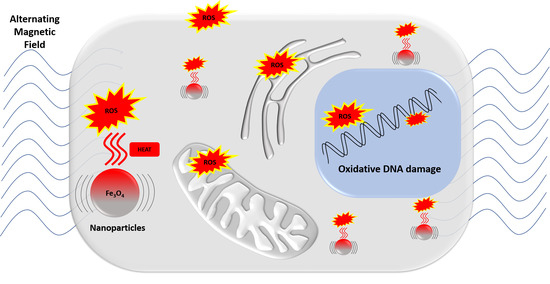
Magnetic Hyperthermia and Oxidative Damage to DNA of Human Hepatocarcinoma Cells
Abstract
:1. Introduction
2. Results
2.1. Fe3O4-Nanoparticles and Exocyclic M1dG Adducts
2.2. Fe3O4-Nanoparticles and 8-oxodG Adducts
2.3. Magnetic Nanoparticles, Alternating Magnetic Field and Exocyclic M1dG Adducts
2.4. Magnetic Nanoparticles, Alternating Magnetic Field and 8-oxodG Adducts
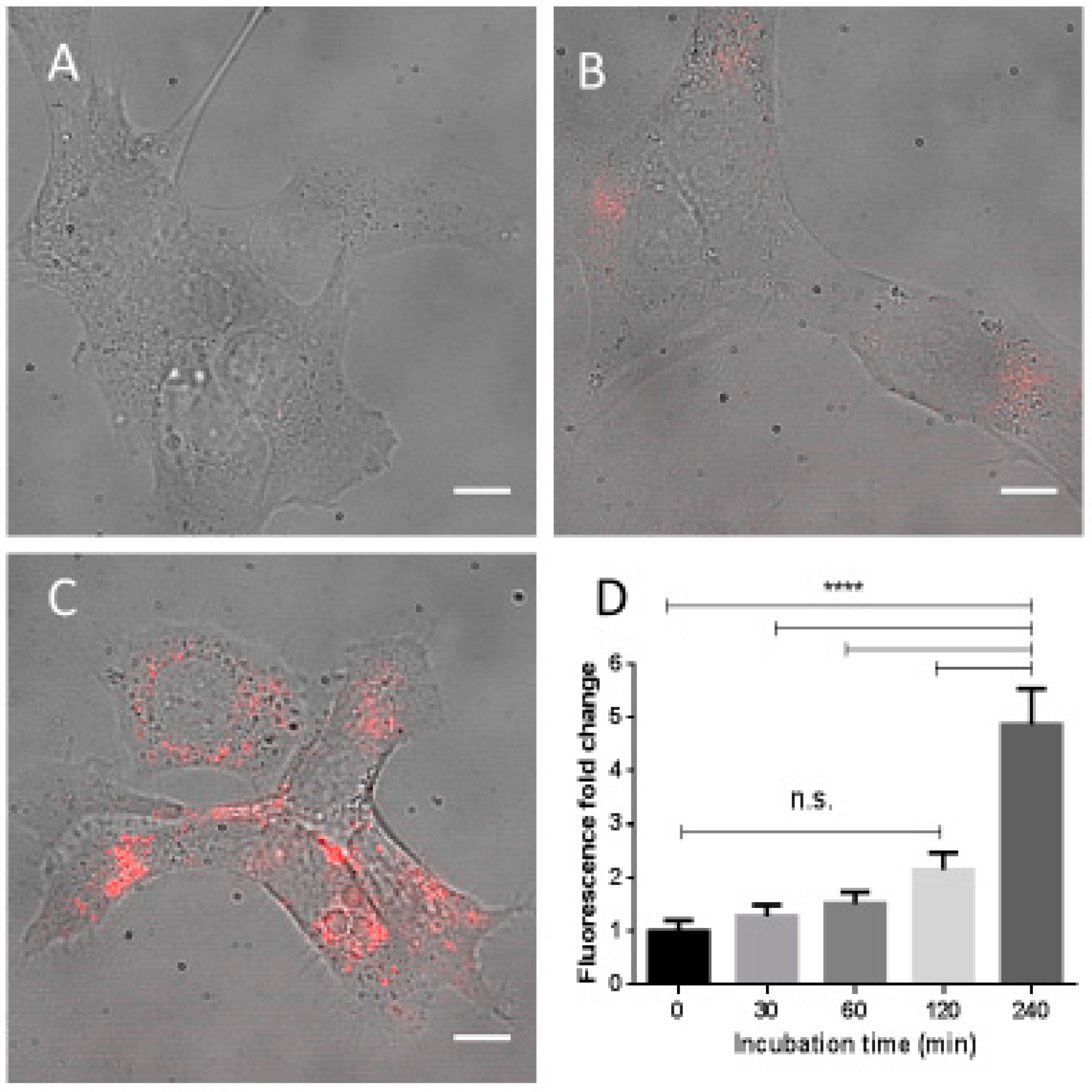
2.5. Fe3O4-Nanoparticle Internalization
3. Discussion
4. Material and Methods
4.1. Fe3O4-Nanoparticles, Cell Culture and Cell Treatments in Presence or Absence of Alternating Magnetic Field
4.2. Preparation of M1dG Reference Adduct Standard
4.3. Mass Spectrometry Analysis
4.4. Reference M1dG Standard by 32P-Postlabeling and Mass Spectrometry
4.5. DNA Isolation and Hydrolysis
4.6. Exocyclic M1dG Adduct Analysis
4.7. 8-oxodG Adduct Analysis
4.8. Fe3O4-Nanoparticle Internalization
4.9. Statistical Analysis
5. Conclusions
Acknowledgment
Author Contributions
Conflicts of Interest
References
- Guo, M.; Sun, Y.; Zhang, X.-D. Enhanced radiation therapy of gold nanoparticles in liver cancer. Appl. Sci. 2017, 7, 232. [Google Scholar] [CrossRef]
- Ariga, K.; Minami, K.; Ebara, M.; Nakanishi, J. What are the emerging concepts and challenges in NANO? Nanoarchitectonics, hand-operating nanotechnology and mechanobiology. Polym. J. 2016, 48, 371–389. [Google Scholar] [CrossRef]
- Chen, G.; Roy, I.; Yang, C.; Prasad, P.N. Nanochemistry and nanomedicine for nanoparticle-based diagnostics and therapy. Chem. Rev. 2016, 116, 2826–2885. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, E.; Kuroda, K. Colloidal Mesoporous Silica Nanoparticles. Bull. Chem. Soc. Jpn. 2016, 89, 501–539. [Google Scholar] [CrossRef]
- Nakanishi, W.; Minami, K.; Shrestha, L.K.; Ji, Q.; Hill, J.P.; Ariga, K. Bioactive nanocarbon assemblies: Nanoarchitectonics and applications. Nano Today 2014, 9, 378–394. [Google Scholar] [CrossRef]
- Wicki, A.; Witzigmann, D.; Balasubramanian, V.; Huwyler, J. Nanomedicine in cancer therapy: Challenges, opportunities, and clinical applications. J. Control. Release 2015, 200, 138–157. [Google Scholar] [CrossRef] [PubMed]
- Karimi, M.; Ghasemi, A.; Sahandi, Z.P.; Rahighi, R.; Moosavi, B.S.M.; Mirshekari, H.; Amiri, M.; Shafaei, P.Z.; Aslani, A.B.M.; Ghosh, D.; et al. Smart micro/nanoparticles in stimulus-responsive drug/gene delivery systems. Chem. Soc. Rev. 2016, 45, 1457–1501. [Google Scholar] [CrossRef] [PubMed]
- Koedrith, P.; Boonprasert, R.; Kwon, J.Y.; Kim, I.; Seo, Y.R. Recent toxicological investigations of metal or metal oxide nanoparticles in mammalian models in vitro and in vivo: DNA damaging potential, and relevant physicochemical characteristics. Mol. Toxicol. 2014, 10, 107–126. [Google Scholar] [CrossRef]
- Watanabe, M.; Yoneda, M.; Morohashi, A.; Hori, Y.; Okamoto, D.; Sato, A.; Kurioka, D.; Nittami, T.; Hirokawa, Y.; Shiraishi, T.; et al. Effects of Fe3O4 magnetic nanoparticles on A549 cells. Int. J. Mol. Sci. 2013, 14, 15546–15560. [Google Scholar] [CrossRef] [PubMed]
- Wydra, R.J.; Oliver, C.E.; Anderson, K.W.; Dziubla, T.D.; Hilt, J.Z. Accelerated generation of free radicals by iron oxide nanoparticles in the presence of an alternating magnetic field. RSC Adv. 2015, 5, 18888–18893. [Google Scholar] [CrossRef] [PubMed]
- Dissanayake, N.M.; Current, K.M.; Obare, S.O. Mutagenic effects of iron oxide nanoparticles on biological cells. Int. J. Mol. Sci. 2015, 16, 23482–23516. [Google Scholar] [CrossRef] [PubMed]
- Marnett, L.J. Oxyradicals and DNA damage. Carcinogenesis 2000, 21, 361–370. [Google Scholar] [CrossRef] [PubMed]
- Singh, N.; Jenkins, G.J.; Asadi, R.; Doak, S.H. Potential toxicity of superparamagnetic iron oxide nanoparticles (SPION). Nano Rev. 2010, 1, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Jeong, Y.C.; Swenberg, J.A. Formation of M1G-dR from endogenous and exogenous ROS-inducing chemicals. Free Radic. Biol. Med. 2005, 39, 1021–1029. [Google Scholar] [CrossRef] [PubMed]
- Zhou, X.; Taghizadeh, K.; Dedon, P.C. Chemical and biological evidence for base propenals as the major source of the endogenous M1dG adduct in cellular DNA. J. Biol. Chem. 2005, 280, 25377–25382. [Google Scholar] [CrossRef] [PubMed]
- Cooke, M.S.; Evans, M.D.; Dizdaroglu, M.; Lunec, J. Oxidative DNA damage: Mechanisms, mutation, and disease. FASEB J. 2003, 17, 1195–1214. [Google Scholar] [CrossRef] [PubMed]
- Brancato, B.; Munnia, A.; Cellai, F.; Ceni, E.; Mello, T.; Bianchi, S.; Catarzi, S.; Risso, G.G.; Galli, A.; Peluso, M.E. 8-Oxo-7,8-dihydro-2′-deoxyguanosine and other lesions along the coding strand of the exon 5 of the tumour suppressor gene P53 in a breast cancer case-control study. DNA Res. 2016, 23, 395–402. [Google Scholar] [CrossRef] [PubMed]
- Bargonetti, J.; Manfredi, J.J. Multiple roles of the tumor suppressor p53. Curr. Opin. Oncol. 2002, 14, 86–91. [Google Scholar] [CrossRef]
- Loft, S.; Moller, P. Oxidative DNA damage and human cancer: Need for cohort studies. Antioxid. Redox Signal. 2006, 8, 1021–1031. [Google Scholar] [CrossRef] [PubMed]
- Peluso, M.; Munnia, A.; Risso, G.G.; Catarzi, S.; Piro, S.; Ceppi, M.; Giese, R.W.; Brancato, B. Breast fine-needle aspiration malondialdehyde deoxyguanosine adduct in breast cancer. Free Radic. Res. 2011, 45, 477–482. [Google Scholar] [CrossRef] [PubMed]
- Munnia, A.; Amasio, M.E.; Peluso, M. Exocyclic malondialdehyde and aromatic DNA adducts in larynx tissues. Free Radic. Biol. Med. 2004, 37, 850–858. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Dhingra, K.; Hittelman, W.N.; Liehr, J.G.; de Andrade, M.; Li, D. Lipid peroxidation-induced putative malondialdehyde—DNA adducts in human breast tissues. Cancer Epidemiol. Biomark. Prev. 1996, 5, 705–710. [Google Scholar]
- Sorensen, M.; Autrup, H.; Moller, P.; Hertel, O.; Jensen, S.S.; Vinzents, P.; Knudsen, L.E.; Loft, S. Linking exposure to environmental pollutants with biological effects. Mutat. Res. 2003, 544, 255–271. [Google Scholar] [CrossRef] [PubMed]
- Vanhees, K.; van Schooten, F.J.; van Doorn-Khosrovani, S.B.; van Helden, S.; Munnia, A.; Peluso, M.; Briede, J.J.; Haenen, G.R.; Godschalk, R.W. Intrauterine exposure to flavonoids modifies antioxidant status at adulthood and decreases oxidative stress-induced DNA damage. Free Radic. Biol. Med. 2013, 57, 154–161. [Google Scholar] [CrossRef] [PubMed]
- Izzotti, A.; Balansky, R.M.; Dagostini, F.; Bennicelli, C.; Myers, S.R.; Grubbs, C.J.; Lubet, R.A.; Kelloff, G.J.; de Flora, S. Modulation of biomarkers by chemopreventive agents in smoke-exposed rats. Cancer Res. 2001, 61, 2472–2479. [Google Scholar] [PubMed]
- Izzotti, A.; Cartiglia, C.; De Flora, S.; Sacca, S. Methodology for evaluating oxidative DNA damage and metabolic genotypes in human trabecular meshwork. Toxicol. Mech. Methods 2003, 13, 161–168. [Google Scholar] [CrossRef] [PubMed]
- Beik, J.; Abed, Z.; Ghoreishi, F.S.; Hosseini-Nami, S.; Mehrzadi, S.; Shakeri-Zadeh, A.; Kamrava, S.K. Nanotechnology in hyperthermia cancer therapy: From fundamental principles to advanced applications. J. Control. Release 2016, 235, 205–221. [Google Scholar] [CrossRef] [PubMed]
- Link, E.M.; Riley, P.A. Role of hydrogen peroxide in the cytotoxicity of the xanthine/xanthine oxidase system. Biochem. J. 1988, 249, 391–399. [Google Scholar] [CrossRef] [PubMed]
- European Standards Committee on Oxidative DNA Damage (ESCODD). Comparative analysis of baseline 8-oxo-7,8-dihydroguanine in mammalian cell DNA, by different methods in different laboratories: An approach to consensus. Carcinogenesis 2003, 23, 2129–2133. [Google Scholar]
- Reimer, P.; Balzer, T. Ferucarbotran (Resovist): A new clinically approved RES-specific contrast agent for contrast-enhanced MRI of the liver: Properties, clinical development, and applications. Eur. Radiol. 2003, 13, 1266–1276. [Google Scholar] [PubMed]
- Kim, J.S.; Yoon, T.J.; Yu, K.N.; Kim, B.G.; Park, S.J.; Kim, H.W.; Lee, K.H.; Park, S.B.; Lee, J.K.; Cho, M.H. Toxicity and tissue distribution of magnetic nanoparticles in mice. Toxicol. Sci. 2006, 89, 338–347. [Google Scholar] [CrossRef] [PubMed]
- Malvindi, M.A.; De Matteis, V.; Galeone, A.; Brunetti, V.; Anyfantis, G.C.; Athanassiou, A.; Cingolani, R.; Pompa, P.P. Toxicity assessment of silica coated iron oxide nanoparticles and biocompatibility improvement by surface engineering. PLoS ONE 2014, 9, e85835. [Google Scholar] [CrossRef] [PubMed]
- Valdiglesias, V.; Fernández-Bertólez, N.; Kiliç, G.; Costa, C.; Costa, S.; Fraga, S.; Bessa, M.J.; Pásaro, E.; Teixeira, J.P.; Laffon, B. Are iron oxide nanoparticles safe? Current knowledge and future perspectives. J. Trace Elem. Med. Biol. 2016, 38, 53–63. [Google Scholar] [CrossRef] [PubMed]
- Lin, X.L.; Zhao, S.H.; Zhang, L.; Hu, G.Q.; Sun, Z.W.; Yang, W.S. Dose-dependent cytotoxicity and oxidative stress induced by “naked” Fe3O4 Nanoparticles in human hepatocyte. Chem. Res. Chin. Univ. 2012, 28, 114–118. [Google Scholar]
- Sadeghi, L.; Tanwir, F.; Yousefi Babadi, V. In vitro toxicity of iron oxide nanoparticle: Oxidative damages on Hep G2 cells. Exp. Toxicol. Pathol. 2015, 67, 197–203. [Google Scholar] [CrossRef] [PubMed]
- Ma, P.; Luo, Q.; Chen, J.; Gan, Y.; Du, J.; Ding, S.; Xi, Z.; Yang, X. Intraperitoneal injection of magnetic Fe3O4-nanoparticle induces hepatic and renal tissue injury via oxidative stress in mice. Int. J. Nanomed. 2012, 7, 4809–4818. [Google Scholar]
- Shaw, J.; Raja, S.O.; Dasgupta, A.K. Modulation of cytotoxic and genotoxic effects of nanoparticles in cancer cells by external magnetic field. Cancer Nanotechnol. 2014, 5, 2. [Google Scholar] [CrossRef] [PubMed]
- Van Helden, Y.G.; Keijer, J.; Heil, S.G.; Pico, C.; Palou, A.; Oliver, P.; Munnia, A.; Briede, J.J.; Peluso, M.; Franssen-van Hal, N.L.; et al. β-carotene affects oxidative stress-related DNA damage in lung epithelial cells and in ferret lung. Carcinogenesis 2009, 30, 2070–2076. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.H.; Kim, J.; Choi, W. Effect of magnetic field on the zero valent iron induced oxidation reaction. J. Hazard. Mater. 2011, 192, 928–931. [Google Scholar] [CrossRef] [PubMed]
- Kadiiska, M.B.; Burkitt, M.J.; Xiang, Q.H.; Mason, R.P. Iron supplementation generates hydroxyl radical in vivo. An ESR spin-trapping investigation. J. Clin. Investig. 1995, 96, 1653–1657. [Google Scholar] [CrossRef] [PubMed]
- Mello, T.; Zanieri, F.; Ceni, E.; Galli, A. Oxidative Stress in the Healthy and Wounded Hepatocyte: A Cellular Organelles Perspective. Oxid. Med. Cell. Longev. 2016, 2016, 832741. [Google Scholar] [CrossRef] [PubMed]
- Orlando, A.; Colombo, M.; Prosperi, D.; Gregori, M.; Panariti, A.; Rivolta, I.; Masserini, M.; Cazzaniga, E. Iron oxide nanoparticles surface coating and cell uptake affect biocompatibility and inflammatory responses of endothelial cells and macrophages. J. Nanopart. Res. 2015, 17, 351. [Google Scholar] [CrossRef]
- Gungor, N.; Knaapen, A.M.; Munnia, A.; Peluso, M.; Haenen, G.R.; Chiu, R.K.; Godschalk, R.W.; van Schooten, F.J. Genotoxic effects of neutrophils and hypochlorous acid. Mutagenesis 2010, 25, 149–154. [Google Scholar] [CrossRef] [PubMed]
- Gungor, N.; Pennings, J.L.; Knaapen, A.M.; Chiu, R.K.; Peluso, M.; Godschalk, R.W.; van Schooten, F.J. Transcriptional profiling of the acute pulmonary inflammatory response induced by LPS: Role of neutrophils. Respir. Res. 2010, 11, 24. [Google Scholar] [CrossRef] [PubMed]
- Parka, E.J.; Choia, D.H.; Kimb, H.; Leec, E.W.; Songc, J.; Chod, M.H.; Kima, J.H.; Kim, S.W. Magnetic iron oxide nanoparticles induce autophagy preceding apoptosis through mitochondrial damage and ER stress in RAW264.7 cells. Toxicol. In Vitro 2014, 1402–1412. [Google Scholar] [CrossRef] [PubMed]
- Filomeni, G.; De Zio, D.; Cecconi, F. Oxidative stress and autophagy: The clash between damage and metabolic needs. Cell Death Differ. 2015, 22, 377–388. [Google Scholar] [CrossRef] [PubMed]
- Khan, M.; Mohammad, A.; Patil, G.; Naqvi, S.A.; Chauhan, L.K.; Ahmad, I. Induction of ROS, mitochondrial damage and autophagy in lung epithelial cancer cells by iron oxide nanoparticles. Biomaterials 2012, 33, 1477–1488. [Google Scholar] [CrossRef] [PubMed]
- Alarifi, S.; Ali, D.; Alkahtani, S.; Alhader, M.S. Iron oxide nanoparticles induce oxidative stress, DNA damage, and caspase activation in the human breast cancer cell line. Biol. Trace Elem. Res. 2014, 159, 416–424. [Google Scholar] [CrossRef] [PubMed]
- Bono, R.; Romanazzi, V.; Munnia, A.; Piro, S.; Allione, A.; Ricceri, F.; Guarrera, S.; Pignata, C.; Matullo, G.; Wang, P.; et al. Malondialdehyde-deoxyguanosine adduct formation in workers of pathology wards: The role of air formaldehyde exposure. Chem. Res. Toxicol. 2010, 23, 1342–1348. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Gao, J.; Li, G.; Shimelis, O.; Giese, R.W. Nontargeted analysis of DNA adducts by mass-tag MS: Reaction of p-benzoquinone with DNA. Chem. Res. Toxicol. 2012, 25, 2737–2743. [Google Scholar] [CrossRef] [PubMed]
- Peluso, M.; Castegnaro, M.; Malaveille, C.; Talaska, G.; Vineis, P.; Kadlubar, F.; Bartsch, H. 32P-postlabelling analysis of DNA adducted with urinary mutagens from smokers of black tobacco. Carcinogenesis 1990, 11, 1307–1311. [Google Scholar] [CrossRef] [PubMed]
- Peluso, M.; Srivatanakul, P.; Munnia, A.; Jedpiyawongse, A.; Ceppi, M.; Sangrajrang, S.; Piro, S.; Boffetta, P. Malondialdehyde-deoxyguanosine adducts among workers of a Thai industrial estate and nearby residents. Environ. Health Perspect. 2010, 118, 55–59. [Google Scholar] [CrossRef] [PubMed]
- Munnia, A.; Saletta, F.; Allione, A.; Piro, S.; Confortini, M.; Matullo, G.; Peluso, M. 32P-Post-labelling method improvements for aromatic compound-related molecular epidemiology studies. Mutagenesis 2007, 22, 381–385. [Google Scholar] [CrossRef] [PubMed]
- Valavanidis, A.; Vlachogianni, T.; Fiotakis, C. 8-hydroxy-2′-deoxyguanosine (8-OHdG): A critical biomarker of oxidative stress and carcinogenesis. J. Environ. Sci. Health. Part C Environ. Carcinog. Ecotoxicol. Rev. 2009, 27, 120–139. [Google Scholar] [CrossRef] [PubMed]
- Izzotti, A.; De Flora, S.; Cartiglia, C.; Are, B.M.; Longobardi, M.; Camoirano, A.; Mura, I.; Dore, M.P.; Scanu, A.M.; Rocca, P.C.; et al. Interplay between Helicobacter pylori and host gene polymorphisms in inducing oxidative DNA damage in the gastric mucosa. Carcinogenesis 2007, 28, 892–898. [Google Scholar] [CrossRef] [PubMed]
- Balansky, R.; Izzotti, A.; D’Agostini, F.; Longobardi, M.; Micale, R.T.; La Maestra, S.; Camoirano, A.; Ganchev, G.; Iltcheva, M.; Steele, V.E.; et al. Assay of lapatinib in murine models of cigarette smoke carcinogenesis. Carcinogenesis 2014, 35, 2300–2307. [Google Scholar] [CrossRef] [PubMed]
- Micale, R.T.; La Maestra, S.; Di Pietro, A.; Visalli, G.; Baluce, B.; Balansky, R.; Steele, V.E.; de Flora, S. Oxidative stress in the lung of mice exposed to cigarette smoke either early in life or in adulthood. Arch. Toxicol. 2013, 87, 915–918. [Google Scholar] [CrossRef] [PubMed]
- Devanaboyina, U.; Gupta, R.C. Sensitive detection of 8-hydroxy-2′deoxyguanosine in DNA by 32P-postlabeling assay and the basal levels in rat tissues. Carcinogenesis 1996, 17, 917–924. [Google Scholar] [CrossRef] [PubMed]
- Schindelin, J.; Arganda-Carreras, I.; Frise, E.; Kaynig, V.; Longair, M.; Pietzsch, T.; Preibisch, S.; Rueden, C.; Saalfeld, S.; Schmid, B.; et al. Fiji: An open-source platform for biological-image analysis. Nat. Methods 2012, 9, 676–682. [Google Scholar] [CrossRef] [PubMed]
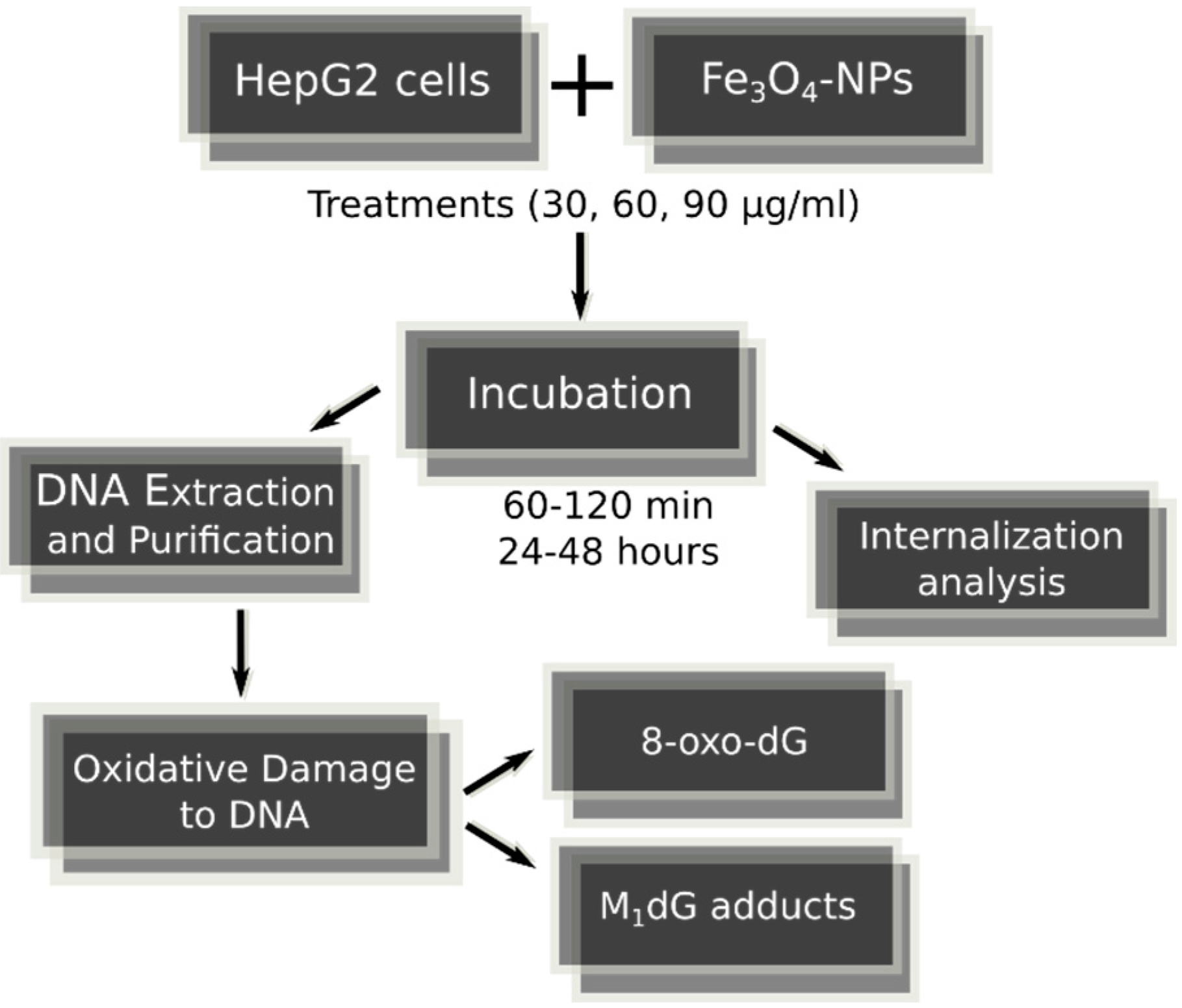
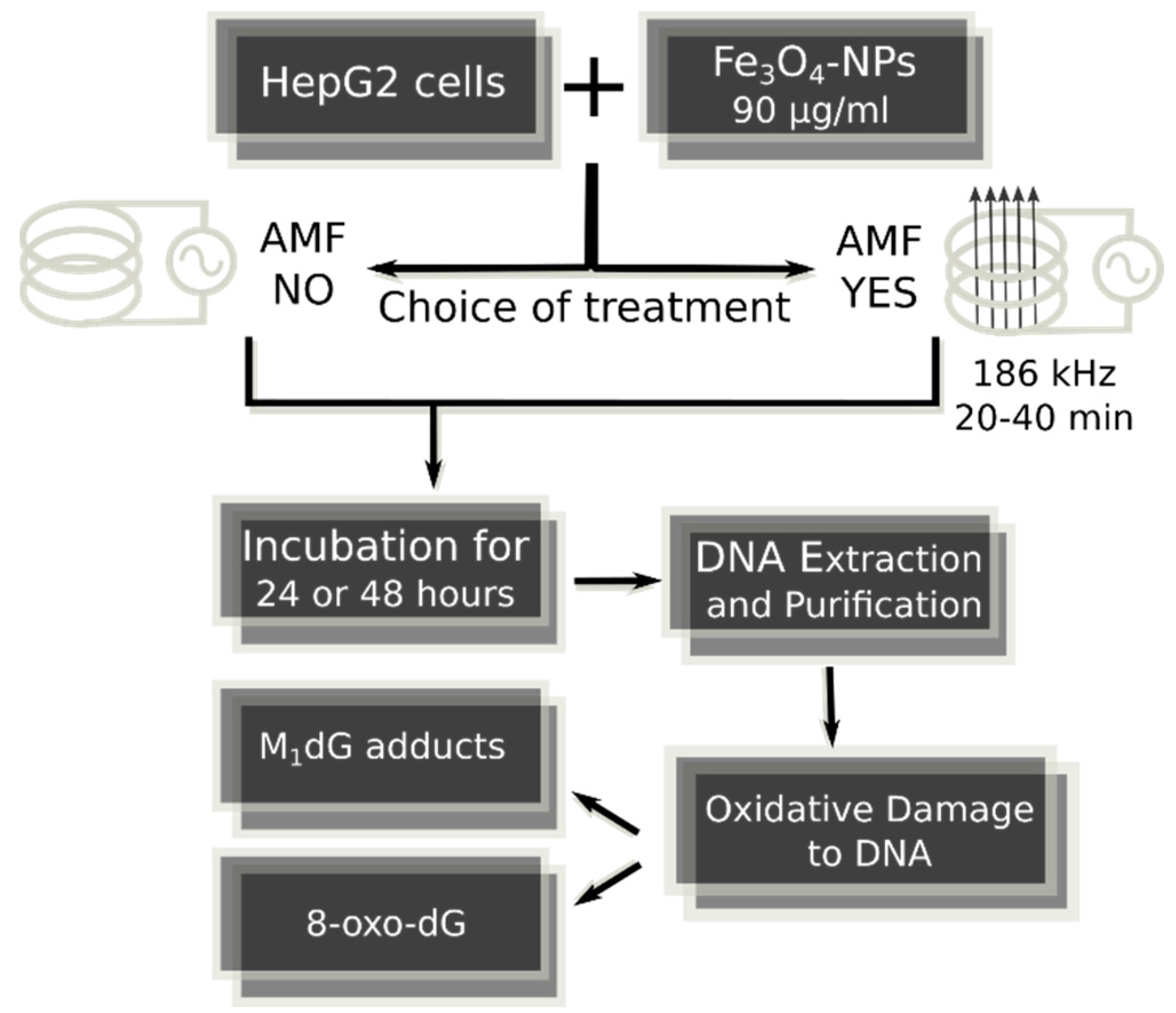
| Exocyclic M1dG and 8-oxodG Lesions | |||||
|---|---|---|---|---|---|
| N | M1dG ± SE | p-Value | 8-oxodG ± SE | p-Value | |
| Adduct background | |||||
| Control cells | |||||
| Incubation times | |||||
| 24 h a | 10 | 0.4 ± 0.1 | 3.2 ± 0.2 | ||
| 48 h a | 10 | 0.4 ± 0.1 | 3.1 ± 0.2 | ||
| Adduct levels of magnetic nanoparticle treated cells | |||||
| Treated cells | |||||
| 30 µg/mL | |||||
| Incubation times | |||||
| 24 h | 10 | 0.5 ± 0.1 | 0.057 b | 11.2 ± 0.8 | <0.05 b |
| 48 h | 10 | 0.6 ± 0.1 | <0.05 b | 14.7 ± 0.4 | <0.05 b |
| 60 µg/mL | |||||
| Incubation times | |||||
| 24 h | 10 | 0.8 ± 0.1 | <0.05 b | 18.4 ± 2.8 | <0.05 b |
| 48 h | 10 | 0.9 ± 0.1 | <0.05 b | 19.4 ± 1.4 | <0.05 b |
| 90 µg/mL | |||||
| Incubation times | |||||
| 24 h | 10 | 0.7 ± 0.1 | <0.05 b | 17.1 ± 1.7 | <0.05 b |
| 48 h | 10 | 0.8 ± 0.1 | <0.05 b | 18.0 ± 1.8 | <0.05 b |
| Adduct levels of positive internal control | |||||
| Free radical-generating system | |||||
| 0.2 mM xanthine/1.0 mU xanthine oxidase | |||||
| Incubation times | |||||
| 24 h | 10 | 1.9 ± 0.1 | <0.05 b | 4.6 ± 0.2 | <0.05 b |
| 48 h | 10 | 2.2 ± 0.1 | <0.05 b | 5.4 ± 0.2 | <0.05 b |
| 0.2 mM xanthine/5.0 mU xanthine oxidase | |||||
| Incubation times | |||||
| 24 h | 10 | 2.9 ± 0.6 | <0.05 b | 8.3 ± 0.2 | <0.05 b |
| 48 h | 10 | 2.7 ± 0.4 | <0.05 b | 9.3 ± 0.4 | <0.05 b |
| Exocyclic M1dG and 8-oxodG Adducts | |||||
|---|---|---|---|---|---|
| N | M1dG ± SE | p-Value | 8-oxodG ± SE | p-Value | |
| Adduct background in presence or absence of AMF exposures | |||||
| Control cells | |||||
| Nonexposed to AMF | |||||
| Incubation times | |||||
| 24 h a | 10 | 0.4 ± 0.1 | 3.2 ± 0.2 | ||
| 48 a | 10 | 0.4 ± 0.1 | 3.1 ± 0.2 | ||
| Exposed to AMF (20 min) | |||||
| Incubation times | |||||
| 24 h a | 10 | 0.5 ± 0.1 | 3.2 ± 0.2 | ||
| 48 a | 10 | 0.5 ± 0.1 | 3.3 ± 0.3 | ||
| Exposed to AMF (40 min) | |||||
| Incubation times | |||||
| 24 h a | 10 | 0.5 ± 0.1 | 3.4 ± 0.2 | ||
| 48 a | 10 | 0.5 ± 0.1 | 3.9 ± 0.3 | ||
| Adduct levels of treated cells in presence or absence of AMF exposures | |||||
| NP treated cells | |||||
| Nonexposed to AMF | |||||
| Incubation times | |||||
| 24 h a | 10 | 0.7 ± 0.1 | <0.05 b | 16.7 ± 1.7 | <0.05 b |
| 48 a | 10 | 0.8 ± 0.1 | <0.05 b | 21.2 ± 1.9 | <0.05 b |
| Exposed to AMF (20 min) | |||||
| Incubation times | |||||
| 24 h a | 10 | 4.6 ± 0.5 | <0.05 b | 40.3 ± 3.3 | <0.05 b |
| 48 a | 10 | 6.9 ± 0.6 | <0.05 b | 69.3 ± 1.4 | <0.05 b |
| Exposed to AMF (40 min) | |||||
| Incubation times | |||||
| 24 h a | 10 | 6.4 ± 1.4 | <0.05 b | 36.5 ± 4.0 | <0.05 b |
| 48 a | 10 | 15.9 ± 6.7 | <0.05 b | 79.3 ± 1.1 | <0.05 b |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cellai, F.; Munnia, A.; Viti, J.; Doumett, S.; Ravagli, C.; Ceni, E.; Mello, T.; Polvani, S.; Giese, R.W.; Baldi, G.; et al. Magnetic Hyperthermia and Oxidative Damage to DNA of Human Hepatocarcinoma Cells. Int. J. Mol. Sci. 2017, 18, 939. https://doi.org/10.3390/ijms18050939
Cellai F, Munnia A, Viti J, Doumett S, Ravagli C, Ceni E, Mello T, Polvani S, Giese RW, Baldi G, et al. Magnetic Hyperthermia and Oxidative Damage to DNA of Human Hepatocarcinoma Cells. International Journal of Molecular Sciences. 2017; 18(5):939. https://doi.org/10.3390/ijms18050939
Chicago/Turabian StyleCellai, Filippo, Armelle Munnia, Jessica Viti, Saer Doumett, Costanza Ravagli, Elisabetta Ceni, Tommaso Mello, Simone Polvani, Roger W. Giese, Giovanni Baldi, and et al. 2017. "Magnetic Hyperthermia and Oxidative Damage to DNA of Human Hepatocarcinoma Cells" International Journal of Molecular Sciences 18, no. 5: 939. https://doi.org/10.3390/ijms18050939
APA StyleCellai, F., Munnia, A., Viti, J., Doumett, S., Ravagli, C., Ceni, E., Mello, T., Polvani, S., Giese, R. W., Baldi, G., Galli, A., & Peluso, M. E. M. (2017). Magnetic Hyperthermia and Oxidative Damage to DNA of Human Hepatocarcinoma Cells. International Journal of Molecular Sciences, 18(5), 939. https://doi.org/10.3390/ijms18050939