RETRACTED: Microenvironment Stimuli HGF and Hypoxia Differently Affected miR-125b and Ets-1 Function with Opposite Effects on the Invasiveness of Bone Metastatic Cells: A Comparison with Breast Carcinoma Cells
Abstract
1. Introduction
2. Results
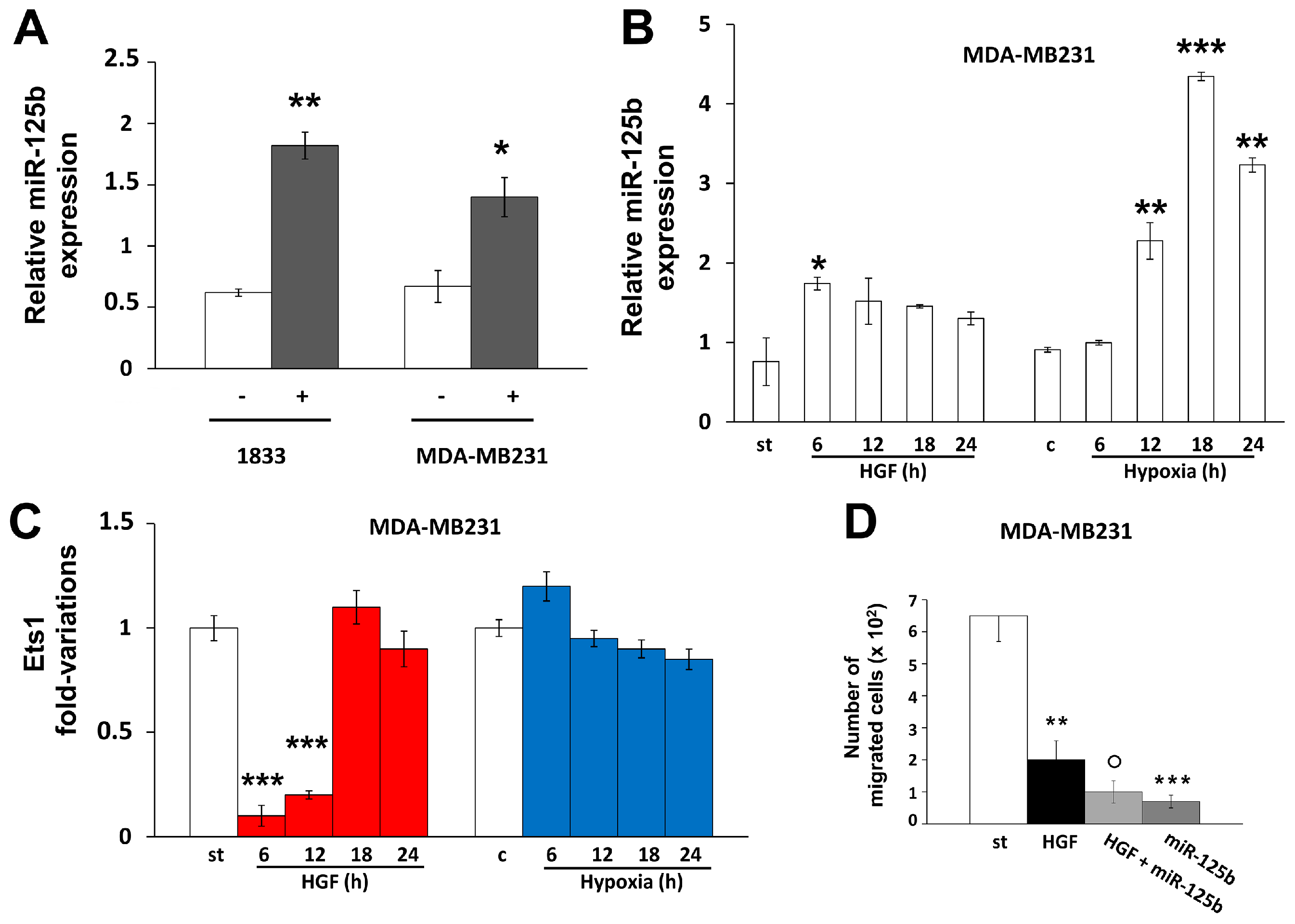
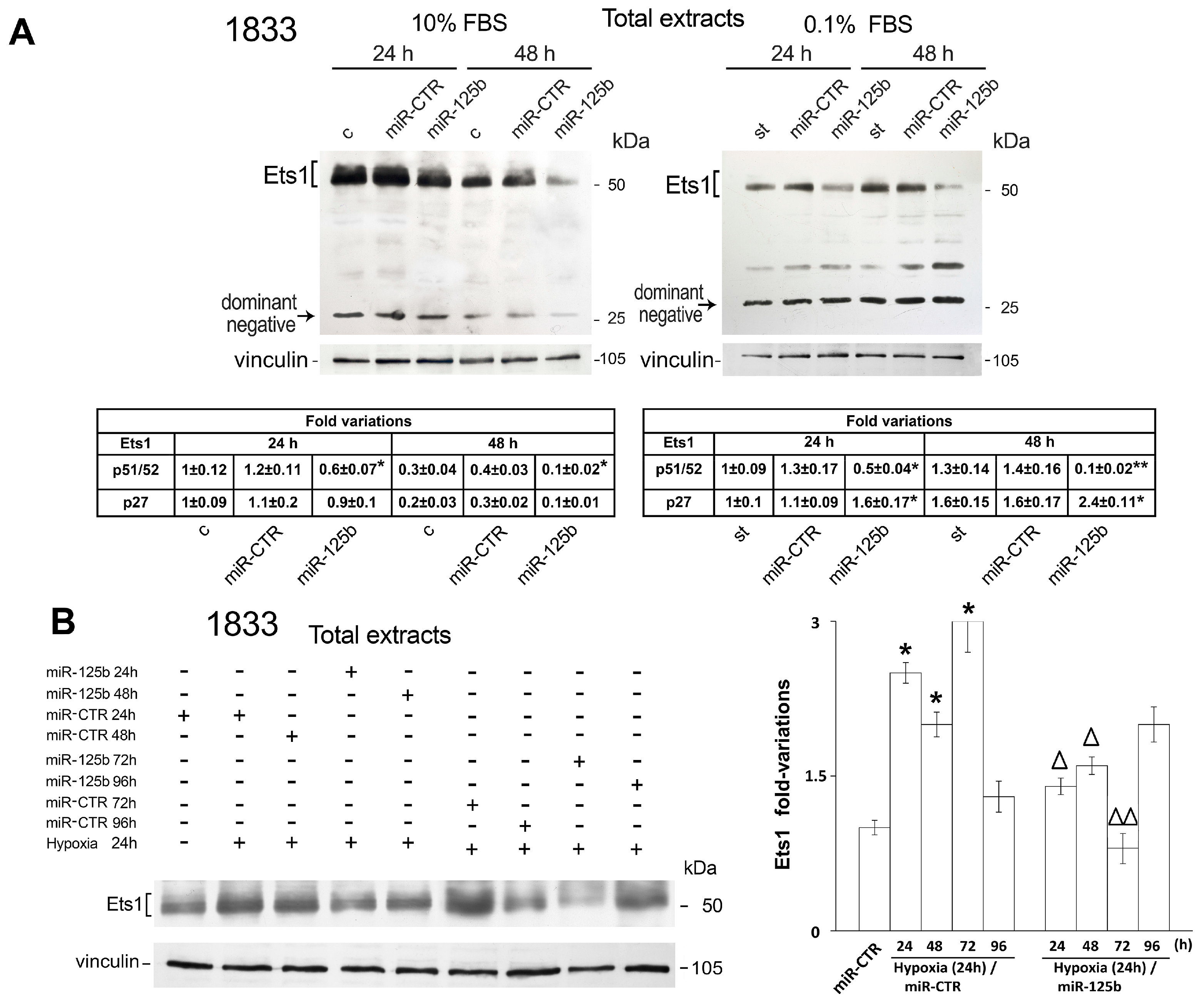
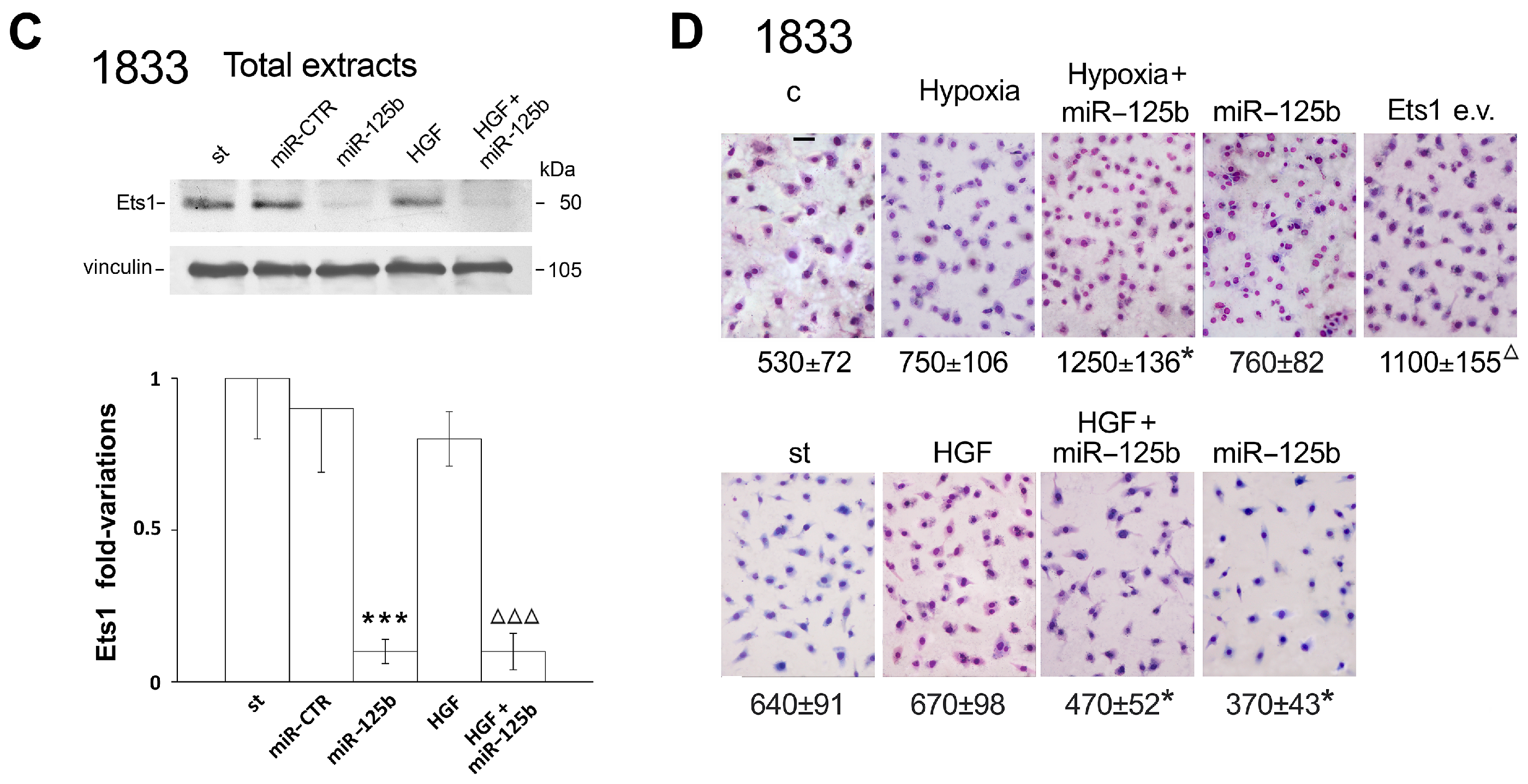
2.1. Different Effects of Stimuli of Bone Metastasis Microenvironment on the Expression of miR-125b, the Ets1 Protein Levels and Invasiveness in MDA-MB231 and 1833 Cells
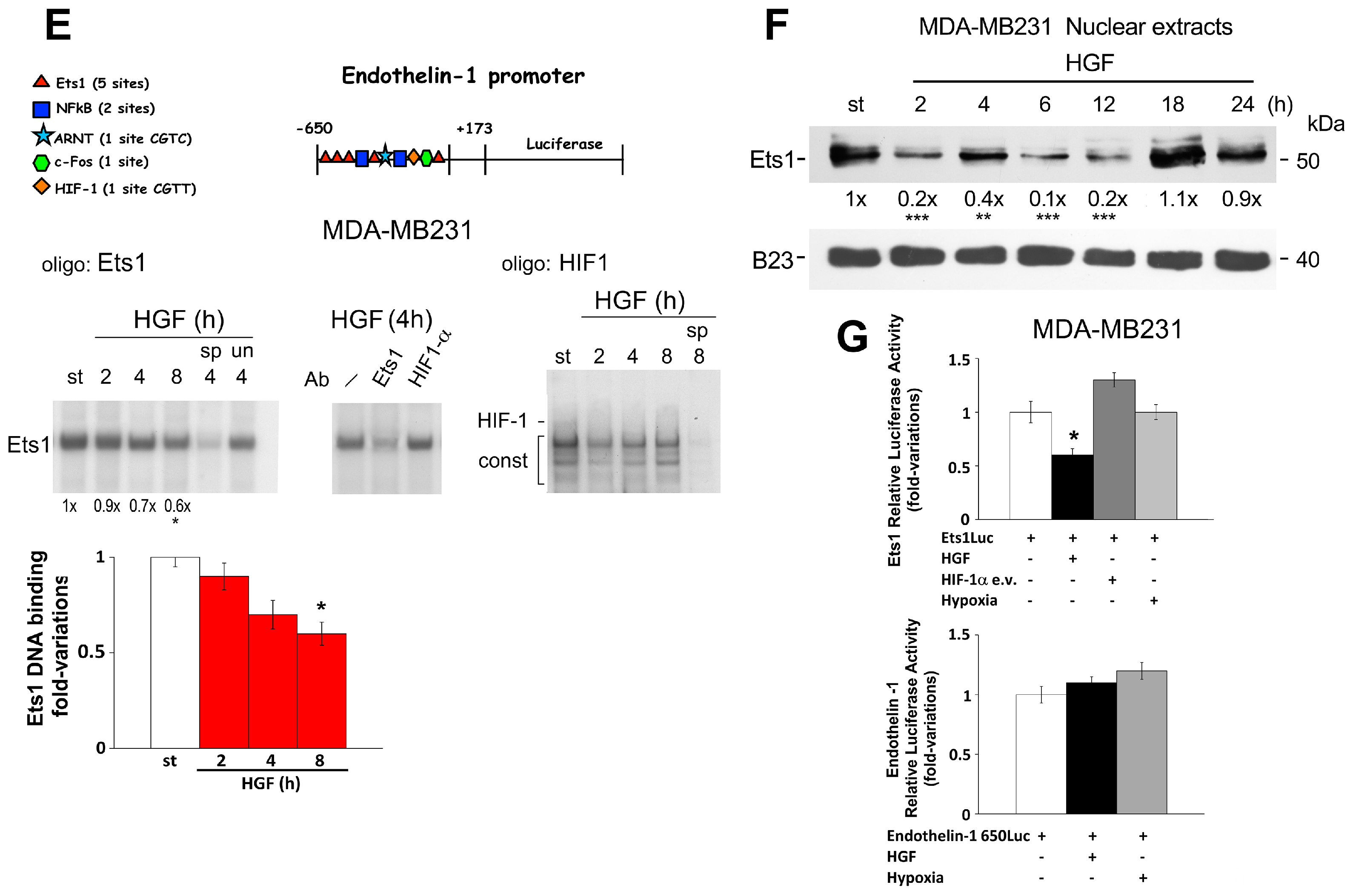
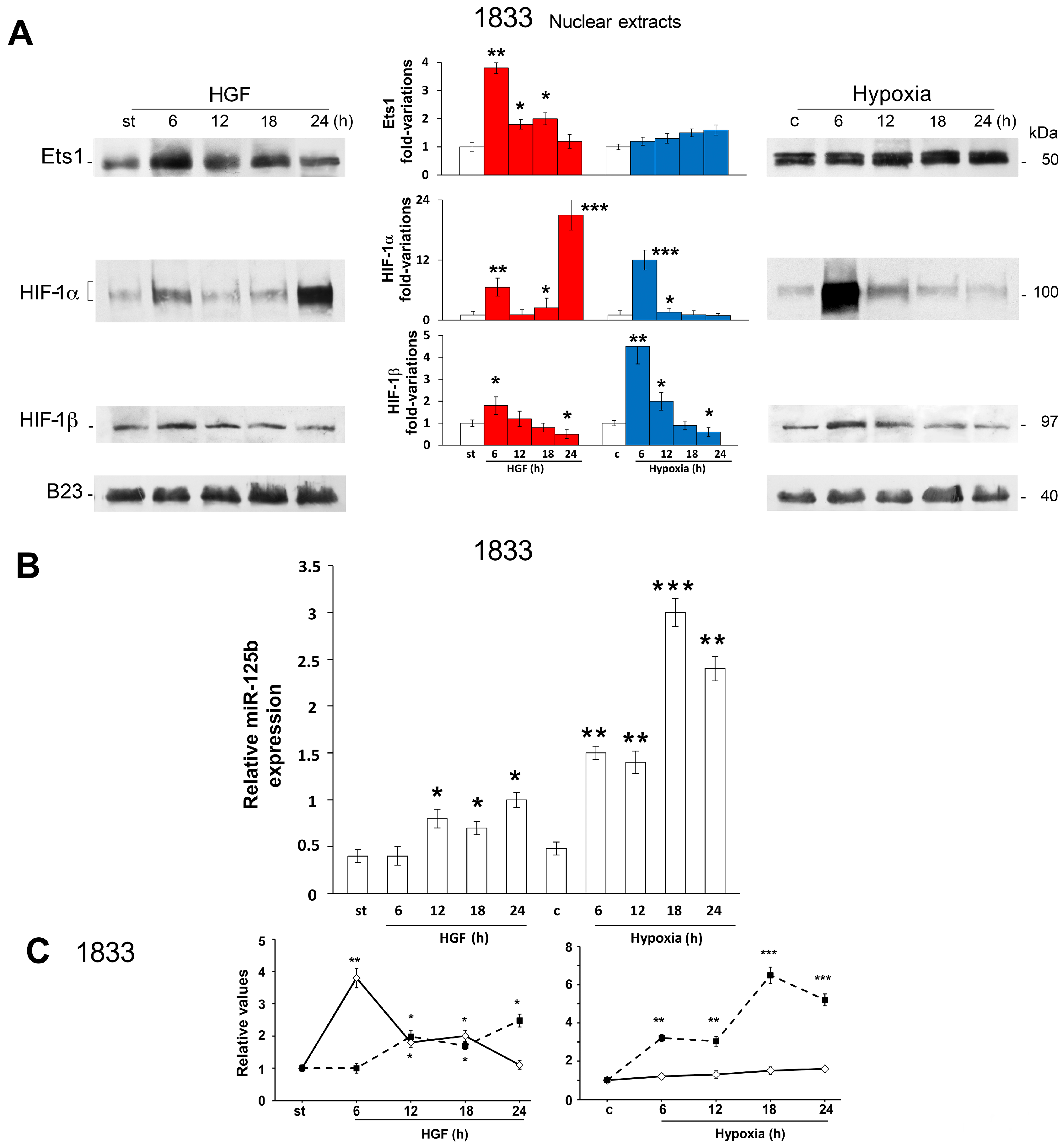
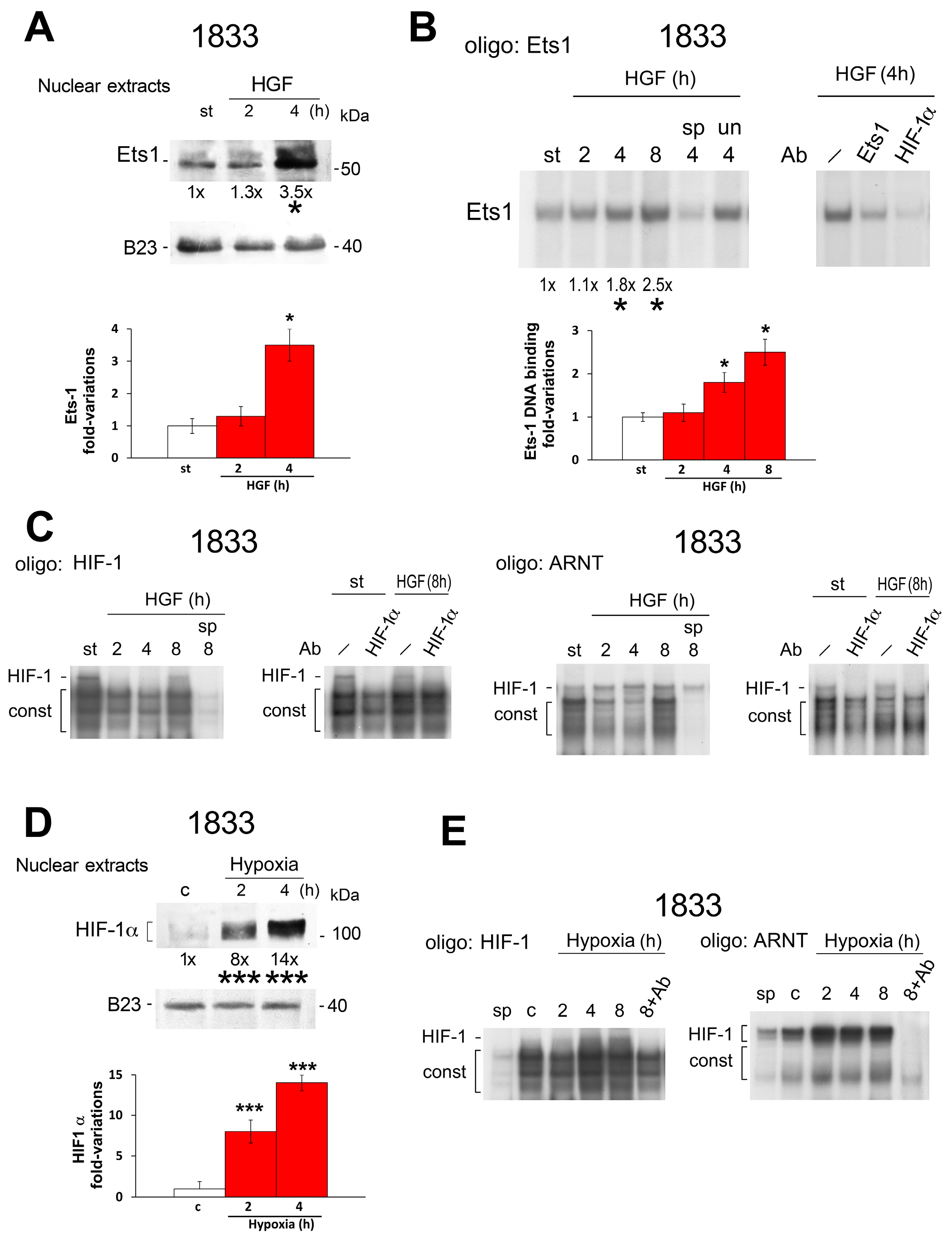
2.2. Reciprocal Regulation of Ets-1 and HIF-1α Transactivating Activities in 1833 Cells
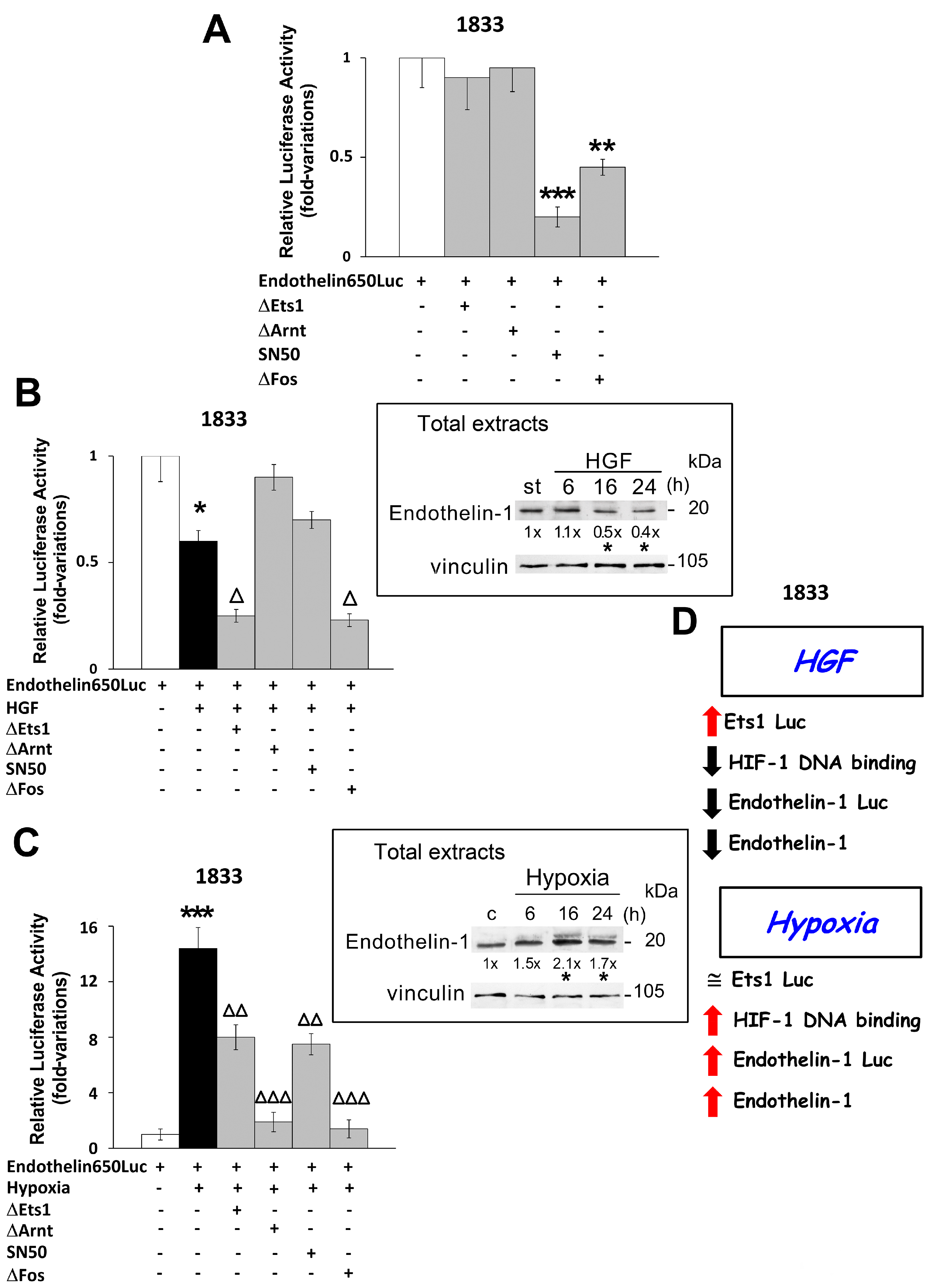
2.3. Regulation of Endothelin-1 Expression by Microenvironment Stimuli in 1833 Cells
3. Discussion
4. Materials and Methods
4.1. Cell Cultures and miR-125b Transfection
4.2. Western Blot Analysis
4.3. Purification of Small RNA Species (miRNAs), and Quantification of miR-125b
4.4. Matrigel Invasion Assay
4.5. Constructs and Cell Transfections
4.6. EMSA Analysis
4.7. Statistical Analysis
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| HGF | hepatocyte growth factor |
| HIF | hypoxia inducible factor |
| HRE | HIF-1 responsive element |
| MET | mesenchymal–epithelial transition |
References
- Croucher, P.I.; McDonald, M.M.; Martin, T.J. Bone metastasis: The importance of the neighbourhood. Nat. Rev. Cancer 2016, 16, 373–386. [Google Scholar] [CrossRef] [PubMed]
- Bendinelli, P.; Maroni, P.; Matteucci, E.; Desiderio, M.A. Cell and signal components of the microenvironment of bone metastasis are affected by hypoxia. Int. J. Mol. Sci. 2016, 17, 706. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Yan, L.X.; Wu, Q.N.; Du, Z.M.; Chen, J.; Liao, D.Z.; Huang, M.Y.; Hou, J.H.; Wu, Q.L.; Zeng, M.S.; et al. miR-125b is methylated and functions as a tumor suppressor by regulating the ETS1 proto-oncogene in human invasive breast cancer. Cancer Res. 2011, 71, 3552–3562. [Google Scholar] [CrossRef] [PubMed]
- Bendinelli, P.; Maroni, P.; Matteucci, E.; Desiderio, M.A. Epigenetic regulation of HGF/Met receptor axis is critical for the outgrowth of bone metastasis from breast carcinoma. Cell Death Dis. 2017, 8, e2578. [Google Scholar] [CrossRef] [PubMed]
- Maroni, P.; Puglisi, R.; Mattia, G.; Carè, A.; Matteucci, E.; Bendinelli, P.; Desiderio, M.A. In bone metastasis miR-34a-5p absence inversely correlates with Met expression, while Met oncogene is unaffected by miR-34a-5p in non-metastatic and metastatic breast carcinomas. Carcinogenesis 2017, 38, 492–503. [Google Scholar] [CrossRef] [PubMed]
- Vallet, S.; Bashari, M.H.; Fan, F.J.; Malvestiti, S.; Schneeweiss, A.; Wuchter, P.; Jäger, D.; Podar, K. Pre-osteoblasts stimulate migration of breast cancer cells via the HGF/MET pathway. PLoS ONE 2016, 11, e0150507. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.-T.; Tsou, H.-K.; Chang, C.-H.; Tang, C.-H. Hepatocyte growth factor increases osteopontin expression in human osteoblasts through PI3K, Akt, c-Src, and AP-1 signaling pathway. PLoS ONE 2012, 7, e38378. [Google Scholar] [CrossRef] [PubMed]
- Maroni, P.; Matteucci, E.; Luzzati, A.; Perrucchini, G.; Bendinelli, P.; Desiderio, M.A. Nuclear co-localization and functional interaction of COX-2 and HIF-1α characterize bone metastasis of human breast carcinoma. Breast Cancer Res. Treat. 2011, 129, 433–450. [Google Scholar] [CrossRef] [PubMed]
- Schito, L.; Semenza, G.L. Hypoxia-inducible factors: Master regulators of cancer progression. Trends Cancer 2016, 2, 758–770. [Google Scholar] [CrossRef] [PubMed]
- Gilkes, D.M. Implications of hypoxia in breast cancer metastasis to bone. Int. J. Mol.Sci. 2016, 17, 1669. [Google Scholar] [CrossRef] [PubMed]
- Furlan, A.; Vercamer, C.; Bouali, F.; Damour, I.; Chotteau-Lelievre, A.; Wernert, N.; Desbiens, X.; Pourtier, A. Ets-1 controls breast cancer cell balance between invasion and growth. Int. J. Cancer 2014, 135, 2317–2328. [Google Scholar] [CrossRef] [PubMed]
- Oikawa, T. ETS transcription factors: Possible targets for cancer therapy. Cancer Sci. 2004, 95, 626–633. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Wu, D.; Aldarouish, M.; Yin, X.; Li, C.; Wang, C. ETS-1: A potential target of glycolysis for metabolic therapy by regulating glucose metabolism in pancreatic cancer. Int. J. Oncol. 2017, 50, 232–240. [Google Scholar] [CrossRef] [PubMed]
- Bendinelli, P.; Maroni, P.; Matteucci, E.; Desiderio, M.A. HGF and TGFβ1 differently influenced Wwox regulatory function on Twist program for mesenchymal-epithelial transition in bone metastatic versus parental breast carcinoma cells. Mol. Cancer 2015, 14, 112. [Google Scholar] [CrossRef] [PubMed]
- Bendinelli, P.; Maroni, P.; Matteucci, E.; Luzzati, A.; Perrucchini, G.; Desiderio, M.A. Microenvironmental stimuli affect Endothelin-1 signaling responsible for invasiveness and osteomimicry of bone metastasis from breast cancer. Biochim. Biophys. Acta 2014, 1843, 815–826. [Google Scholar] [CrossRef] [PubMed]
- Maroni, P.; Bendinelli, P.; Morelli, D.; Drago, L.; Luzzati, A.; Perrucchini, G.; Bonini, C.; Matteucci, E.; Desiderio, M.A. High SPARC expression starting from dysplasia, associated with breast carcinoma, is predictive for bone metastasis without enhancement of plasma levels. Int. J. Mol. Sci. 2015, 16, 28108–28122. [Google Scholar] [CrossRef] [PubMed]
- Yin, H.; Sun, Y.; Wang, X.; Park, J.; Zhang, Y.; Li, M.; Yin, J.; Liu, Q.; Wei, M. Progress on the relationship between miR-125 family and tumorigenesis. Exp. Cell Res. 2015, 339, 252–260. [Google Scholar] [CrossRef] [PubMed]
- Pronina, I.V.; Loginov, V.I.; Burdennyy, A.M.; Fridman, M.V.; Senchenko, V.N.; Kazubskaya, T.P.; Kushlinskii, N.E.; Dmitriev, A.A.; Braga, E.A. DNA methylation contributes to deregulation of 12 cancer-associated microRNAs and breast cancer progression. Gene 2017, 604, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Al Rifai, O.; Chow, J.; Lacombe, J.; Julien, C.; Faubert, D.; Susan-Resiga, D.; Essalmani, R.; Creemers, J.W.; Seidah, N.G.; Ferron, M. Proprotein convertase furin regulates osteocalcin and bone endocrine function. J. Clin. Investig. 2017, 127, 4104–4117. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Huang, R.; Fan, W.; Prasadam, I.; Crawford, R.; Xiao, Y. Mesenchymal stromal cells regulate the cell mobility and the immune response during osteogenesis through secretion of vascular endothelial growth factor A. J. Tissue Eng. Regen. Med. 2017. [Google Scholar] [CrossRef] [PubMed]
- Dallas, S.L.; Park-Snyder, S.; Miyazono, K.; Twardzik, D.; Mundy, G.R.; Bonewald, L.F. Characterization and autoregulation of latent transforming growth factor β (TGFβ) complexes in osteoblast-like cell lines. J. Biol. Chem. 1994, 269, 6815–6822. [Google Scholar] [PubMed]
- Xie, Y.; Chen, Y.; Zhang, L.; Ge, W.; Tang, P. The roles of bone-derived exosomes and exosomal microRNAs in regulating bone remodelling. J. Cell. Mol. Med. 2017, 21, 1033–1041. [Google Scholar] [CrossRef] [PubMed]
- Nanduri, J.; Semenza, G.L.; Prabhakar, N.R. Epigenetic changes by DNA methylation in chronic and intermittent hypoxia. Am. J. Physiol. Lung Cell. Mol. Physiol. 2017, 313, 1096–1100. [Google Scholar] [CrossRef] [PubMed]
- Kvietikova, I.; Wenger, R.H.; Marti, H.H.; Gassmann, M. The transcription factors ATF-1 and CREB-1 bind constitutively to the hypoxia-inducible factor-1 (HIF-1) DNA recognition site. Nucleic Acids Res. 1995, 23, 4542–4550. [Google Scholar] [CrossRef] [PubMed]
- Tacchini, L.; Dansi, P.; Matteucci, E.; Desiderio, M.A. Hepatocyte growth factor signalling stimulates hypoxia inducible factor-1 (HIF-1) activity in HepG2 hepatoma cells. Carcinogenesis 2001, 22, 1363–1371. [Google Scholar] [CrossRef] [PubMed]
- Laitem, C.; Leprivier, G.; Choul-Li, S.; Begue, A.; Monte, D.; Larsimont, D.; Dumont, P.; Duterque-Coquillaud, M.; Aumercier, M. Ets-1 p27: A novel Ets-1 isoform with dominant-negative effects on the transcriptional properties and the subcellular localization of Ets-1 p51. Oncogene 2009, 28, 2087–2099. [Google Scholar] [CrossRef] [PubMed]
- Mo, A.; Zhao, Y.; Shi, Y.; Qian, F.; Hao, Y.; Chen, J.; Yang, S.; Jiang, Y.; Luo, Z.; Yu, P. Association between polymorphisms of thymidylate synthase gene 5′- and 3′-UTR and gastric cancer risk: Meta-analysis. Biosci. Rep. 2016, 36, e00429. [Google Scholar] [CrossRef] [PubMed]
- Ridolfi, E.; Matteucci, E.; Maroni, P.; Desiderio, M.A. Inhibitory effect of HGF on invasiveness of aggressive MDA-MB231 breast carcinoma cells, and role of HDACs. Br. J. Cancer 2008, 99, 1623–1634. [Google Scholar] [CrossRef] [PubMed]
- Gegonne, A.; Punyammalee, B.; Rabault, B.; Bosselut, R.; Seneca, S.; Crabeel, M.; Ghysdael, J. Analysis of the DNA binding and transcriptional activation properties of the Ets1 oncoprotein. New Biol. 1992, 4, 512–519. [Google Scholar] [PubMed]
- Bendinelli, P.; Maroni, P.; Matteucci, E.; Luzzati, A.; Perrucchini, G.; Desiderio, M.A. Hypoxia inducible factor-1 is activated by transcriptional co-activator with PDZ-binding motif (TAZ) versus WWdomain-containing oxidoreductase (WWOX) in hypoxic microenvironment of bone metastasis from breast cancer. Eur. J. Cancer 2013, 49, 2608–2618. [Google Scholar] [CrossRef] [PubMed]
- Fry, C.J.; Farnham, P.J. Context-dependent transcriptional regulation. J. Biol. Chem. 1999, 274, 29583–29586. [Google Scholar] [CrossRef] [PubMed]
- Kanamori, Y.; Murakami, M.; Matsui, T.; Funaba, M. The regulation of hepcidin expression by serum treatment: Requirements of the BMP response element and STAT- and AP-1-binding sites. Gene 2014, 551, 119–126. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Zhang, K.; Chen, J.; Chen, L.; Wang, R.; Chu, X. MicroRNAs as regulators and mediators of forkhead box transcription factors function in human cancers. Oncotarget 2017, 8, 12433–12450. [Google Scholar] [CrossRef] [PubMed]
- Dittmer, J. The role of the transcription factor Ets1 in carcinoma. Semin. Cancer Biol. 2015, 35, 20–38. [Google Scholar] [CrossRef] [PubMed]
- Rupaimoole, R.; Wu, S.Y.; Pradeep, S.; Ivan, C.; Pecot, C.V.; Gharpure, K.M.; Nagaraja, A.S.; Armaiz-Pena, G.N.; McGuire, M.; Zand, B.; et al. Hypoxia-mediated downregulation of miRNA biogenesis promotes tumour progression. Nat. Commun. 2014, 5, 5202. [Google Scholar] [CrossRef] [PubMed]
- Peinado, H.; Zhang, H.; Matei, I.R.; Costa-Silva, B.; Hoshino, A.; Rodrigues, G.; Psaila, B.; Kaplan, R.N.; Bromberg, J.F.; Kang, Y.; et al. Pre-metastatic niches: Organ-specific homes for metastases. Nat. Rev. Cancer 2017, 17, 302–317. [Google Scholar] [CrossRef] [PubMed]
- Maroni, P.; Matteucci, E.; Drago, L.; Banfi, G.; Bendinelli, P.; Desiderio, M.A. Hypoxia induced E-cadherin involving regulators of Hippo pathway due to HIF-1α stabilization/nuclear translocation in bone metastasis from breast carcinoma. Exp. Cell Res. 2015, 330, 287–299. [Google Scholar] [CrossRef] [PubMed]
- Matteucci, E.; Maroni, P.; Luzzati, A.; Perrucchini, G.; Bendinelli, P.; Desiderio, M.A. Bone metastatic process of breast cancer involves methylation state affecting E-cadherin expression through TAZ and WWOX nuclear effectors. Eur. J. Cancer 2013, 49, 231–244. [Google Scholar] [CrossRef] [PubMed]
- Biswas, S.; Rao, C.M. Epigenetics in cancer: Fundamentals and beyond. Pharmacol. Ther. 2017, 173, 118–134. [Google Scholar] [CrossRef] [PubMed]
- Dittmer, J. The biology of the Ets1 proto-oncogene. Mol. Cancer 2003, 2, 29. [Google Scholar] [CrossRef] [PubMed]
- Van der Groep, P.; Bouter, A.; Menko, F.H.; van der Wall, E.; van Diest, P.J. High frequency of HIF-1α overexpression in BRCA1 related breast cancer. Breast Cancer Res. Treat. 2008, 111, 475–480. [Google Scholar] [CrossRef] [PubMed]
- Dang, C.V.; Semenza, G.L. Oncogenic alterations of metabolism. Trends Biochem. Sci. 1999, 24, 68–72. [Google Scholar] [CrossRef]
- Gunasinghe, N.P.A.D.; Wells, A.; Thompson, E.W.; Hugo, H.J. Mesenchymal-epithelial transition (MET) as a mechanism for metastatic colonisation in breast cancer. Cancer Metastasis Rev. 2012, 31, 469–478. [Google Scholar] [CrossRef] [PubMed]
- Bill, R.; Christofori, G. The relevance of EMT in breast cancer metastasis: Correlation or causality? FEBS Lett. 2015, 589, 1577–1587. [Google Scholar] [CrossRef] [PubMed]
- Prieto-García, E.; Díaz-García, C.V.; García-Ruiz, I.; Agulló-Ortuño, M.T. Epithelial-to-mesenchymal transition in tumor progression. Med. Oncol. 2017, 34, 122. [Google Scholar] [CrossRef] [PubMed]
- Hoppe, T.; Göser, V.; Kraus, D.; Probstmeier, R.; Frentzen, M.; Wenghofer, M.; Jepsen, S.; Winter, J. Response of MG63 osteoblasts on bacterial challenge is dependent on the state of differentiation. Mol. Oral Microbiol. 2017. [Google Scholar] [CrossRef] [PubMed]
- Yamashita, K.; Discher, D.J.; Hu, J.; Bishopric, N.H.; Webster, K.A. Molecular regulation of the endothelin-1 gene by hypoxia. Contributions of hypoxia-inducible factor-1, activator protein-1, GATA-2, and p300/CBP. J. Biol. Chem. 2001, 276, 12645–12653. [Google Scholar] [CrossRef] [PubMed]


© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Matteucci, E.; Maroni, P.; Nicassio, F.; Ghini, F.; Bendinelli, P.; Desiderio, M.A. RETRACTED: Microenvironment Stimuli HGF and Hypoxia Differently Affected miR-125b and Ets-1 Function with Opposite Effects on the Invasiveness of Bone Metastatic Cells: A Comparison with Breast Carcinoma Cells. Int. J. Mol. Sci. 2018, 19, 258. https://doi.org/10.3390/ijms19010258
Matteucci E, Maroni P, Nicassio F, Ghini F, Bendinelli P, Desiderio MA. RETRACTED: Microenvironment Stimuli HGF and Hypoxia Differently Affected miR-125b and Ets-1 Function with Opposite Effects on the Invasiveness of Bone Metastatic Cells: A Comparison with Breast Carcinoma Cells. International Journal of Molecular Sciences. 2018; 19(1):258. https://doi.org/10.3390/ijms19010258
Chicago/Turabian StyleMatteucci, Emanuela, Paola Maroni, Francesco Nicassio, Francesco Ghini, Paola Bendinelli, and Maria Alfonsina Desiderio. 2018. "RETRACTED: Microenvironment Stimuli HGF and Hypoxia Differently Affected miR-125b and Ets-1 Function with Opposite Effects on the Invasiveness of Bone Metastatic Cells: A Comparison with Breast Carcinoma Cells" International Journal of Molecular Sciences 19, no. 1: 258. https://doi.org/10.3390/ijms19010258
APA StyleMatteucci, E., Maroni, P., Nicassio, F., Ghini, F., Bendinelli, P., & Desiderio, M. A. (2018). RETRACTED: Microenvironment Stimuli HGF and Hypoxia Differently Affected miR-125b and Ets-1 Function with Opposite Effects on the Invasiveness of Bone Metastatic Cells: A Comparison with Breast Carcinoma Cells. International Journal of Molecular Sciences, 19(1), 258. https://doi.org/10.3390/ijms19010258





