17.1 a |
| (−Pi) | 8.31 ± 1.51 ab | 0.12 ± 0.024 a | 106.5 ± 19.5 b | 49.7 ± 8.7 a | |
| Scout 66 | (+Pi) | 5.62 ± 1.45 b | 0.10 ± 0.044 a | 189.2 ± 39.0 a | 68.2 ± 9.8 a |
| (−Pi) | 12.41 ± 4.25 a | 0.12 ± 0.007 a | 114.5 ± 1.5 b | 60.6 ± 4.3 a | |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Karim, M.R.; Dong, X.; Zheng, L.; Shen, R.; Lan, P. Can Aluminum Tolerant Wheat Cultivar Perform Better under Phosphate Deficient Conditions? Int. J. Mol. Sci. 2018, 19, 2964. https://doi.org/10.3390/ijms19102964
Karim MR, Dong X, Zheng L, Shen R, Lan P. Can Aluminum Tolerant Wheat Cultivar Perform Better under Phosphate Deficient Conditions? International Journal of Molecular Sciences. 2018; 19(10):2964. https://doi.org/10.3390/ijms19102964
Chicago/Turabian StyleKarim, Mohammad Rezaul, Xiaoying Dong, Lu Zheng, Renfang Shen, and Ping Lan. 2018. "Can Aluminum Tolerant Wheat Cultivar Perform Better under Phosphate Deficient Conditions?" International Journal of Molecular Sciences 19, no. 10: 2964. https://doi.org/10.3390/ijms19102964
APA StyleKarim, M. R., Dong, X., Zheng, L., Shen, R., & Lan, P. (2018). Can Aluminum Tolerant Wheat Cultivar Perform Better under Phosphate Deficient Conditions? International Journal of Molecular Sciences, 19(10), 2964. https://doi.org/10.3390/ijms19102964






