Plasma LncRNA-ATB, a Potential Biomarker for Diagnosis of Patients with Coal Workers’ Pneumoconiosis: A Case-Control Study
Abstract
:1. Introduction
2. Results
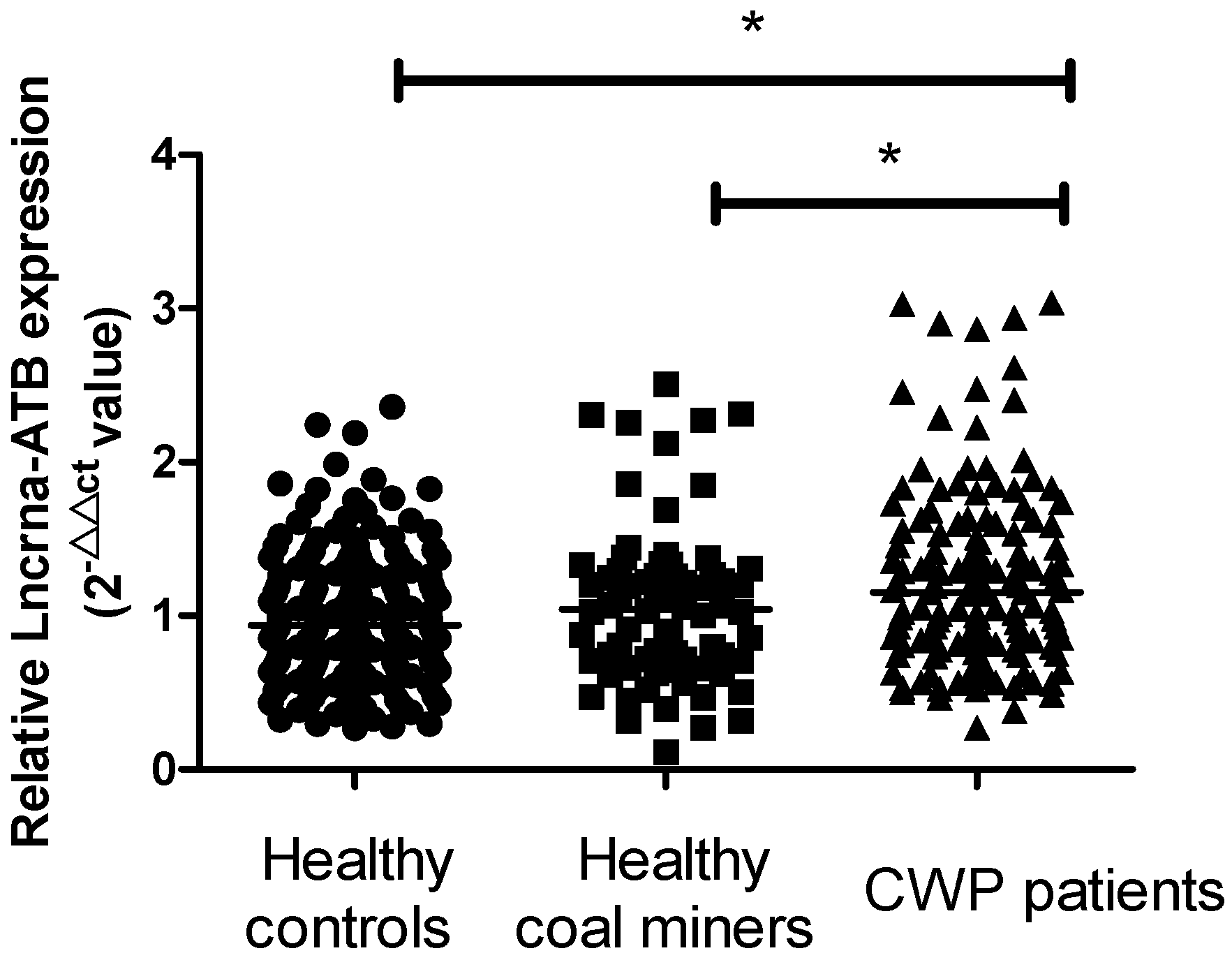
2.1. LncRNA-ATB Expression in the Plasma of All Participants
2.2. Association between LncRNA-ATB Expression and Coal Workers’ Pneumoconiosis (CWP)
2.3. Relationship between LncRNA-ATB Expression and Clinical/Biological Features in CWP Patients
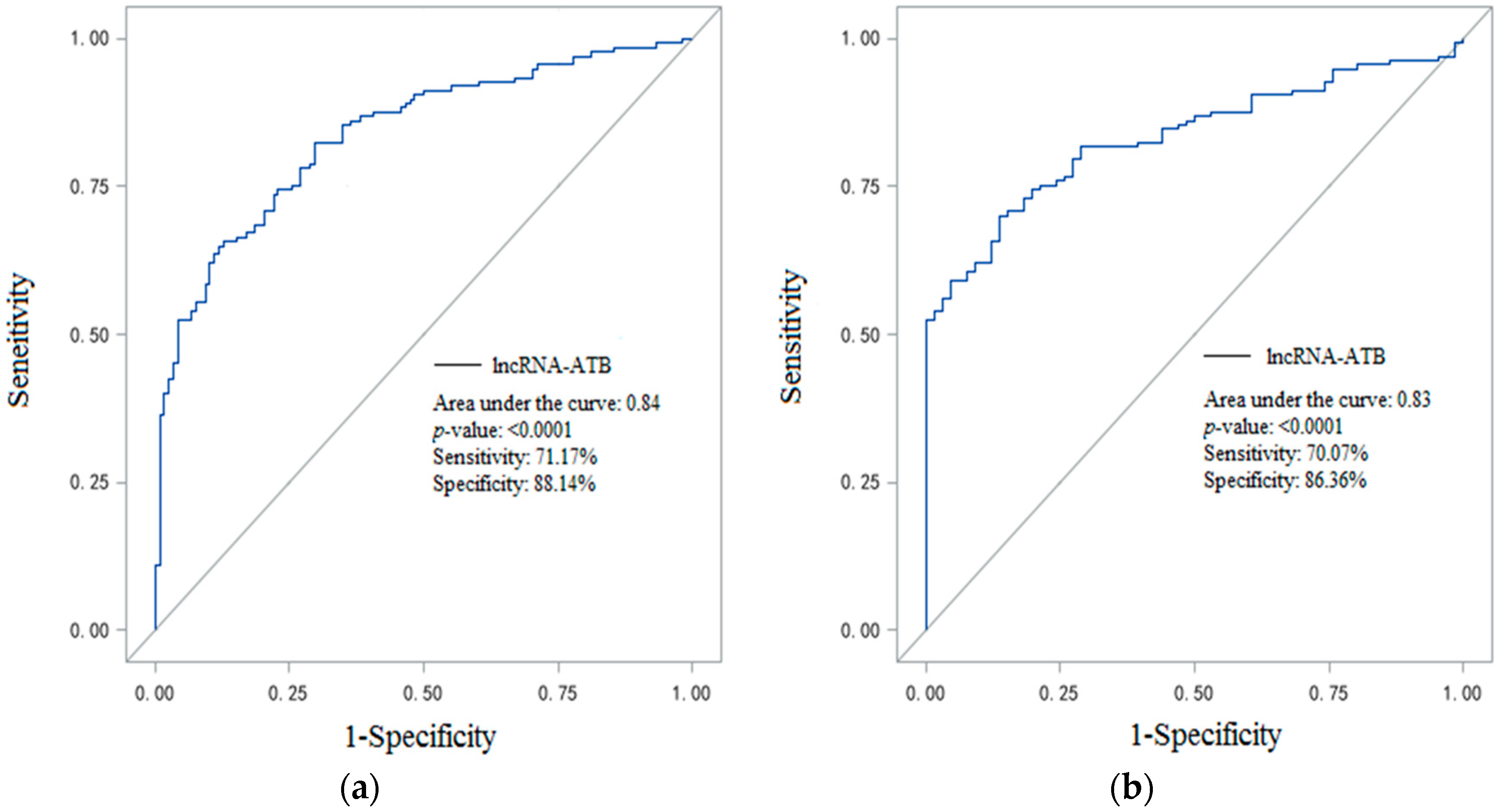
2.4. Plasma LncRNA-ATB Expression Can Be a Potential CWP Biomarker
3. Discussion
4. Materials and Methods
4.1. Study Population
4.2. Total RNA Extraction and Quantitative Real-Time Polymerase Chain Reaction (qRT-PCR)
4.3. Enzyme-Linked Immunosorbent Assay for Plasma Measurements (ELISA)
4.4. Statistical Analysis
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Pascolo, L.; Borelli, V.; Canzonieri, V.; Gianoncelli, A.; Birarda, G.; Bedolla, D.E.; Salome, M.; Vaccari, L.; Calligaro, C.; Cotte, M.; et al. Differential protein folding and chemical changes in lung tissues exposed to asbestos or particulates. Sci. Rep. 2015, 5, 12129. [Google Scholar] [CrossRef] [PubMed]
- Shen, C.H.; Lin, T.Y.; Huang, W.Y.; Chen, H.J.; Kao, C.H. Pneumoconiosis increases the risk of peripheral arterial disease: A nationwide population-based study. Medicine 2015, 94, e911. [Google Scholar] [CrossRef] [PubMed]
- Castranova, V.; Vallyathan, V. Silicosis and coal workers’ pneumoconiosis. Environ. Health Perspect. 2000, 108 (Suppl. S4), 675–684. [Google Scholar] [CrossRef] [PubMed]
- Leung, C.C.; Yu, I.T.; Chen, W. Silicosis. Lancet 2012, 379, 2008–2018. [Google Scholar] [CrossRef]
- Ponting, C.P.; Oliver, P.L.; Reik, W. Evolution and functions of long noncoding RNAs. Cell 2009, 136, 629–641. [Google Scholar] [CrossRef] [PubMed]
- Guttman, M.; Amit, I.; Garber, M.; French, C.; Lin, M.F.; Feldser, D.; Huarte, M.; Zuk, O.; Carey, B.W.; Cassady, J.P.; et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 2009, 458, 223–227. [Google Scholar] [CrossRef] [PubMed]
- Ji, P.; Diederichs, S.; Wang, W.; Boing, S.; Metzger, R.; Schneider, P.M.; Tidow, N.; Brandt, B.; Buerger, H.; Bulk, E.; et al. Malat-1, a novel noncoding RNA, and thymosin β4 predict metastasis and survival in early-stage non-small cell lung cancer. Oncogene 2003, 22, 8031–8041. [Google Scholar] [CrossRef] [PubMed]
- Cao, G.; Zhang, J.; Wang, M.; Song, X.; Liu, W.; Mao, C.; Lv, C. Differential expression of long non-coding RNAs in bleomycin-induced lung fibrosis. Int. J. Mol. Med. 2013, 32, 355–364. [Google Scholar] [PubMed]
- Nakagawa, T.; Endo, H.; Yokoyama, M.; Abe, J.; Tamai, K.; Tanaka, N.; Sato, I.; Takahashi, S.; Kondo, T.; Satoh, K. Large noncoding RNA hotair enhances aggressive biological behavior and is associated with short disease-free survival in human non-small cell lung cancer. Biochem. Biophys. Res. Commun. 2013, 436, 319–324. [Google Scholar] [CrossRef] [PubMed]
- Kondo, M.; Suzuki, H.; Ueda, R.; Osada, H.; Takagi, K.; Takahashi, T.; Takahashi, T. Frequent loss of imprinting of the h19 gene is often associated with its overexpression in human lung cancers. Oncogene 1995, 10, 1193–1198. [Google Scholar] [PubMed]
- Song, X.; Cao, G.; Jing, L.; Lin, S.; Wang, X.; Zhang, J.; Wang, M.; Liu, W.; Lv, C. Analysing the relationship between lncRNA and protein-coding gene and the role of lncRNA as ceRNA in pulmonary fibrosis. J. Cell. Mol. Med. 2014, 18, 991–1003. [Google Scholar] [CrossRef] [PubMed]
- Xie, Z.; Chen, X.; Li, J.; Guo, Y.; Li, H.; Pan, X.; Jiang, J.; Liu, H.; Wu, B. Salivary HOTAIR and PVT1 as novel biomarkers for early pancreatic cancer. Oncotarget 2016, 7. [Google Scholar] [CrossRef] [PubMed]
- Tian, Z.Z.; Guo, X.J.; Zhao, Y.M.; Fang, Y. Decreased expression of long non-coding RNA MEG3 acts as a potential predictor biomarker in progression and poor prognosis of osteosarcoma. Int. J. Clin. Exp. Pathol. 2015, 8, 15138–15142. [Google Scholar] [PubMed]
- Iguchi, T.; Uchi, R.; Nambara, S.; Saito, T.; Komatsu, H.; Hirata, H.; Ueda, M.; Sakimura, S.; Takano, Y.; Kurashige, J.; et al. A long noncoding RNA, lncRNA-ATB, is involved in the progression and prognosis of colorectal cancer. Anticancer Res. 2015, 35, 1385–1388. [Google Scholar] [PubMed]
- Qu, S.; Yang, X.; Song, W.; Sun, W.; Li, X.; Wang, J.; Zhong, Y.; Shang, R.; Ruan, B.; Zhang, Z.; et al. Downregulation of lncRNA-ATB correlates with clinical progression and unfavorable prognosis in pancreatic cancer. Tumour Biol. 2015, 37, 3933–3988. [Google Scholar] [CrossRef] [PubMed]
- Yuan, J.H.; Yang, F.; Wang, F.; Ma, J.Z.; Guo, Y.J.; Tao, Q.F.; Liu, F.; Pan, W.; Wang, T.T.; Zhou, C.C.; et al. A long noncoding RNA activated by TGF-β promotes the invasion-metastasis cascade in hepatocellular carcinoma. Cancer Cell 2014, 25, 666–681. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.H.; Kim, K.W.; Lee, K.E.; Lee, M.J.; Kim, S.K.; Kim, S.H.; Shim, H.S.; Lee, C.Y.; Kim, M.J.; Sohn, M.H.; et al. Transforming growth factor-β 1 in humidifier disinfectant-associated children’s interstitial lung disease. Pediatr. Pulmonol. 2016, 51, 173–182. [Google Scholar] [CrossRef] [PubMed]
- Liu, M.; Chen, J.; Huang, Y.; Ke, J.; Li, L.; Huang, D.; Wu, W. Triptolide alleviates isoprenaline-induced cardiac remodeling in rats via TGF-β1/Smad3 and p38 MAPK signaling pathway. Die Pharm. 2015, 70, 244–250. [Google Scholar]
- Roy, S.; Benz, F.; Vargas Cardenas, D.; Vucur, M.; Gautheron, J.; Schneider, A.; Hellerbrand, C.; Pottier, N.; Alder, J.; Tacke, F.; et al. Mir-30c and mir-193 are a part of the TGF-β-dependent regulatory network controlling extracellular matrix genes in liver fibrosis. J. Dig. Dis. 2015, 16, 513–524. [Google Scholar] [CrossRef] [PubMed]
- Thakur, S.; Viswanadhapalli, S.; Kopp, J.B.; Shi, Q.; Barnes, J.L.; Block, K.; Gorin, Y.; Abboud, H.E. Activation of AMP-activated protein kinase prevents TGF-β1-induced epithelial-mesenchymal transition and myofibroblast activation. Am. J. Pathol. 2015, 185, 2168–2180. [Google Scholar] [CrossRef] [PubMed]
- Rong, Y.; Shen, Y.; Zhang, Z.; Cui, X.; Xiao, L.; Liu, Y.; Luo, X.; Chen, W. Blocking TGF-β expression inhibits silica particle-induced epithelial-mesenchymal transition in human lung epithelial cells. Environ. Toxicol. Pharmacol. 2015, 40, 861–869. [Google Scholar] [CrossRef] [PubMed]
- Guo, L.; Ji, X.; Yang, S.; Hou, Z.; Luo, C.; Fan, J.; Ni, C.; Chen, F. Genome-wide analysis of aberrantly expressed circulating mirnas in patients with coal workers’ pneumoconiosis. Mol. Biol. Rep. 2013, 40, 3739–3747. [Google Scholar] [CrossRef] [PubMed]
- Guttman, M.; Rinn, J.L. Modular regulatory principles of large non-coding RNAs. Nature 2012, 482, 339–346. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Gejman, R.; Mahta, A.; Zhong, Y.; Rice, K.A.; Zhou, Y.; Cheunsuchon, P.; Louis, D.N.; Klibanski, A. Maternally expressed gene 3, an imprinted noncoding RNA gene, is associated with meningioma pathogenesis and progression. Cancer Res. 2010, 70, 2350–2358. [Google Scholar] [CrossRef] [PubMed]
- Gutschner, T.; Diederichs, S. The hallmarks of cancer: A long non-coding RNA point of view. RNA Biol. 2012, 9, 703–719. [Google Scholar] [CrossRef] [PubMed]
- Saito, T.; Kurashige, J.; Nambara, S.; Komatsu, H.; Hirata, H.; Ueda, M.; Sakimura, S.; Uchi, R.; Takano, Y.; Shinden, Y.; et al. A long non-coding RNA activated by transforming growth factor-β is an independent prognostic marker of gastric cancer. Ann. Surg. Oncol. 2015, 22 (Suppl. S3), 915–922. [Google Scholar] [CrossRef] [PubMed]
- Xiong, J.; Liu, Y.; Jiang, L.; Zeng, Y.; Tang, W. High expression of long non-coding RNA lncRNA-ATB is correlated with metastases and promotes cell migration and invasion in renal cell carcinoma. Jpn. J. Clin. Oncol. 2016, 46, 378–384. [Google Scholar] [CrossRef] [PubMed]
- Yue, B.; Qiu, S.; Zhao, S.; Liu, C.; Zhang, D.; Yu, F.; Peng, Z.; Yan, D. LncRNA-ATB mediated E-cadherin repression promotes the progression of colon cancer and predicts poor prognosis. J. Gastroenterol. Hepatol. 2016, 31, 595–603. [Google Scholar] [CrossRef] [PubMed]
- Tang, H.; Gao, L.; Mao, J.; He, H.; Liu, J.; Cai, X.; Lin, H.; Wu, T. Salidroside protects against bleomycin-induced pulmonary fibrosis: Activation of NRF2-antioxidant signaling, and inhibition of NF-κB and TGF-β1/Smad-2/-3 pathways. Cell Stress Chaperones 2016, 21, 239–249. [Google Scholar] [CrossRef] [PubMed]
- Han, Y.Y.; Shen, P.; Chang, W.X. Involvement of epithelial-to-mesenchymal transition and associated transforming growth factor-β/Smad signaling in paraquat-induced pulmonary fibrosis. Mol. Med. Rep. 2015, 12, 7979–7984. [Google Scholar] [CrossRef] [PubMed]
- Zhu, H.Y.; Bai, W.D.; Li, C.; Zheng, Z.; Guan, H.; Liu, J.Q.; Yang, X.K.; Han, S.C.; Gao, J.X.; Wang, H.T.; et al. Knockdown of lncRNA-ATB suppresses autocrine secretion of TGF-β2 by targeting ZNF217 via miR-200c in keloid fibroblasts. Sci. Rep. 2016, 6, 24728. [Google Scholar] [CrossRef] [PubMed]
- The 57,000 Chinese Coal Miners Suffer from Lung Disease Annually. Available online: http://english.peopledaily.com.cn/90001/90782/7196279.html (accessed on 11 November 2010).
- Ministry of Health. Chinese Annual Health Statistical Report in 2010; Ministry of Health of the People’s Republic of China: Beijing, China, 2010.
- Zhou, Y.; Sun, H.; Xie, J.; Song, Y.; Liu, Y.; Huang, X.; Zhou, T.; Rong, Y.; Wu, T.; Yuan, J.; et al. Urinary polycyclic aromatic hydrocarbon metabolites and altered lung function in Wuhan, China. Am. J. Respir. Crit. Care Med. 2016, 193, 835–846. [Google Scholar] [CrossRef] [PubMed]
- Shan, Z.; Bao, W.; Zhang, Y.; Rong, Y.; Wang, X.; Jin, Y.; Song, Y.; Yao, P.; Sun, C.; Hu, F.B.; et al. Interactions between zinc transporter-8 gene (slc30a8) and plasma zinc concentrations for impaired glucose regulation and type 2 diabetes. Diabetes 2014, 63, 1796–1803. [Google Scholar] [CrossRef] [PubMed]
| Variables | Normal Controls (n = 168) | Healthy Coal Miners (n = 72) | CWP Patients (n = 137) | p-Value |
|---|---|---|---|---|
| Age (years, mean ± SD) | 58.83 ± 10.63 | 57.69 ± 10.86 | 59.27 ± 7.88 | 0.65 a |
| Age (n, %) | ||||
| <55 | 45 (26.79) | 54 (75.00) | 40 (29.19) | <0.05 b |
| ≥55 | 123 (73.21) | 18 (25.00) | 97 (70.81) | |
| BMI (kg/m2, mean ± SD) | 24.61 ± 3.33 | 25.50 ± 2.88 | 22.36 ± 3.46 #,* | <0.05 a |
| Net year in dust (n, %) | ||||
| ≤30 | / | 62 (86.11) | 90 (65.69) | <0.05 b |
| >30 | / | 10 (13.89) | 47 (34.31) | |
| Smoking status (n, %) | ||||
| Non-smoking | 74 (44.05) | 27 (37.50) | 40 (29.20) | <0.05 b |
| Smoking | 94 (55.95) | 45 (62.50) | 97 (70.80) | |
| Blood pressure systolic (mm Hg, mean ± SD) | 127.86 ± 21.51 | n.d. | 140.20 ± 23.76 | <0.05 d |
| Blood pressure diastolic (mm Hg, mean ± SD) | 82.85 ± 16.67 | n.d. | 76.20 ± 13.06 | <0.05 d |
| FVC (% PRED FVC) | 90.65 (64.90–120.30) | 93.55 (71.40–109.40) | 66.20 (28.20–104.50) #,* | <0.05 c |
| FEV1 (% PRED FEV1) | 97.25 (67.10–95.44) | 98.65 (89.00–111.20) | 62.00 (17.70–108.40) #,* | <0.05 c |
| FEV1/FVC (% PRED) | 80.75 (68.10–86.83) | 83.94 (63.11–97.15) | 74.81 (43.83–98.64) #,* | <0.05 c |
| TGF-β1 (pg/mL) | 303.41 ± 28.38 | 425.64 ± 33.98 # | 569.99 ± 64.13 #,* | <0.05 a |
| Col-3 (ng/mL) | 67.28 ± 7.79 | 70.26 ± 8.17 | 112.15 ± 9.16 #,* | <0.05 a |
| MMP2 (ng/mL) | 142.27 (114.16–172.00) | 166.73 (157.68–183.63) # | 206.32 (186.32–219.16) #,* | <0.05 c |
| MMP9 (ng/mL) | 55.84 (45.03–68.81) | 98.88 (75.03–112.32) # | 123.68 (105.16–143.27) #,* | <0.05 c |
| Col-1 (ng/mL) | 28.65 (26.53–29.53) | 29.90 (27.28–32.25) | 34.15 (31.90.90–36.53) #,* | <0.05 c |
| Variables | LncRNA-ATB Expression Levels (Fold Change) | Per 1 Log-Unit Increment | p-Value * | ||
|---|---|---|---|---|---|
| First Group <0.7559 | Second Group 0.7559–1.1433 | Third Group >1.1433 | |||
| Healthy controls vs. CWP | |||||
| No. of cases/control subjects | 26/55 | 39/55 | 72/55 | ||
| Model 1 (OR: 95% CI) | 1.00 (referent) | 1.47 (0.79–2.74) | 2.67 (1.49–4.78) | 2.75 (1.69–4.48) | <0.05 |
| Model 2 (OR: 95% CI) | 1.00 (referent) | 1.48 (0.80–2.75) | 2.68 (1.50–4.80) | 2.76 (1.70–4.49) | <0.05 |
| Model 3 (OR: 95% CI) | 1.00 (referent) | 1.43 (0.75–2.74) | 2.34 (1.28–4.30) | 2.56 (1.54–4.27) | <0.05 |
| Model 4 (OR: 95% CI) | 1.00 (referent) | 1.41 (0.73–2.72) | 2.39 (1.29–4.42) | 2.57 (1.52–4.33) | <0.05 |
| Healthy coal miners vs. CWP | <0.7542 | 0.7542–1.2161 | >1.2161 | ||
| No. of cases/control subjects | 49/23 | 75/24 | 85/25 | ||
| Model 1 (OR: 95% CI) | 1.00 (referent) | 1.77 (0.85–3.70) | 2.25 (1.08–4.68) | 1.82 (1.05–3.17) | <0.05 |
| Model 2 (OR: 95% CI) | 1.00 (referent) | 1.86 (0.84–4.15) | 2.25 (1.02–4.97) | 1.83 (1.01–3.35) | <0.05 |
| Model 3 (OR: 95% CI) | 1.00 (referent) | 1.54 (0.64–3.70) | 1.78 (0.74–4.31) | 1.90 (0.97–3.72) | 0.06 |
| Model 4 (OR: 95% CI) | 1.00 (referent) | 1.47 (0.59–3.65) | 2.05 (0.81–5.19) | 2.25 (1.07–4.71) | <0.05 |
| Model 5 (OR: 95% CI) | 1.00 (referent) | 1.65 (0.65–4.18) | 1.90 (0.74–4.89) | 2.17 (1.04–4.53) | <0.05 |
| Variables | LncRNA-ATB | TGF-β1 | MMP-2 | MMP-9 | Col-1 | Col-3 | % PRED VC | % PRED FVC | % PRED FEV1 | FEV1/FVC |
|---|---|---|---|---|---|---|---|---|---|---|
| LncRNA-ATB | 1.00 | 0.30 * | 0.07 | −0.03 | 0.09 | 0.02 | −0.18 * | −0.18 * | −0.10 | 0.05 |
| TGF-β1 | 1.00 | −0.04 | 0.09 | 0.18 | −0.06 | 0.47 * | −0.55 * | −0.50 * | −0.30 * | |
| MMP-2 | 1.00 | 0.04 | 0.09 | −0.03 | −0.02 | −0.04 | −0.08 | −0.06 | ||
| MMP-9 | 1.00 | 0.29 | 0.07 | 0.23 * | −0.20 | −0.29 * | −0.31 * | |||
| Col-1 | 1.00 | 0.14 | 0.28 * | −0.25 * | −0.31 * | −0.20 | ||||
| Col-3 | 1.00 | −0.15 | −0.08 | −0.13 | −0.07 | |||||
| % PRED VC | 1.00 | 0.93 * | 0.88 * | 0.47 * | ||||||
| % PRED FVC | 1.00 | 0.90* | 0.42 * | |||||||
| % PRED FEV1 | 1.00 | 0.75 * | ||||||||
| FEV1/FVC | 1.00 |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ma, J.; Cui, X.; Rong, Y.; Zhou, Y.; Guo, Y.; Zhou, M.; Xiao, L.; Chen, W. Plasma LncRNA-ATB, a Potential Biomarker for Diagnosis of Patients with Coal Workers’ Pneumoconiosis: A Case-Control Study. Int. J. Mol. Sci. 2016, 17, 1367. https://doi.org/10.3390/ijms17081367
Ma J, Cui X, Rong Y, Zhou Y, Guo Y, Zhou M, Xiao L, Chen W. Plasma LncRNA-ATB, a Potential Biomarker for Diagnosis of Patients with Coal Workers’ Pneumoconiosis: A Case-Control Study. International Journal of Molecular Sciences. 2016; 17(8):1367. https://doi.org/10.3390/ijms17081367
Chicago/Turabian StyleMa, Jixuan, Xiuqing Cui, Yi Rong, Yun Zhou, Yanjun Guo, Min Zhou, Lili Xiao, and Weihong Chen. 2016. "Plasma LncRNA-ATB, a Potential Biomarker for Diagnosis of Patients with Coal Workers’ Pneumoconiosis: A Case-Control Study" International Journal of Molecular Sciences 17, no. 8: 1367. https://doi.org/10.3390/ijms17081367
APA StyleMa, J., Cui, X., Rong, Y., Zhou, Y., Guo, Y., Zhou, M., Xiao, L., & Chen, W. (2016). Plasma LncRNA-ATB, a Potential Biomarker for Diagnosis of Patients with Coal Workers’ Pneumoconiosis: A Case-Control Study. International Journal of Molecular Sciences, 17(8), 1367. https://doi.org/10.3390/ijms17081367







