High-Density Linkage Map Construction and Mapping of Salt-Tolerant QTLs at Seedling Stage in Upland Cotton Using Genotyping by Sequencing (GBS)
Abstract
:1. Introduction
2. Results
2.1. Quality Control and Mapping to Reference Genome
2.2. SNP Detection and Annotation
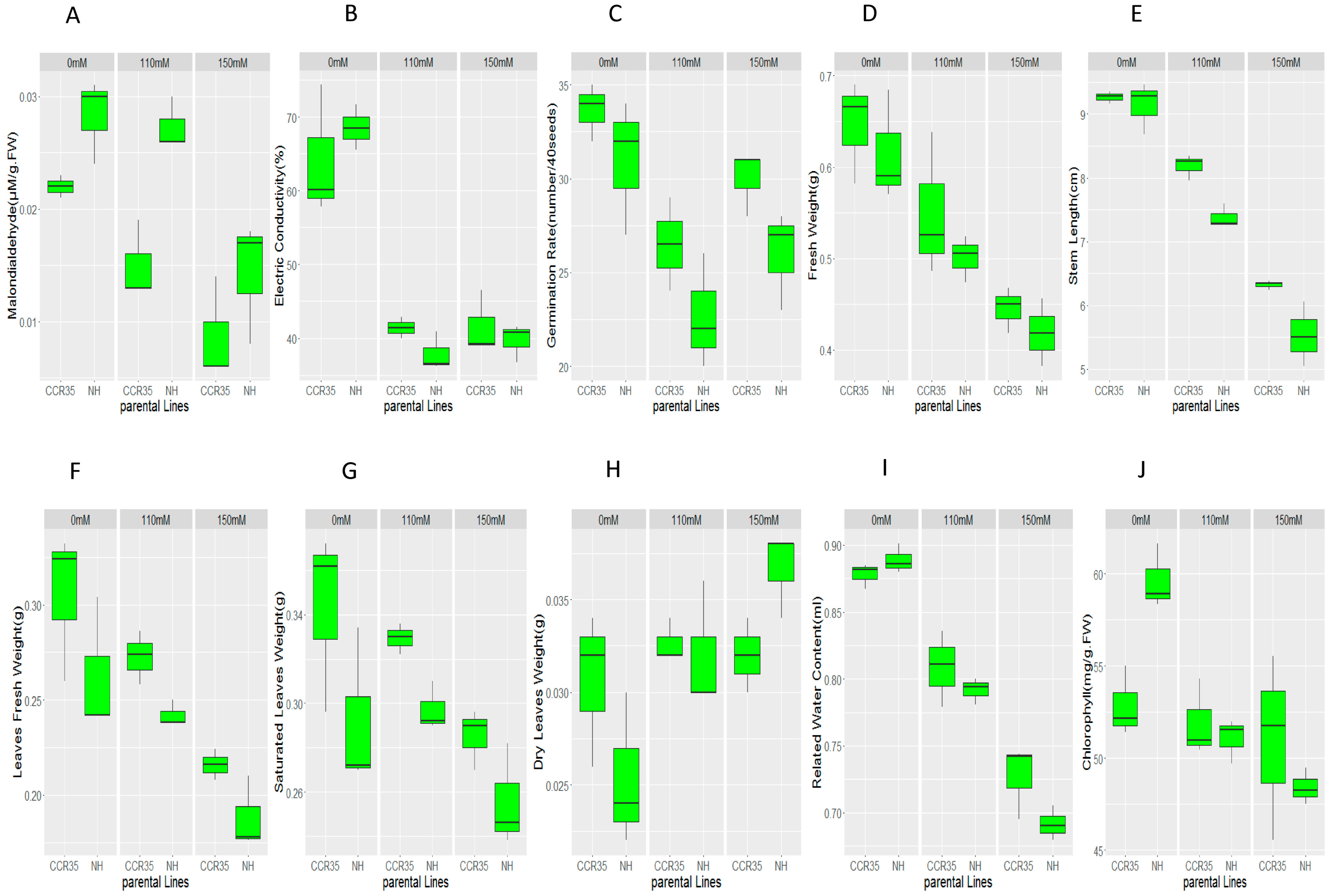
2.3. Phenotypic Variation between Parents
2.4. ANOVA and Heritability Analysis of Salt Tolerance Traits for the Two Parents and the Progeny
2.5. Correlation Analysis
2.5.1. Salt Treatment (110 mM)
2.5.2. Salt Treatment (150 mM)
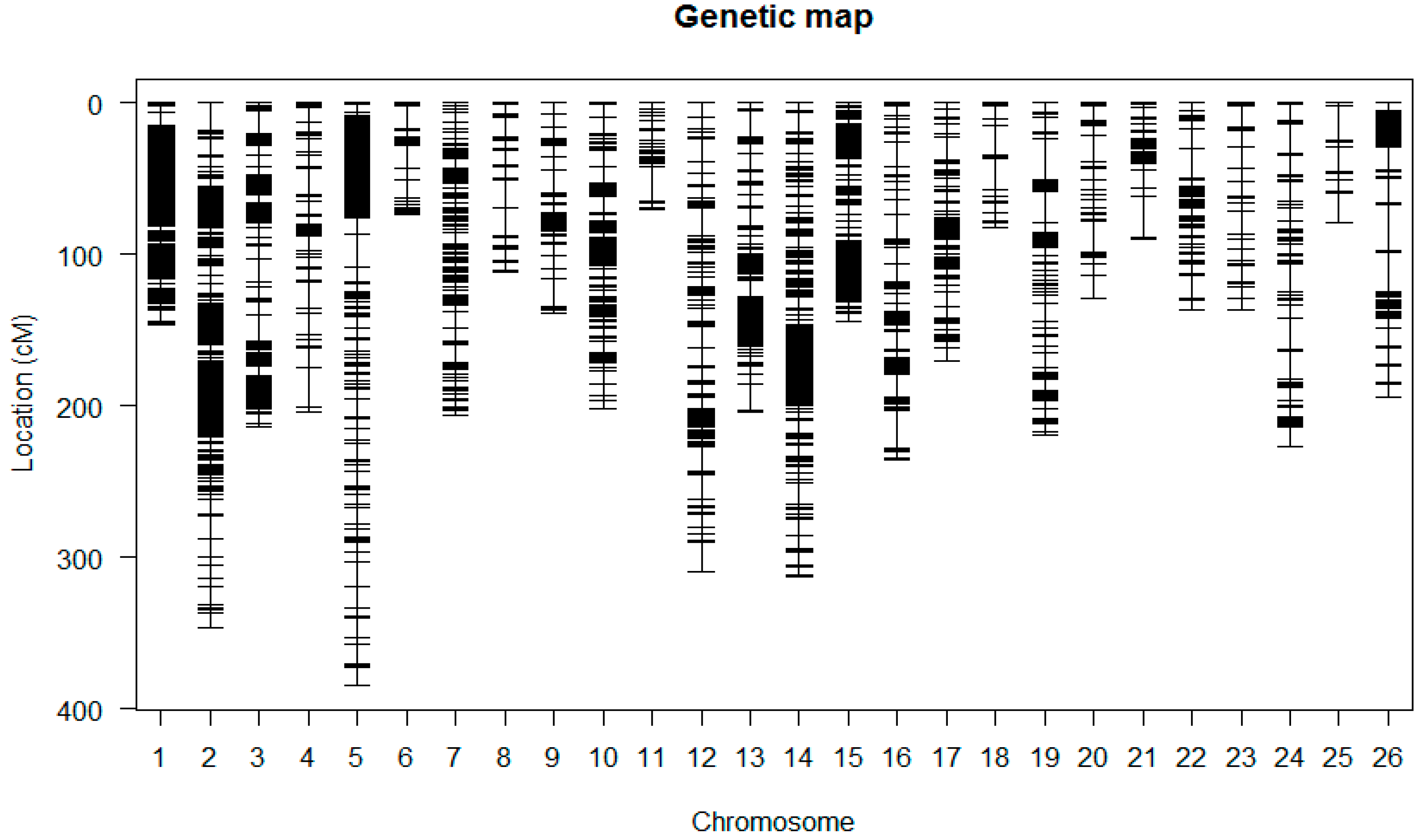
2.6. Construction of the Linkage Maps
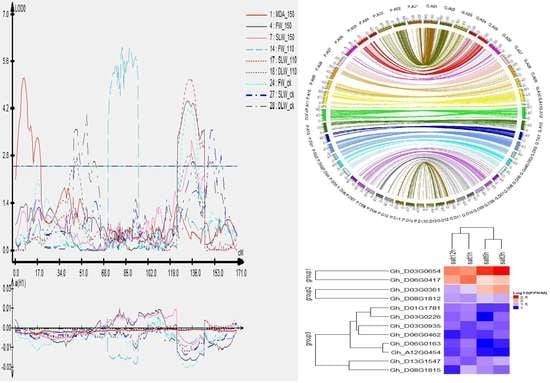
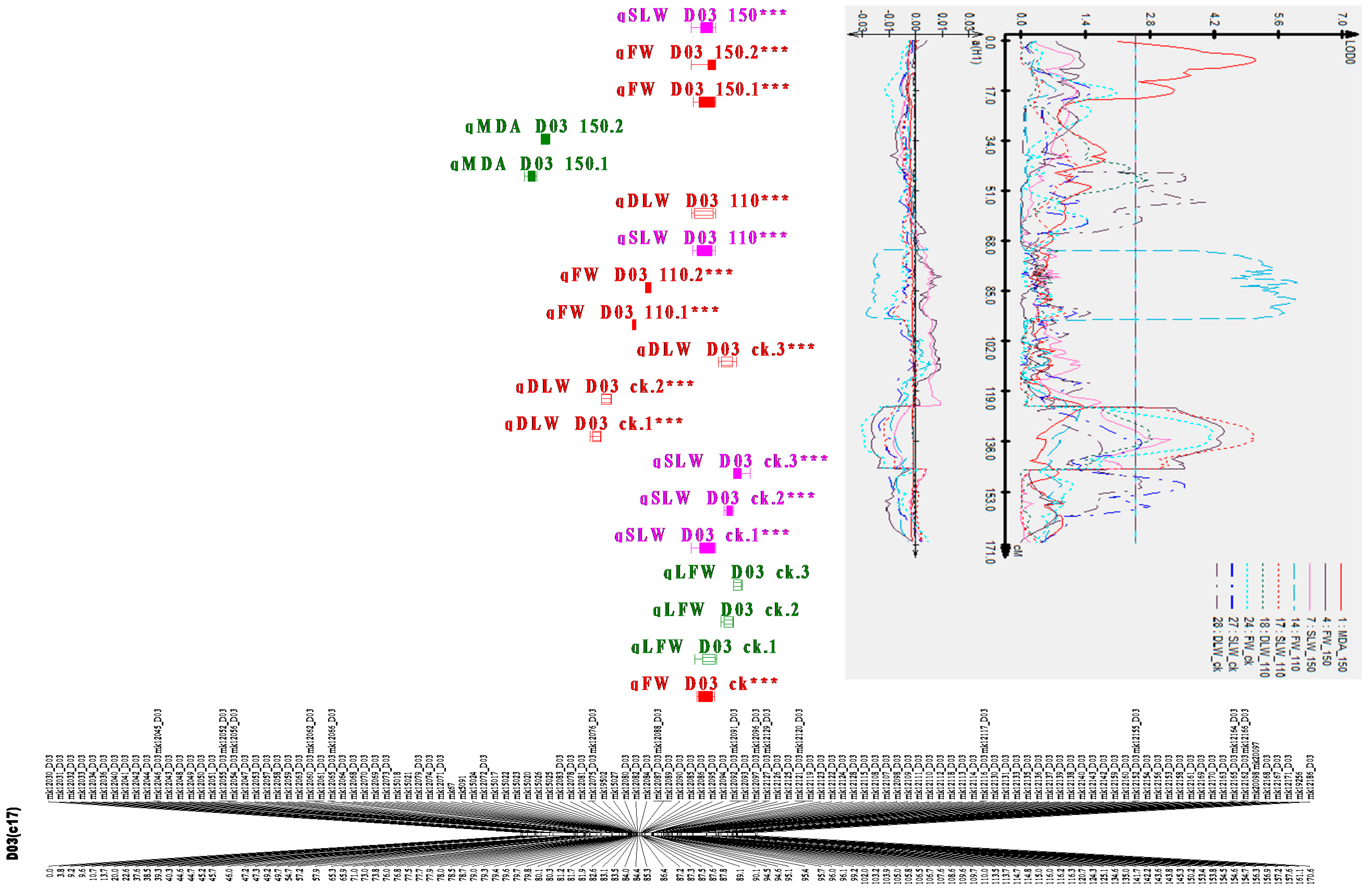
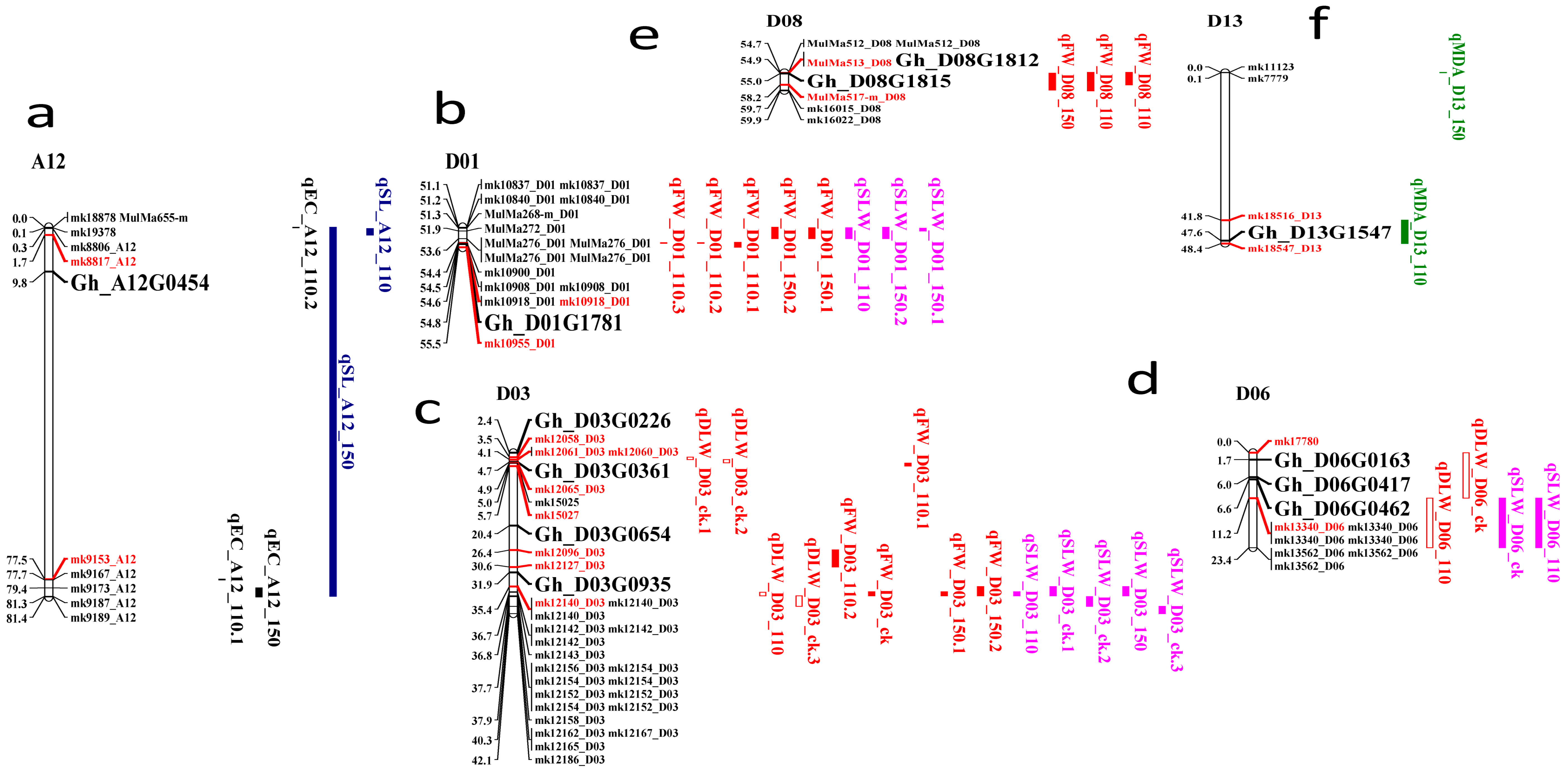
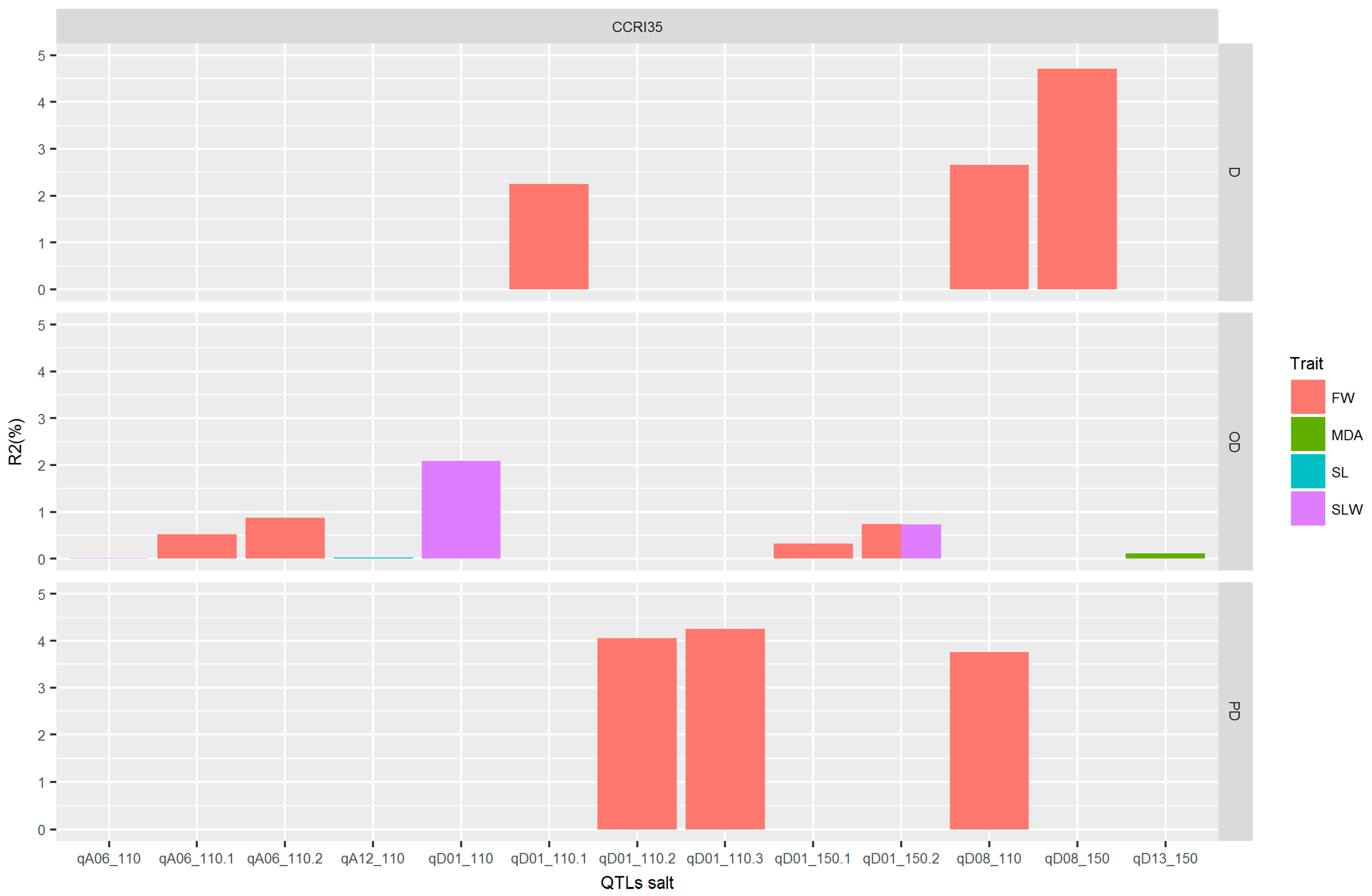
2.7. Clustered QTLs Region
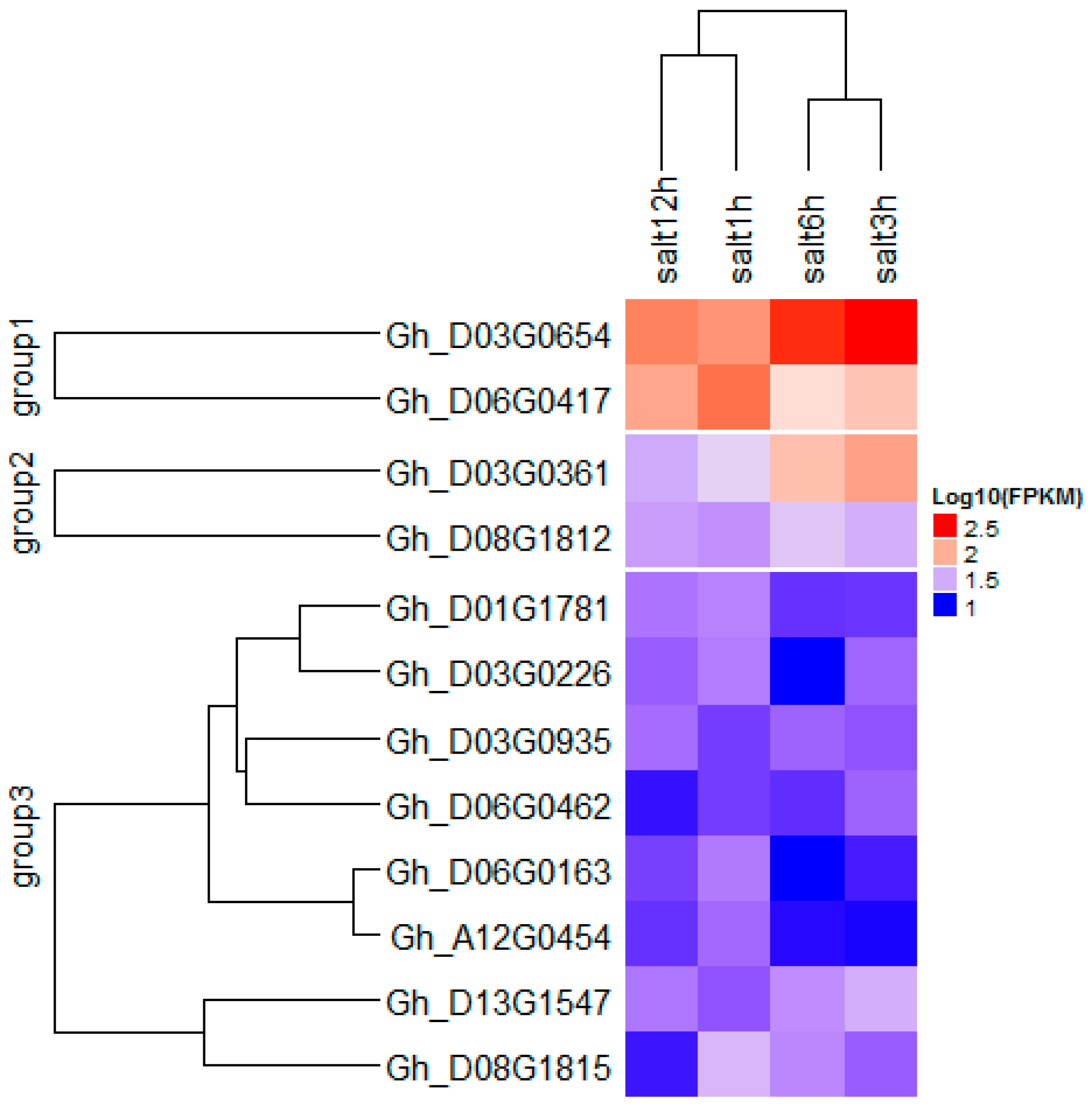
2.8. Identification of Candidate Genes
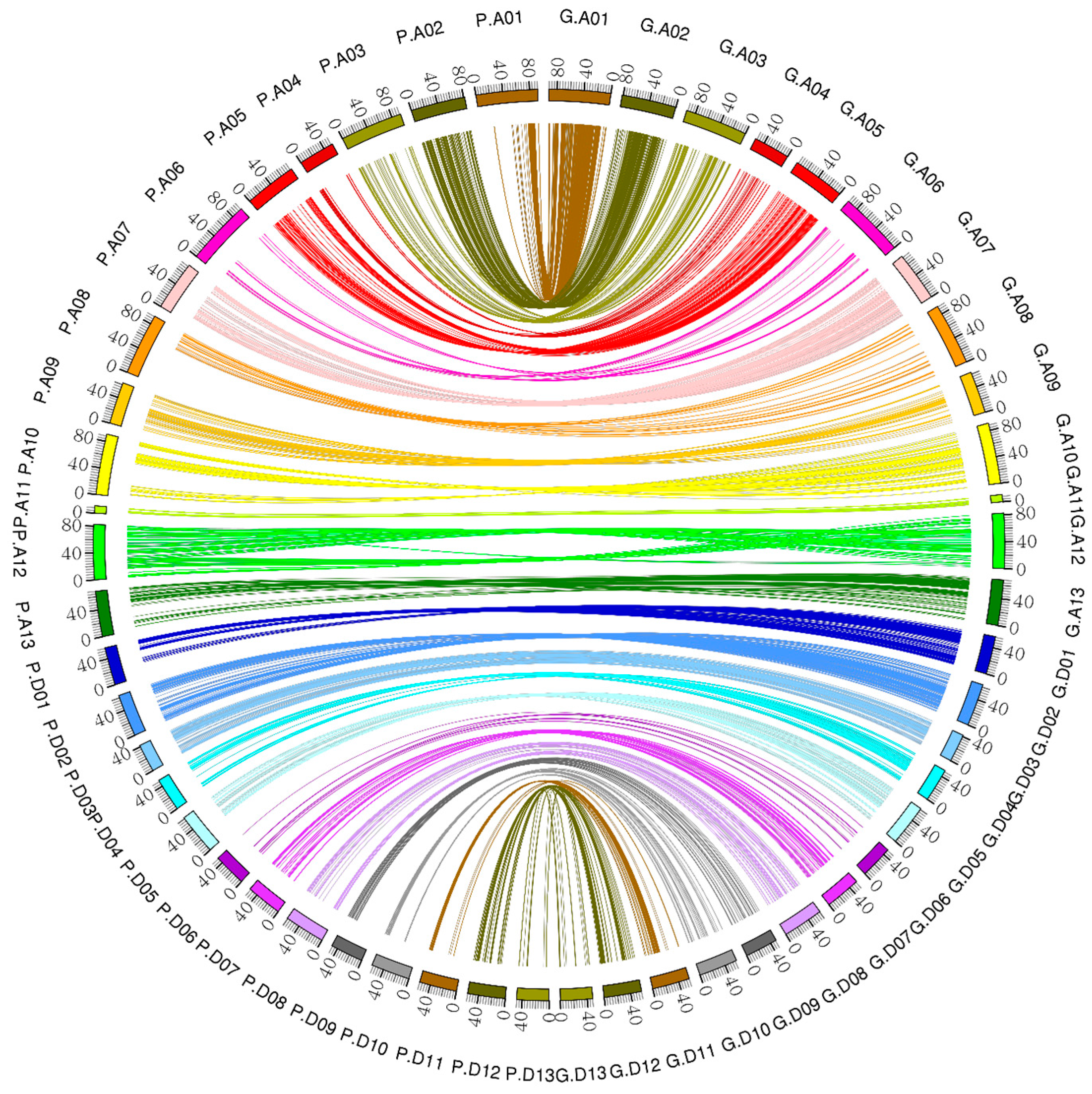
2.9. Collinearity Analysis
3. Discussion
4. Materials and Methods
4.1. Greenhouse Experiment
4.2. Sample Collection, Library Preparation, Sequencing, and SNP Genotyping
4.2.1. DNA Quantification and Qualification
4.2.2. GBS Library Preparation, Sequencing, and SNP Genotyping
4.3. Construction of the Linkage Maps and Data Analysis
4.4. Collinearity Analysis
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| Trait | Meaning | Unit |
| MDA | Malondialdehyde | µM/g.FW |
| EC | Electric Conductivity | % |
| GR | Germination Rate | number/40 seeds |
| FW | Fresh Weight | g |
| SL | Stem Length | cm |
| LFW | Leaf Fresh Weight | g |
| SLW | Saturated Leaf Weight | g |
| DLW | Dry Leaf Weight | g |
| RWC | Related Water Content | mL |
| CHL | Chlorophyll | mg/g.FW |
| NH | Nan Dan Ba Di Da Hua | |
| CCRI35 | Zhong35 |
References
- Munns, R. Genes and salt tolerance: Bringing them together. New Phytol. 2005, 167, 645–663. [Google Scholar] [CrossRef] [PubMed]
- Saeed, M.; Dahab, A.H.A.; Guo, W.Z.; Zhang, T.Z. Acascade of recently discovered molecular mechanisms involved in abiotic stress tolerance of plants. OMICS 2012, 16, 188–199. [Google Scholar] [CrossRef] [PubMed]
- Yue, G.H. Recent advances of genome mapping and marker-assisted selection in aquaculture. Fish Fish. 2014, 15, 376–396. [Google Scholar] [CrossRef]
- Goddard, M.E.; Hayes, B.J. Mapping genes for complex traits in domestic animals and their use in breeding programmes. Nat. Rev. Genet. 2009, 10, 381–391. [Google Scholar] [CrossRef] [PubMed]
- Meuwissen, T.H.E.; Hayes, B.J.; Goddard, M.E. Prediction of total genetic value using genome-wide dense marker maps. Genetics 2001, 157, 1819–1829. [Google Scholar] [PubMed]
- He, J.; Zhao, X.; Laroche, A.; Lu, Z.X.; Liu, H.; Li, Z. Genotyping-by-sequencing (GBS), an ultimate marker-assisted selection (MAS) tool to accelerate plant breeding. Front. Plant Sci. 2014, 5. [Google Scholar] [CrossRef] [PubMed]
- Thomson, M.; Zhao, K.; Wright, M.; McNally, K.; Rey, J.; Tung, C.W.; Reyn-olds, A.; Scheffler, B.; Eizenga, G.; McClung, A.K.H.; et al. High-throughput single nucleotide polymorphism genotyping for breeding applications in rice using the BeadXpress platform. Mol. Breed. 2012, 29, 875–886. [Google Scholar] [CrossRef]
- Poland, J.A.; Rife, T.W. Genotyping-by-sequencing for plant breeding and genetics. Plant Genome 2012, 5, 92–102. [Google Scholar] [CrossRef]
- Muhammad, A.; Majid, R.F. Crop breeding for salt tolerance in the era of molecular markers and marker-assisted selection. Plant Breed. 2013, 132, 10–20. [Google Scholar]
- Quesada, V.; García-Martínez, S.; Piqueras, P.; Ponce, M.; Micol, J.L. Genetic architecture of NaCl tolerance in Arabidopsis thaliana. Plant Physiol. 2002, 130, 951–963. [Google Scholar] [CrossRef] [PubMed]
- Takehisa, H.; Shimodate, T.; Fukuta, Y. Identification of quantitative trait loci for plant growth of rice in paddy field flooded with salt water. Field Crop Res. 2004, 89, 85–95. [Google Scholar] [CrossRef]
- Mano, Y.; Takeda, K. Mapping quantitative trait loci for salt tolerance at germination and the seedling stage in barley (Hordeum vulgare L.). Euphytica 1997, 94, 263–272. [Google Scholar] [CrossRef]
- Villalta, I.; Reina-Sa´nchez, A.; Boları´n, M.C. Genetic analysis of Na+ and K+ concentrations in leaf and stem as physiological components of salt tolerance in Tomato. Theor. Appl. Genet. Int. J. Plant Breed. Res. 2008, 116, 869–880. [Google Scholar] [CrossRef] [PubMed]
- Saeed, M.; Wangzhen, G.; Tianzhen, Z. Association mapping for salinity tolerance in cotton (Gossypium hirsutum L.) germplasm from US and diverse regions of China. Aust. J. Crop Sci. 2014, 8, 338–346. [Google Scholar]
- Penf, Z.; He, S.; Gong, W.; Sun, J.; Pan, Z.; Xu, F.; Lu, Y.; Xiongming, D. Comprehensive analysis of differentially expressed genes and transcriptional regulation induced by salt stress in two contrasting cotton genotypes. BMC Genom. 2014, 15, 760. [Google Scholar]
- Jin, L.; Li, H.; Liu, J.Y. Molecular characterization of three ethylene responsive element binding factor genes from cotton. J. Integr. Plant Biol. 2010, 52, 485–495. [Google Scholar] [CrossRef] [PubMed]
- Champion, A.; Hébrard, E.; Parra, B.; Bournaud, C.; Marmey, P.; Tranchant, C.; Nicole, M. Molecular diversity and gene expression of cotton ERF transcription factors reveal that group IXa members are responsive to jasmonate, ethylene and Xanthomonas. Mol. Plant Pathol. 2009, 10, 471–485. [Google Scholar] [CrossRef] [PubMed]
- Guo, Y.; Yu, Y.P.; Wang, D.; Wu, C.A.; Yang, G.D.; Huang, J.G.; Zheng, C.C. GhZFP1, a novel CCCH-type zinc finger protein from cotton, enhances salt stress tolerance and fungal disease resistance in transgenic tobacco by interacting with GZIRD21A and GZIPR5. New Phytol. 2009, 183, 62–75. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.A.; Yang, G.D.; Meng, Q.W.; Zheng, C.C. The cotton GhNHX1 gene encoding a novel putative tonoplast Na+/H+ antiporter plays an important role in salt stress. Plant Cell Physiol. 2004, 45, 600–607. [Google Scholar] [CrossRef] [PubMed]
- Meng, C.; Cai, C.; Zhang, T.; Guo, W. Characterization of six novel NAC genes and their responses to abiotic stresses in Gossypium hirsutum L. Plant Sci. Int. J. Exp. Plant Biol. 2009, 176, 352–359. [Google Scholar] [CrossRef]
- Xue, T.; Li, X.; Zhu, W.; Wu, C.; Yang, G.; Zheng, C. Cotton metallothionein GhMT3a, a reactive oxygen species scavenger, increased tolerance against abiotic stress in transgenic tobacco and yeast. J. Exp. Biol. 2009, 60, 339–349. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Xi, D.; Li, S.; Gao, Z.; Zhao, S.; Shi, J.; Wu, C.; Guo, X. A cotton group C MAP kinase gene, GhMPK2, positively regulates salt and drought tolerance in tobacco. Plant Mol. Biol. 2011, 77, 17–31. [Google Scholar] [CrossRef] [PubMed]
- Lu, W.; Chu, X.; Li, Y.; Wang, C.; Guo, X. Cotton GhMKK1 Induces the Tolerance of Salt and Drought Stress, and Mediates Defence Responses to Pathogen Infection in Transgenic Nicotianabenthamiana. PLoS ONE 2013. [Google Scholar] [CrossRef]
- Gao, S.Q.; Chen, M.; Xia, L.Q.; Xiu, H.J.; Xu, Z.S.; Li, L.-C.; Zhao, C.P.; Cheng, X.G.; Ma, Y.Z. A cotton (Gossypium hirsutum) DRE-binding transcription factor gene, GhDREB, confers enhanced tolerance to drought, high salt, and freezing stresses in transgenic wheat. Plant Cell Rep. 2009, 28, 301–311. [Google Scholar] [CrossRef] [PubMed]
- Oluoch, G.; Zheng, J.; Wang, X.; Khan, M.K.R.; Zhou, Z.; Cai, X.; Wang, C.; Wang, Y.; Li, X.; Wang, H.; et al. QTL mapping for salt tolerance at seedling stage in the interspecific cross of Gossypium tomentosum with Gossypium hirsutum. Euphytica 2016, 209, 223–235. [Google Scholar] [CrossRef]
- Khorsandi, F.; Anagholi, A. Reproductive compensation of cotton after salt stress relief at different growth stages. J. Agron. Crop Sci. 2009, 195, 278–283. [Google Scholar] [CrossRef]
- Munns, R. Utilizing genetic resources to enhance productivity of salt-prone land. CAB Rev. 2007, 2, 1–11. [Google Scholar] [CrossRef]
- Wang, S.; Zeng, Z.B. Department of Statistics; Windows QTL Cartographer 2.5; North Carolina State University: Raleigh, NC, USA, 2007. [Google Scholar]
- Li, F.; Fan, G.; Lu, C.; Xiao, G.; Zou, C.; Kohel, R.J.; Ma, Z.; Shang, H.; Ma, X.; Wu, J.; et al. Genome sequence of cultivated Upland cotton (Gossypium hirsutum TM-1) provides insights into genome evolution. Nat. Biotechnol. 2015, 33, 524–530. [Google Scholar] [CrossRef] [PubMed]
- Rong, J.; Abbey, C.; Bowers, J.E. A 3347-locus genetic recombination map of sequence-tagged sites reveals features of genome organization, transmission and evolution of cotton (Gossypium). Genet. Mol. Biol. 2004, 166, 389–417. [Google Scholar] [CrossRef]
- Singh, U.M.; Yadav, S.; Dixit, S.; Ramayya, P.J.; Devi, M.N.; Raman, K.A.; Kumar, A. QTL Hotspots for Early Vigor and Related Traits under Dry Direct-Seeded System in Rice (Oryza sativa L.). Front. Plant Sci. 2017, 8. [Google Scholar] [CrossRef] [PubMed]
- Foolad, M.R.; Jones, R.A. Mapping salt-tolerance genes in tomato (Lycopersiconesculentum) using trait-based marker analysis. Theor. Appl. Genet. Int. J. Plant Breed. Res. 1993, 87, 184–192. [Google Scholar]
- García-Sánchez, F.; Syvertsen, J.P. Substrate Type and Salinity Affect Growth Allocation, Tissue Ion Concentrations, and Physiological Responses of Carrizo Citrange Seedlings. Hortscience 2009, 44, 1432–1437. [Google Scholar]
- Esfandiari, E.; Enayati, V.; Abbasi, A. Biochemical and Physiological Changes in Response to Salinity in Two Durum Wheat (Triticumturgidum L.) Genotypes. Bot. Hort. Agrobot. 2011, 39, 165–170. [Google Scholar]
- Abeer Ahmad, R.; Ali Farghaly, F.; Hamada, A.M. Physiological and biochemical responses of salt-tolerant. J. Biol. Earth Sci. 2013, 3, B72–B88. [Google Scholar]
- Koca, H.; Bor, M.; Özdemir, F.; Türkan, I. The effect of salt stress on lipid peroxidation, antioxidative enzymes and proline content of sesame cultivars. Environ. Exp. Bot. 2007, 60, 344–351. [Google Scholar] [CrossRef]
- Ksouri, R.; Megdiche, W.; Debez, A.; Falleh, H.; Grignon, C. Salinity effects on polyphenol content and antioxidant activities in leaves of the halophyte Cakile maritime. Plant Physiol. Biochem. 2007, 45, 244–249. [Google Scholar] [CrossRef] [PubMed]
- Kaur, N.; Dhawan, M.; Sharma, I.; Pati, P.K. Interdependency of Reactive Oxygen Species generating and scavenging system in salt sensitive and salt tolerant cultivars of rice. BMC Plant Biol. 2016, 16, 131. [Google Scholar] [CrossRef] [PubMed]
- André, D.; José, T.; Joaquim, E.; Carlos, E.; Enéas, G. Effect of salt stress on antioxidative enzymes and lipid peroxidation in leaves and roots of salt-tolerant and salt-sensitive maize genotypes. Environ. Exp. Bot. 2006, 56, 87–94. [Google Scholar]
- Sharma, I.; Ching, E.; Saini, S.; Bhardwaj, R.; Pati, P.K. Exogenous application of brassinosteroid offers tolerance to salinity by altering stress responses in rice variety Pusa Basmati-1. Plant Physiol. Biochem. 2013, 69, 17–26. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Hu, L.; Zhang, L.; Pan, X.; Hu, X. Exogenous spermidine is enhancing tomato tolerance to salinity–alkalinity stress by regulating chloroplast antioxidant system and chlorophyll metabolism. BMC Plant Biol. 2015. [Google Scholar] [CrossRef] [PubMed]
- Stepien, P.; Johnson, G.N. Contrasting responses of photosynthesis to salt stress in the glycophyte Arabidopsis and the halophyte Thellungiella: Role of the plastid terminal oxidase as an alternative electron sink. Plant Physiol. 2009, 149, 1154–1165. [Google Scholar] [CrossRef] [PubMed]
- Ashraf, M.; Harris, P.J.C. Photosynthesis under stressful environments: An overview. Photosynthetica 2013, 51, 163–190. [Google Scholar] [CrossRef]
- Gossett, D.R.; Millhollon, E.P.; Lucas, M.C. Antiox-idant response to NaCl stress in salt-tolerant and salt-sensitive cultivars of cotton. Crop Sci. 1994, 34, 706–714. [Google Scholar] [CrossRef]
- Sairam, R.; Veerabhadra Rao, K.; Srivastava, G.C. Differential response of wheat genotypes to long term salinity stress in relation to oxidative stress, antioxidant activity and osmolyte concentration. Plant Sci. Int. J. Exp. Plant Biol. 2002, 163, 1037–1046. [Google Scholar] [CrossRef]
- Muhammad, A. Some important physiological selection criteria for salt tolerance in plants. Flora 2004, 199, 361–376. [Google Scholar]
- Lauchli, A.; Grattan, S.R. Plant growth and development under salinity stress. Springer 2007. [Google Scholar] [CrossRef]
- Gossett, D.R.; Millhollon, E.P.; Caldwell, W.D.; Mundy, S. Isozyme variation among salt tolerant and salt sensitive varieties of cotton. In Proceedings of the Beltwide Cotton Production Research Conference, Nashnille, TN, USA, 6–10 January 1992; pp. 556–559. [Google Scholar]
- Bhatti, M.A.; Azhar, F.M. Salt tolerance of nine Gossypium hirsutum L. varieties to NaCl salinity at early stage of plant development. Int. J. Agric. Biol. 2002, 4, 544–546. [Google Scholar]
- Lang, L.; Xu, A.; Ding, J.; Zhang, Y.; Zhao, N.; Tian, Z.; Liu, Y.; Wang, Y.; Liu, X.; Liang, F.; et al. Quantitative Trait Locus Mapping of Salt Tolerance and Identification of Salt-Tolerant Genes in Brassica napus L. Front. Plant Sci. 2017, 8. [Google Scholar] [CrossRef] [PubMed]
- Saba, J.; Moghaddam, M.; Ghassemi, K.; Nishabouri, M.R. Genetic properties of drought resistance indices. J. Agric. Sci. Technol. 2001, 3, 43–49. [Google Scholar]
- Hanif, M.; Noor, E.; Murtaza, N. Assessment of variability for salt tolerance at seedling stage in Gossypium hirsutum L. J. Food Agric. Environ. 2008, 6, 134–138. [Google Scholar]
- Nadarajan, N.G.M. Quantitative Genetics and Biometrical Techniques in Plant Breeding; Kalyani Publication: New Delhi, India, 2005. [Google Scholar]
- Rachid Al-Chaarani, G.; Gentzbittel, L.; Huang, X.; Sarrafi, A. Genotypic variation and identification of QTLs for agronomic traits using AFLP and SSR in recombinant inbred lines of sunflower (Helianthus annuus L.). Theor. Appl. Genet. Theor. Angew. Genet. 2004, 109, 1353–1360. [Google Scholar] [CrossRef] [PubMed]
- Loudet, O.; Sylvain, C.; Patricia, M.; Joël, T.; Françoise, D.-V. Quantitative trait loci analysis of water and anion contents in interaction with nitrogen availability in Arabidopsis thaliana. Genetics 2003, 163, 711–722. [Google Scholar] [PubMed]
- Loudet, O.; Sylvain, C.; Patricia, M.; Joël, T.; Françoise, D.-V. Quantitative trait loci analysis of nitrogen use efficiency in Arabidopsis. Plant Physiol. 2003, 131, 345–358. [Google Scholar] [CrossRef] [PubMed]
- Jamshed, M.; Jia, F.; Gong, J.; Palanga, K.K.; Shi, Y.; Li, J.; Shang, H.; Liu, A.; Chen, T.; Zhang, Z.; et al. Identification of stable quantitative trait loci (QTLs) for fiber quality traits across multiple environments in Gossypium hirsutum recombinant inbred line population. BMC Genom. 2016, 17, 197. [Google Scholar] [CrossRef] [PubMed]
- Cho, S.K.; Ryu, M.Y.; Song, C.; Kwak, J.M.; Kima, W.T. Arabidopsis PUB22 and PUB23 are homologous U-Box E3 ubiquitin ligases that play combinatory roles in response to drought stress. Plant Cell 2008, 20, 1899–1914. [Google Scholar] [CrossRef] [PubMed]
- Du, X.-M.; Sun, J.-L.; Zhou, Z.-L.; Gu, Y.-H.; Pan, Z.-E.; He, S.-P.; Pang, B.-Y.; Wang, L.-R. Current Situation and the Future in Collection, Preservation, Evaluation. Plant Genet. Resour. 2012, 13, 163–168. [Google Scholar]
- Hoagland, D.R.; Arnon, D.I. The Water-Culture Method for Growing Plants without Soil; California Experiment Station Circular, The College of Agriculture, University of California: Berkeley, CA, USA, 1950. [Google Scholar]
- Barr, H.D.; Weatherley, P.E. A re-examination of the relative turgidity technique for estimating water deficit in leaves. Aust. J. Biol. Sci. 1962, 15, 413–428. [Google Scholar] [CrossRef]
- Madhava Rao, K.V.; Sresty, T.V. Antioxidative parameters in the seedlings of pigeonpea (Cajanuscajan L. Millspaugh) in response to Zn and Ni stresses. Plant Sci. 2000, 157, 113–128. [Google Scholar] [CrossRef]
- Paterson, A.H.; Brubaker, C.; Wendel, J.F. A rapid method for extraction of cotton (Gossypium spp.) genomic DNA suitable for RFLP or PCR analysis. Plant Mol. Biol. Rep. 1993, 11, 122–127. [Google Scholar] [CrossRef]
- Krizman, M.J.J.; Baricevic, D.; Jakše, J.; Prosek, M. Robust CTAB-activated charcoal protocol for plant DNA extraction. Acta Agric. Slov. 2006, 87, 427–433. [Google Scholar]
- Wilfinger, W.W.; Chomczynski, M.K. P 260/280 and 260/230 Ratios NanoDrop® ND-1000 and ND-8000 8-Sample Spectrophotometers. BioTechniques 1997, 22, 474–481. [Google Scholar] [PubMed]
- Kante, M.; Rattunde, H.F.W.; Leiser, W.L.; Nebié, B.; Diallo, B.; Diallo, A.; Touré, A.O.; Weltzien, E.; Haussmann, B.I.G. Can Tall Guinea-Race Sorghum Hybrids Deliver Yield Advantage to Smallholder Farmers in West and Central Africa? Crop Sci. 2017, 57. [Google Scholar] [CrossRef]
- Elshire, R.J.; Glaubitz, G.J.; Sun, Q.; Jesse, A.; Kawamoto, K.; Buckler, E.S.; Mitchell, S.E. A robust, simple genotyping-by-sequencing (GBS) approach for high diversity species. PLoS ONE 2011, 6. [Google Scholar] [CrossRef] [PubMed]
- Glaubitz, J.C.; Casstevens, T.M.; Lu, F.; Harriman, J.; Elshire, R.J.; Sun, Q.; Buck, E.S. TASSEL-GBS: A High Capacity Genotyping by Sequencing Analysis Pipeline. PLoS ONE 2014, 9. [Google Scholar] [CrossRef] [PubMed]
- Paten, B.; Novak, A.; Haussler, D. Mapping to a Reference Genome Structure. arXiv, 2014; 1–26arXiv:1404.5010. [Google Scholar]
- Li, H.; Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar] [CrossRef] [PubMed]
- Danecek, P.; Auton, A.; Abecasis, G.; Albers, C.A.; Banks, E.; DePristo, M.A.; Handsaker, R.E.; Lunter, G.; Marth, G.T.; Sherry, S.T.; et al. The variant call format and VCFtools. Bioinformatics 2011, 27, 2156–2158. [Google Scholar] [CrossRef] [PubMed]
- Bradbury, P.J.; Zhang, Z.; Kroon, D.E.; Casstevens, T.M.; Ramdoss, Y.; Buckler, E.S. TASSEL: Software for association mapping of complex traits in diverse samples. Bioinformatics 2007, 23, 2633–2635. [Google Scholar] [CrossRef] [PubMed]
- Stam, P.; van Ooijen, J.W. JoinMapTM Version 2.0: Software for the Calculation of Genetic Linkage Maps; Plant Research International B.V. and Kyazma B.V.: Wageningen, The Netherlands, 1995. [Google Scholar]
- The R Development Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2008. [Google Scholar]
- Silva, M.J.R.; Ferreira, T.E.; Domiciano, S.; Paula, A.; Paiva, M.; Tecchio, M.A.; Leonel, S. Phenology, yield and fruit quality of four persimmon (Diospyros kaki L.) cultivars in So Paulos Midwest countryside, Brazil. Afr. J. Agric. Res. 2016, 11, 5171–5177. [Google Scholar] [CrossRef]
- Lander, E.; Kruglyak, L. Genetic dissection of complex traits: Guidelines for interpreting and reporting linkage results. Nat. Genet. 1995, 1, 1241–1247. [Google Scholar] [CrossRef] [PubMed]
- Liang, Q.; Hu, C.; Hua, H.; Li, Z.; Hua, J. Construction of a linkage map and QTL mapping for fiber quality traits in upland cotton (Gossypium hirsutum L.). Chin. Sci. Bull. 2013, 58, 3233–3243. [Google Scholar] [CrossRef]
- Stuber, C.; Edwards, M.D.; Wendel, J.F. Molecular marker facilitated investigations of quantitative trait loci in maize. II. Factors influencing yield and its component traits. Crop Sci. Rep. 1987, 27, 639–648. [Google Scholar] [CrossRef]
- Voorrips, R.E. MapChart: Software for the graphical presentation of linkage maps and QTLs. J. Hered. 2002, 93, 77–78. [Google Scholar] [CrossRef] [PubMed]
- Zhang, T.; Hu, Y.; Jiang, W.; Fang, L.; Guan, X.; Chen, J.; Zhang, J.; Saski, C.A.; Scheffler, B.E.; Stelly, D.M.; et al. Sequencing of allotetraploid cotton (Gossypium hirsutum L. acc. TM-1) provides a resource for fiber improvement. Nat. Biotechnol. 2015, 33, 531–537. [Google Scholar] [CrossRef] [PubMed]

| F2:3 | |||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Trait a | Env b | Source | Df | Mean Square | F Value | Pr > F | HB (%) | NH = P2 | CCRI35 = P1 | P1–P2 | Mean | Var | SD | Min | Max | Skew | Kurt |
| MDA | CK | e | 2 | 0.009 | 199.712 | <0.0001 | 73.4 | 0.028 | 0.022 | −0.006 | 0.042 | 0 | 0.011 | 0 | 0.085 | 0.052 | 0.753 |
| 1 | g | 276 | 0.001 | 14.401 | <0.0001 | 0.027 | 0.015 | −0.012 | 0.037 | 0 | 0.011 | 0.004 | 0.066 | −0.05 | 0.159 | ||
| 2 | g * e | 543 | 0.0003 | 6.11 | <0.0001 | 0.014 | 0.009 | −0.005 | 0.036 | 0 | 0.016 | 0 | 0.098 | 0.198 | 0.081 | ||
| EC | CK | e | 2 | 29,102.2 | 978.452 | <0.0001 | 57.5 | 68.505 | 64.085 | −4.42 | 45.469 | 109.207 | 10.45 | 22.754 | 92.32 | 0.72 | 0.762 |
| 1 | g | 276 | 246.072 | 8.273 | <0.0001 | 37.882 | 41.406 | 3.524 | 35.448 | 41.403 | 6.435 | 18.824 | 59.902 | 0.63 | 0.645 | ||
| 2 | g * e | 543 | 125.338 | 4.214 | <0.0001 | 39.695 | 41.617 | 1.922 | 34.601 | 75.858 | 8.71 | 16.713 | 67.505 | 0.966 | 0.883 | ||
| GR | CK | e | 2 | 8313.01 | 427.302 | <0.0001 | 68.4 | 31 | 33.667 | 2.667 | 32.922 | 33.583 | 5.795 | 7 | 40 | −1.17 | 1.364 |
| 1 | g | 276 | 191.801 | 9.859 | <0.0001 | 22.667 | 26.5 | 3.833 | 26.677 | 61.173 | 7.821 | 6 | 40 | −0.533 | −0.535 | ||
| 2 | g * e | 543 | 65.441 | 3.364 | <0.0001 | 26 | 30 | 4 | 28.355 | 54.829 | 7.405 | 4 | 40 | −0.774 | 0.058 | ||
| FW | CK | e | 2 | 8.543 | 7435 | <0.0001 | 87.4 | 0.615 | 0.646 | 0.031 | 0.668 | 0.007 | 0.086 | 0.412 | 0.894 | −0.055 | −0.394 |
| 1 | g | 276 | 0.033 | 28.453 | <0.0001 | 0.501 | 0.55 | 0.049 | 0.513 | 0.005 | 0.071 | 0.318 | 0.7 | 0.053 | −0.343 | ||
| 2 | g * e | 543 | 0.009 | 7.621 | <0.0001 | 0.419 | 0.445 | 0.026 | 0.472 | 0.007 | 0.085 | 0.284 | 0.79 | 0.444 | −0.029 | ||
| SL | CK | e | 2 | 971.272 | 6639.29 | <0.0001 | 83.1 | 9.14 | 9.267 | 0.127 | 8.082 | 0.512 | 0.716 | 5.46 | 10.54 | −0.269 | 0.407 |
| 1 | g | 276 | 3.988 | 27.261 | <0.0001 | 7.38 | 8.187 | 0.807 | 7.06 | 0.882 | 0.939 | 4.12 | 10.44 | 0.107 | 0.194 | ||
| 2 | g * e | 543 | 1.833 | 12.526 | <0.0001 | 5.533 | 6.32 | 0.787 | 5.852 | 1.575 | 1.255 | 2.9 | 9.58 | 0.172 | −0.388 | ||
| LFW | CK | e | 2 | 1.103 | 3127.67 | <0.0001 | 87.3 | 0.263 | 0.305 | 0.042 | 0.292 | 0.002 | 0.042 | 0.168 | 0.416 | −0.019 | −0.21 |
| 1 | g | 276 | 0.01 | 27.156 | <0.0001 | 0.242 | 0.273 | 0.031 | 0.258 | 0.002 | 0.042 | 0.138 | 0.396 | 0.205 | 0.064 | ||
| 2 | g * e | 543 | 0.002 | 6.484 | <0.0001 | 0.188 | 0.216 | 0.028 | 0.217 | 0.002 | 0.046 | 0.062 | 0.39 | 0.449 | 0.385 | ||
| SLW | CK | e | 2 | 0.778 | 1641.26 | <0.0001 | 86.8 | 0.292 | 0.343 | 0.051 | 0.311 | 0.002 | 0.044 | 0.178 | 0.432 | −0.035 | −0.152 |
| 1 | g | 276 | 0.012 | 25.249 | <0.0001 | 0.297 | 0.329 | 0.032 | 0.319 | 0.002 | 0.049 | 0.18 | 0.494 | 0.147 | 0.037 | ||
| 2 | g * e | 543 | 0.003 | 5.455 | <0.0001 | 0.255 | 0.285 | 0.03 | 0.26 | 0.003 | 0.05 | 0.076 | 0.478 | 0.398 | 0.657 | ||
| DLW | CK | e | 2 | 0.001 | 86.178 | <0.0001 | 82.6 | 0.025 | 0.031 | 0.006 | 0.032 | 0 | 0.005 | 0.016 | 0.046 | −0.015 | −0.053 |
| 1 | g | 276 | 0.0001 | 17.117 | <0.0001 | 0.032 | 0.033 | 0.001 | 0.034 | 0 | 0.005 | 0.018 | 0.05 | 0.044 | 0.113 | ||
| 2 | g * e | 543 | 2.18 × 10−5 | 2.826 | <0.0001 | 0.037 | 0.032 | −0.005 | 0.034 | 0 | 0.005 | 0.018 | 0.052 | 0.145 | 0.02 | ||
| RWC | CK | e | 2 | 5.091 | 7581.42 | <0.0001 | 42.3 | 0.889 | 0.878 | −0.011 | 0.934 | 0.002 | 0.042 | 0.524 | 0.994 | −3.565 | 0.22 |
| 1 | g | 276 | 0.006 | 8.758 | <0.0001 | 0.792 | 0.809 | 0.017 | 0.786 | 0.001 | 0.038 | 0.608 | 0.941 | −0.138 | 1.348 | ||
| 2 | g * e | 543 | 0.004 | 6.559 | <0.0001 | 0.691 | 0.727 | 0.036 | 0.807 | 0.003 | 0.057 | 0.615 | 0.987 | −0.141 | 0.969 | ||
| CHL | CK | e | 2 | 15,495.4 | 2024.51 | <0.0001 | 51 | 59.633 | 52.86 | −6.773 | 54.091 | 15.969 | 3.996 | 39.58 | 65.82 | −0.173 | 0.345 |
| 1 | g | 276 | 67.297 | 8.793 | <0.0001 | 51.073 | 51.907 | 0.834 | 57.583 | 14.45 | 3.801 | 41.44 | 66.78 | −0.486 | 0.467 | ||
| 2 | g * e | 543 | 43.419 | 5.673 | <0.0001 | 48.413 | 50.94 | 2.527 | 48.721 | 38.069 | 6.17 | 29.74 | 63.46 | −0.53 | 0.126 | ||
| Group | Marker_Number | Map_Length (cM) | Av_Distance (cM) | Max_Gap (cM) | <5 cM | 5–10 cM | 10–20 cM | >20 cM | Ratio |
|---|---|---|---|---|---|---|---|---|---|
| A01(c1) | 448 | 146.704 | 0.33 | 8.505 | 445 | 2 | 0 | 0 | 1 |
| A02(c2) | 705 | 346.314 | 0.49 | 17.848 | 687 | 12 | 5 | 0 | 0.98 |
| A03(c3) | 323 | 213.937 | 0.66 | 17.145 | 309 | 10 | 3 | 0 | 0.96 |
| A04(c4) | 106 | 203.891 | 1.92 | 26.598 | 91 | 8 | 5 | 1 | 0.87 |
| A05(c5) | 378 | 385.092 | 1.02 | 21.198 | 354 | 11 | 11 | 1 | 0.94 |
| A06(c6) | 58 | 73.063 | 1.26 | 15.032 | 53 | 1 | 3 | 0 | 0.93 |
| A07(c7) | 279 | 205.892 | 0.74 | 11.622 | 272 | 4 | 2 | 0 | 0.98 |
| A08(c8) | 69 | 112.137 | 1.63 | 18.894 | 58 | 7 | 3 | 0 | 0.85 |
| A09(c9) | 98 | 138.501 | 1.41 | 19.234 | 86 | 9 | 2 | 0 | 0.89 |
| A10(c10) | 292 | 202.134 | 0.69 | 10.551 | 280 | 7 | 4 | 0 | 0.96 |
| A11(c11) | 51 | 70.548 | 1.38 | 23.241 | 47 | 2 | 0 | 1 | 0.94 |
| A12(c12) | 244 | 309.608 | 1.27 | 19.593 | 224 | 12 | 7 | 0 | 0.92 |
| A13(c13) | 262 | 203.61 | 0.78 | 17.425 | 249 | 7 | 5 | 0 | 0.95 |
| At sub-genome | 3313 | 2611.43 | 0.79 | 26.598 | 3155 | 92 | 50 | 3 | 0.94 |
| D01(c15) | 319 | 144.092 | 0.45 | 6.351 | 316 | 2 | 0 | 0 | 0.99 |
| D02(c14) | 454 | 313.268 | 0.69 | 14.541 | 438 | 12 | 3 | 0 | 0.97 |
| D03(c17) | 133 | 170.555 | 1.28 | 14.993 | 124 | 7 | 1 | 0 | 0.94 |
| D04(c22) | 114 | 136.228 | 1.19 | 20.275 | 107 | 3 | 2 | 1 | 0.95 |
| D05(c19) | 153 | 218.788 | 1.43 | 27.062 | 141 | 7 | 2 | 2 | 0.93 |
| D06(c25) | 16 | 79.084 | 4.94 | 22.389 | 11 | 1 | 2 | 1 | 0.73 |
| D07(c16) | 169 | 235.366 | 1.39 | 26.041 | 154 | 7 | 6 | 1 | 0.92 |
| D08(c24) | 118 | 226.688 | 1.92 | 20.878 | 103 | 6 | 7 | 1 | 0.88 |
| D09(c23) | 40 | 136.744 | 3.42 | 14.48 | 28 | 5 | 6 | 0 | 0.72 |
| D10(c20) | 80 | 129.051 | 1.61 | 20.539 | 71 | 5 | 2 | 1 | 0.9 |
| D11(c21) | 98 | 89.782 | 0.92 | 27.564 | 93 | 2 | 1 | 1 | 0.96 |
| D12(c26) | 143 | 194.735 | 1.36 | 30.082 | 133 | 2 | 5 | 2 | 0.94 |
| D13(c18) | 28 | 82.286 | 2.94 | 20.917 | 22 | 3 | 1 | 1 | 0.81 |
| Dt sub-genome | 1865 | 2156.67 | 1.156 | 30.082 | 1741 | 62 | 38 | 11 | 0.9 |
| Total (At + Dt) | 5178 | 4768.1 | 0.92 | 30.082 | 4896 | 154 | 88 | 14 | 0.92 |
| Cluster | Trait | QTLs | Chr | Start Marker | End Marker | Start Marker (bp) | End Marker (bp) | Length of CI Markers (Mb) | Position (cM) | LOD | Ae | De | |d/a| | R2 (%) | GA | DPE |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | EC | qEC_A02_ck | A02 | mk1046_A02 | mk1107_A02 | 4,445,557 | 7,584,839 | 3.14 | 2.64 | −1.3522 | 4.3713 | 3.23 | 3.65 | OD | NH | |
| qEC_A02_150 | A02 | MulMa26-m_A02 | mk1778_A02 | 76,134,216 | 81,766,125 | 5.63 | 20.81 | 2.78 | −1.3853 | −2.2243 | 1.61 | 1.73 | OD | NH | ||
| qEC_A02_110.2 | A02 | mk19866 | mk14259 | 1876 | 2247 | 0.0004 | 197.41 | 2.71 | −7.1631 | −2.5385 | 0.35 | 1.14 | PD | NH | ||
| qEC_A02_110.3 | A02 | mk1293_A02 | mk1512_A02 | 23,817,680 | 40,364,786 | 16.55 | 220.01 | 3.1 | −9.7207 | −2.9306 | 0.3 | 0 | PD | NH | ||
| qEC_A02_110.1 | A02 | mk1340_A02 | mk1443_A02 | 25,911,715 | 35,098,219 | 9.19 | 134.51 | 2.85 | −8.6005 | −3.3756 | 0.39 | 0 | PD | NH | ||
| 2 | FW | qFW_A06_150.1 | A06 | mk4886_A06 | mk4999_A06 | 92,310,429 | 97,063,193 | 4.75 | 0.31 | 3.71 | −0.0093 | 0.0504 | 5.42 | 3.39 | OD | NH |
| qFW_A06_150.2 | A06 | mk4886_A06 | mk4999_A06 | 92,310,429 | 97,063,193 | 4.75 | 0.81 | 4.5 | −0.0041 | 0.0325 | 7.93 | 3.14 | OD | NH | ||
| qFW_A06_110.1 | A06 | mk4886_A06 | mk4999_A06 | 92,310,429 | 97,063,193 | 4.75 | 1.31 | 3.32 | 0.0019 | 0.0432 | 22.74 | 0.52 | OD | CCRI35 | ||
| qFW_A06_110.2 | A06 | mk4998_A06 | mk5001_A06 | 97,063,190 | 97,172,480 | 0.11 | 1.31 | 4.75 | 0.0004 | 0.0294 | 73.5 | 0.87 | OD | CCRI35 | ||
| SLW | qSLW_A06_110 | A06 | mk4886_A06 | mk4999_A06 | 92,310,429 | 97,063,193 | 4.75 | 1.31 | 3.05 | 0.0035 | 0.0277 | 7.91 | 0.01 | OD | CCRI35 | |
| qSLW_A06_150 | A06 | mk4999_A06 | mk5000_A06 | 97,063,193 | 97,137,676 | 0.07 | 0.81 | 4.5 | −0.0038 | 0.0359 | 9.45 | 2.76 | OD | NH | ||
| 3 | EC | qEC_A12_110.1 | A12 | mk9153_A12 | mk9167_A12 | 77,455,444 | 77,705,877 | 0.25 | 57.51 | 4.35 | −1.4649 | 2.4503 | 1.67 | 7.91 | OD | NH |
| qEC_A12_150 | A12 | mk9173_A12 | mk9189_A12 | 79,355,806 | 81,365,007 | 2.01 | 29.51 | 3.01 | −1.9851 | −0.6644 | 0.33 | 4.47 | PD | NH | ||
| qEC_A12_110.2 | A12 | MulMa655-m | mk19378 | 23,988 | 92,678 | 0.07 | 75.21 | 5.28 | −1.2555 | 4.3961 | 3.5 | 8.29 | OD | NH | ||
| SL | qSL_A12_150 | A12 | mk18878 | mk9187_A12 | 173 | 81,262,301 | 81.26 | 46.51 | 2.68 | −0.2876 | −0.0593 | 0.21 | 4.18 | PD | NH | |
| qSL_A12_110 | A12 | mk8806_A12 | mk8817_A12 | 253,901 | 1,660,995 | 1.41 | 287.81 | 2.64 | 0.065 | 0.456 | 7.02 | 0.03 | OD | CCRI35 | ||
| 4 | FW | qFW_D01_110.3 | D01 | mk10908_D01 | mk10918_D01 | 54,507,760 | 54,630,017 | 0.12 | 55.41 | 3.95 | 0.0097 | 0.0064 | 0.66 | 4.25 | PD | CCRI35 |
| qFW_D01_110.2 | D01 | mk10908_D01 | mk10918_D01 | 54,507,760 | 54,630,017 | 0.12 | 55.41 | 3.32 | 0.0157 | 0.0072 | 0.46 | 4.06 | PD | CCRI35 | ||
| qFW_D01_110.1 | D01 | mk10900_D01 | mk10955_D01 | 54,444,280 | 55,458,818 | 1.01 | 50.21 | 2.51 | 0.013 | 0.0127 | 0.98 | 2.25 | D | CCRI35 | ||
| qFW_D01_150.2 | D01 | mk10837_D01 | MulMa276_D01 | 51,102,990 | 53,566,034 | 2.46 | 82.31 | 2.72 | 0.0065 | 0.019 | 2.92 | 0.74 | OD | CCRI35 | ||
| qFW_D01_150.1 | D01 | mk10840_D01 | MulMa276_D01 | 51,199,333 | 53,566,034 | 2.37 | 83.31 | 2.69 | 0.0097 | 0.0385 | 3.97 | 0.32 | OD | CCRI35 | ||
| SLW | qSLW_D01_110 | D01 | mk10840_D01 | MulMa276_D01 | 51,199,333 | 53,566,034 | 2.37 | 80.71 | 3.37 | 0.0095 | 0.0193 | 2.03 | 2.09 | OD | CCRI35 | |
| qSLW_D01_150.2 | D01 | mk10837_D01 | MulMa276_D01 | 51,102,990 | 53,566,034 | 2.46 | 82.31 | 2.84 | 0.0072 | 0.0218 | 3.03 | 0.73 | OD | CCRI35 | ||
| qSLW_D01_150.1 | D01 | MulMa268-m_D01 | MulMa272_D01 | 51,320,058 | 51,944,113 | 0.62 | 2.53 | −0.01 | 0.0227 | 2.27 | 4.94 | OD | NH | |||
| 5 | DLW | qDLW_D03_ck.1 | D03 | mk12058_D03 | mk12060_D03 | 3,459,294 | 4,072,981 | 0.61 | 54.71 | 4.04 | −0.001 | 0.0016 | 1.6 | 6.92 | OD | NH |
| qDLW_D03_ck.2 | D03 | mk12061_D03 | mk12065_D03 | 4,073,116 | 4,904,755 | 0.83 | 60.91 | 2.65 | −0.0007 | 0.0018 | 2.57 | 4.29 | OD | NH | ||
| qDLW_D03_110 | D03 | mk12143_D03 | mk12152_D03 | 36,784,173 | 37,665,167 | 0.88 | 135.01 | 2.88 | −0.0011 | 0.0001 | 0.09 | 4.29 | A | NH | ||
| qDLW_D03_ck.3 | D03 | mk12156_D03 | mk12167_D03 | 37,695,156 | 40,261,578 | 2.57 | 149.31 | 2.66 | −0.0006 | 0.0019 | 3.17 | 3.34 | OD | NH | ||
| FW | qFW_D03_110.2 | D03 | mk12096_D03 | mk12127_D03 | 26,385,562 | 30,628,951 | 4.24 | 92.11 | 5.76 | −0.0231 | 0.0067 | 0.29 | 9.87 | PD | NH | |
| qFW_D03_ck | D03 | mk12142_D03 | mk12152_D03 | 36,697,656 | 37,665,167 | 0.97 | 132.21 | 4.19 | −0.0227 | 0.0241 | 1.06 | 8 | D | NH | ||
| qFW_D03_110.1 | D03 | mk15025 | mk15027 | 5,017,089 | 5,704,231 | 0.69 | 81.91 | 6.05 | −0.0219 | −0.0093 | 0.42 | 7.24 | PD | NH | ||
| qFW_D03_150.1 | D03 | mk12142_D03 | mk12154_D03 | 36,697,656 | 37,676,414 | 0.98 | 132.21 | 4.45 | −0.0241 | −0.014 | 0.58 | 5.46 | PD | NH | ||
| qFW_D03_150.2 | D03 | mk12140_D03 | mk12154_D03 | 35,382,974 | 37,676,414 | 2.29 | 136.01 | 2.79 | −0.0103 | −0.0047 | 0.46 | 3.63 | PD | NH | ||
| SLW | qSLW_D03_110 | D03 | mk12142_D03 | mk12152_D03 | 36,697,656 | 37,665,167 | 0.97 | 135.01 | 5.12 | −0.0143 | −0.0002 | 0.01 | 7.6 | A | NH | |
| qSLW_D03_ck.1 | D03 | mk12140_D03 | mk12154_D03 | 35,382,974 | 37,676,414 | 2.29 | 136.01 | 2.64 | −0.0074 | 0.0168 | 2.27 | 4.83 | OD | NH | ||
| qSLW_D03_ck.2 | D03 | mk12158_D03 | mk12165_D03 | 37,938,158 | 40,254,875 | 2.32 | 151.21 | 3.58 | −0.0053 | 0.0223 | 4.21 | 4.28 | OD | NH | ||
| qSLW_D03_150 | D03 | mk12140_D03 | mk12154_D03 | 35,382,974 | 37,676,414 | 2.29 | 135.01 | 3.27 | −0.0117 | −0.007 | 0.6 | 3.69 | PD | NH | ||
| qSLW_D03_ck.3 | D03 | mk12162_D03 | mk12186_D03 | 40,254,814 | 42,120,184 | 1.87 | 157.61 | 2.81 | −0.0034 | 0.0203 | 5.97 | 2.72 | OD | NH | ||
| 6 | DLW | qDLW_D06_110 | D06 | mk13340_D06 | mk13562_D06 | 11,196,129 | 23,393,584 | 12.2 | 59.21 | 4.21 | −0.0008 | 0.002 | 2.5 | 5.85 | OD | NH |
| qDLW_D06_ck | D06 | mk17780 | mk13340_D06 | 982 | 11,196,129 | 11.2 | 59.21 | 3.25 | −0.0006 | 0.002 | 3.33 | 4.29 | OD | NH | ||
| SLW | qSLW_D06_ck | D06 | mk13340_D06 | mk13562_D06 | 11,196,129 | 23,393,584 | 12.2 | 59.21 | 2.72 | −0.0071 | 0.0146 | 2.06 | 4.4 | OD | NH | |
| qSLW_D06_110 | D06 | mk13340_D06 | mk13562_D06 | 11,196,129 | 23,393,584 | 12.2 | 59.21 | 3.14 | −0.006 | 0.0215 | 3.58 | 3.79 | OD | NH | ||
| 7 | FW | qFW_D08_150 | D08 | MulMa513_D08 | mk16015_D08 | 54,937,777 | 59,655,801 | 4.72 | 200.81 | 2.66 | 0.0095 | −0.0096 | 1.01 | 4.71 | D | CCRI35 |
| qFW_D08_110 | D08 | MulMa512_D08 | mk16022_D08 | 54,681,233 | 59,878,825 | 5.2 | 191.61 | 3.29 | 0.0171 | 0.0115 | 0.67 | 3.76 | PD | CCRI35 | ||
| qFW_D08_110 | D08 | MulMa512_D08 | MulMa517-m_D08 | 54,681,233 | 58,216,493 | 3.54 | 187.61 | 3.12 | 0.0087 | 0.0088 | 1.01 | 2.66 | D | CCRI35 | ||
| 8 | MDA | qMDA_D13_110 | D13 | mk18516_D13 | mk18547_D13 | 41,759,681 | 48,359,057 | 6.6 | 1.31 | 2.78 | −0.0021 | 0.0034 | 1.62 | 4.79 | OD | NH |
| qMDA_D13_150 | D13 | mk11123 | mk7779 | 7462 | 58,175 | 0.05 | 80.21 | 2.93 | 0.001 | 0.0093 | 9.3 | 0.11 | OD | CCRI35 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Diouf, L.; Pan, Z.; He, S.-P.; Gong, W.-F.; Jia, Y.H.; Magwanga, R.O.; Romy, K.R.E.; Or Rashid, H.; Kirungu, J.N.; Du, X. High-Density Linkage Map Construction and Mapping of Salt-Tolerant QTLs at Seedling Stage in Upland Cotton Using Genotyping by Sequencing (GBS). Int. J. Mol. Sci. 2017, 18, 2622. https://doi.org/10.3390/ijms18122622
Diouf L, Pan Z, He S-P, Gong W-F, Jia YH, Magwanga RO, Romy KRE, Or Rashid H, Kirungu JN, Du X. High-Density Linkage Map Construction and Mapping of Salt-Tolerant QTLs at Seedling Stage in Upland Cotton Using Genotyping by Sequencing (GBS). International Journal of Molecular Sciences. 2017; 18(12):2622. https://doi.org/10.3390/ijms18122622
Chicago/Turabian StyleDiouf, Latyr, Zhaoe Pan, Shou-Pu He, Wen-Fang Gong, Yin Hua Jia, Richard Odongo Magwanga, Kimbembe Romesh Eric Romy, Harun Or Rashid, Joy Nyangasi Kirungu, and Xiongming Du. 2017. "High-Density Linkage Map Construction and Mapping of Salt-Tolerant QTLs at Seedling Stage in Upland Cotton Using Genotyping by Sequencing (GBS)" International Journal of Molecular Sciences 18, no. 12: 2622. https://doi.org/10.3390/ijms18122622
APA StyleDiouf, L., Pan, Z., He, S.-P., Gong, W.-F., Jia, Y. H., Magwanga, R. O., Romy, K. R. E., Or Rashid, H., Kirungu, J. N., & Du, X. (2017). High-Density Linkage Map Construction and Mapping of Salt-Tolerant QTLs at Seedling Stage in Upland Cotton Using Genotyping by Sequencing (GBS). International Journal of Molecular Sciences, 18(12), 2622. https://doi.org/10.3390/ijms18122622