Arid1b Haploinsufficiency Causes Abnormal Brain Gene Expression and Autism-Related Behaviors in Mice
Abstract
:1. Introduction
2. Results
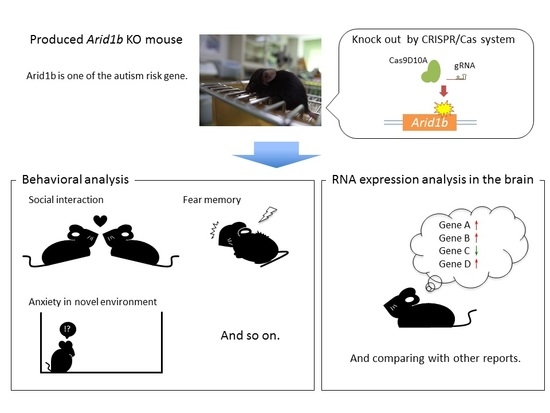
2.1. Generation of Arid1b Heterozygous Knockout Mice and Assessment of Neurological and Sensory Functions
2.2. ASD-Like Behaviours in Arid1b hKO Mice
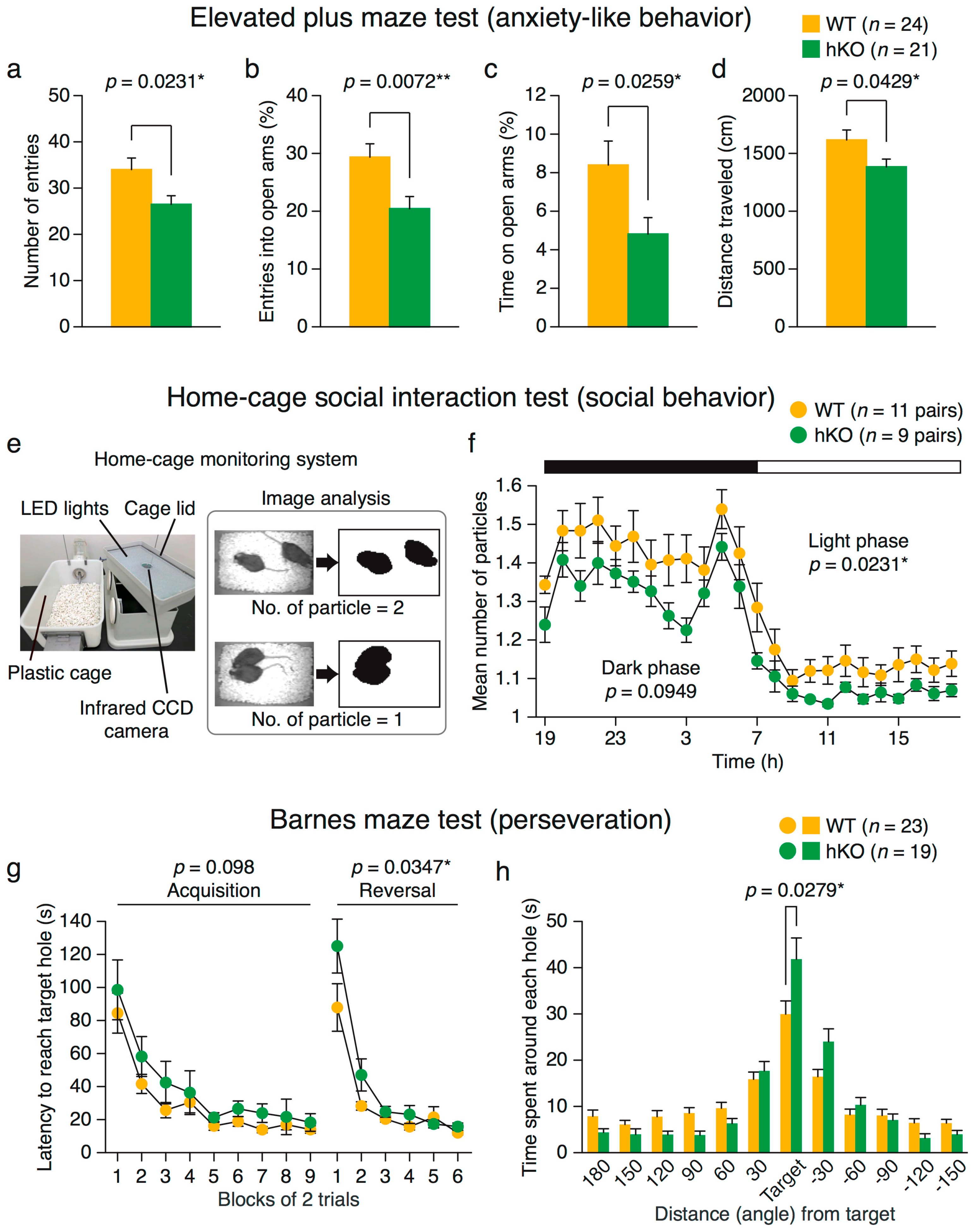
2.2.1. Increased Anxiety-Like Behavior in Arid1b hKO Mice
2.2.2. Altered Social Behavior in Arid1b hKO Mice
2.2.3. Perseveration (Behavioral Inflexibility) in Arid1b hKO Mice
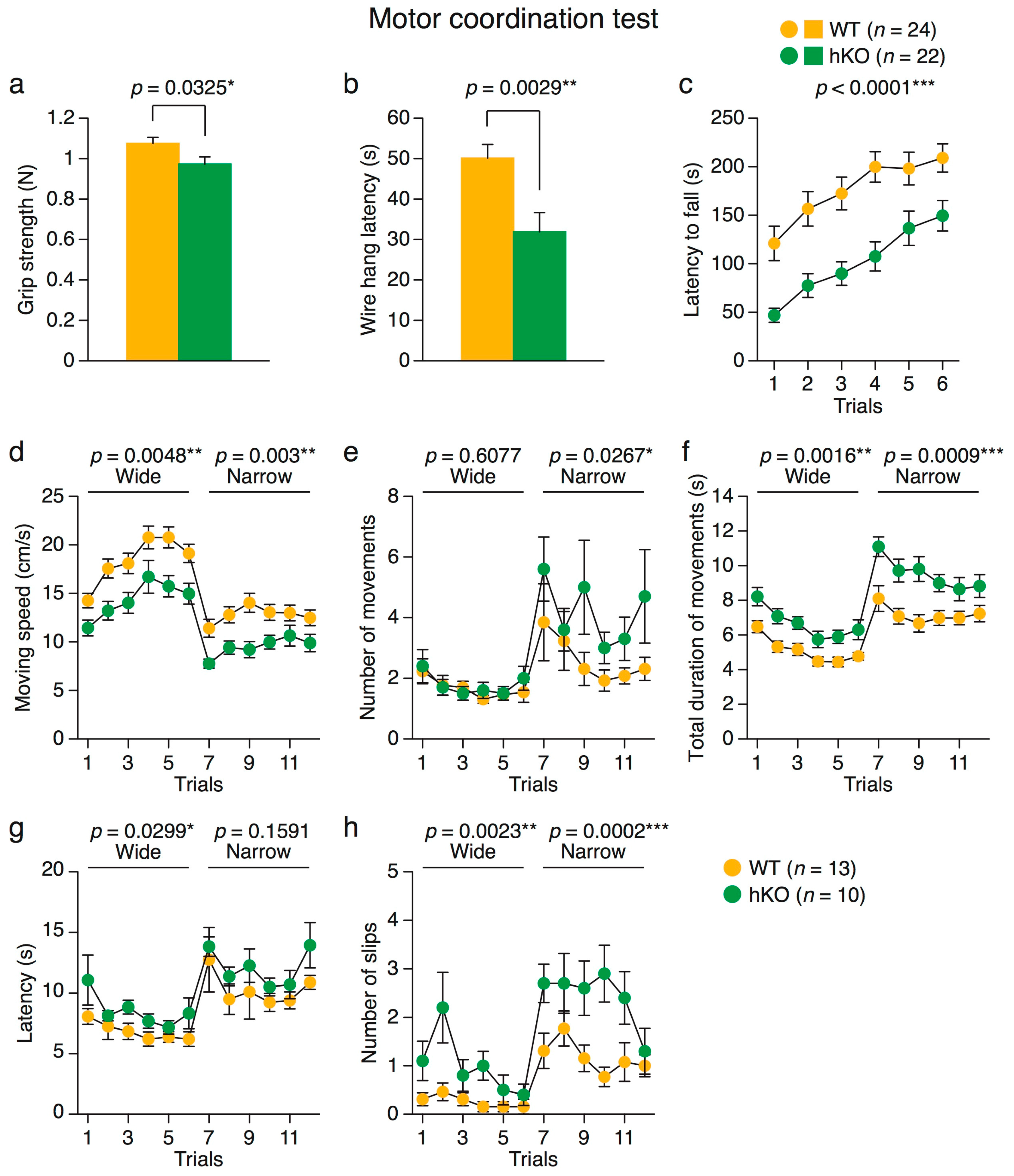
2.2.4. Impaired Motor Coordination in Arid1b hKO Mice
2.3. Other Behavioural Features in Arid1b hKO Mice
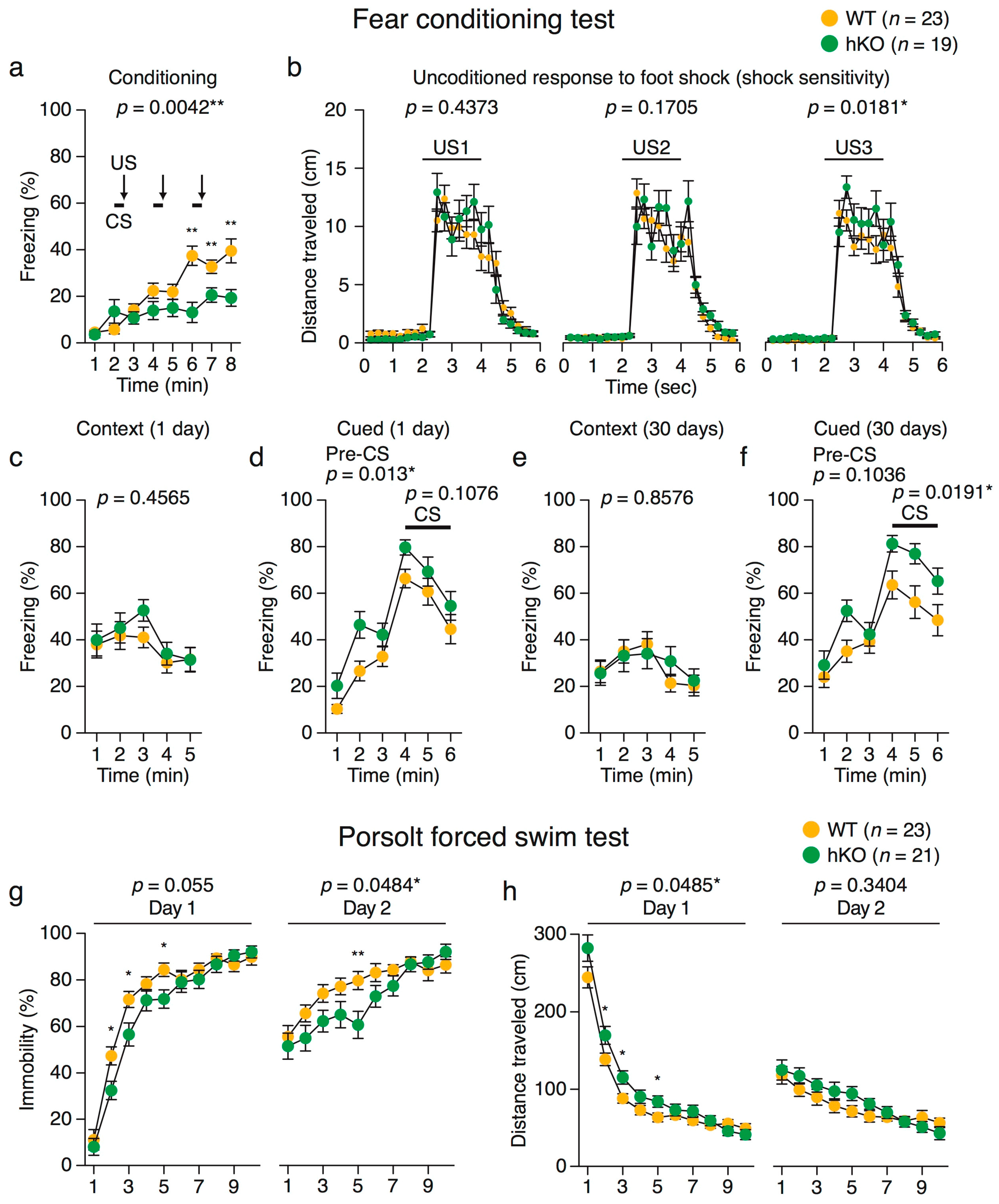
2.3.1. Heightened Response to Intense Aversive Stimuli in Arid1b hKO Mice
2.3.2. Enhanced Fear Generalization and Long-Term Fear Memory in Arid1b hKO Mice
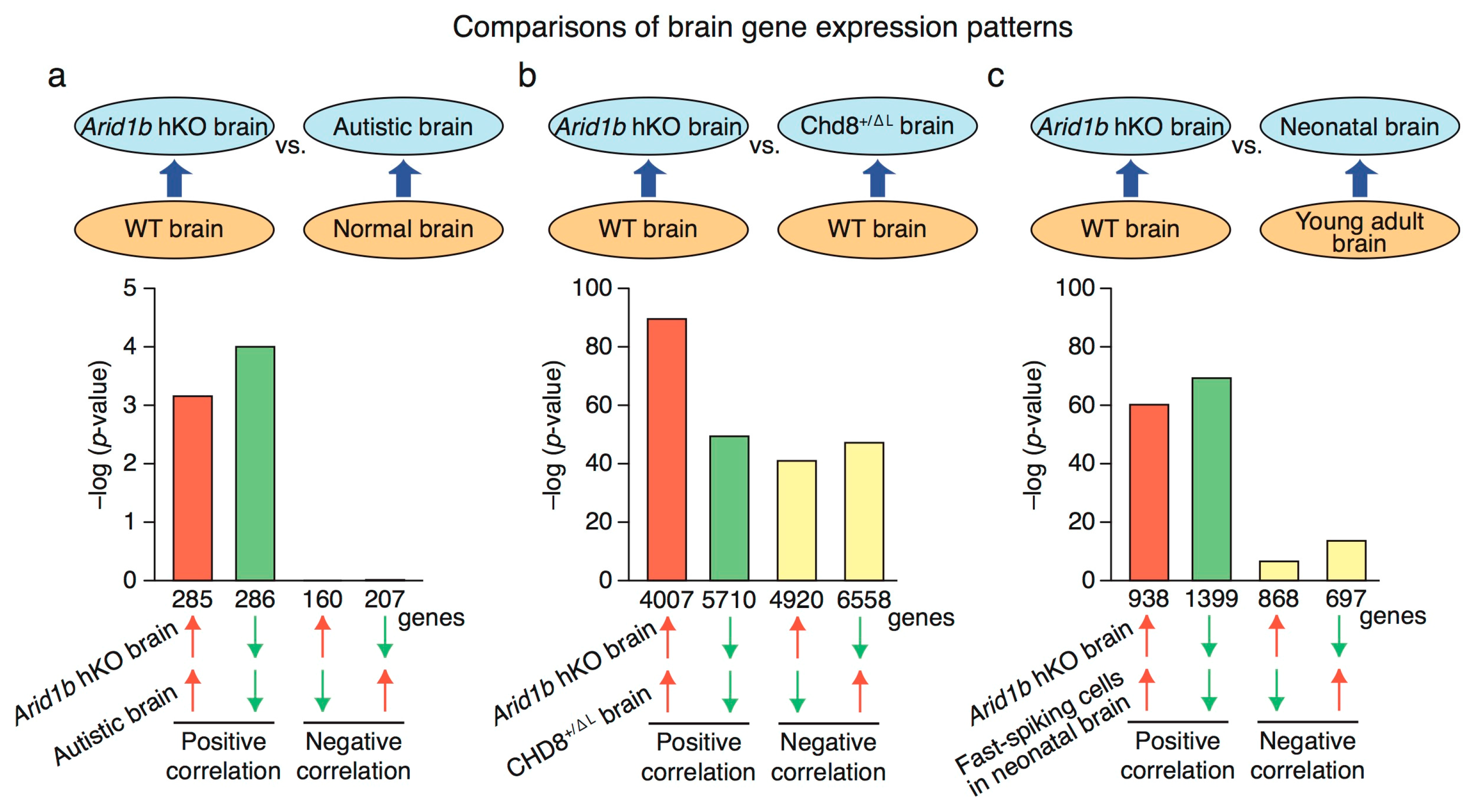
2.4. Genome-Wide RNA Expression Profile in Arid1b hKO Mice
2.5. Hydrocephalus in Arid1b hKO Mice
3. Discussion
4. Materials and Methods
4.1. Animals
4.2. General Procedures for Behavioral Tests
4.3. Motor Function Tests
4.4. Hot Plate Test
4.5. Open Field Test
4.6. Light/Dark Transition Test
4.7. Elevated Plus Maze Test
4.8. Social Interaction Test in a Novel Environment
4.9. Three-Chamber Social Approach Test
4.10. Home-Cage Social Interaction Test
4.11. Barnes Maze Test
4.12. Startle Response and Prepulse Inhibition Tests
4.13. Porsolt Forced Swim Test
4.14. Fear Conditioning Test
4.15. RNA-Seq Analysis
4.16. Gene Expression Pattern Analysis
4.17. Quantitive Reverse Transcription Polymerase Chain-Reaction (RT-PCR) Analysis
4.18. Western Blot Assay
4.19. Histology of Arid1b hKO Mouse Brain
4.20. Statistical Analyses
4.21. Data Availability
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| ASD | Autism spectrum disorder |
| hKO | Hetero knockout |
References
- Wing, L. Language, social, and cognitive impairments in autism and severe mental retardation. J. Autism Dev. Disord. 1981, 11, 31–44. [Google Scholar] [CrossRef] [PubMed]
- Wing, L.; Gould, J. Severe impairments of social interaction and associated abnormalities in children: Epidemiology and classification. J. Autismaal Dev. Disord. 1979, 9, 11–29. [Google Scholar] [CrossRef]
- Turner, A.H.; Greenspan, K.S.; van Erp, T.G. Pallidum and lateral ventricle volume enlargement in autism spectrum disorder. Psychiatry Res. 2016, 252, 40–45. [Google Scholar] [CrossRef] [PubMed]
- Leary, M.R.; Hill, D.A. Moving on: Autism and movement disturbance. Ment. Retard. 1996, 34, 39–53. [Google Scholar] [PubMed]
- Noor, A.; Whibley, A.; Marshall, C.R.; Gianakopoulos, P.J.; Piton, A.; Carson, A.R.; Orlic-Milacic, M.; Lionel, A.C.; Sato, D.; Pinto, D.; et al. Disruption at the PTCHD1 Locus on Xp22.11 in Autism spectrum disorder and intellectual disability. Sci. Transl. Med. 2010, 2, 49–68. [Google Scholar] [CrossRef] [PubMed]
- Purpura, G.; Fulceri, F.; Puglisi, V.; Masoni, P.; Contaldo, A. Motor coordination impairment in children with autism spectrum disorder: A pilot study using Movement Assessment Battery for Children-2 Checklist. Minerva. Pediatr. 2016. Available online: https://www.minervamedica.it/en/journals/minerva-pediatrica/article.php?cod=R15Y9999N00A16101202. [PubMed]
- Christensen, D.L.; Baio, J.; van Naarden Braun, K.; Bilder, D.; Charles, J.; Constantino, J.N.; Daniels, J.; Durkin, M.S.; Fitzgerald, R.T.; Kurzius-Spencer, M.; et al. Prevalence and Characteristics of Autism Spectrum Disorder Among Children Aged 8 Years—Autism and Developmental Disabilities Monitoring Network, 11 Sites, United States, 2012. MMWR Surveill. Summ. 2016, 65, 1–23. [Google Scholar] [CrossRef] [PubMed]
- Holmboe, K.; Rijsdijk, F.V.; Hallett, V.; Happé, F.; Plomin, R.; Ronald, A. Strong genetic influences on the stability of autistic traits in childhood. J. Am. Acad. Child. Adolesc. Psychiatry 2014, 53, 221–230. [Google Scholar] [CrossRef] [PubMed]
- Taylor, M.J.; Gillberg, C.; Linchtenstein, P.; Lundstrom, S. Etiological influences on the stability of autistic traits from childhood to early adulthood: Evidence from a twin study. Mol. Autism 2017, 8, 5. [Google Scholar] [CrossRef] [PubMed]
- Nakatani, J.; Tamada, K.; Hatanaka, F.; Ise, S.; Ohta, H.; Inoue, K.; Tomonaga, S.; Watanabe, Y.; Chung, Y.J.; Banerjee, R.; et al. Abnormal behavior in a chromosome-engineered mouse model for human 15q11-13 duplication seen in Autism. Cell 2009, 137, 1235–1246. [Google Scholar] [CrossRef] [PubMed]
- Jaramillo, T.C.; Speed, H.E.; Xuan, Z.; Reimers, J.M.; Liu, S.; Powell, C.M. Altered striatal synaptic function and abnormal behaviour in shank3 Exon4-9 deletion mouse model of Autism. Autism Res. 2016, 9, 350–375. [Google Scholar] [CrossRef] [PubMed]
- Katayama, Y.; Nishiyama, M.; Shoji, H.; Ohkawa, Y.; Kawamura, A.; Sato, T.; Suyama, M.; Takumi, T.; Miyakawa, T.; Nakayama, K. CHD8 haploinsufficiency results in autistic-like phenotypes in mice. Nature 2016, 537, 675–679. [Google Scholar] [CrossRef] [PubMed]
- Durak, O.; Gao, F.; Kaeser-Woo, Y.J.; Rueda, R.; Martorell, A.J.; Nott, A.; Liu, C.Y.; Watson, L.A.; Tsai, L.H. Chd8 mediates cortical neurogenesis via transcriptional regulation of cell cycle and Wnt signaling. Nat. Neurosci. 2016, 19, 1477–1488. [Google Scholar] [CrossRef] [PubMed]
- Gompers, A.L.; Su-Feher, L.; Ellegood, J.; Copping, N.A.; Riyadh, M.A.; Stradleigh, T.W.; Pride, M.C.; Schaffler, M.D.; Wade, A.A.; Catta-Preta, R.; et al. Germline Chd8 haploinsufficiency alters brain development in mouse. Nat. Neurosci. 2017, 20, 1062–1073. [Google Scholar] [CrossRef] [PubMed]
- Crawley, J.N. Designing mouse behavioral tasks relevant to autistic-like behaviors. Ment. Retard. Dev. Disabil. Res. Rev. 2004, 10, 248–258. [Google Scholar] [CrossRef] [PubMed]
- De Rubeis, S.; He, X.; Goldberg, A.P.; Poultney, C.S.; Samocha, K.; Cicek, A.E.; Kou, Y.; Liu, L.; Fromer, M.; Walker, S.; et al. Synaptic, transcriptional and chromatin genes disrupted in autism. Nature 2014, 515, 209–215. [Google Scholar] [CrossRef] [PubMed]
- O’Roak, B.J.; Vives, L.; Girirajan, S.; Karakoc, E.; Krumm, N.; Coe, B.P.; Levy, R.; Ko, A.; Lee, C.; Smith, J.D.; et al. Sporadic autism exomes reveal a highly interconnected protein network of de novo mutations. Nature 2012, 485, 246–250. [Google Scholar] [CrossRef] [PubMed]
- Neale, B.M.; Kou, Y.; Liu, L.; Ma’ayan, A.; Samocha, K.E.; Sabo, A.; Lin, C.F.; Stevens, C.; Wang, L.S.; Makarov, V.; et al. Patterns and rates of exonic de novo mutations in autism spectrum disorders. Nature 2012, 485, 242–245. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Talkowski, M.E.; Rosenfeld, J.A.; Blumenthal, I.; Pillalamarri, V.; Chiang, C.; Heilbut, A.; Ernst, C.; Hanscom, C.; Rossin, E.; Lindgren, A.M.; et al. Sequencing chromosomal abnormalities reveals neurodevelopmental loci that confer risk across diagnostic boundaries. Cell 2012, 149, 525–537. [Google Scholar] [CrossRef] [PubMed]
- O’Roak, B.J.; Vives, L.; Fu, W.; Egertson, J.D.; Stanaway, I.B.; Phelps, I.G.; Carvill, G.; Kumar, A.; Lee, C.; Ankenman, K.; et al. Multiplex targeted sequencing identifies recurrently mutated genes in autism spectrum disorders. Science 2012, 338, 1619–1622. [Google Scholar] [CrossRef] [PubMed]
- Bernier, R.; Golzio, C.; Xiong, B.; Stessman, H.A.; Coe, B.P.; Penn, O.; Witherspoon, K.; Gerdts, J.; Baker, C.; Vulto-van Silfhout, A.T.; et al. Disruptive CHD8 mutations define a subtype of autism early in development. Cell 2014, 158, 263–276. [Google Scholar] [CrossRef] [PubMed]
- D’Gama, A.M.; Pochareddy, S.; Li, M.; Jamuar, S.S.; Reiff, R.E.; Lam, A.T.; Sestan, N.; Walsh, C.A. Targeted DNA sequencing from Autism spectrum disorder brains implicates multiple genetic mechanisms. Neuron 2015, 88, 910–917. [Google Scholar] [CrossRef] [PubMed]
- Nord, A.S.; Roeb, W.; Dickel, D.E.; Walsh, T.; Kusenda, M.; O’Connor, K.L.; Malhotra, D.; McCarthy, S.E.; Stray, S.M.; Taylor, S.M.; et al. Reduced transcript expression of genes affected by inherited and de novo CNVs in Autism. Eur. J. Hum. Genet. 2011, 19, 727–731. [Google Scholar] [CrossRef] [PubMed]
- Halgren, C.; Kjaergaard, S.; Bak, M.; Hansen, C.; El-Schich, Z.; Anderson, C.M.; Henriksen, K.F.; Hjalgrim, H.; Kirchhoff, M.; Bijlsma, E.K.; et al. Corpus callosum abnormalities, intellectual disability, speech impairment, and autism in patients with haploinsufficiency of ARID1B. Clin. Genet. 2012, 82, 248–255. [Google Scholar] [CrossRef] [PubMed]
- Santen, G.W.; Aten, E.; Sun, Y.; Almomani, R.; Gilissen, C.; Nielsen, M.; Kant, S.G.; Snoeck, I.N.; Peeters, E.A.; Hilhorst-Hofstee, Y.; et al. Mutations in SWI/SNF chromatin remodeling complex gene ARID1B cause Coffin-Siris syndrome. Nat. Genet. 2012, 44, 379–380. [Google Scholar] [CrossRef] [PubMed]
- Kanner, L. Autistic disturbances of affective contact. Nerv. Child. 1943, 2, 217–250. [Google Scholar]
- Gandal, M.J.; Nesbitt, A.M.; McCurdy, R.M.; Alter, M.D. Measuring the maturity of the fast-spiking interneuron transcriptional program in Autism, Schizophrenia, and bipolar disorder. PLoS ONE 2012, 7, e41215. [Google Scholar] [CrossRef] [PubMed]
- Asada, K.; Tojo, Y.; Osanai, H.; Saito, A.; Hasegawa, T.; Kumagaya, S. Reduced personal space in individuals with Autism spectrum disorder. PLoS ONE 2016, 11, e0146306. [Google Scholar] [CrossRef] [PubMed]
- Matson, J.L.; Dempsey, T.; Lovullo, S.V.; Wilkins, J. The effects of intellectual functioning on the range of core symptoms of autism spectrum disorders. Res. Dev. Disabil. 2008, 29, 341–350. [Google Scholar] [CrossRef] [PubMed]
- McCarthy, S.E.; Gillis, J.; Kramer, M.; Lihm, J.; Yoon, S.; Berstein, Y.; Mistry, M.; Pavlidis, P.; Solomon, R.; et al. De novo mutations in schizophrenia implicate chromatin remodeling and support a genetic overlap with autism and intellectual disability. Mol. Psychiatry 2014, 19, 652–658. [Google Scholar] [CrossRef] [PubMed]
- Srivastava, A.K.; Schwartz, C.E. Intellectual disability and Autism spectrum disorders: Causal genes and molecular mechanisms. Neurosci. Biobehav. Rev. 2014, 46, 161–174. [Google Scholar] [CrossRef] [PubMed]
- Celen, C.; Chuang, J.C.; Luo, X.; Nijem, N.; Walker, A.K.; Chen, F.; Zhang, S.; Chung, A.S.; Nguyen, L.H.; Nassour, I.; et al. Arid1b haploinsufficient mice reveal neuropsychiatric phenotypes and reversible causes of growth impairment. Elife 2017, 6, e25730. [Google Scholar] [CrossRef] [PubMed]
- Markram, K.; Rinaldi, T.; La Mendola, D.; Sandi, C.; Markram, H. Abnormal fear conditioning and amygdala processing in an animal model of Autism. Neuropsychopharmacology 2008, 33, 901–912. [Google Scholar] [CrossRef] [PubMed]
- Horii, T.; Arai, Y.; Yamazaki, M.; Morita, S.; Kimura, M.; Itoh, M.; Abe, Y.; Hatada, I. Validation of microinjection methods for generating knockout mice by CRISPR/Cas-mediated genome engineering. Sci. Rep. 2014, 4, 4513. [Google Scholar] [CrossRef] [PubMed]
- Ran, F.A.; Hsu, P.D.; Lin, C.Y.; Gootenberg, J.S.; Konermann, S.; Trevino, A.E.; Scott, D.A.; Inoue, A.; Matoba, S.; Zhang, Y.; et al. Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity. Cell 2013, 154, 1380–1389. [Google Scholar] [CrossRef] [PubMed]
- Shoji, H.; Takao, K.; Hattori, S.; Miyakawa, T. Contextual and cued fear conditioning test using a video analyzing system in mice. J. Vis. Exp. 2014, 1, 85. [Google Scholar] [CrossRef] [PubMed]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [PubMed]
- Langmead, B.; Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 2012, 9, 357–359. [Google Scholar] [CrossRef] [PubMed]
- Robinson, M.D.; Oshlack, A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome. Biol. 2010, 11, R25. [Google Scholar] [CrossRef] [PubMed]
- Hagihara, H.; Ohira, K.; Takao, K.; Miyakawa, T. Transcriptomic evidence for immaturity of the prefrontal cortex in patients with schizophrenia. Mol. Brain 2014, 7, 41. [Google Scholar] [CrossRef] [PubMed]
- Murano, T.; Koshimizu, H.; Hagihara, H.; Miyakawa, T. Transcriptomic immaturity of the hippocampus and prefrontal cortex in patients with alcoholism. Sci. Rep. 2017, 15, 44531. [Google Scholar] [CrossRef] [PubMed]
- Ginsberg, M.R.; Rubin, R.A.; Falcone, T.; Ting, A.H.; Natowicz, M.R. Brain transcriptional and epigenetic associations with Autism. PLoS ONE 2012, 7, e44736. [Google Scholar] [CrossRef] [PubMed]
- Okaty, B.W.; Miller, M.N.; Sugino, K.; Hempel, C.M.; Nelson, S.B. Transcriptional and electrophysiological maturation of neocortical fast-spiking GABAergic interneurons. J. Neurosci. 2009, 29, 7040–7052. [Google Scholar] [CrossRef] [PubMed]
- Kupershmidt, I.; Su, Q.J.; Grewal, A.; Sundaresh, S.; Halperin, I.; Flynn, J.; Shekar, M.; Wang, H.; Park, J.; Cui, W.; et al. Ontology-based meta-analysis of global collections of high-throughput public data. PLoS ONE 2010, 5, e13066. [Google Scholar] [CrossRef] [PubMed]
| Features | Behavioral Test | Our Study | Celen et al. [32] |
|---|---|---|---|
| Intellectual/cognitive disability | Morris water maze test | Unknown | No |
| Barnes maze test | No | Unknown | |
| T-maze spontaneous alternation test | No | Unknown | |
| Fear conditioning test | Altered | No | |
| Growth retardation | Yes | Yes | |
| Coarse facial features | Unknown | Unknown | |
| Brachydactyly, hypoplastic nail/finger | Unknown | Unknown | |
| Muscle hypotonia | Yes | Yes | |
| Hydrocephalus | Yes (5.5%) | Yes (6.6%) | |
| Abnormal vocalization | Unknown | Yes | |
| Anxiety | Open field test | Not clear | Yes |
| Elevated plus maze test | Yes | Yes | |
| Light/dark transition test | No | Yes | |
| Social behavior deficit | Social interaction test in novel environment | No | Yes |
| Three-chamber social approach test | No | Unknown | |
| Home-cage social interaction test | Yes | Unknown | |
| Perseveration | Barnes maze test | Yes | Unknown |
| Repetitive behavior | Grooming test | No | Yes |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shibutani, M.; Horii, T.; Shoji, H.; Morita, S.; Kimura, M.; Terawaki, N.; Miyakawa, T.; Hatada, I. Arid1b Haploinsufficiency Causes Abnormal Brain Gene Expression and Autism-Related Behaviors in Mice. Int. J. Mol. Sci. 2017, 18, 1872. https://doi.org/10.3390/ijms18091872
Shibutani M, Horii T, Shoji H, Morita S, Kimura M, Terawaki N, Miyakawa T, Hatada I. Arid1b Haploinsufficiency Causes Abnormal Brain Gene Expression and Autism-Related Behaviors in Mice. International Journal of Molecular Sciences. 2017; 18(9):1872. https://doi.org/10.3390/ijms18091872
Chicago/Turabian StyleShibutani, Mihiro, Takuro Horii, Hirotaka Shoji, Sumiyo Morita, Mika Kimura, Naomi Terawaki, Tsuyoshi Miyakawa, and Izuho Hatada. 2017. "Arid1b Haploinsufficiency Causes Abnormal Brain Gene Expression and Autism-Related Behaviors in Mice" International Journal of Molecular Sciences 18, no. 9: 1872. https://doi.org/10.3390/ijms18091872