Predicting Amyloidogenic Proteins in the Proteomes of Plants
Abstract
1. Introduction
2. Results
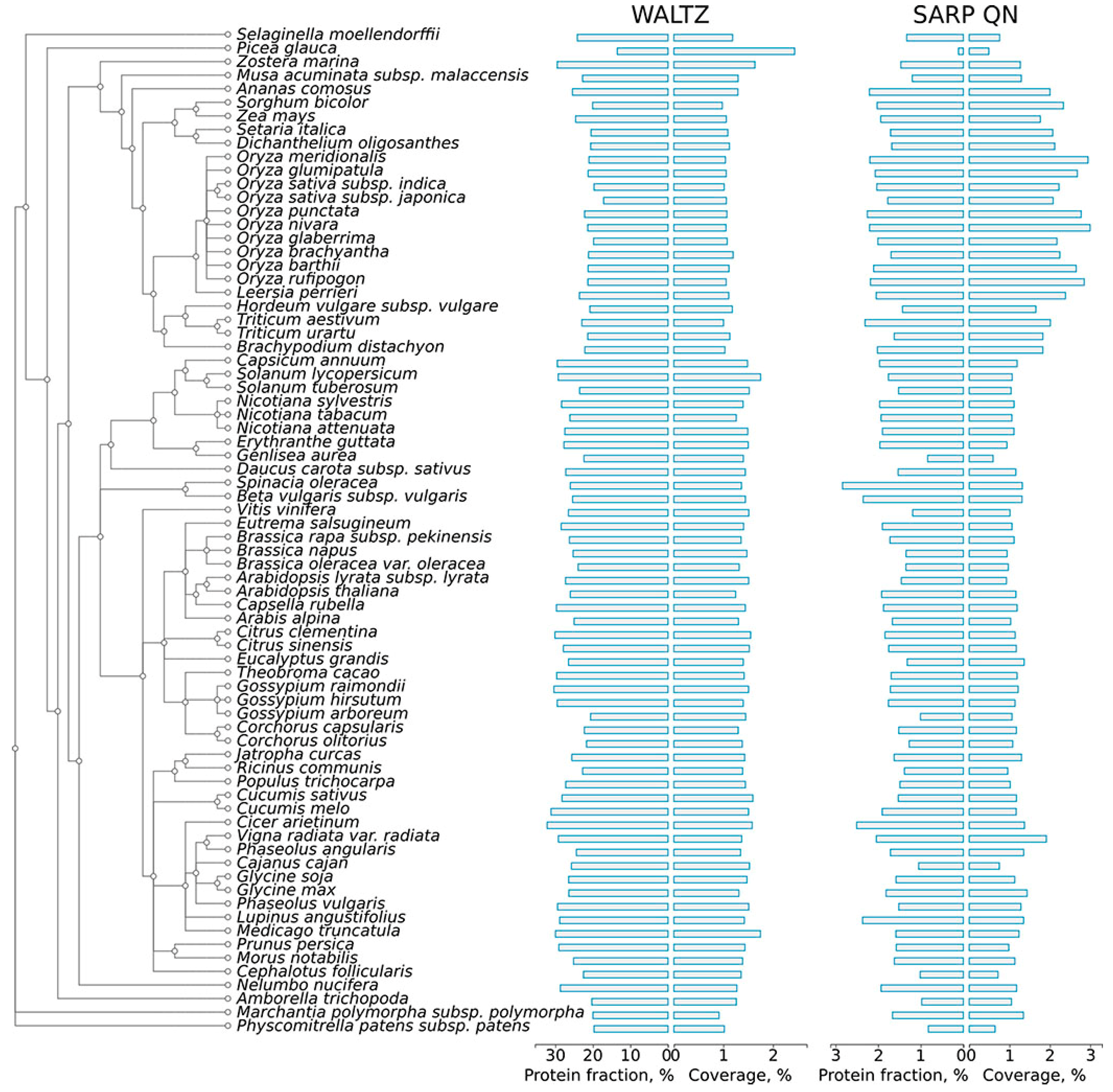
2.1. Abundance of Potentially Amyloidogenic Proteins in the Proteomes of Plants
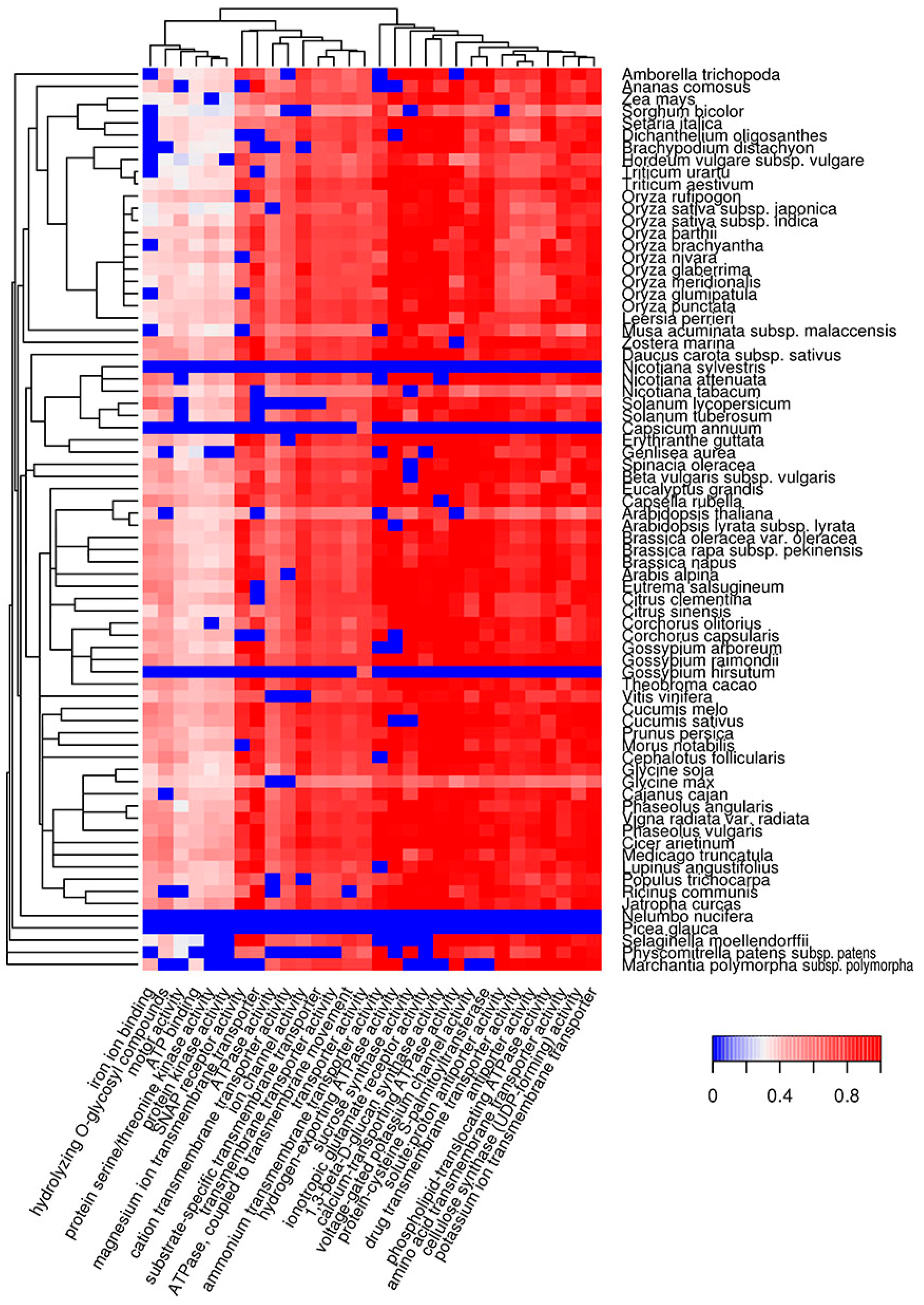
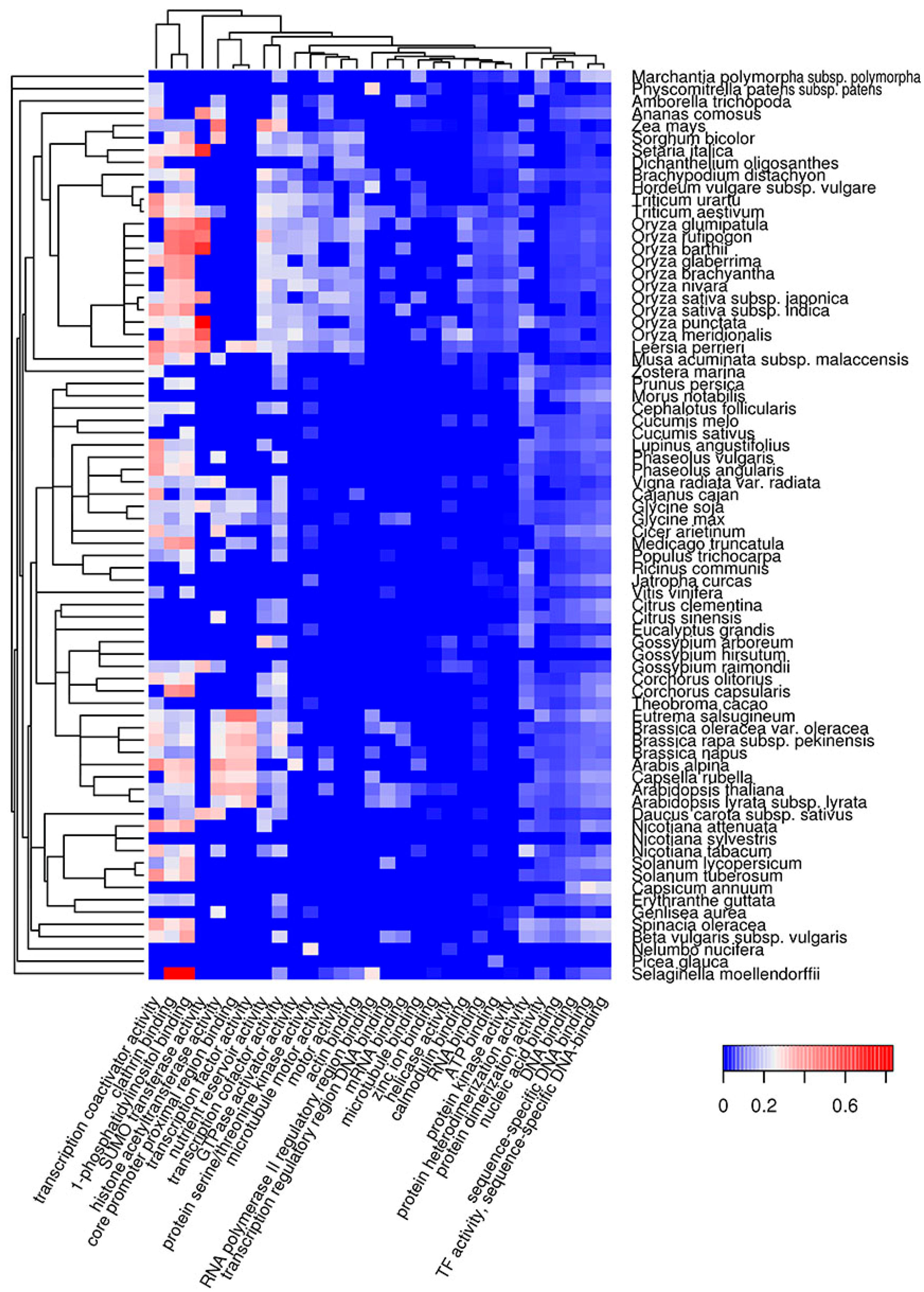
2.2. Molecular Functions of Potentially Amyloidogenic Proteins of Plants
2.3. Subcellular Localization of Potentially Amyloidogenic Proteins of Plants
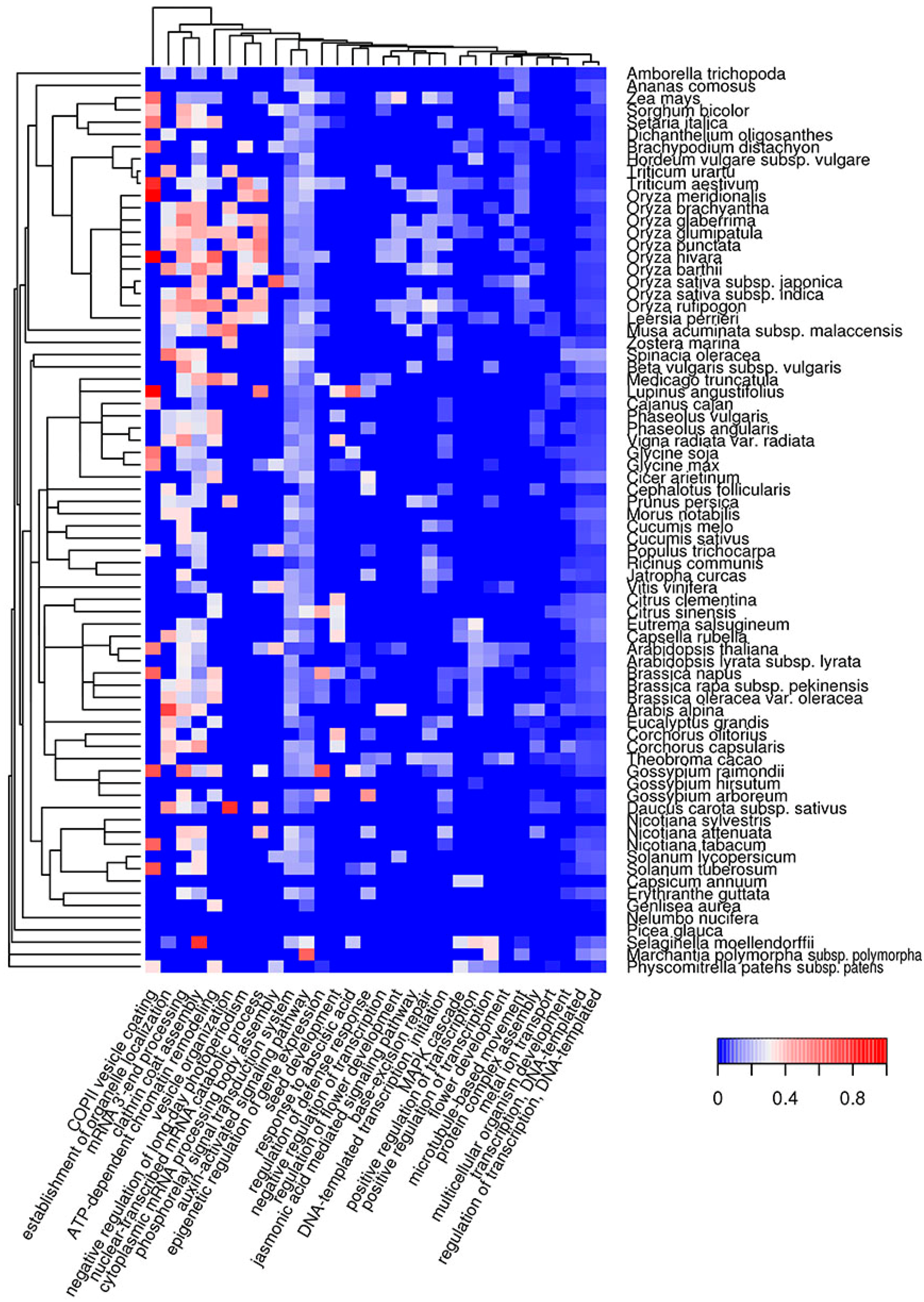
2.4. Biological Processes Implementing Potentially Amyloidogenic Proteins of Plants
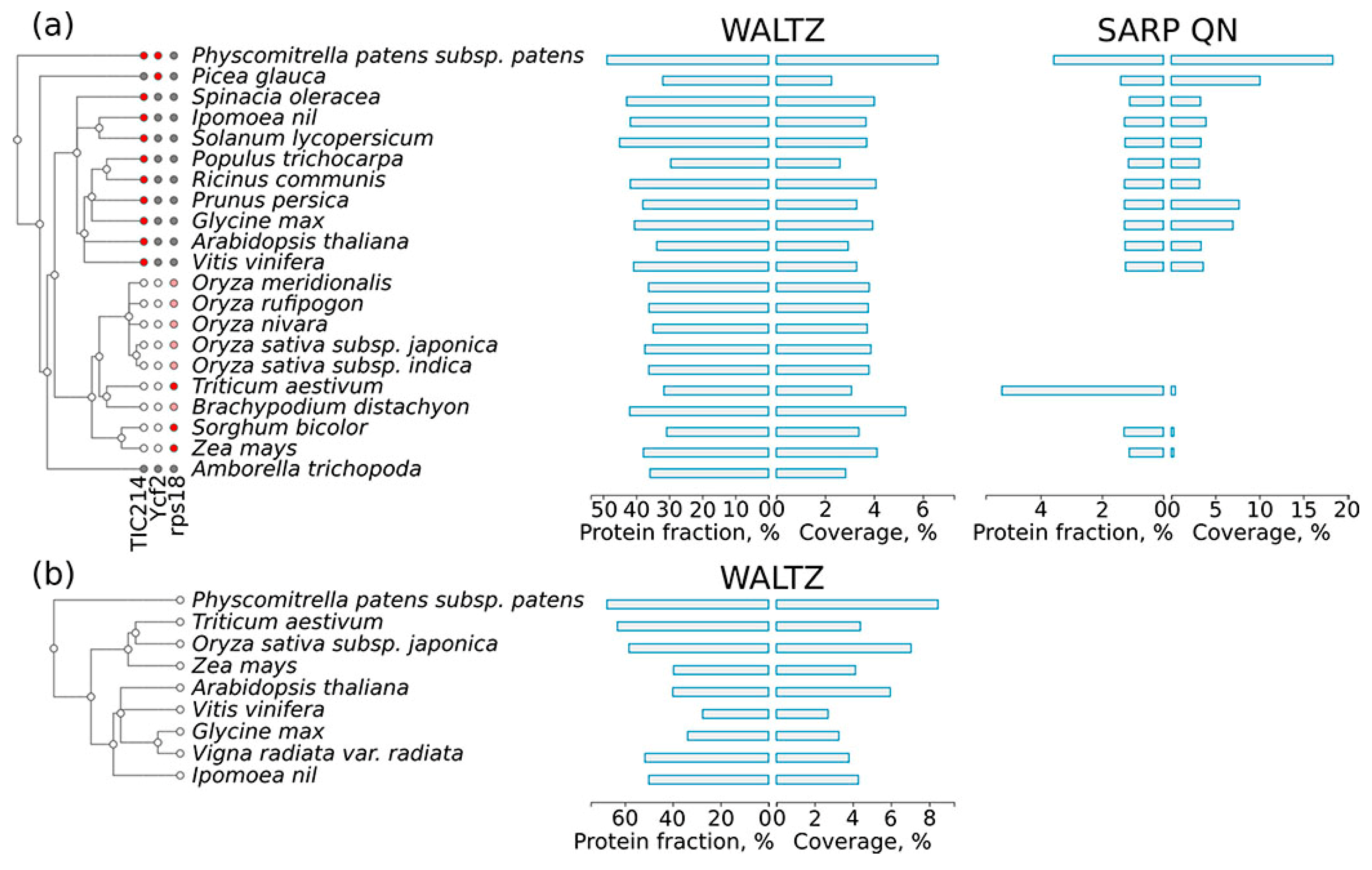
2.5. Amyloidogenic Proteins in the Chloroplast and Mitochondrial Proteomes of Different Plant Species
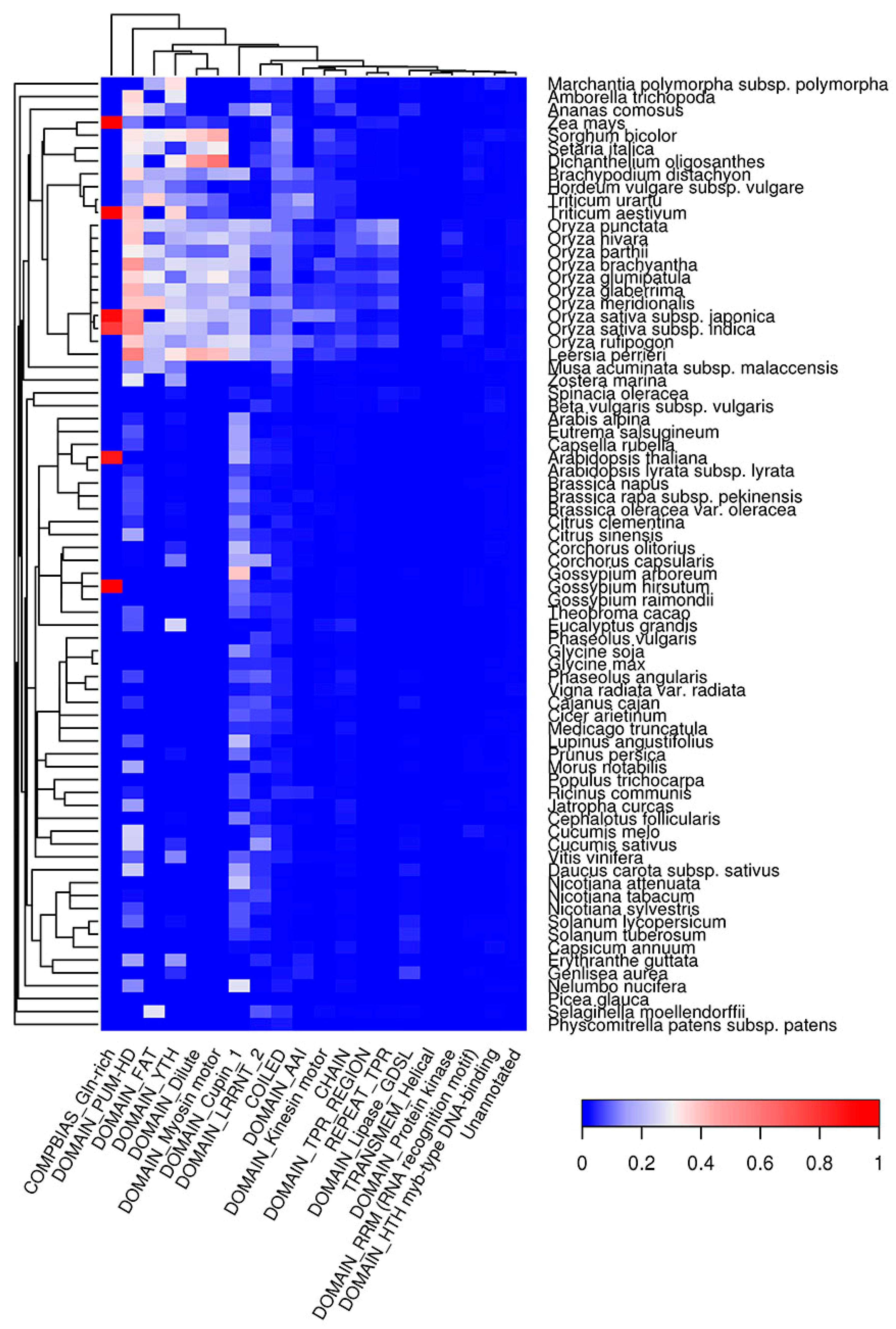
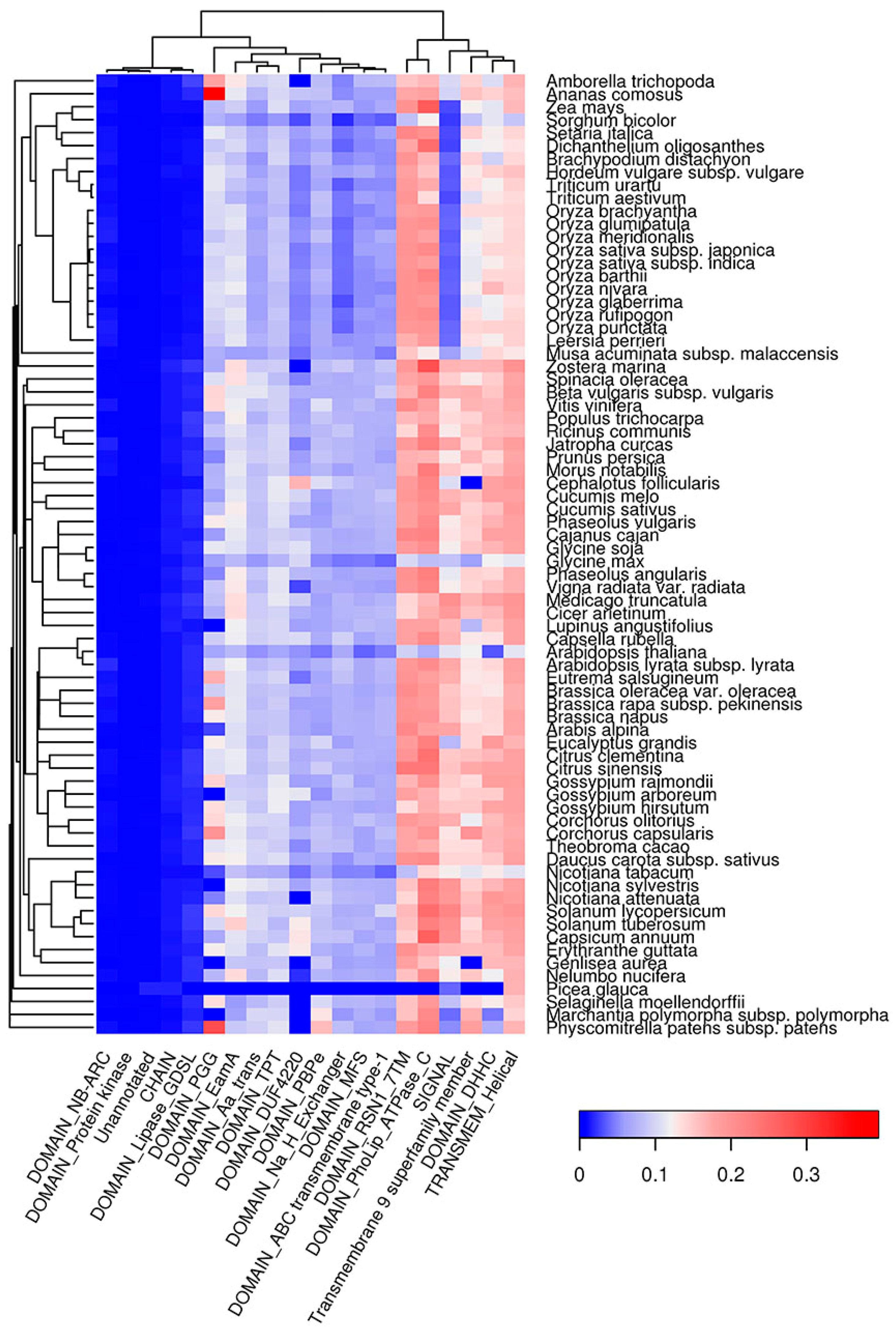
2.6. Co-Occurrence of Potentially Amyloidogenic Regions with the Structural Features of Proteins
3. Discussion
4. Materials and Methods
4.1. Datasets
4.2. Prediction of Amyloidogenic Regions
4.3. Prediction of Compositionally Biased Regions
4.4. GO Term Enrichment Test
4.5. Identification of Potentially Amyloidogenic Proteins in the Proteomes of Organelles
4.6. Analysis of the Association between Amyloidogenic Regions and Different Protein Features
4.7. Analysis of the Abundance of the PFAM Domains among Proteins Containing CBRs
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| AR | Amyloidogenic region |
| SARP | Sequence Analysis Based on the Ranking of Probabilities |
| CBR | compositionally biased region |
References
- Sipe, J.D.; Cohen, A.S. Review: History of the amyloid fibril. J. Struct. Biol. 2000, 130, 88–98. [Google Scholar] [CrossRef] [PubMed]
- Eanes, E.D.; Glenner, G.G. X-ray diffraction studies on amyloid filaments. J. Histochem. Cytochem. 1968, 16, 673–677. [Google Scholar] [CrossRef] [PubMed]
- Tycko, R.; Wickner, R.B. Molecular structures of amyloid and prion fibrils: Consensus versus controversy. Acc. Chem. Res. 2013, 46, 1487–1496. [Google Scholar] [CrossRef] [PubMed]
- Selkoe, D.J.; Ihara, Y.; Salazar, F.J. Alzheimer’s disease: Insolubility of partially purified paired helical filaments in sodium dodecyl sulfate and urea. Science 1982, 215, 1243–1245. [Google Scholar] [CrossRef] [PubMed]
- Hazeki, N.; Tukamoto, T.; Goto, J.; Kanazawa, I. Formic acid dissolves aggregates of an N-terminal huntingtin fragment containing an expanded polyglutamine tract: Applying to quantification of protein components of the aggregates. Biochem. Biophys. Res. Commun. 2000, 277, 386–393. [Google Scholar] [CrossRef] [PubMed]
- Bolton, D.C.; McKinley, M.P.; Prusiner, S.B. Identification of a protein that purifies with the scrapie prion. Science 1982, 218, 1309–1311. [Google Scholar] [CrossRef] [PubMed]
- Kyle, R.A. Amyloidosis: A convoluted story. Br. J. Haematol. 2001, 114, 529–538. [Google Scholar] [CrossRef] [PubMed]
- Virchow, R. Ueber eine im Gehirn und Ruckenmark des Menschen aufgefunde Substanz mit der chemishen Reaction der Cellulose. Virchows Arch. Path. Anat. Physiol. 1854, 6, 135–138. [Google Scholar] [CrossRef]
- Friedreich, N.; Kekule, F.A. Zur Amyloidfrage. Virchows Arch. Path. Anat. Physiol. 1859, 16, 50–65. [Google Scholar] [CrossRef]
- Buxbaum, J.N.; Linke, R.P. A molecular history of the amyloidoses. J. Mol. Biol. 2012, 421, 142–159. [Google Scholar] [CrossRef] [PubMed]
- Chiti, F.; Dobson, C.M. Protein Misfolding, Amyloid Formation, and Human Disease: A Summary of Progress Over the Last Decade. Annu. Rev. Biochem. 2017, 86, 27–68. [Google Scholar] [CrossRef] [PubMed]
- Sipe, J.D.; Benson, M.D.; Buxbaum, J.N.; Ikeda, S.; Merlini, G.; Saraiva, M.J.; Westermark, P. Nomenclature 2014: Amyloid fibril proteins and clinical classification of the amyloidosis. Amyloid 2014, 21, 221–224. [Google Scholar] [CrossRef] [PubMed]
- Pham, C.L.L.; Kwan, A.H.; Sunde, M. Functional amyloid: Widespread in nature, diverse in purpose. Essays Biochem. 2014, 56, 207–219. [Google Scholar] [CrossRef] [PubMed]
- Kelly, J.W.; Balch, W.E. Amyloid as a natural product. J. Cell Biol. 2003, 161, 461–462. [Google Scholar] [CrossRef] [PubMed]
- Chapman, M.R.; Robinson, L.S.; Pinkner, J.S.; Roth, R.; Heuser, J.; Hammar, M.; Normark, S.; Hultgren, S.J. Role of Escherichia coli curli operons in directing amyloid fiber formation. Science 2002, 295, 851–855. [Google Scholar] [CrossRef] [PubMed]
- Bieler, S.; Estrada, L.; Lagos, R.; Baeza, M.; Castilla, J.; Soto, C. Amyloid formation modulates the biological activity of a bacterial protein. J. Biol. Chem. 2005, 280, 26880–26885. [Google Scholar] [CrossRef] [PubMed]
- Claessen, D.; Rink, R.; De Jong, W.; Siebring, J.; De Vreugd, P.; Boersma, F.G.H.; Dijkhuizen, L.; Wosten, H.A.B. A novel class of secreted hydrophobic proteins is involved in aerial hyphae formation in Streptomyces coelicolor by forming amyloid-like fibrils. Genes Dev. 2003, 17, 1714–1726. [Google Scholar] [CrossRef] [PubMed]
- Chimileski, S.; Franklin, M.J.; Papke, R.T. Biofilms formed by the archaeon Haloferax volcanii exhibit cellular differentiation and social motility, and facilitate horizontal gene transfer. BMC Biol. 2014, 12, 65. [Google Scholar] [CrossRef] [PubMed]
- Dueholm, M.S.; Larsen, P.; Finster, K.; Stenvang, M.R.; Christiansen, G.; Vad, B.S.; Boggild, A.; Otzen, D.E.; Nielsen, P.H. The tubular sheaths encasing methanosaeta thermophila filaments are functional amyloids. J. Biol. Chem. 2015, 290, 20590–20600. [Google Scholar] [CrossRef] [PubMed]
- Coustou, V.; Deleu, C.; Saupe, S.; Begueret, J. The protein product of the het-s heterokaryon incompatibility gene of the fungus Podospora anserina behaves as a prion analog. Proc. Natl. Acad. Sci. USA 1997, 94, 9773–9778. [Google Scholar] [CrossRef] [PubMed]
- Holmes, D.L.; Lancaster, A.K.; Lindquist, S.; Halfmann, R. Heritable remodeling of yeast multicellularity by an environmentally responsive prion. Cell 2013, 153, 153–165. [Google Scholar] [CrossRef] [PubMed]
- Gebbink, M.F.B.G.; Claessen, D.; Bouma, B.; Dijkhuizen, L.; Wösten, H.A.B. Amyloids—A functional coat for microorganisms. Nat. Rev. Microbiol. 2005, 3, 333–341. [Google Scholar] [CrossRef] [PubMed]
- Si, K.; Giustetto, M.; Etkin, A.; Hsu, R.; Janisiewicz, A.M.; Miniaci, M.C.; Kim, J.H.; Zhu, H.; Kandel, E.R. A neuronal isoform of cpeb regulates local protein synthesis and stabilizes synapse-specific long-term facilitation in aplysia. Cell 2003, 115, 893–904. [Google Scholar] [CrossRef]
- Majumdar, A.; Cesario, W.C.; White-Grindley, E.; Jiang, H.; Ren, F.; Khan, M.R.; Li, L.; Choi, E.M.L.; Kannan, K.; Guo, F.; et al. Critical role of amyloid-like oligomers of Drosophila Orb2 in the persistence of memory. Cell 2012, 148, 515–529. [Google Scholar] [CrossRef] [PubMed]
- Fowler, D.M.; Koulov, A.V.; Alory-Jost, C.; Marks, M.S.; Balch, W.E.; Kelly, J.W. Functional amyloid formation within mammalian tissue. PLoS Biol. 2006, 4, e6. [Google Scholar] [CrossRef] [PubMed]
- Maji, S.K.; Perrin, M.H.; Sawaya, M.R.; Jessberger, S.; Vadodaria, K.; Rissman, R.A.; Singru, P.S.; Nilsson, K.P.; Simon, R.; Schubert, D.; et al. Functional amyloids as natural storage of peptide hormones in pituitary secretory granules. Science 2009, 325, 328–332. [Google Scholar] [CrossRef] [PubMed]
- Carneiro, K.M.M.; Zhai, H.; Zhu, L.; Horst, J.A.; Sitlin, M.; Nguyen, M.; Wagner, M.; Simpliciano, C.; Milder, M.; Chen, C.-L.; et al. Amyloid-like ribbons of amelogenins in enamel mineralization. Sci. Rep. 2016, 6, 23105. [Google Scholar] [CrossRef] [PubMed]
- Cai, X.; Chen, J.; Xu, H.; Liu, S.; Jiang, Q.X.; Halfmann, R.; Chen, Z.J. Prion-like polymerization underlies signal transduction in antiviral immune defense and inflammasome activation. Cell 2014, 156, 1207–1222. [Google Scholar] [CrossRef] [PubMed]
- Teng, P.K.; Eisenberg, D. Short protein segments can drive a non-fibrillizing protein into the amyloid state. Protein Eng. Des. Sel. 2009, 22, 531–536. [Google Scholar] [CrossRef] [PubMed]
- Von Bergen, M.; Friedhoff, P.; Biernat, J.; Heberle, J.; Mandelkow, E.M.; Mandelkow, E. Assembly of tau protein into Alzheimer paired helical filaments depends on a local sequence motif ((306)VQIVYK(311)) forming β structure. Proc. Natl. Acad. Sci. USA 2000, 97, 5129–5134. [Google Scholar] [CrossRef] [PubMed]
- López de la Paz, M.; Serrano, L. Sequence determinants of amyloid fibril formation. Proc. Natl. Acad. Sci. USA 2004, 101, 87–92. [Google Scholar] [CrossRef] [PubMed]
- Esteras-Chopo, A.; Serrano, L.; López de la Paz, M. The amyloid stretch hypothesis: Recruiting proteins toward the dark side. Proc. Natl. Acad. Sci. USA 2005, 102, 16672–16677. [Google Scholar] [CrossRef] [PubMed]
- Kadnar, M.L.; Articov, G.; Derkatch, I.L. Distinct type of transmission barrier revealed by study of multiple prion determinants of Rnq1. PLoS Genet. 2010, 6, e1000824. [Google Scholar] [CrossRef] [PubMed]
- Das, S.; Pal, U.; Das, S.; Bagga, K.; Roy, A.; Mrigwani, A.; Maiti, N.C. Sequence complexity of amyloidogenic regions in intrinsically disordered human proteins. PLoS ONE 2014, 9. [Google Scholar] [CrossRef] [PubMed]
- Das, A.K.; Pandit, R.; Maiti, S. Effect of amyloids on the vesicular machinery: Implications for somatic neurotransmission. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2015, 370. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, A.B.; Kajava, A.V. Breaking the amyloidogenicity code: Methods to predict amyloids from amino acid sequence. FEBS Lett. 2013, 587, 1089–1095. [Google Scholar] [CrossRef] [PubMed]
- Maurer-Stroh, S.; Debulpaep, M.; Kuemmerer, N.; Lopez de la Paz, M.; Martins, I.C.; Reumers, J.; Morris, K.L.; Copland, A.; Serpell, L.; Serrano, L.; et al. Exploring the sequence determinants of amyloid structure using position-specific scoring matrices. Nat. Methods 2010, 7, 237–242. [Google Scholar] [CrossRef] [PubMed]
- Nizhnikov, A.A.; Antonets, K.S.; Bondarev, S.A.; Inge-Vechtomov, S.G.; Derkatch, I.L. Prions, amyloids, and RNA: Pieces of a puzzle. Prion 2016, 10, 182–206. [Google Scholar] [CrossRef] [PubMed]
- Scherzinger, E.; Lurz, R.; Turmaine, M.; Mangiarini, L.; Hollenbach, B.; Hasenbank, R.; Bates, G.P.; Davies, S.W.; Lehrach, H.; Wanker, E.E. Huntingtin-encoded polyglutamine expansions form amyloid-like protein aggregates in vitro and in vivo. Cell 1997, 90, 549–558. [Google Scholar] [CrossRef]
- Michelitsch, M.D.; Weissman, J.S. A census of glutamine/asparagine-rich regions: Implications for their conserved function and the prediction of novel prions. Proc. Natl. Acad. Sci. USA 2000, 97, 11910–11915. [Google Scholar] [CrossRef] [PubMed]
- Colaco, M.; Park, J.; Blanch, H. The kinetics of aggregation of poly-glutamic acid based polypeptides. Biophys. Chem. 2008, 136, 74–86. [Google Scholar] [CrossRef] [PubMed]
- Harrison, P.M.; Gerstein, M. A method to assess compositional bias in biological sequences and its application to prion-like glutamine/asparagine-rich domains in eukaryotic proteomes. Genome Biol. 2003, 4, R40. [Google Scholar] [CrossRef] [PubMed]
- Antonets, K.S.; Nizhnikov, A.A. SARP: A novel algorithm to assess compositional biases in protein sequences. Evol. Bioinform. Online 2013, 9, 263–273. [Google Scholar] [CrossRef] [PubMed]
- Beerten, J.; Van Durme, J.; Gallardo, R.; Capriotti, E.; Serpell, L.; Rousseau, F.; Schymkowitz, J. WALTZ-DB: A benchmark database of amyloidogenic hexapeptides. Bioinformatics 2014, 31, 1698–1700. [Google Scholar] [CrossRef] [PubMed]
- Alberti, S.; Halfmann, R.; King, O.; Kapila, A.; Lindquist, S. A systematic survey identifies prions and illuminates sequence features of prionogenic proteins. Cell 2009, 137, 146–158. [Google Scholar] [CrossRef] [PubMed]
- Chakrabortee, S.; Kayatekin, C.; Newby, G.A.; Mendillo, M.L.; Lancaster, A.; Lindquist, S. Luminidependens (LD) is an Arabidopsis protein with prion behavior. Proc. Natl. Acad. Sci. USA 2016, 113, 6065–6070. [Google Scholar] [CrossRef] [PubMed]
- Yang, W.; Willemse, J.; Sawyer, E.B.; Lou, F.; Gong, W.; Zhang, H.; Gras, S.L.; Claessen, D.; Perrett, S. The propensity of the bacterial rodlin protein RdlB to form amyloid fibrils determines its function in Streptomyces coelicolor. Sci. Rep. 2017, 7, 42867. [Google Scholar] [CrossRef] [PubMed]
- Macindoe, I.; Kwan, A.H.; Ren, Q.; Morris, V.K.; Yang, W.; Mackay, J.P.; Sunde, M. Self-assembly of functional, amphipathic amyloid monolayers by the fungal hydrophobin EAS. Proc. Natl. Acad. Sci. USA 2012, 109, 5152–5153. [Google Scholar] [CrossRef] [PubMed]
- Gour, S.; Kaushik, V.; Kumar, V.; Bhat, P.; Yadav, S.C.; Yadav, J.K. Antimicrobial peptide (Cn-AMP2) from liquid endosperm of Cocos nucifera forms amyloid-like fibrillar structure. J. Pept. Sci. 2016, 22, 201–207. [Google Scholar] [CrossRef] [PubMed]
- Berthelot, K.; Lecomte, S.; Coulary-Salin, B.; Bentaleb, A.; Peruch, F. Hevea brasiliensis prohevein possesses a conserved C-terminal domain with amyloid-like properties in vitro. Biochim. Biophys. Acta 2016, 1864, 388–399. [Google Scholar] [CrossRef] [PubMed]
- Antonets, K.S.; Nizhnikov, A.A. Amyloids and prions in plants: Facts and perspectives. Prion 2017, 11, 300–312. [Google Scholar] [CrossRef] [PubMed]
- Wendel, J.F. New World tetraploid cottons contain Old World cytoplasm. Proc. Natl. Acad. Sci. USA 1989, 86, 4132–4136. [Google Scholar] [CrossRef] [PubMed]
- Wickner, R.B.; Shewmaker, F.P.; Bateman, D.A.; Edskes, H.K.; Gorkovskiy, A.; Dayani, Y.; Bezsonov, E.E. Yeast prions: Structure, biology, and prion-handling systems. Microbiol. Mol. Biol. Rev. 2015, 79, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Nizhnikov, A.A.; Antonets, K.S.; Inge-Vechtomov, S.G. Amyloids: From pathogenesis to function. Biochemistry 2015, 80, 1127–1144. [Google Scholar] [CrossRef] [PubMed]
- Fowler, D.M.; Koulov, A.V.; Balch, W.E.; Kelly, J.W. Functional amyloid—From bacteria to humans. Trends Biochem. Sci. 2007, 32, 217–224. [Google Scholar] [CrossRef] [PubMed]
- Balakireva, A.V.; Zamyatnin, A.A. Properties of gluten intolerance: Gluten structure, evolution, pathogenicity and detoxification capabilities. Nutrients 2016, 8, 644. [Google Scholar] [CrossRef] [PubMed]
- Jackson, P.; Boulter, D.; Thurman, D.A. A comparison of some properties of vicilin and legumin isolated from seeds of Pisum sativum, Vicia faba and Cicer arietinum. New Phytol. 1969, 68, 25–33. [Google Scholar] [CrossRef]
- Kikuchi, S.; Bédard, J.; Hirano, M.; Hirabayashi, Y.; Oishi, M.; Imai, M.; Takase, M.; Ide, T.; Nakai, M. Uncovering the protein translocon at the chloroplast inner envelope membrane. Science 2013, 339, 571–574. [Google Scholar] [CrossRef] [PubMed]
- De Vries, J.; Sousa, F.L.; Bölter, B.; Soll, J.; Gould, S.B. YCF1: A Green TIC? Plant Cell 2015, 27, 1827–1833. [Google Scholar] [CrossRef] [PubMed]
- Kobe, B.; Kajava, A.V. The leucine-rich repeat as a protein recognition motif. Curr. Opin. Struct. Biol. 2001, 11, 725–732. [Google Scholar] [CrossRef]
- Finn, R.D.; Coggill, P.; Eberhardt, R.Y.; Eddy, S.R.; Mistry, J.; Mitchell, A.L.; Potter, S.C.; Punta, M.; Qureshi, M.; Sangrador-Vegas, A.; et al. The Pfam protein families database: Towards a more sustainable future. Nucleic Acids Res. 2016, 44, D279–D285. [Google Scholar] [CrossRef] [PubMed]
- Danoff, E.J.; Fleming, K.G. Aqueous, Unfolded OmpA forms amyloid-like fibrils upon self-association. PLoS ONE 2015, 10. [Google Scholar] [CrossRef] [PubMed]
- Joseph Sahaya Rajan, J.; Chinnappan Santiago, T.; Singaravel, R.; Ignacimuthu, S. Outer membrane protein C (OmpC) of Escherichia coli induces neurodegeneration in mice by acting as an amyloid. Biotechnol. Lett. 2016, 38, 689–700. [Google Scholar] [CrossRef] [PubMed]
- Nawrot, R.; Barylski, J.; Nowicki, G.; Broniarczyk, J.; Buchwald, W.; Goździcka-Józefiak, A. Plant antimicrobial peptides. Folia Microbiol. 2014, 59, 181–196. [Google Scholar] [CrossRef] [PubMed]
- Garvey, M.; Meehan, S.; Gras, S.L.; Schirra, H.J.; Craik, D.J.; van der Weerden, N.L.; Anderson, M.A.; Gerrard, J.A.; Carver, J.A. A radish seed antifungal peptide with a high amyloid fibril-forming propensity. Biochim. Biophys. Acta Proteins Proteom. 2013, 1834, 1615–1623. [Google Scholar] [CrossRef] [PubMed]
- Wiggins, R.C. Prion Stability and infectivity in the environment. Neurochem. Res. 2009, 34, 158–168. [Google Scholar] [CrossRef] [PubMed]
- Munialo, C.D.; Martin, A.H.; van der Linden, E.; de Jongh, H.H.J. Fibril formation from pea protein and subsequent gel formation. J. Agric. Food Chem. 2014, 62, 2418–2427. [Google Scholar] [CrossRef] [PubMed]
- Tang, C.H.; Wang, C.S. Formation and characterization of amyloid-like fibrils from soy β-conglycinin and glycinin. J. Agric. Food Chem. 2010, 58, 11058–11066. [Google Scholar] [CrossRef] [PubMed]
- Ridgley, D.M.; Ebanks, K.C.; Barone, J.R. Peptide mixtures can self-assemble into large amyloid fibers of varying size and morphology. Biomacromolecules 2011, 12, 3770–3779. [Google Scholar] [CrossRef] [PubMed]
- Podrabsky, J.E.; Carpenter, J.F.; Hand, S.C. Survival of water stress in annual fish embryos: Dehydration avoidance and egg envelope amyloid fibers. Am. J. Physiol. Regul. Integr. Comp. Physiol. 2001, 280, R123–R131. [Google Scholar] [PubMed]
- Wolf, P.G.; Der, J.P.; Duffy, A.M.; Davidson, J.B.; Grusz, A.L.; Pryer, K.M. The evolution of chloroplast genes and genomes in ferns. Plant Mol. Biol. 2011, 76, 251–261. [Google Scholar] [CrossRef] [PubMed]
- Antonets, K.S.; Volkov, K.V.; Maltseva, A.L.; Arshakian, L.M.; Galkin, A.P.; Nizhnikov, A.A. Proteomic analysis of Escherichia coli protein fractions resistant to solubilization by ionic detergents. Biochemistry 2016, 81, 34–46. [Google Scholar] [CrossRef] [PubMed]
- Nizhnikov, A.A.; Alexandrov, A.I.; Ryzhova, T.A.; Mitkevich, O.V.; Dergalev, A.A.; Ter-Avanesyan, M.D.; Galkin, A.P. Proteomic screening for amyloid proteins. PLoS ONE 2014, 9, e116003. [Google Scholar] [CrossRef] [PubMed]
- Nizhnikov, A.A.; Ryzhova, T.A.; Volkov, K.V.; Zadorsky, S.P.; Sopova, J.V.; Inge-Vechtomov, S.G.; Galkin, A.P. Interaction of Prions Causes Heritable Traits in Saccharomyces cerevisiae. PLOS Genet. 2016, 12, e1006504. [Google Scholar] [CrossRef] [PubMed]
- Kryndushkin, D.; Pripuzova, N.; Burnett, B.G.; Shewmaker, F. Non-targeted identification of prions and amyloid-forming proteins from yeast and mammalian cells. J. Biol. Chem. 2013, 288, 27100–27111. [Google Scholar] [CrossRef] [PubMed]
- Wan, S.; Duan, Y.; Zou, Q. HPSLPred: An Ensemble Multi-Label Classifier for Human Protein Subcellular Location Prediction with Imbalanced Source. Proteomics 2017, 17. [Google Scholar] [CrossRef] [PubMed]
- Liao, Z.; Wang, X.; Zeng, Y.; Zou, Q. Identification of DEP domain-containing proteins by a machine learning method and experimental analysis of their expression in human HCC tissues. Sci. Rep. 2016, 6, 39655. [Google Scholar] [CrossRef] [PubMed]
- Nightingale, A.; Antunes, R.; Alpi, E.; Bursteinas, B.; Gonzales, L.; Liu, W.; Luo, J.; Qi, G.; Turner, E.; Martin, M. The Proteins API: Accessing key integrated protein and genome information. Nucleic Acids Res. 2017, 45, W539–W544. [Google Scholar] [CrossRef] [PubMed]
- Fisher, R.A. The logic of inductive inference. J. R. Stat. Soc. 1932, 98, 39–82. [Google Scholar] [CrossRef]
- Benjamini, Y.; Hochberg, Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. J. R. Stat. Soc. Ser. B 1995, 57, 289–300. [Google Scholar]
- Alexa, A.; Rahnenfuhrer, J. topGO: Enrichment Analysis for Gene Ontology. R package version 2.28.0. Bioconductor 2016. [Google Scholar] [CrossRef]

| PFAM id | Domain Family Description | Number of PFAM Proteins * | Number of Species with PFAM Proteins | Number of QN-Rich Proteins | Number of Species with QN-Rich PFAM Proteins | Percentage of QN-Rich PFAM Proteins | Percentage of Species with QN-Rich PFAM Proteins |
|---|---|---|---|---|---|---|---|
| PF00190 | Cupin | 3973 | 70 | 302 | 54 | 7.60 | 77.14 |
| PF01559 | Zein seed storage protein | 161 | 3 | 119 | 3 | 73.91 | 100.00 |
| PF13016 | Cys-rich Gliadin N-terminal | 133 | 16 | 121 | 15 | 90.98 | 93.75 |
| PF00234 | Protease inhibitor/seed storage/LTP family | 113 | 28 | 52 | 15 | 46.02 | 53.57 |
| PF01535 | PPR repeat | 18 | 8 | 13 | 5 | 72.22 | 62.50 |
| PF13041 | PPR repeat family | 17 | 8 | 13 | 5 | 76.47 | 62.50 |
| PF04702 | Vicilin N terminal region | 17 | 13 | 13 | 9 | 76.47 | 69.23 |
| PF12854 | PPR repeat | 16 | 7 | 13 | 5 | 81.25 | 71.43 |
| PF03157 | High molecular weight glutenin subunit | 9 | 3 | 7 | 2 | 77.78 | 66.67 |
| PF13639 | Ring finger domain | 7 | 6 | 7 | 6 | 100.00 | 100.00 |
| PF03330 | Lytic transglycolase | 2 | 1 | 1 | 1 | 50.00 | 100.00 |
| PF01357 | Pollen allergen | 2 | 1 | 1 | 1 | 50.00 | 100.00 |
| PF13446 | A repeated domain in UCH-protein | 1 | 1 | 1 | 1 | 100.00 | 100.00 |
| PF03145 | Seven in absentia protein family | 1 | 1 | 1 | 1 | 100.00 | 100.00 |
| Total | 4487 | 70 | 601 | 59 | 13.39 | 84.29 | |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Antonets, K.S.; Nizhnikov, A.A. Predicting Amyloidogenic Proteins in the Proteomes of Plants. Int. J. Mol. Sci. 2017, 18, 2155. https://doi.org/10.3390/ijms18102155
Antonets KS, Nizhnikov AA. Predicting Amyloidogenic Proteins in the Proteomes of Plants. International Journal of Molecular Sciences. 2017; 18(10):2155. https://doi.org/10.3390/ijms18102155
Chicago/Turabian StyleAntonets, Kirill S., and Anton A. Nizhnikov. 2017. "Predicting Amyloidogenic Proteins in the Proteomes of Plants" International Journal of Molecular Sciences 18, no. 10: 2155. https://doi.org/10.3390/ijms18102155
APA StyleAntonets, K. S., & Nizhnikov, A. A. (2017). Predicting Amyloidogenic Proteins in the Proteomes of Plants. International Journal of Molecular Sciences, 18(10), 2155. https://doi.org/10.3390/ijms18102155






