Physiological and Transcriptomic Responses of Chinese Cabbage (Brassica rapa L. ssp. Pekinensis) to Salt Stress
Abstract
1. Introduction
2. Results and Discussion
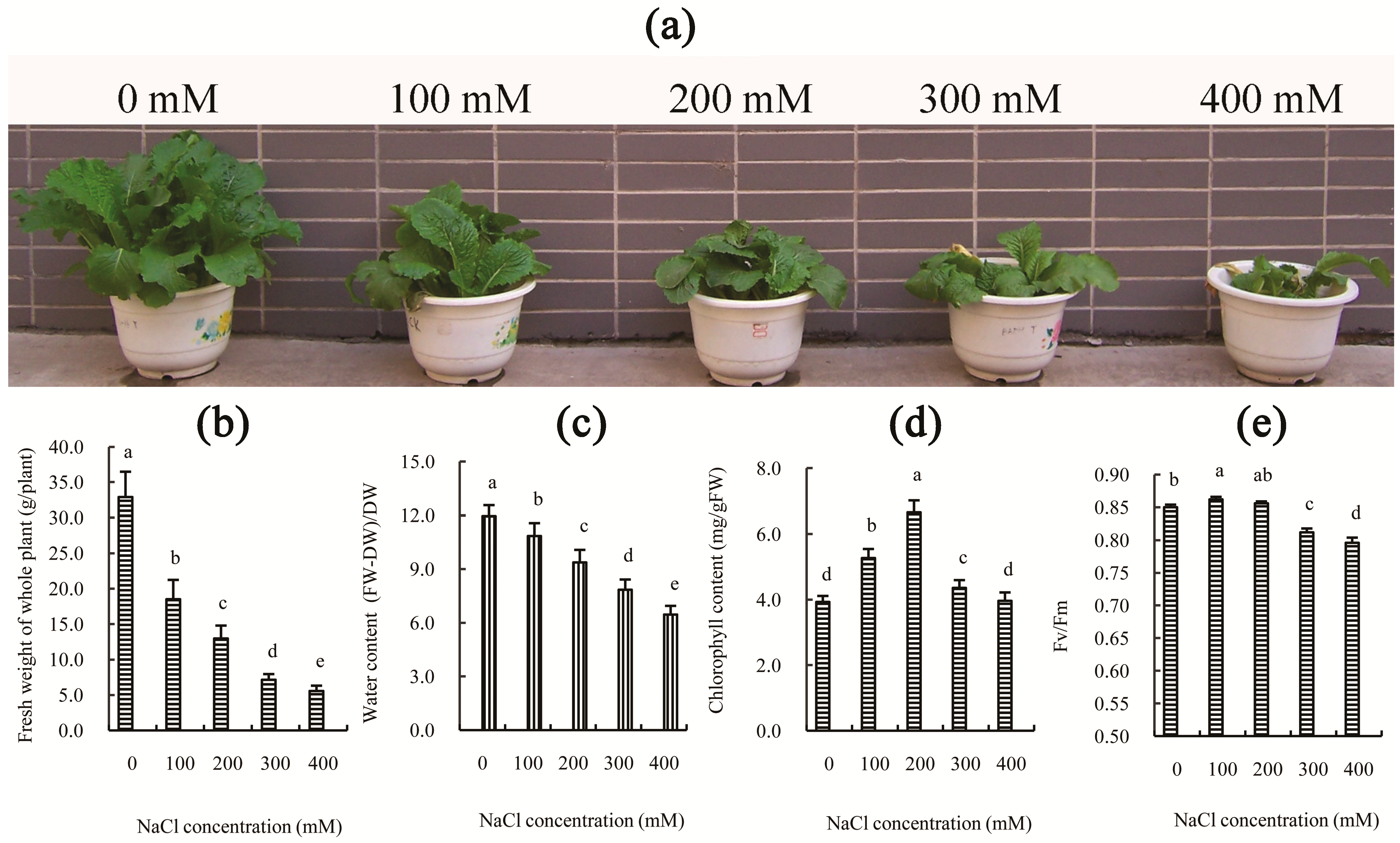
2.1. Physiological Influence of Salt Treatment on Chinese Cabbage
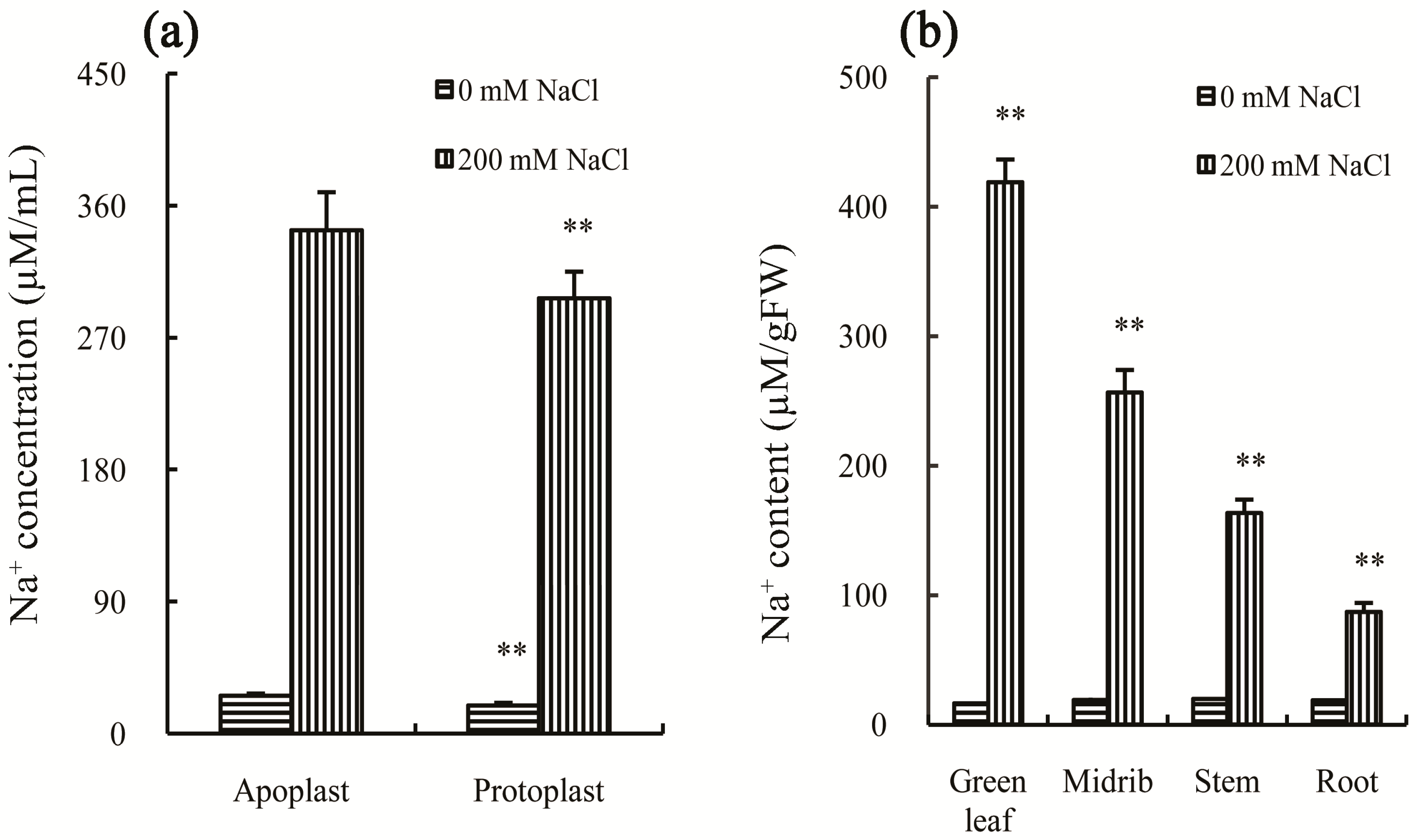
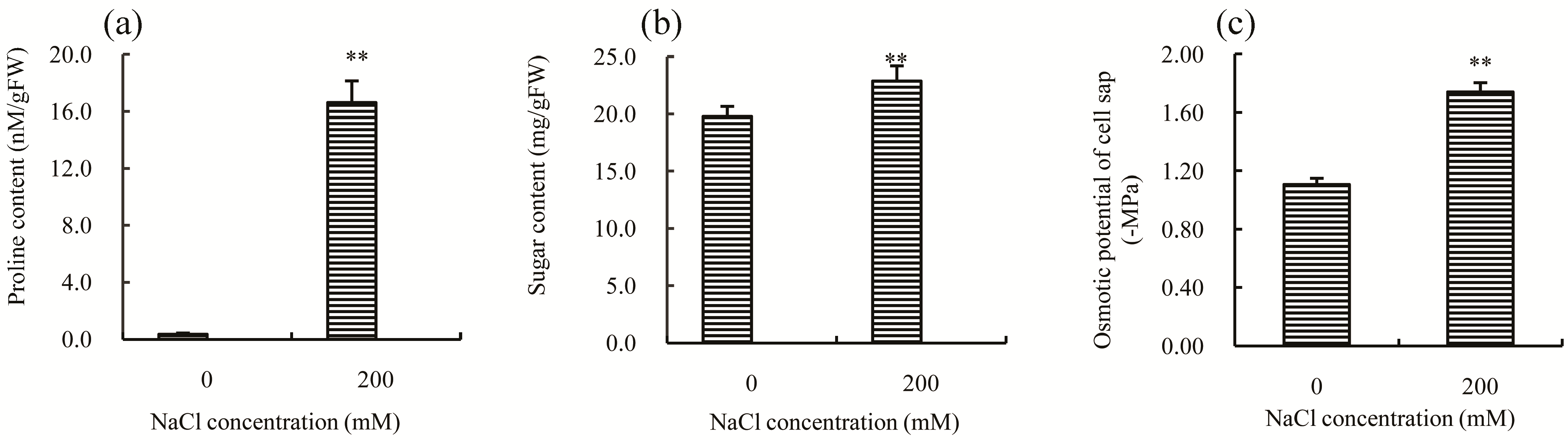
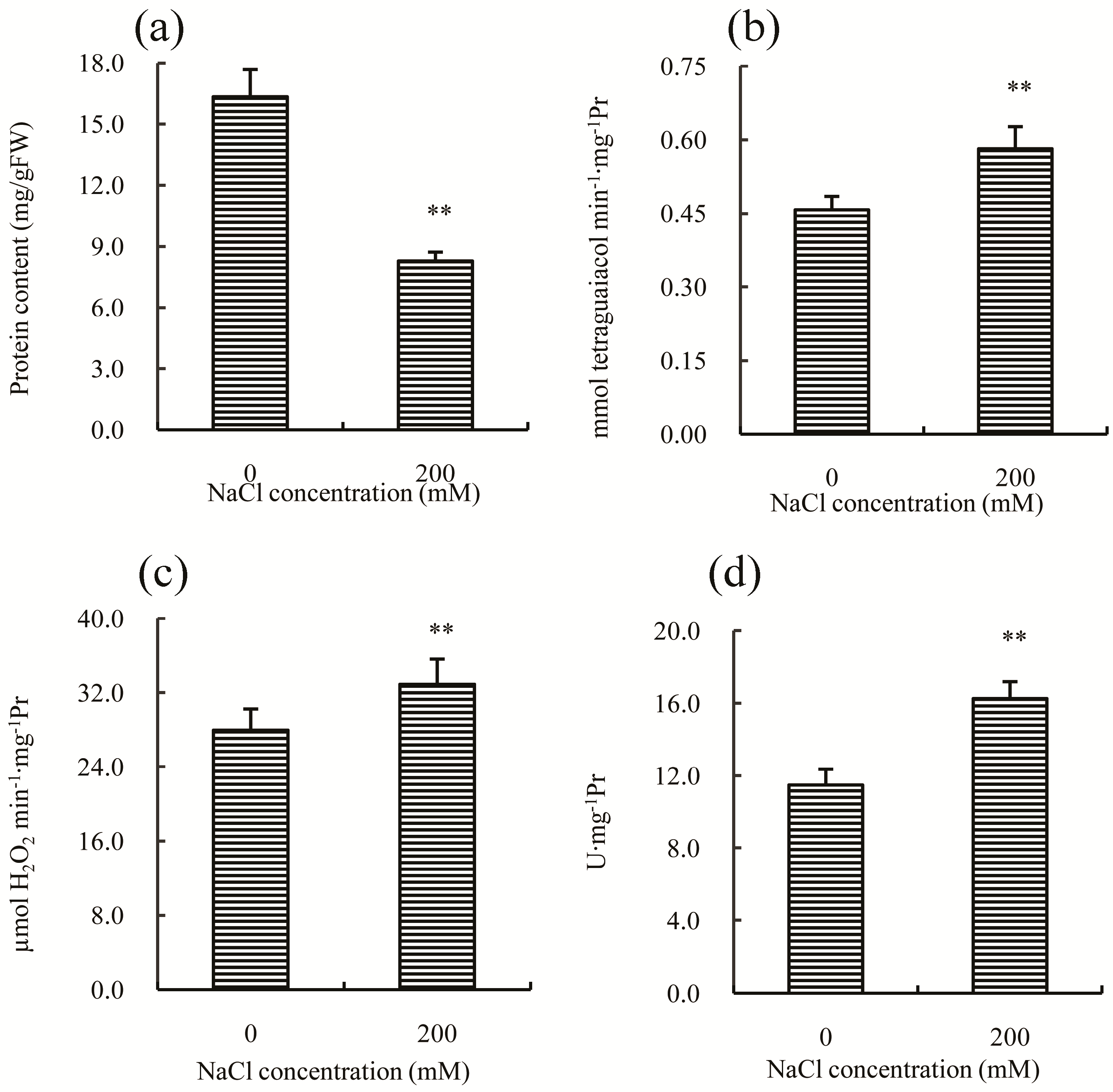
2.2. Physiological Responses of Chinese Cabbage to Salt Stress
2.3. Transcriptomic Responses of Chinese Cabbage to Salt Stress
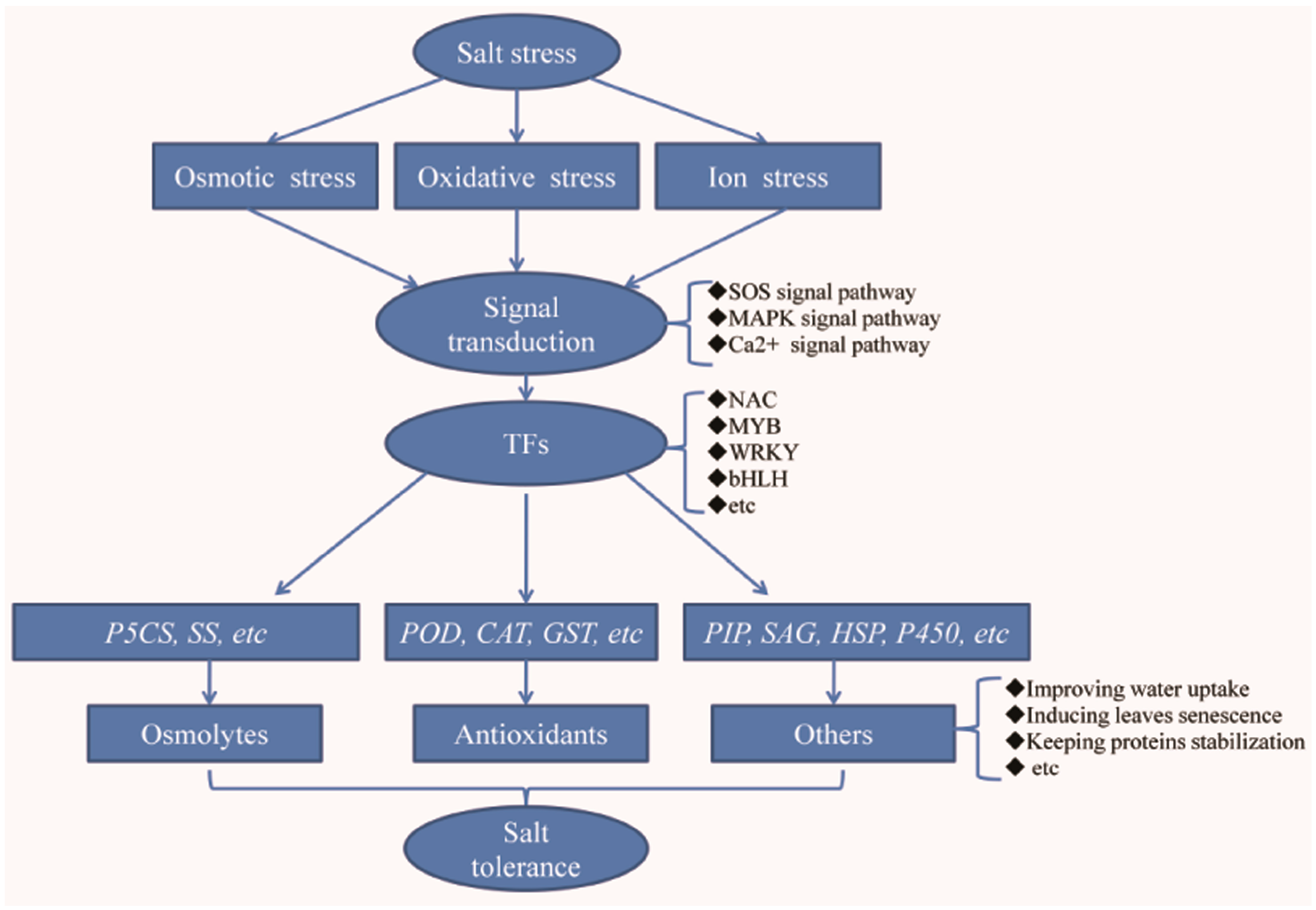
2.4. Up-Regulated Genes Involved in Signal Transduction
2.5. Up-Regulated Genes Encoding Transcription Factors
2.6. Up-Regulated Genes Encoding Proteins Related to Osmolyte Synthesis
2.7. Up-Regulated Genes Encoding Antioxidant Proteins
2.8. Other Genes Induced by Salt Treatment
2.9. Down-Regulated Genes Involved in Salt Stress Response
3. Materials and Methods
3.1. Plant Materials and Salt Treatment
3.2. Determination of Fresh Weight, Water Content, and Chlorophyll Content
3.3. Determination of Na+ and K+ Concentrations
3.4. Determination of the Maximal Efficiency of PSII Photochemistry (Fv/Fm) and the Photosynthetic Gas Exchange Indexes
3.5. Determination of Proline and Soluble Sugars Content and Osmotic Potential (Ψs)
3.6. Determination of Soluble Protein Content and Antioxidant Enzyme Assays
3.7. Identification of Differentially Expressed Genes Using DGE
3.8. Real-Time Quantitative PCR Validation of the DEGs
3.9. Statistical and Geostatistical Analyses
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Wang, W.X.; Vinocur, B.; Altman, A. Plant responses to drought, salinity and extreme temperatures: Towards genetic engineering for stress tolerance. Planta 2003, 218, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.K. Plant salt tolerance. Trends Plant Sci. 2001, 6, 66–71. [Google Scholar] [CrossRef]
- Zhang, J.L.; Shi, H. Physiological and molecular mechanisms of plant salt tolerance. Photosynth. Res. 2013, 115, 1–22. [Google Scholar] [CrossRef] [PubMed]
- Cabot, C.; Sibole, J.V.; Barceló, J.; Poschenrieder, C. Lessons from crop plants struggling with salinity. Plant Sci. 2014, 226, 2–13. [Google Scholar] [CrossRef] [PubMed]
- Deinlein, U.; Stephan, A.B.; Horie, T.; Luo, W.; Xu, G.; Schroeder, J.I. Plant salt-tolerance mechanisms. Trends Plant Sci. 2014, 19, 371–379. [Google Scholar] [CrossRef] [PubMed]
- Munns, R. Genes and salt tolerance: Bringing them together. New Phytol. 2005, 167, 645–663. [Google Scholar] [CrossRef] [PubMed]
- Jamil, A.; Riaz, S.; Ashraf, M.; Foolad, M.R. Gene expression profiling of plants under salt stress. Crit. Rev. Plant Sci. 2011, 30, 435–458. [Google Scholar] [CrossRef]
- Qiu, N.W.; Liu, Q.; Wang, F.D.; Zhao, N.; Sun, K.Y.; Miao, X.M.; Zhao, L.J.; Li, L.L.; Gao, J.W. Long-term observation on salt tolerance of Chinese cabbage at different development stages. Plant Physiol. J. 2015, 51, 1597–1603, (In Chinese with English Abstract). [Google Scholar]
- Kumar, M.; Choi, J.Y.; Kumari, N.; Pareek, A.; Kim, S.R. Molecular breeding in Brassica for salt tolerance: Importance of microsatellite (SSR) markers for molecular breeding in Brassica. Front. Plant Sci. 2015, 6, 688. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.L.; Li, H.R.; Guo, S.Y.; Wang, S.M.; Shi, H.Z.; Han, Q.Q.; Bao, A.K.; Ma, Q. Research advances in higher plant adaptation to salt stress. Acta Pratacult. Sin. 2015, 24, 220–236, (In Chinese with English Abstract). [Google Scholar]
- Türkan, I.; Demiral, T. Recent developments in understanding salinity tolerance. Environ. Exp. Bot. 2009, 67, 2–9. [Google Scholar] [CrossRef]
- Gulen, H.; Turhan, E.; Eris, A. Changes in peroxidase activities and soluble proteins in strawberry varieties under salt-stress. Acta Physiol. Plant 2006, 28, 109–116. [Google Scholar] [CrossRef]
- Doganlar, Z.B.; Demi̇R, K.; Basak, H.; Gul, I. Effects of salt stress on pigment and total soluble protein contents of three different tomato cultivars. Afr. J. Agric. Res. 2010, 5, 2056–2065. [Google Scholar]
- Fariduddin, Q.; Mir, B.A.; Ahmad, A. Physiological and biochemical traits as tools to screen sensitive and resistant varieties of tomatoes exposed to salt stress. Braz. J. Plant Physiol. 2012, 24, 281–292. [Google Scholar] [CrossRef]
- Morant-Avice, A.; Pradier, E.; Houchi, R. Osmotic adjustment in triticales grown in presence of NaCl. Biol. Plant. 1998, 41, 227–234. [Google Scholar] [CrossRef]
- Lee, D.H.; Kim, Y.S.; Lee, C.B. The inductive responses of the antioxidant enzymes by salt stress in the rice (Oryza sativa L.). J. Plant Physiol. 2001, 158, 737–745. [Google Scholar] [CrossRef]
- Rasool, S.; Ahmad, A.; Siddiqi, T.O.; Ahmad, P. Changes in growth, lipid peroxidation and some key antioxidant enzymes in chickpea genotypes under salt stress. Acta Physiol. Plant. 2013, 35, 1039–1050. [Google Scholar] [CrossRef]
- Wang, W.B.; Kim, Y.H.; Lee, H.S.; Kim, K.Y.; Deng, X.P.; Kwak, S.S. Analysis of antioxidant enzyme activity during germination of alfalfa under salt and drought stresses. Plant Physiol. Biochem. 2009, 47, 570–577. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.L.; Flowers, T.J.; Wang, S.M. Differentiation of low-affinity Na+ uptake pathways and kinetics of the effects of K+ on Na+ uptake in the halophyte Suaeda maritime. Plant Soil 2013, 368, 629–640. [Google Scholar] [CrossRef]
- Zhang, J.L.; Flowers, T.J.; Wang, S.M. Mechanisms of sodium uptake by roots of higher plants. Plant Soil 2010, 326, 45–60. [Google Scholar] [CrossRef]
- Hörtensteiner, S.; Feller, U. Nitrogen metabolism and remobilization during senescence. J. Exp. Bot. 2002, 53, 927–937. [Google Scholar] [CrossRef] [PubMed]
- Qiu, Q.S.; Barkla, B.J.; Vera-Estrella, R.; Zhu, J.K.; Schumaker, K.S. Na+/H+ exchange activity in the plasma membrane of Arabidopsis. Plant Physiol. 2003, 132, 1041–1052. [Google Scholar] [CrossRef] [PubMed]
- Gao, D.; Knight, M.R.; Trewavas, A.J.; Sattelmacher, B.; Plieth, C. Self-reporting Arabidopsis expressing pH and Ca2+ indicators unveil ion dynamics in the cytoplasm and in the apoplast under abiotic stress. Plant Physiol. 2004, 134, 898–908. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Z.; Zhu, S.; Ye, R.; Xue, Y.; Chen, A.; An, L.; Pei, Z.M. Relationship between NaCl− and H2O2− induced cytosolic Ca2+ increases in response to stress in Arabidopsis. PLoS ONE 2013, 8, e76130. [Google Scholar] [CrossRef] [PubMed]
- Gong, D.; Guo, Y.; Schumaker, K.S.; Zhu, J.K. The SOS3 family of calcium sensors and SOS2 family of protein kinases in Arabidopsis. Plant Physiol. 2004, 134, 919–926. [Google Scholar] [CrossRef] [PubMed]
- Kolukisaoglu, U.; Weinl, S.; Blazevic, D.; Batistic, O.; Kudla, J. Calcium sensors and their interacting protein kinases: Genomics of the Arabidopsis and rice CBL-CIPK signaling networks. Plant Physiol. 2004, 134, 43–58. [Google Scholar] [CrossRef] [PubMed]
- Hashimoto, K.; Kudla, J. Calcium decoding mechanisms in plants. Biochimie 2011, 93, 2054–2059. [Google Scholar] [CrossRef] [PubMed]
- Xu, G.Y.; Rocha, P.S.; Wang, M.L.; Xu, M.L.; Cui, Y.C.; Li, L.Y.; Zhu, Y.X.; Xia, X. A novel rice calmodulin-like gene, OsMSR2, enhances drought and salt tolerance and increases ABA sensitivity in Arabidopsis. Planta 2011, 234, 47–59. [Google Scholar] [CrossRef] [PubMed]
- Magnan, F.; Ranty, B.M.; Sotta, B.; Galaud, J.P.; Aldon, D. Mutations in AtCML9, a calmodulin-like protein from Arabidopsis thaliana, alter plant responses to abiotic stress and abscisic acid. Plant J. 2008, 56, 575–589. [Google Scholar] [CrossRef] [PubMed]
- Miransari, M.; Rangbar, B.; Khajeh, K.; Tehranchi, M.M.; Rusta Azad, R.; Nagafi, F.; Rahnemaie, R. Salt Stress and MAPK Signaling in Plants in Salt Stress in Plants; Springer: New York, NY, USA, 2013; Volume 1389, pp. 157–173. [Google Scholar]
- Rodriguez, M.C.; Petersen, M.; Mundy, J. Mitogen-activated protein kinase signaling in plants. Annu. Rev. Plant Biol. 2010, 61, 621–649. [Google Scholar] [PubMed]
- Ding, H.D.; Zhang, X.H.; Xu, S.C.; Sun, L.L.; Jiang, M.Y.; Zhang, A.Y.; Jin, Y.G. Induction of protection against paraquat-induced oxidative damage by abscisic acid in maize leaves is mediated through mitogen-activated protein kinase. J. Integr. Plant Biol. 2009, 51, 961–972. [Google Scholar] [CrossRef] [PubMed]
- Lu, K.; Guo, W.J.; Lu, J.X.; Yu, H.; Qu, C.M.; Tang, Z.L.; Li, J.; Chai, Y.; Liang, Y. Genome-wide survey and expression profile analysis of the mitogen-activated protein kinase (MAPK) gene family in Brassica rapa. PLoS ONE 2015, 10, e0132051. [Google Scholar]
- Song, Q.; Li, D.; Dai, Y.; Liu, S.; Huang, L.; Hong, Y.; Zhang, H.; Song, F. Characterization, expression patterns and functional analysis of the MAPK and MAPKK genes in watermelon (Citrullus lanatus). BMC Plant Biol. 2015, 15, 298. [Google Scholar] [CrossRef] [PubMed]
- Xu, H.; Li, K.; Yang, F.; Shi, Q.; Wang, X. Overexpression of CsNMAPK in tobacco enhanced seed germination under salt and osmotic stresses. Mol. Biol. Rep. 2010, 37, 3157–3163. [Google Scholar] [CrossRef] [PubMed]
- Matsuoka, D.; Sasayama, D.; Furuya, T.; Nanmori, T. Over-expression of MAP3Kδ4, an ABA-inducible Raf-like MAP3K that confers salt tolerance in Arabidopsis. Plant Biotechnol. 2013, 30, 111–118. [Google Scholar]
- Hoang, X.L.T.; Thu, N.B.A.; Thao, N.P.; Tran, L.S.P. Transcription factors in abiotic stress responses: Their potentials in crop improvement. In Improvement of Crops in the Era of Climatic Changes; Springer-Verlag: New York, NY, USA, 2014; pp. 337–366. [Google Scholar]
- Malhotra, S.; Sowdhamini, R. Interactions among plant transcription factors regulating expression of stress-responsive genes. Bioinform. Biol. Insights 2014, 8, 193–198. [Google Scholar] [PubMed]
- Hu, H.H.; Dai, M.Q.; Yao, J.L.; Xiao, B.Z.; Li, X.H.; Zhang, Q.F.; Xiong, L. Overexpressing a NAM, ATAF, and CUC (NAC) transcription factor enhances drought resistance and salt tolerance in rice. Proc. Natl. Acad. Sci. USA 2006, 103, 12987–12992. [Google Scholar] [CrossRef] [PubMed]
- Shingote, P.R.; Kawar, P.G.; Pagariya, M.C.; Kuhikar, R.S.; Thorat, A.S.; Babu, K.H. SoMYB18, a sugarcane MYB transcription factor improves salt and dehydration tolerance in tobacco. Acta Physiol. Plant. 2015, 37, 1–12. [Google Scholar] [CrossRef]
- Shen, Z.; Yao, J.; Sun, J.; Chang, L.; Wang, S.; Ding, M.; Qian, Z.; Zhang, H.; Zhao, N.; Sa, G. Populus euphratica HSF binds the promoter of WRKY1 to enhance salt tolerance. Plant Sci. 2015, 235, 89–100. [Google Scholar] [CrossRef] [PubMed]
- Liu, Q.L.; Xu, K.D.; Pan, Y.Z.; Jiang, B.B.; Liu, G.L.; Jia, Y.; Zhang, H.Q. Functional analysis of a novel chrysanthemum WRKY transcription factor gene involved in salt tolerance. Plant Mol. Biol. Rep. 2014, 32, 282–289. [Google Scholar] [CrossRef]
- Babitha, K.C.; Vemanna, R.S.; Nataraja, K.N.; Udayakumar, M. Overexpression of EcbHLH57 transcription factor from Eleusine coracana L. in tobacco confers tolerance to salt, oxidative and drought stress. PLoS ONE 2015, 10, e0137098. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.; Huang, W.; Chen, H.; Wu, G.; Yuan, H.; Song, X.; Kang, Q.; Zhao, D.; Jiang, W.; Liu, Y. Identification of differentially expressed genes in flax (Linum usitatissimum L.) under saline-alkaline stress by digital gene expression. Gene 2014, 549, 113–122. [Google Scholar] [CrossRef] [PubMed]
- Shen, X.; Wang, Z.; Song, X.; Xu, J.; Jiang, C.; Zhao, Y.; Ma, C.; Zhang, H. Transcriptomic profiling revealed an important role of cell wall remodeling and ethylene signaling pathway during salt acclimation in Arabidopsis. Plant Mol. Biol. 2014, 86, 303–317. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.C.; Xu, L.; Wang, Y.; Luo, X.B.; Zhu, X.W.; Kinuthia, K.B.; Nie, S.; Feng, H.; Li, C.; Liu, L. Transcriptome-based gene expression profiling identifies differentially expressed genes critical for salt stress response in radish (Raphanus sativus L.). Plant Cell Rep. 2016, 35, 329–346. [Google Scholar] [CrossRef] [PubMed]
- Shavrukov, Y. Salt stress or salt shock: Which genes are we studying? J. Exp. Bot. 2013, 64, 119–127. [Google Scholar] [CrossRef] [PubMed]
- Hmida-Sayari, A.; Gargouri-Bouzid, R.; Bidani, A.; Jaoua, L.; Savouré, A.; Jaoua, S. Overexpression of Δ1-pyrroline-5-carboxylate synthetase increases proline production and confers salt tolerance in transgenic potato plants. Plant Sci. 2005, 169, 746–752. [Google Scholar] [CrossRef]
- Guerzoni, J.T.S.; Belintani, N.G.; Moreira, R.M.P.; Hoshino, A.A.; Domingues, D.S.; Filho, J.C.B.; Vieira, L.G.E. Stress-induced Δ1-pyrroline-5-carboxylate synthetase (P5CS) gene confers tolerance to salt stress in transgenic sugarcane. Acta Physiol. Plant 2014, 36, 2309–2319. [Google Scholar] [CrossRef]
- Mittova, V.; Tal, M.; Volokita, M.; Guy, M. Salt stress induces up-regulation of an efficient chloroplast antioxidant system in the salt-tolerant wild tomato species Lycopersicon pennellii but not in the cultivated species. Physiol. Plant. 2002, 115, 393–400. [Google Scholar] [CrossRef] [PubMed]
- Rios-Gonzalez, K.; Erdei, L.; Lips, S.H. The activity of antioxidant enzymes in maize and sunflower seedlings as affected by salinity and different nitrogen sources. Plant Sci. 2002, 162, 923–930. [Google Scholar] [CrossRef]
- Gao, X.; Ren, Z.; Zhao, Y.; Zhang, H. Overexpression of SOD2 increases salt tolerance of Arabidopsis. Plant Physiol. 2004, 133, 1873–1881. [Google Scholar] [CrossRef] [PubMed]
- Luo, X.L.; Wu, J.H.; Li, Y.B.; Nan, Z.R.; Guo, X.; Wang, Y.X.; Zhang, A.; Wang, Z.; Xia, G.; Tian, Y. Synergistic effects of GhSOD1 and GhCAT1 overexpression in cotton chloroplasts on enhancing tolerance to methyl viologen and salt stresses. PLoS ONE 2013, 8, e54002. [Google Scholar] [CrossRef] [PubMed]
- Roxas, V.P.; Smith, R.K.; Allen, E.R.; Allen, R.D. Overexpression of glutathione S-transferase/glutathione peroxidase enhances the growth of transgenic tobacco seedlings during stress. Nat. Biotechnol. 1997, 15, 988–991. [Google Scholar] [CrossRef] [PubMed]
- Puhakainen, T.; Hess, M.W.; Mäkelä, P.; Svensson, J.; Heino, P.; Palva, E.T. Overexpression of multiple dehydrin genes enhances tolerance to freezing stress in Arabidopsis. Plant Mol. Biol. 2004, 54, 743–753. [Google Scholar] [CrossRef] [PubMed]
- Duan, J.; Cai, W. OsLEA3-2, an abiotic stress induced gene of rice plays a key role in salt and drought tolerance. PLoS ONE 2012, 7, e45117. [Google Scholar] [CrossRef] [PubMed]
- Grelet, J.; Benamar, A.; Teyssier, E.; Avelange-Macherel, M.H.; Grunwald, D.; Macherel, D. Identification in pea seed mitochondria of a late-embryogenesis abundant protein able to protect enzymes from drying. Plant Physiol. 2005, 137, 157–167. [Google Scholar] [CrossRef] [PubMed]
- Kosová, K.; Vítámvás, P.; Prášil, I.T. The role of dehydrins in plant response to cold. Biol. Plant. 2007, 51, 601–617. [Google Scholar] [CrossRef]
- Tolleter, D.; Hincha, D.K.; Macherel, D. A mitochondrial late embryogenesis abundant protein stabilizes model membranes in the dry state. Biochim. Biophys. Acta 2010, 1798, 1926–1933. [Google Scholar] [CrossRef] [PubMed]
- Ingram, J.; Bartels, D. The Molecular basis of dehydration tolerance in plants. Annu. Rev. Plant Physiol. Plant Mol. Biol. 1996, 47, 377–403. [Google Scholar] [CrossRef] [PubMed]
- Boursiac, Y.; Chen, S.; Luu, D.T.; Sorieul, M.; van den Dries, N.; Maurel, C. Early effects of salinity on water transport in Arabidopsis roots. Molecular and cellular features of aquaporin expression. Plant Physiol. 2005, 139, 790–805. [Google Scholar] [CrossRef] [PubMed]
- Aroca, R.; Porcel, R.; Ruiz-Lozano, J.M. Regulation of root water uptake under abiotic stress conditions. J. Exp. Bot. 2012, 63, 43–57. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.L.; Chen, A.P.; Zhong, N.Q.; Liu, N.; Wu, X.M.; Wang, F.; Yang, C.L.; Romero, M.F.; Xia, G.X. The Thellungiella salsuginea tonoplast aquaporin TsTIP1; 2 functions in protection against multiple abiotic stresses. Plant Cell Physiol. 2014, 55, 148–161. [Google Scholar] [CrossRef] [PubMed]
- Fricke, W.; Akhiyarova, G.; Wei, W.; Alexandersson, E.; Miller, A.; Kjellbom, P.O.; Richardson, A.; Wojciechowski, T.; Schreiber, L.; Veselov, D. The short-term growth response to salt of the developing barley leaf. J. Exp. Bot. 2006, 57, 1079–1095. [Google Scholar] [CrossRef] [PubMed]
- Gepstein, S.; Sabehi, G.; Carp, M.J.; Hajouj, T.; Nesher, M.F.; Yariv, I.; Dor, C.; Bassani, M. Large-scale identification of leaf senescence-associated genes. Plant J. 2003, 36, 629–642. [Google Scholar] [CrossRef] [PubMed]
- Seo, P.J.; Park, J.M.; Kang, S.K.; Kim, S.G.; Park, C.M. An Arabidopsis senescence-associated protein SAG29 regulates cell viability under high salinity. Planta 2011, 233, 189–200. [Google Scholar] [CrossRef] [PubMed]
- Bahieldin, A.; Atef, A.; Sabir, J.S.; Gadalla, N.O.; Edris, S.; Alzohairy, A.M.; Radhwan, N.A.; Baeshen, M.N.; Ramadan, A.M.; Eissa, H.F.; et al. RNA-Seq analysis of the wild barley (H. spontaneum) leaf transcriptome under salt stress. C. R. Biol. 2015, 338, 285–297. [Google Scholar] [CrossRef] [PubMed]
- Guo, J.; Shi, G.; Guo, X.; Zhang, L.; Xu, W.; Wang, Y.; Su, Z.; Hua, J. Transcriptome analysis reveals that distinct metabolic pathways operate in salt-tolerant and salt-sensitive upland cotton varieties subjected to salinity stress. Plant Sci. 2015, 238, 33–45. [Google Scholar] [CrossRef] [PubMed]
- Ziaf, K.; Loukehaich, R.; Gong, P.J.; Liu, H.; Han, Q.Q.; Wang, T.T.; Li, H.; Ye, Z. A multiple stress-responsive gene ERD15 from Solanum pennellii confers stress tolerance in tobacco. Plant Cell Physiol. 2011, 52, 1055–1067. [Google Scholar] [CrossRef] [PubMed]
- Zou, J.; Liu, C.F.; Liu, A.; Zou, D.; Chen, X.B. Overexpression of OsHsp17.0 and OsHsp23.7 enhances drought and salt tolerance in rice. J. Plant Physiol. 2012, 169, 628–635. [Google Scholar] [CrossRef] [PubMed]
- Kurotani, K.; Hayashi, K.; Hatanaka, S.; Toda, Y.; Ogawa, D.; Ichikawa, H.; Ishimaru, Y.; Tashita, R.; Suzuki, T.; Ueda, M.; et al. Elevated levels of CYP94 family gene expression alleviate the jasmonate response and enhance salt tolerance in rice. Plant Cell Physiol. 2015, 56, 779–789. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Guo, S.; Zhao, Y.; Chen, D.; Chong, K.; Xu, Y. Overexpression of a homopeptide repeat-containing bHLH protein gene (OrbHLH001) from Dongxiang Wild Rice confers freezing and salt tolerance in transgenicArabidopsis. Plant Cell Rep. 2010, 29, 977–986. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Lu, G.; Hao, Y.; Guo, H.; Zhao, J.; Cheng, H. ABP9, a maize bZIP transcription factor, enhances tolerance to salt and drought in transgenic cotton. Planta 2017, 246, 453–469. [Google Scholar] [CrossRef] [PubMed]
- Hoagland, D.R.; Arnon, D.I. The water-culture method for growing plants without soil. Calif. Agric. Exp. Stn. Circ. 1950, 347, 357–359. [Google Scholar]
- Porra, R.J. The chequered history of the development and use of simultaneous equations for the accurate determination of chlorophylls a and b. Photosyn. Res. 2002, 73, 149–156. [Google Scholar] [CrossRef] [PubMed]
- Qiu, N.W.; Zhou, F.; Wang, Y.; Peng, X.Y.; Hua, C. The strategy of Na+ compartmentation and growth of Atriplex centralasiatica in adaptation to saline environments. Russ. J. Plant Physl. 2014, 61, 238–245. [Google Scholar] [CrossRef]
- Bates, L.S.; Waldren, R.P.; Teare, I.D. Rapid determination of free proline for water stress studies. Plant Soil 1973, 39, 205–207. [Google Scholar] [CrossRef]
- Jermyn, M.A. Increasing the sensitivity of the anthrone method for carbohydrate. Anal. Biochem. 1975, 68, 332–335. [Google Scholar] [CrossRef]
- Bradford, M.M. A rapid and sensitive method for the quantization of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 1976, 72, 248–254. [Google Scholar] [CrossRef]
- Knöraer, O.C.; Durner, J.; Böger, P. Alterations in the antioxidative system of suspension-cultured soybean cells (Glycine max) induced by oxidatice stress. Physiol. Plant. 1996, 97, 388–396. [Google Scholar] [CrossRef]
- Chance, B.; Maehly, A.C. Assay of catalase and peroxidases. In Methods in Enzymology; Elsevier: Amsterdam, The Netherlands, 1955; pp. 764–775. [Google Scholar]
- Beyer, W.F.; Fridovich, I. Assaying for superoxide dismutase activity: Some large consequences of minor changes in condition. Anal. Biochem. 1987, 161, 559–566. [Google Scholar] [CrossRef]
- Li, H.; Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar] [CrossRef] [PubMed]
- Langmead, B.; Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 2012, 9, 357–359. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Dewey, C.N. RSEM: Accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform. 2011, 12, 93–99. [Google Scholar] [CrossRef] [PubMed]
- Tarazona, S.; Garcia-Alcalde, F.; Dopazo, J.; Ferrer, A.; Conesa, A. Differential expression in RNA-seq: A matter of depth. Genome Res. 2011, 21, 2213–2223. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCt method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]


| NaCl Concentration | Pn/µmol (CO2) m−2∙s−1 | Gs/mmol (H2O) m−2∙s−1 | Ci/µmol∙mol−1 | Tr/mmol (H2O) m−2∙s−1 |
|---|---|---|---|---|
| 0 mM | 15.72 ± 1.02 a | 141.7 ± 19.0 a | 193.8 ± 17.9 a | 2.30 ± 0.14 a |
| 100 mM | 11.92 ± 0.89 b | 84.7 ± 8.5 b | 151.7 ± 13.1 b | 1.48 ± 0.12 b |
| 200 mM | 9.58 ± 0.87 c | 58.0 ± 5.6 c | 114.5 ± 14.8 c | 1.17 ± 0.11 c |
| 300 mM | 5.20 ± 1.23 d | 26.2 ± 6.4 d | 88.2 ± 12.7 d | 0.57 ± 0.10 d |
| 400 mM | 2.13 ± 0.34 e | 10.5 ± 3.8 e | 67.8 ± 7.6 e | 0.26 ± 0.07 e |
| Gene ID | log2 Ratio (Treatment/CK) | Probability | Annotation (BlastX to A. thaliana) |
|---|---|---|---|
| Bra015388 | 5.58 | 0.90 | mitogen-activated protein kinase kinase kinase 18 |
| Bra009003 | 2.95 | 0.87 | CBL-interacting protein kinase 5 |
| Bra023777 | 1.27 | 0.81 | CBL-interacting protein kinase 7 |
| Bra011236 | 1.97 | 0.87 | CBL-interacting protein kinase 6 |
| Bra022794 | 1.94 | 0.83 | CBL-interacting protein kinase 11 |
| Bra010263 | 1.65 | 0.84 | CBL-interacting protein kinase 6 |
| Bra016962 | 1.74 | 0.83 | Protein kinase superfamily protein |
| Bra014542 | 2.52 | 0.84 | Protein kinase superfamily protein |
| Bra028679 | 2.36 | 0.86 | Diacylglycerol kinase1 |
| Bra011033 | 1.87 | 0.80 | Leucine-rich receptor-like protein kinase family protein |
| Bra005168 | 1.69 | 0.83 | Receptor lectin kinase |
| Bra028545 | 1.64 | 0.84 | Pyruvate kinase family protein |
| Bra006864 | 1.63 | 0.83 | Phosphofructokinase 7 |
| Bra022870 | 1.36 | 0.80 | Leucine-rich repeat protein kinase family protein |
| Bra033745 | 3.16 | 0.81 | Calmodulin like 43 |
| Bra012889 | 2.79 | 0.88 | Calmodulin-like 41 |
| Bra025654 | 2.21 | 0.86 | Calcium-binding EF-hand family protein |
| Bra017927 | 1.58 | 0.82 | Calcium-binding EF-hand family protein |
| Bra000430 | 1.56 | 0.84 | Calcium-binding EF-hand family protein |
| Bra016936 | 1.54 | 0.84 | Calcium-binding EF-hand family protein |
| Bra004620 | 1.51 | 0.84 | Calcium-binding EF-hand family protein |
| Bra039661 | 1.51 | 0.84 | Sodium/calcium exchanger family protein/calcium-binding EF hand |
| Bra028148 | 1.38 | 0.81 | Calcium-dependent lipid-binding (CaLB domain) family protein |
| Gene ID | log2 Ratio (Treatment/CK) | Probability | Annotation (BlastX to A. thaliana) |
|---|---|---|---|
| Bra007637 | 4.10 | 0.89 | Homeobox 12 |
| Bra014417 | 1.83 | 0.85 | Homeobox 12 |
| Bra039116 | 1.44 | 0.83 | Homeobox 1 |
| Bra039265 | 3.75 | 0.90 | Homeobox 7 |
| Bra025658 | 3.47 | 0.83 | NAC domain containing protein 6 |
| Bra019052 | 3.38 | 0.89 | NAC (No Apical Meristem) domain transcriptional regulator superfamily protein |
| Bra026353 | 3.11 | 0.88 | NAC (No Apical Meristem) domain transcriptional regulator superfamily protein |
| Bra004385 | 2.90 | 0.86 | NAC-like, activated by AP3/PI |
| Bra018998 | 2.28 | 0.87 | NAC domain containing protein 19 |
| Bra008849 | 1.70 | 0.80 | NAC domain containing protein 83 |
| Bra006186 | 1.44 | 0.81 | NAC domain containing protein 83 |
| Bra016441 | 3.12 | 0.85 | PLATZ transcription factor family protein |
| Bra017670 | 3.10 | 0.86 | GATA transcription factor 3 |
| Bra040092 | 3.09 | 0.87 | Integrase-type DNA-binding superfamily protein |
| Bra020017 | 3.06 | 0.84 | Phytochrome interacting factor 3 |
| Bra001806 | 2.93 | 0.83 | Nuclear factor Y, subunit A9 |
| Bra010049 | 2.85 | 0.86 | Heat shock transcription factor B2A |
| Bra013253 | 4.19 | 0.88 | Heat shock transcription factor C1 |
| Bra039022 | 2.52 | 0.87 | CCCH-type zinc finger family protein |
| Bra011087 | 2.08 | 0.87 | Zinc finger C-x8-C-x5-C-x3-H type family protein |
| Bra003500 | 1.81 | 0.82 | Basic region/leucine zipper motif 53 |
| Bra001752 | 1.92 | 0.84 | Zinc-finger protein 2 |
| Bra009464 | 2.18 | 0.80 | Zinc finger of Arabidopsis thaliana 6 |
| Bra011485 | 2.47 | 0.88 | Abscisic acid responsive elements-binding factor 3 |
| Bra019645 | 1.88 | 0.86 | A20/AN1-like zinc finger family protein |
| Bra040260 | 2.12 | 0.85 | Abscisic acid responsive elements-binding factor 2 |
| Bra011545 | 2.27 | 0.87 | G-box binding factor 6 |
| Bra004550 | 2.10 | 0.84 | G-box binding factor 3 |
| Bra017664 | 1.56 | 0.81 | G-box binding factor 6 |
| Bra012337 | 2.27 | 0.85 | Myb domain protein 3 |
| Bra024526 | 1.99 | 0.82 | Myb domain protein 3 |
| Bra028707 | 2.19 | 0.82 | WRKY DNA-binding protein 26 |
| Bra010231 | 1.81 | 0.82 | WRKY DNA-binding protein 11 |
| Bra039409 | 1.79 | 0.84 | AtBS1(activation-tagged BRI1 suppressor 1)-interacting factor 1 |
| Bra036061 | 1.66 | 0.84 | cooperatively regulated by ethylene and jasmonate 1 |
| Bra039658 | 1.66 | 0.82 | Ethylene response factor 8 |
| Bra021200 | 1.33 | 0.81 | Ethylene-responsive element binding protein |
| Bra011700 | 1.52 | 0.82 | GRAS family transcription factor |
| Bra018896 | 2.43 | 0.87 | Basic helix-loop-helix (bHLH) DNA-binding superfamily protein |
| Gene ID | log2 Ratio (Treatment/CK) | Probability | Annotation (BlastX to A. thaliana) |
|---|---|---|---|
| Bra017051 | 5.19 | 0.86 | Δ1-pyrroline-5-carboxylate synthase 1 |
| Bra005012 | 4.04 | 0.92 | Δ1-pyrroline-5-carboxylate synthase 1 |
| Bra007179 | 2.34 | 0.87 | Δ1-pyrroline-5-carboxylate synthase 2 |
| Bra036282 | 3.24 | 0.84 | Sucrose synthase 3 |
| Gene ID | log2 Ratio (Treatment/CK) | Probability | Annotation (BlastX to A. thaliana) |
|---|---|---|---|
| Bra016127 | 1.43 | 0.82 | Peroxidase superfamily protein |
| Bra013576 | 1.23 | 0.81 | Peroxidase superfamily protein |
| Bra009105 | 3.56 | 0.83 | Peroxidase superfamily protein |
| Bra039816 | 2.59 | 0.88 | Peroxidase superfamily protein |
| Bra029933 | 4.02 | 0.91 | peroxidase CB |
| Bra012239 | 1.69 | 0.81 | Catalase 1 |
| Bra025995 | 1.74 | 0.82 | Glutathione S-transferase TAU 24 |
| Bra018543 | 1.74 | 0.85 | Glutathione S-transferase F3 |
| Bra000875 | 1.50 | 0.81 | Glutathione S-transferase F3 |
| Bra008915 | 1.65 | 0.84 | Glutathione S-transferase |
| Bra024820 | 1.33 | 0.81 | Glutathione S-transferase zeta 1 |
| Bra005677 | 2.68 | 0.89 | Ferretin 1 |
| Bra011408 | 1.49 | 0.83 | Thioredoxin superfamily protein |
| Bra007718 | 1.48 | 0.81 | Thioredoxin superfamily protein |
| Bra004455 | 1.45 | 0.82 | Thioredoxin family protein |
| Gene ID | log2 Ratio (Treatment/CK) | Probability | Annotation (BlastX to A. thaliana) |
|---|---|---|---|
| Bra008242 | 1.61 | 0.84 | Dehydrin family protein |
| Bra012230 | 2.80 | 0.89 | Dehydrin family protein |
| Bra015779 | 2.36 | 0.88 | Dehydrin family protein |
| Bra025819 | 1.97 | 0.87 | Dehydrin family protein |
| Bra037177 | 6.82 | 0.99 | Dehydrin family protein |
| Bra031809 | 6.89 | 0.98 | Dehydrin family protein |
| Bra027219 | 12.22 | 0.98 | Late embryogenesis abundant (LEA) protein |
| Bra001603 | 10.30 | 0.91 | Late embryogenesis abundant (LEA) protein |
| Bra021457 | 13.63 | 0.99 | Late embryogenesis abundant (LEA) protein |
| Bra021436 | 6.75 | 0.97 | Late embryogenesis abundant (LEA) protein |
| Bra022221 | 8.59 | 0.99 | Late embryogenesis abundant (LEA) protein |
| Bra039946 | 2.11 | 0.87 | Late embryogenesis abundant (LEA) protein |
| Bra005353 | 9.83 | 0.88 | Late embryogenesis abundant (LEA) protein |
| Bra039956 | 6.66 | 0.93 | Late embryogenesis abundant (LEA) protein |
| Bra007054 | 2.77 | 0.87 | Late embryogenesis abundant (LEA) protein |
| Bra003039 | 1.44 | 0.80 | Late embryogenesis abundant (LEA) protein |
| Bra009225 | 8.91 | 0.99 | Late embryogenesis abundant (LEA) protein |
| Bra005911 | 5.92 | 0.97 | Late embryogenesis abundant (LEA) protein |
| Bra030494 | 2.79 | 0.88 | Late embryogenesis abundant (LEA) protein |
| Bra029121 | 10.09 | 0.90 | Low temperature induced 65 |
| Bra022584 | 4.93 | 0.94 | Low temperature induced 65 |
| Bra022585 | 2.64 | 0.88 | Low temperature induced65 |
| Bra002594 | 5.21 | 0.94 | Highly ABA-induced PP2C gene 1 |
| Bra015579 | 4.80 | 0.91 | Highly ABA-induced PP2C gene 2 |
| Bra031574 | 3.73 | 0.83 | Highly ABA-induced PP2C gene 2 |
| Bra040610 | 3.70 | 0.90 | AIG2-like (avirulence induced gene) family protein |
| Bra025365 | 1.68 | 0.82 | AIG2-like (avirulence induced gene) family protein |
| Bra021385 | 1.33 | 0.81 | AIG2-like (avirulence induced gene) family protein |
| Bra025649 | 1.50 | 0.83 | AIG2-like (avirulence induced gene) family protein |
| Bra008661 | 2.85 | 0.89 | Stress-responsive protein (KIN2) |
| Bra009620 | 2.08 | 0.83 | SALT induced serine rich (SIS). |
| Bra000227 | 1.44 | 0.81 | Dehydration-induced protein (ERD15) |
| Bra002217 | 1.41 | 0.83 | Aluminium induced protein with YGL and LRDR motifs |
| Bra002216 | 1.30 | 0.82 | Aluminium induced protein with YGL and LRDR motifs |
| Bra016644 | 2.45 | 0.85 | Heat shock protein 70B |
| Bra015922 | 2.20 | 0.83 | Heat shock protein 101 |
| Bra013774 | 1.27 | 0.81 | Heat shock protein 90-7 |
| Bra001078 | 3.58 | 0.90 | Cytochrome P450, family 87, subfamily A, polypeptide 9 |
| Bra017819 | 1.85 | 0.85 | Cytochrome P450, family 81, subfamily D, polypeptide 8 |
| Bra023394 | 7.62 | 0.98 | Senescence-associated gene 29 |
| Bra008850 | 6.01 | 0.97 | Senescence-associated gene 29 |
| Bra006185 | 6.01 | 0.93 | Senescence-associated gene 29 |
| Bra000111 | 1.60 | 0.82 | Plasma membrane intrinsic protein 2E |
| Bra007100 | 2.50 | 0.85 | Plasma membrane intrinsic protein 2;5 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Qiu, N.; Liu, Q.; Li, J.; Zhang, Y.; Wang, F.; Gao, J. Physiological and Transcriptomic Responses of Chinese Cabbage (Brassica rapa L. ssp. Pekinensis) to Salt Stress. Int. J. Mol. Sci. 2017, 18, 1953. https://doi.org/10.3390/ijms18091953
Qiu N, Liu Q, Li J, Zhang Y, Wang F, Gao J. Physiological and Transcriptomic Responses of Chinese Cabbage (Brassica rapa L. ssp. Pekinensis) to Salt Stress. International Journal of Molecular Sciences. 2017; 18(9):1953. https://doi.org/10.3390/ijms18091953
Chicago/Turabian StyleQiu, Nianwei, Qian Liu, Jingjuan Li, Yihui Zhang, Fengde Wang, and Jianwei Gao. 2017. "Physiological and Transcriptomic Responses of Chinese Cabbage (Brassica rapa L. ssp. Pekinensis) to Salt Stress" International Journal of Molecular Sciences 18, no. 9: 1953. https://doi.org/10.3390/ijms18091953
APA StyleQiu, N., Liu, Q., Li, J., Zhang, Y., Wang, F., & Gao, J. (2017). Physiological and Transcriptomic Responses of Chinese Cabbage (Brassica rapa L. ssp. Pekinensis) to Salt Stress. International Journal of Molecular Sciences, 18(9), 1953. https://doi.org/10.3390/ijms18091953





