Allyl Isothiocyanate Exhibits No Anticancer Activity in MDA-MB-231 Breast Cancer Cells
Abstract
:1. Introduction
2. Results
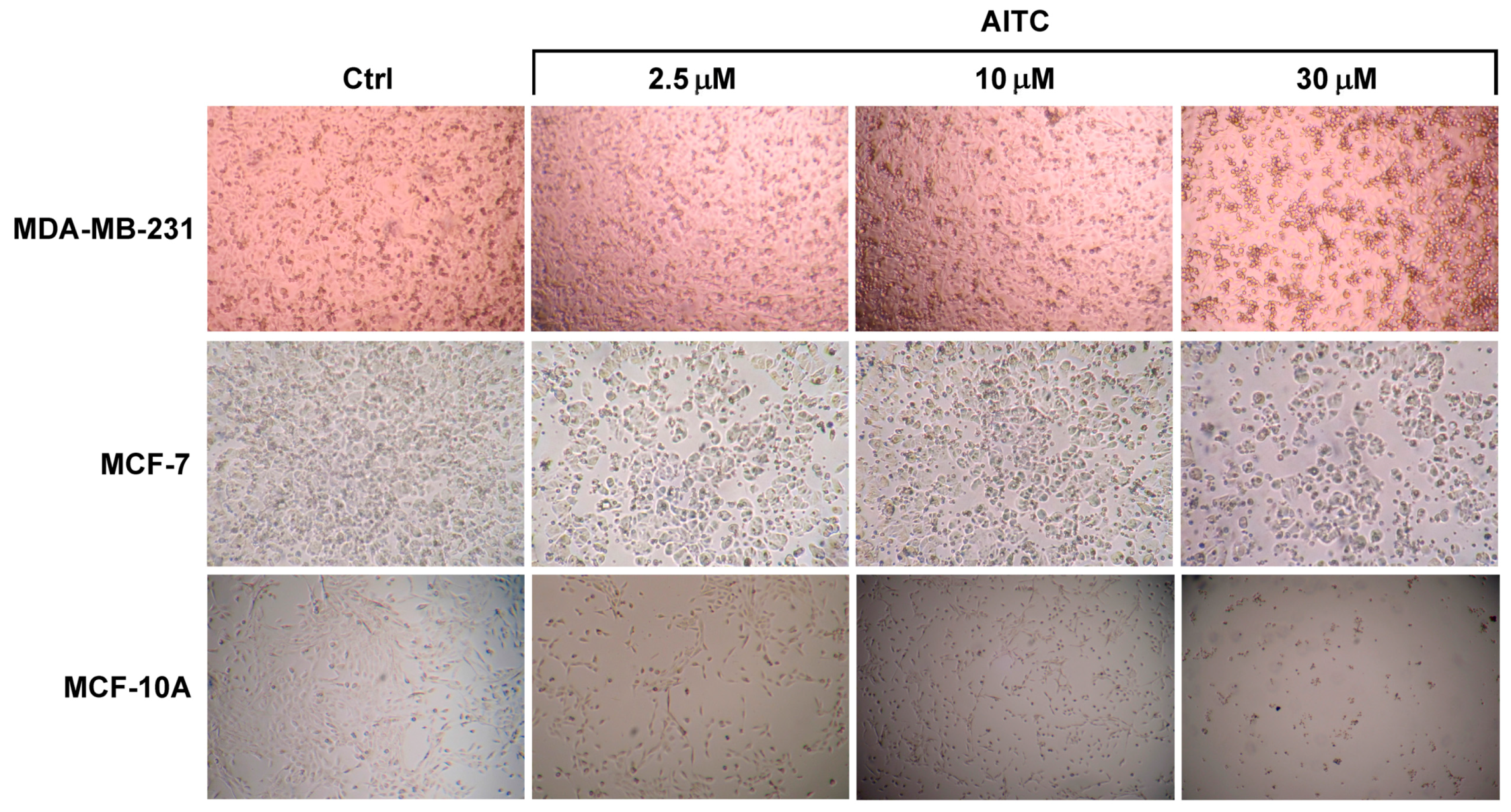
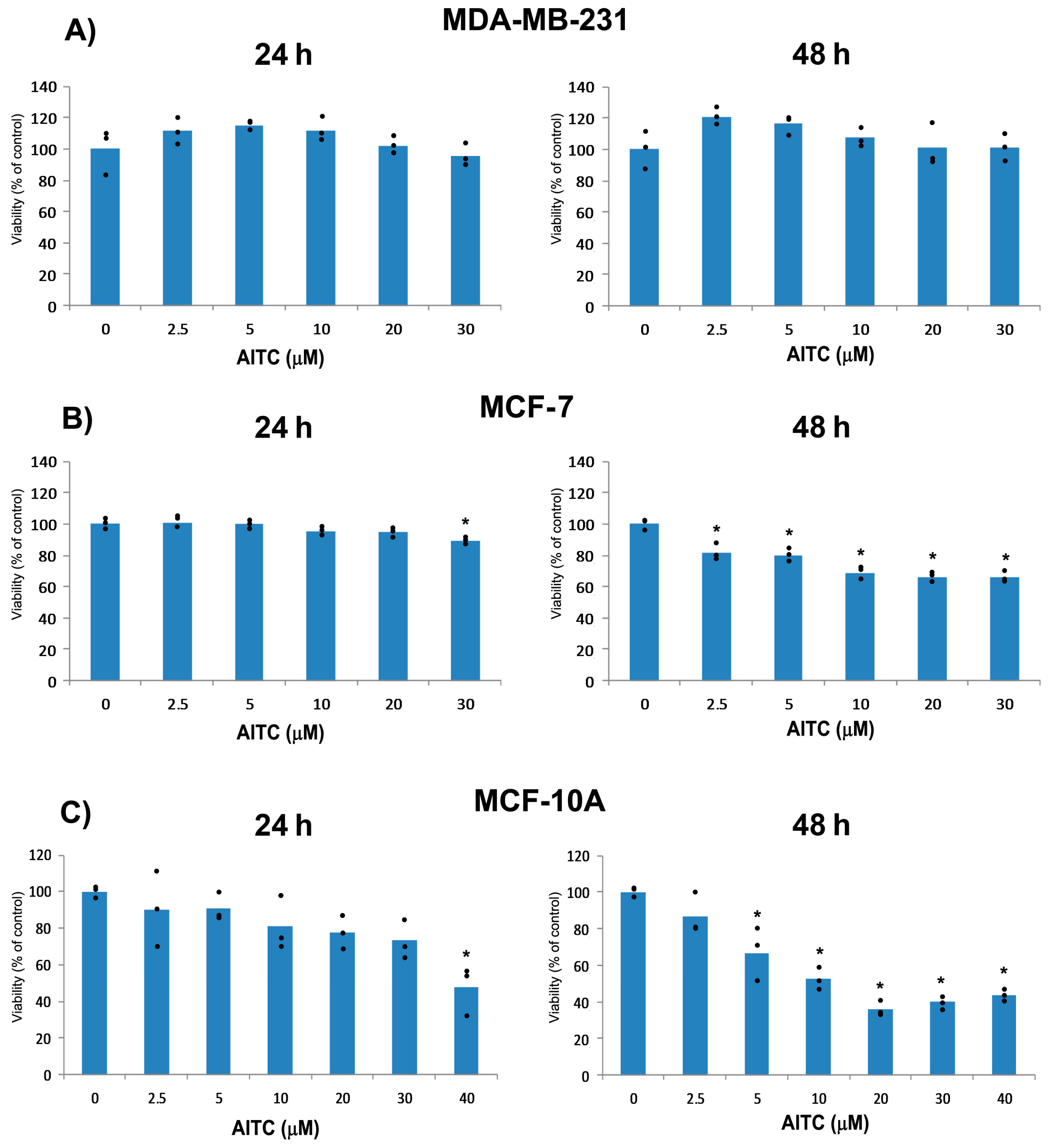
2.1. AITC Did Not Inhibit MDA-MB-231 Cell Proliferation While Affected MCF-7 and MCF-10A Cells
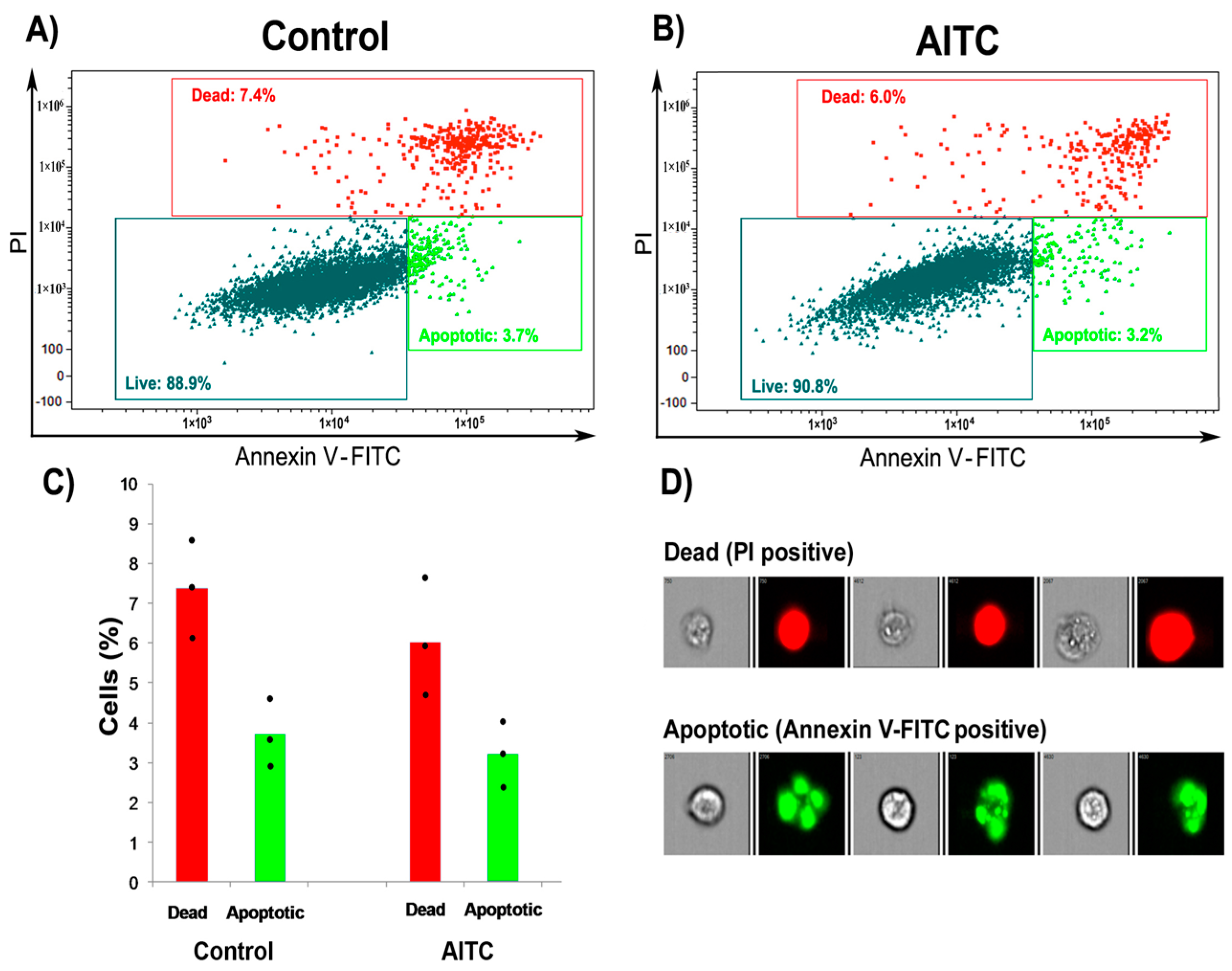
2.2. AITC Did Not Induce Apoptosis and Cell Cycle Arrest
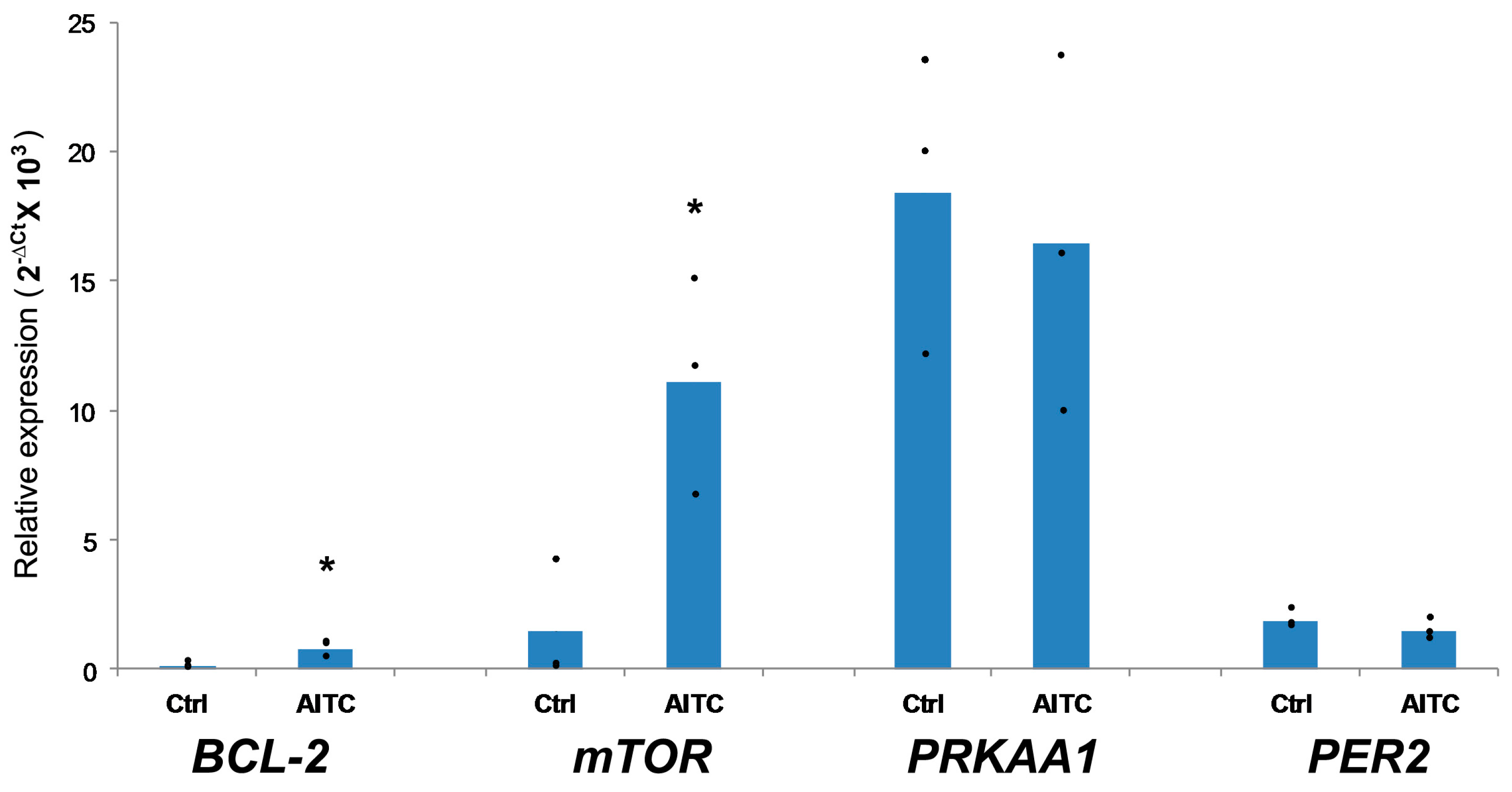
2.3. AITC UpregulatedBCL-2 and mTOR Expression, While Induced No Changes in PRKAA1 and PER2 Expression
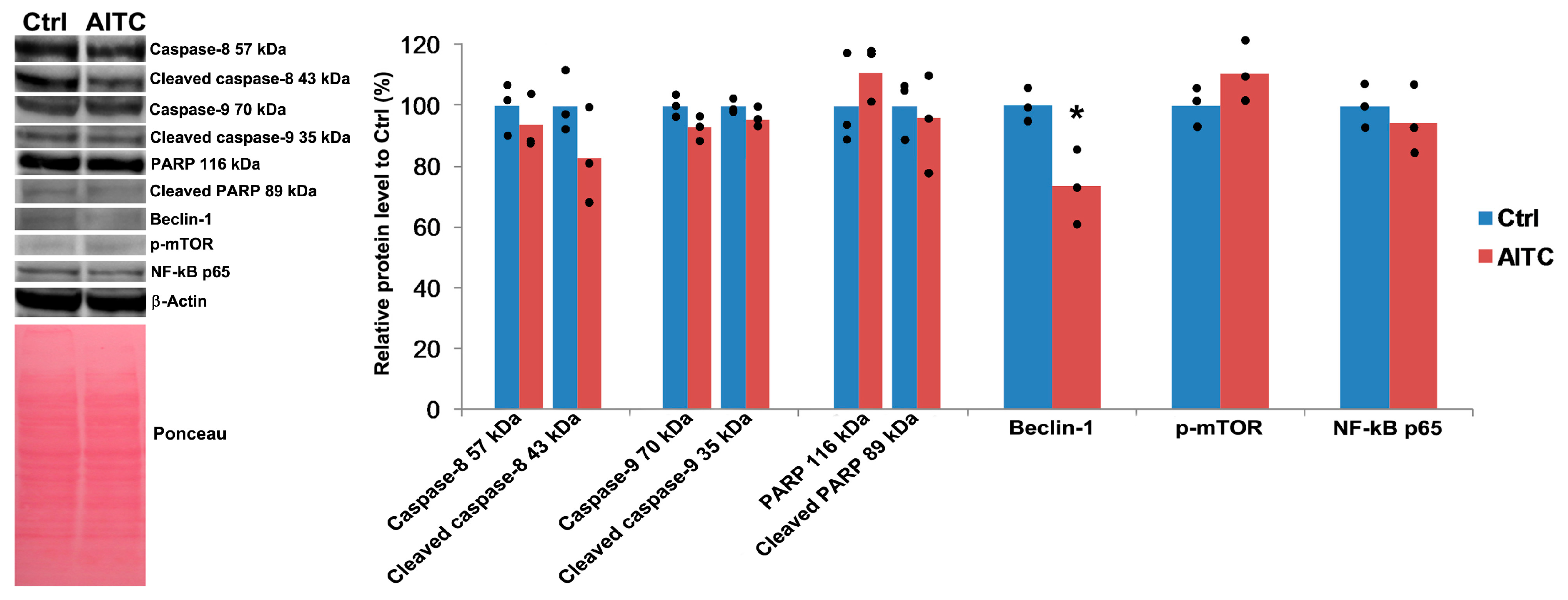
2.4. AITC Did Not Affect Caspase-8, Caspase-9, PARP, p-mTOR, and NF-κB p65 Proteins Level, While Downregulated Beclin-1 Protein Level
3. Discussion
4. Materials and Methods
4.1. Chemicals and Reagents
4.2. Cell Line and Culture Conditions
4.3. Cell Viability Analysis
4.4. Apoptosis Analysis
4.5. Cell Cycle Analysis
4.6. Western Blotting
4.7. RNA Extraction and Real-Time PCR
4.8. Statistical Analysis
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Sayeed, M.A.; Bracci, M.; Lazzarini, R.; Tomasetti, M.; Amati, M.; Lucarini, G.; Di Primio, R.; Santarelli, L. Use of potential dietary phytochemicals to target miRNA: Promising option for breast cancer prevention and treatment? J. Funct. Foods 2017, 28, 177–193. [Google Scholar] [CrossRef]
- Dandawate, P.R.; Subramaniam, D.; Jensen, R.A.; Anant, S. Targeting cancer stem cells and signaling pathways by phytochemicals: Novel approach for breast cancer therapy. Semin. Cancer Biol. 2016, 40–41, 192–208. [Google Scholar] [CrossRef] [PubMed]
- Kushad, M.M.; Brown, A.F.; Kurilich, A.C.; Juvik, J.A.; Klein, B.P.; Wallig, M.A.; Jeffery, E.H. Variation of glucosinolates in vegetable crops of Brassica oleracea. J. Agric. Food Chem. 1999, 47, 1541–1548. [Google Scholar] [CrossRef] [PubMed]
- Hao, G.; Wang, Q.; Liu, B.; Liu, J. Phytochemical profiling of five medicinally active constituents across 14 Eutrema species. Fitoterapia 2016, 110, 83–88. [Google Scholar] [CrossRef] [PubMed]
- Loebers, A.; Muller-Uri, F.; Kreis, W. A young root-specific gene (ArMY2) from horseradish encoding a MYR II myrosinase with kinetic preference for the root-specific glucosinolategluconasturtiin. Phytochemistry 2014, 99, 26–35. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y. Allyl isothiocyanate as a cancer chemopreventive phytochemical. Mol. Nutr. Food Res. 2010, 54, 127–135. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.J.; Lee, D.H.; Ahn, J.; Chung, W.J.; Jang, Y.J.; Seong, K.S.; Moon, J.H.; Ha, T.Y.; Jung, C.H. Pharmacokinetics, Tissue Distribution, and Anti-Lipogenic/Adipogenic Effects of Allyl-Isothiocyanate Metabolites. PLoS ONE 2015, 10, e0132151. [Google Scholar] [CrossRef] [PubMed]
- Encinas-Basurto, D.; Ibarra, J.; Juarez, J.; Burboa, M.G.; Barbosa, S.; Taboada, P.; Troncoso-Rojas, R.; Valdez, M.A. Poly(lactic-co-glycolic acid) nanoparticles for sustained release of allyl isothiocyanate: Characterization, in vitro release and biological activity. J. Microencapsul. 2017, 34, 231–242. [Google Scholar] [CrossRef] [PubMed]
- Lashkari, E.; Wang, H.; Liu, L.; Li, J.; Yam, K. Innovative application of metal-organic frameworks for encapsulation and controlled release of allyl isothiocyanate. Food Chem. 2017, 221, 926–935. [Google Scholar] [CrossRef] [PubMed]
- Ahn, M.; Kim, J.; Bang, H.; Moon, J.; Kim, G.O.; Shin, T. Hepatoprotective effects of allyl isothiocyanate against carbon tetrachloride-induced hepatotoxicity in rat. Chem. Biol. Interact. 2016, 254, 102–108. [Google Scholar] [CrossRef] [PubMed]
- Subedi, L.; Venkatesan, R.; Kim, S.Y. Neuroprotective and Anti-Inflammatory Activities of Allyl Isothiocyanate through Attenuation of JNK/NF-κB/TNF-α Signaling. Int. J. Mol. Sci. 2017, 18, 1423. [Google Scholar] [CrossRef] [PubMed]
- Wagner, A.E.; Boesch-Saadatmandi, C.; Dose, J.; Schultheiss, G.; Rimbach, G. Anti-inflammatory potential of allyl-isothiocyanate—Role of Nrf2, NF-κB and microRNA-155. J. Cell. Mol. Med. 2012, 16, 836–843. [Google Scholar] [CrossRef] [PubMed]
- Matsuda, H.; Ochi, M.; Nagatomo, A.; Yoshikawa, M. Effects of allyl isothiocyanate from horseradish on several experimental gastric lesions in rats. Eur. J. Pharmacol. 2007, 561, 172–181. [Google Scholar] [CrossRef] [PubMed]
- Nowicki, D.; Rodzik, O.; Herman-Antosiewicz, A.; Szalewska-Palasz, A. Isothiocyanates as effective agents against enterohemorrhagic Escherichia coli: Insight to the mode of action. Sci. Rep. 2016, 6, 22263. [Google Scholar] [CrossRef] [PubMed]
- Manyes, L.; Luciano, F.B.; Manes, J.; Meca, G. In vitro antifungal activity of allyl isothiocyanate (AITC) against Aspergillus parasiticus and Penicillium expansum and evaluation of the AITC estimated daily intake. Food Chem. Toxicol. 2015, 83, 293–299. [Google Scholar] [CrossRef] [PubMed]
- Tsai, S.C.; Huang, W.W.; Huang, W.C.; Lu, C.C.; Chiang, J.H.; Peng, S.F.; Chung, J.G.; Lin, Y.H.; Hsu, Y.M.; Amagaya, S.; et al. ERK-modulated intrinsic signaling and G(2)/M phase arrest contribute to the induction of apoptotic death by allyl isothiocyanate in MDA-MB-468 human breast adenocarcinoma cells. Int. J. Oncol. 2012, 41, 2065–2072. [Google Scholar] [CrossRef] [PubMed]
- Bo, P.; Lien, J.C.; Chen, Y.Y.; Yu, F.S.; Lu, H.F.; Yu, C.S.; Chou, Y.C.; Yu, C.C.; Chung, J.G. Allyl Isothiocyanate Induces Cell Toxicity by Multiple Pathways in Human Breast Cancer Cells. Am. J. Chin. Med. 2016, 44, 415–437. [Google Scholar] [CrossRef] [PubMed]
- Savio, A.L.; da Silva, G.N.; Salvadori, D.M. Inhibition of bladder cancer cell proliferation by allyl isothiocyanate (mustard essential oil). Mutat. Res. 2015, 771, 29–35. [Google Scholar] [CrossRef] [PubMed]
- Geng, F.; Tang, L.; Li, Y.; Yang, L.; Choi, K.S.; Kazim, A.L.; Zhang, Y. Allyl isothiocyanate arrests cancer cells in mitosis, and mitotic arrest in turn leads to apoptosis via Bcl-2 protein phosphorylation. J. Biol. Chem. 2011, 286, 32259–32267. [Google Scholar] [CrossRef] [PubMed]
- Chen, N.G.; Chen, K.T.; Lu, C.C.; Lan, Y.H.; Lai, C.H.; Chung, Y.T.; Yang, J.S.; Lin, Y.C. Allyl isothiocyanate triggers G2/M phase arrest and apoptosis in human brain malignant glioma GBM 8401 cells through a mitochondria-dependent pathway. Oncol. Rep. 2010, 24, 449–455. [Google Scholar] [PubMed]
- Lai, K.C.; Lu, C.C.; Tang, Y.J.; Chiang, J.H.; Kuo, D.H.; Chen, F.A.; Chen, I.L.; Yang, J.S. Allyl isothiocyanate inhibits cell metastasis through suppression of the MAPK pathways in epidermal growth factorstimulated HT29 human colorectal adenocarcinoma cells. Oncol. Rep. 2014, 31, 189–196. [Google Scholar] [CrossRef] [PubMed]
- Xiao, D.; Srivastava, S.K.; Lew, K.L.; Zeng, Y.; Hershberger, P.; Johnson, C.S.; Trump, D.L.; Singh, S.V. Allyl isothiocyanate, a constituent of cruciferous vegetables, inhibits proliferation of human prostate cancer cells by causing G2/M arrest and inducing apoptosis. Carcinogenesis 2003, 24, 891–897. [Google Scholar] [CrossRef] [PubMed]
- Srivastava, S.K.; Xiao, D.; Lew, K.L.; Hershberger, P.; Kokkinakis, D.M.; Johnson, C.S.; Trump, D.L.; Singh, S.V. Allyl isothiocyanate, a constituent of cruciferous vegetables, inhibits growth of PC-3 human prostate cancer xenografts in vivo. Carcinogenesis 2003, 24, 1665–1670. [Google Scholar] [CrossRef] [PubMed]
- Xu, K.; Thornalley, P.J. Studies on the mechanism of the inhibition of human leukaemia cell growth by dietary isothiocyanates and their cysteine adducts in vitro. Biochem. Pharmacol. 2000, 60, 221–231. [Google Scholar] [CrossRef]
- Ling, X.; Westover, D.; Cao, F.; Cao, S.; He, X.; Kim, H.R.; Zhang, Y.; Chan, D.C.; Li, F. Synergistic effect of allyl isothiocyanate (AITC) on cisplatin efficacy in vitro and in vivo. Am. J. Cancer Res. 2015, 5, 2516–2530. [Google Scholar] [PubMed]
- Tripathi, K.; Hussein, U.K.; Anupalli, R.; Barnett, R.; Bachaboina, L.; Scalici, J.; Rocconi, R.P.; Owen, L.B.; Piazza, G.A.; Palle, K. Allyl isothiocyanate induces replication-associated DNA damage response in NSCLC cells and sensitizes to ionizing radiation. Oncotarget 2015, 6, 5237–5252. [Google Scholar] [CrossRef] [PubMed]
- Hengartner, M.O. The biochemistry of apoptosis. Nature 2000, 407, 770–776. [Google Scholar] [CrossRef] [PubMed]
- Stennicke, H.R.; Jurgensmeier, J.M.; Shin, H.; Deveraux, Q.; Wolf, B.B.; Yang, X.; Zhou, Q.; Ellerby, H.M.; Ellerby, L.M.; Bredesen, D.; et al. Pro-caspase-3 is a major physiologic target of caspase-8. J. Biol. Chem. 1998, 273, 27084–27090. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Zhu, H.; Xu, C.J.; Yuan, J. Cleavage of BID by caspase 8 mediates the mitochondrial damage in the Fas pathway of apoptosis. Cell 1998, 94, 491–501. [Google Scholar] [CrossRef]
- Oliver, F.J.; de la Rubia, G.; Rolli, V.; Ruiz-Ruiz, M.C.; de Murcia, G.; Murcia, J.M. Importance of poly(ADP-ribose) polymerase and its cleavage in apoptosis. Lesson from an uncleavable mutant. J. Biol. Chem. 1998, 273, 33533–33539. [Google Scholar] [CrossRef] [PubMed]
- Satoh, M.S.; Lindahl, T. Role of poly(ADP-ribose) formation in DNA repair. Nature 1992, 356, 356–358. [Google Scholar] [CrossRef] [PubMed]
- Nicholson, D.W.; Ali, A.; Thornberry, N.A.; Vaillancourt, J.P.; Ding, C.K.; Gallant, M.; Gareau, Y.; Griffin, P.R.; Labelle, M.; Lazebnik, Y.A.; et al. Identification and inhibition of the ICE/CED-3 protease necessary for mammalian apoptosis. Nature 1995, 376, 37–43. [Google Scholar] [CrossRef] [PubMed]
- Zhong, Y.; Wang, Q.J.; Li, X.; Yan, Y.; Backer, J.M.; Chait, B.T.; Heintz, N.; Yue, Z. Distinct regulation of autophagic activity by Atg14L and Rubicon associated with Beclin 1-phosphatidylinositol-3-kinase complex. Nat. Cell Biol. 2009, 11, 468–476. [Google Scholar] [CrossRef] [PubMed]
- Erlich, S.; Mizrachy, L.; Segev, O.; Lindenboim, L.; Zmira, O.; Adi-Harel, S.; Hirsch, J.A.; Stein, R.; Pinkas-Kramarski, R. Differential interactions between Beclin 1 and Bcl-2 family members. Autophagy 2007, 3, 561–568. [Google Scholar] [CrossRef] [PubMed]
- Baeuerle, P.A.; Baltimore, D. NF-κB: Ten years after. Cell 1996, 87, 13–20. [Google Scholar] [CrossRef]
- Gupta, S. Molecular signaling in death receptor and mitochondrial pathways of apoptosis (Review). Int. J. Oncol. 2003, 22, 15–20. [Google Scholar] [CrossRef] [PubMed]
- Laplante, M.; Sabatini, D.M. mTOR signaling in growth control and disease. Cell 2012, 149, 274–293. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Toh, K.L.; Jones, C.R.; Shin, J.Y.; Fu, Y.H.; Ptacek, L.J. Modeling of a human circadian mutation yields insights into clock regulation by PER2. Cell 2007, 128, 59–70. [Google Scholar] [CrossRef] [PubMed]
- Gwinn, D.M.; Shackelford, D.B.; Egan, D.F.; Mihaylova, M.M.; Mery, A.; Vasquez, D.S.; Turk, B.E.; Shaw, R.J. AMPK phosphorylation of raptor mediates a metabolic checkpoint. Mol. Cell 2008, 30, 214–226. [Google Scholar] [CrossRef] [PubMed]
- DeBerardinis, R.J.; Thompson, C.B. Cellular metabolism and disease: What do metabolic outliers teach us? Cell 2012, 148, 1132–1144. [Google Scholar] [CrossRef] [PubMed]
- Xu, H.; Zhou, Y.; Coughlan, K.A.; Ding, Y.; Wang, S.; Wu, Y.; Song, P.; Zou, M.H. AMPKalpha1 deficiency promotes cellular proliferation and DNA damage via p21 reduction in mouse embryonic fibroblasts. Biochim. Biophys. Acta 2015, 1853, 65–73. [Google Scholar] [CrossRef] [PubMed]
- Obba, S.; Hizir, Z.; Boyer, L.; Selimoglu-Buet, D.; Pfeifer, A.; Michel, G.; Hamouda, M.A.; Goncalves, D.; Cerezo, M.; Marchetti, S.; et al. The PRKAA1/AMPKalpha1 pathway triggers autophagy during CSF1-induced human monocyte differentiation and is a potential target in CMML. Autophagy 2015, 11, 1114–1129. [Google Scholar] [CrossRef] [PubMed]
- Singh, B.N.; Singh, H.B.; Singh, A.; Naqvi, A.H.; Singh, B.R. Dietary phytochemicals alter epigenetic events and signaling pathways for inhibition of metastasis cascade: Phytoblockers of metastasis cascade. Cancer Metastasis Rev. 2014, 33, 41–85. [Google Scholar] [CrossRef] [PubMed]
- Sayeed, M.A.; Bracci, M.; Lucarini, G.; Lazzarini, R.; Di Primio, R.; Santarelli, L. Regulation of microRNA using promising dietary phytochemicals: Possible preventive and treatment option of malignant mesothelioma. Biomed. Pharmacother. 2017, 94, 1197–1224. [Google Scholar] [CrossRef] [PubMed]
- Gupta, S.C.; Kim, J.H.; Prasad, S.; Aggarwal, B.B. Regulation of survival, proliferation, invasion, angiogenesis, and metastasis of tumor cells through modulation of inflammatory pathways by nutraceuticals. Cancer Metastasis Rev. 2010, 29, 405–434. [Google Scholar] [CrossRef] [PubMed]
- Hasanpourghadi, M.; Pandurangan, A.K.; Mustafa, M.R. Modulation of oncogenic transcription factors by bioactive natural products in breast cancer. Pharmacol. Res. 2017. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Li, S.; Meng, X.; Gan, R.Y.; Zhang, J.J.; Li, H.B. Dietary Natural Products for Prevention and Treatment of Breast Cancer. Nutrients 2017, 9. [Google Scholar] [CrossRef] [PubMed]
- Pandey, M.K.; Gupta, S.C.; Nabavizadeh, A.; Aggarwal, B.B. Regulation of cell signaling pathways by dietary agents for cancer prevention and treatment. Semin. Cancer Biol. 2017, 46, 158–181. [Google Scholar] [CrossRef] [PubMed]
- Hirose, M.; Yamaguchi, T.; Kimoto, N.; Ogawa, K.; Futakuchi, M.; Sano, M.; Shirai, T. Strong promoting activity of phenylethyl isothiocyanate and benzyl isothiocyanate on urinary bladder carcinogenesis in F344 male rats. Int. J. Cancer 1998, 77, 773–777. [Google Scholar] [CrossRef]
- Green, D.R. Apoptotic pathways: Paper wraps stone blunts scissors. Cell 2000, 102, 1–4. [Google Scholar] [CrossRef]
- Ashkenazi, A.; Dixit, V.M. Death receptors: Signaling and modulation. Science 1998, 281, 1305–1308. [Google Scholar] [CrossRef] [PubMed]
- Lau, W.S.; Chen, T.; Wong, Y.S. Allyl isothiocyanate induces G2/M arrest in human colorectal adenocarcinoma SW620 cells through down-regulation of Cdc25B and Cdc25C. Mol. Med. Rep. 2010, 3, 1023–1030. [Google Scholar] [PubMed]
- Savio, A.L.; da Silva, G.N.; de Camargo, E.A.; Salvadori, D.M. Cell cycle kinetics, apoptosis rates, DNA damage and TP53 gene expression in bladder cancer cells treated with allyl isothiocyanate (mustard essential oil). Mutat. Res. 2014, 762, 40–46. [Google Scholar] [CrossRef] [PubMed]
- Jung, Y.Y.; Lee, Y.K.; Koo, J.S. The potential of Beclin 1 as a therapeutic target for the treatment of breast cancer. Expert Opin. Ther. Targets 2016, 20, 167–178. [Google Scholar] [CrossRef] [PubMed]
- Liang, X.H.; Jackson, S.; Seaman, M.; Brown, K.; Kempkes, B.; Hibshoosh, H.; Levine, B. Induction of autophagy and inhibition of tumorigenesis by beclin 1. Nature 1999, 402, 672–676. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Li, H.Q.; Zhou, F.X.; Yu, J.W.; Sun, L.; Han, Z.H. Targeting the mTOR pathway in breast cancer. Tumor Biol. 2017, 39. [Google Scholar] [CrossRef] [PubMed]
- Davis, N.M.; Sokolosky, M.; Stadelman, K.; Abrams, S.L.; Libra, M.; Candido, S.; Nicoletti, F.; Polesel, J.; Maestro, R.; D’Assoro, A.; et al. Deregulation of the EGFR/PI3K/PTEN/Akt/mTORC1 pathway in breast cancer: Possibilities for therapeutic intervention. Oncotarget 2014, 5, 4603–4650. [Google Scholar] [CrossRef] [PubMed]
- Bommareddy, A.; Hahm, E.R.; Xiao, D.; Powolny, A.A.; Fisher, A.L.; Jiang, Y.; Singh, S.V. Atg5 regulates phenethyl isothiocyanate-induced autophagic and apoptotic cell death in human prostate cancer cells. Cancer Res. 2009, 69, 3704–3712. [Google Scholar] [CrossRef] [PubMed]
- Boreddy, S.R.; Pramanik, K.C.; Srivastava, S.K. Pancreatic tumor suppression by benzyl isothiocyanate is associated with inhibition of PI3K/AKT/FOXO pathway. Clin. Cancer Res. 2011, 17, 1784–1795. [Google Scholar] [CrossRef] [PubMed]
- Shao, W.Y.; Yang, Y.L.; Yan, H.; Huang, Q.; Liu, K.J.; Zhang, S. Phenethyl isothiocyanate suppresses the metastasis of ovarian cancer associated with the inhibition of CRM1-mediated nuclear export and mTOR-STAT3 pathway. Cancer Biol. Ther. 2017, 18, 26–35. [Google Scholar] [CrossRef] [PubMed]
- Wiczk, A.; Hofman, D.; Konopa, G.; Herman-Antosiewicz, A. Sulforaphane, a cruciferous vegetable-derived isothiocyanate, inhibits protein synthesis in human prostate cancer cells. Biochim. Biophys. Acta 2012, 1823, 1295–1305. [Google Scholar] [CrossRef] [PubMed]
- Pan, Y.; Shi, J.; Ni, W.; Liu, Y.; Wang, S.; Wang, X.; Wei, Z.; Wang, A.; Chen, W.; Lu, Y. Cryptotanshinone inhibition of mammalian target of rapamycin pathway is dependent on oestrogen receptor α in breast cancer. J. Cell. Mol. Med. 2017, 21, 2129–2139. [Google Scholar] [CrossRef] [PubMed]
- Suman, S.; Sharma, P.K.; Rai, G.; Mishra, S.; Arora, D.; Gupta, P.; Shukla, Y. Current perspectives of molecular pathways involved in chronic inflammation-mediated breast cancer. Biochem. Biophys. Res. Commun. 2016, 472, 401–409. [Google Scholar] [CrossRef] [PubMed]
- Rajakumar, T.; Pugalendhi, P.; Jayaganesh, R.; Ananthakrishnan, D.; Gunasekaran, K. Effect of allyl isothiocyanate on NF-κB signaling in 7,12-dimethylbenz(a)anthracene and N-methyl-N-nitrosourea-induced mammary carcinogenesis. Breast Cancer 2018, 25, 50–59. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.K.; Park, S.Y.; Kim, Y.M.; Kim, D.C.; Lee, W.S.; Surh, Y.J.; Park, O.J. Suppression of mTOR via Akt-dependent and -independent mechanisms in selenium-treated colon cancer cells: Involvement of AMPKalpha1. Carcinogenesis 2010, 31, 1092–1099. [Google Scholar] [CrossRef] [PubMed]
- Yoon, J.H.; Ahn, S.G.; Lee, B.H.; Jung, S.H.; Oh, S.H. Role of autophagy in chemoresistance: Regulation of the ATM-mediated DNA-damage signaling pathway through activation of DNA-PKcs and PARP-1. Biochem. Pharmacol. 2012, 83, 747–757. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.T.; Choo, K.B.; Hou, M.F.; Yeh, K.T.; Kuo, S.J.; Chang, J.G. Deregulated expression of the PER1, PER2 and PER3 genes in breast cancers. Carcinogenesis 2005, 26, 1241–1246. [Google Scholar] [CrossRef] [PubMed]
- Winter, S.L.; Bosnoyan-Collins, L.; Pinnaduwage, D.; Andrulis, I.L. Expression of the circadian clock genes Per1 and Per2 in sporadic and familial breast tumors. Neoplasia 2007, 9, 797–800. [Google Scholar] [CrossRef] [PubMed]
- Fu, L.; Lee, C.C. The circadian clock: Pacemaker and tumour suppressor. Nat. Rev. Cancer 2003, 3, 350–361. [Google Scholar] [CrossRef] [PubMed]
- Fu, L.; Pelicano, H.; Liu, J.; Huang, P.; Lee, C. The circadian gene Period2 plays an important role in tumor suppression and DNA damage response in vivo. Cell 2002, 111, 41–50. [Google Scholar] [CrossRef]
- Yang, X.; Wood, P.A.; Oh, E.Y.; Du-Quiton, J.; Ansell, C.M.; Hrushesky, W.J. Down regulation of circadian clock gene Period 2 accelerates breast cancer growth by altering its daily growth rhythm. Breast Cancer Res. Treat. 2009, 117, 423–431. [Google Scholar] [CrossRef] [PubMed]
- Wood, P.A.; Yang, X.; Taber, A.; Oh, E.Y.; Ansell, C.; Ayers, S.E.; Al-Assaad, Z.; Carnevale, K.; Berger, F.G.; Pena, M.M.; et al. Period 2 mutation accelerates ApcMin/+ tumorigenesis. Mol. Cancer Res. 2008, 6, 1786–1793. [Google Scholar] [CrossRef] [PubMed]
- Staffolani, S.; Manzella, N.; Strafella, E.; Nocchi, L.; Bracci, M.; Ciarapica, V.; Amati, M.; Rubini, C.; Re, M.; Pugnaloni, A.; et al. Wood dust exposure induces cell transformation through EGFR-mediated OGG1 inhibition. Mutagenesis 2015, 30, 487–497. [Google Scholar] [CrossRef] [PubMed]
- Darzynkiewicz, Z.; Juan, G.; Bedner, E. Determining cell cycle stages by flow cytometry. In Current Protocols in Cell Biology; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2001; Chapter 8. [Google Scholar]
- Manzella, N.; Bracci, M.; Strafella, E.; Staffolani, S.; Ciarapica, V.; Copertaro, A.; Rapisarda, V.; Ledda, C.; Amati, M.; Valentino, M.; et al. Circadian Modulation of 8-Oxoguanine DNA Damage Repair. Sci. Rep. 2015, 4, 13752. [Google Scholar] [CrossRef] [PubMed]
- Tomasetti, M.; Monaco, F.; Manzella, N.; Rohlena, J.; Rohlenova, K.; Staffolani, S.; Gaetani, S.; Ciarapica, V.; Amati, M.; Bracci, M.; et al. MicroRNA-126 induces autophagy by altering cell metabolism in malignant mesothelioma. Oncotarget 2016, 7, 36338–36352. [Google Scholar] [CrossRef] [PubMed]
- Islam, M.S.; Giampieri, F.; Janjusevic, M.; Gasparrini, M.; Forbes-Hernandez, T.Y.; Mazzoni, L.; Greco, S.; Giannubilo, S.R.; Ciavattini, A.; Mezzetti, B.; et al. An anthocyanin rich strawberry extract induces apoptosis and ROS while decreases glycolysis and fibrosis in human uterine leiomyoma cells. Oncotarget 2017, 8, 23575–23587. [Google Scholar] [PubMed]

© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sayeed, M.A.; Bracci, M.; Ciarapica, V.; Malavolta, M.; Provinciali, M.; Pieragostini, E.; Gaetani, S.; Monaco, F.; Lucarini, G.; Rapisarda, V.; et al. Allyl Isothiocyanate Exhibits No Anticancer Activity in MDA-MB-231 Breast Cancer Cells. Int. J. Mol. Sci. 2018, 19, 145. https://doi.org/10.3390/ijms19010145
Sayeed MA, Bracci M, Ciarapica V, Malavolta M, Provinciali M, Pieragostini E, Gaetani S, Monaco F, Lucarini G, Rapisarda V, et al. Allyl Isothiocyanate Exhibits No Anticancer Activity in MDA-MB-231 Breast Cancer Cells. International Journal of Molecular Sciences. 2018; 19(1):145. https://doi.org/10.3390/ijms19010145
Chicago/Turabian StyleSayeed, Md. Abu, Massimo Bracci, Veronica Ciarapica, Marco Malavolta, Mauro Provinciali, Ernesta Pieragostini, Simona Gaetani, Federica Monaco, Guendalina Lucarini, Venerando Rapisarda, and et al. 2018. "Allyl Isothiocyanate Exhibits No Anticancer Activity in MDA-MB-231 Breast Cancer Cells" International Journal of Molecular Sciences 19, no. 1: 145. https://doi.org/10.3390/ijms19010145
APA StyleSayeed, M. A., Bracci, M., Ciarapica, V., Malavolta, M., Provinciali, M., Pieragostini, E., Gaetani, S., Monaco, F., Lucarini, G., Rapisarda, V., Di Primio, R., & Santarelli, L. (2018). Allyl Isothiocyanate Exhibits No Anticancer Activity in MDA-MB-231 Breast Cancer Cells. International Journal of Molecular Sciences, 19(1), 145. https://doi.org/10.3390/ijms19010145