Genome-Wide Identification and Characterization of Carboxypeptidase Genes in Silkworm (Bombyx mori)
Abstract
:1. Introduction
2. Results
2.1. Identification and Characterization of the Carboxypeptidase Family
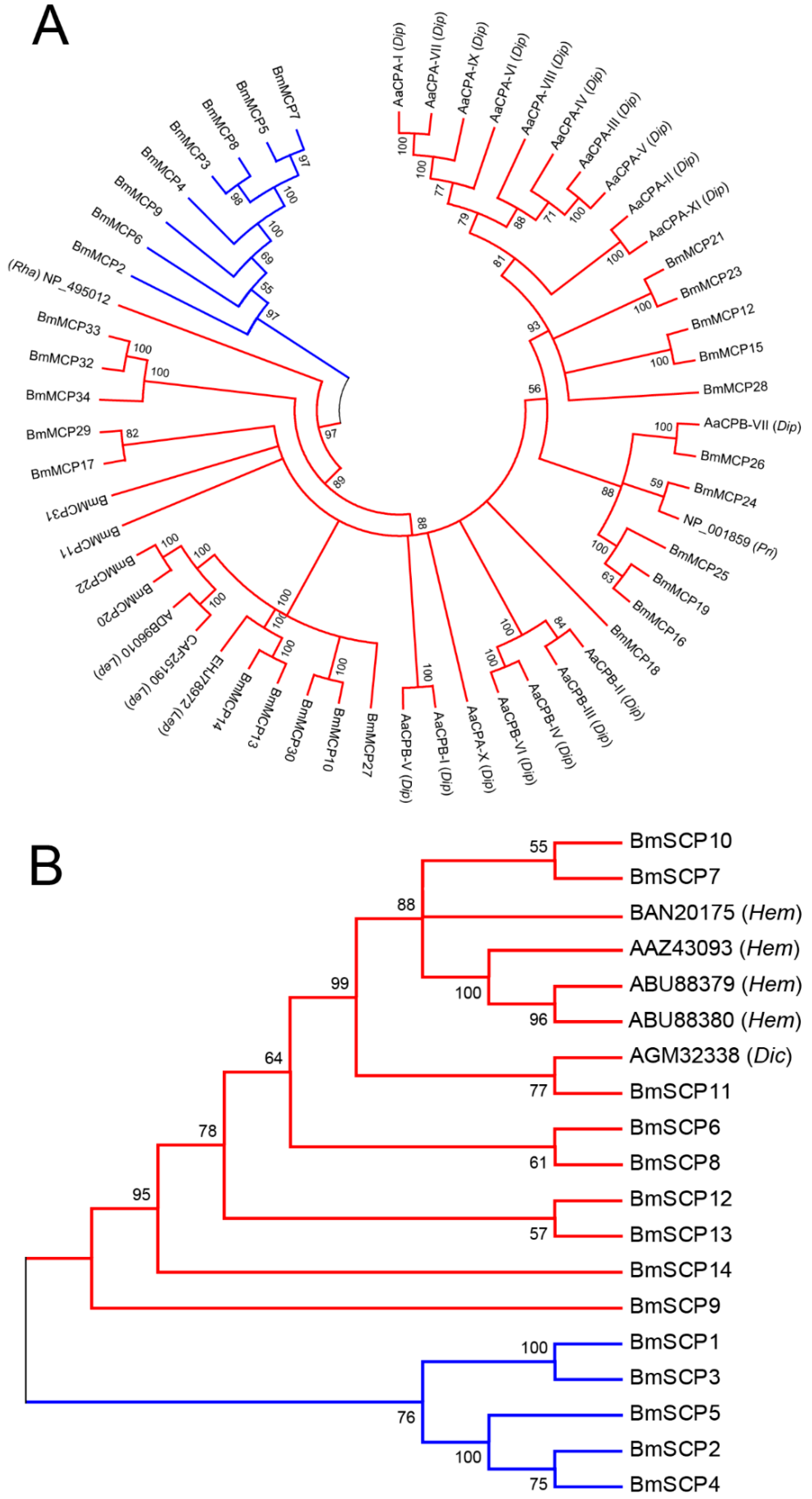
2.2. Phylogenetic Analysis
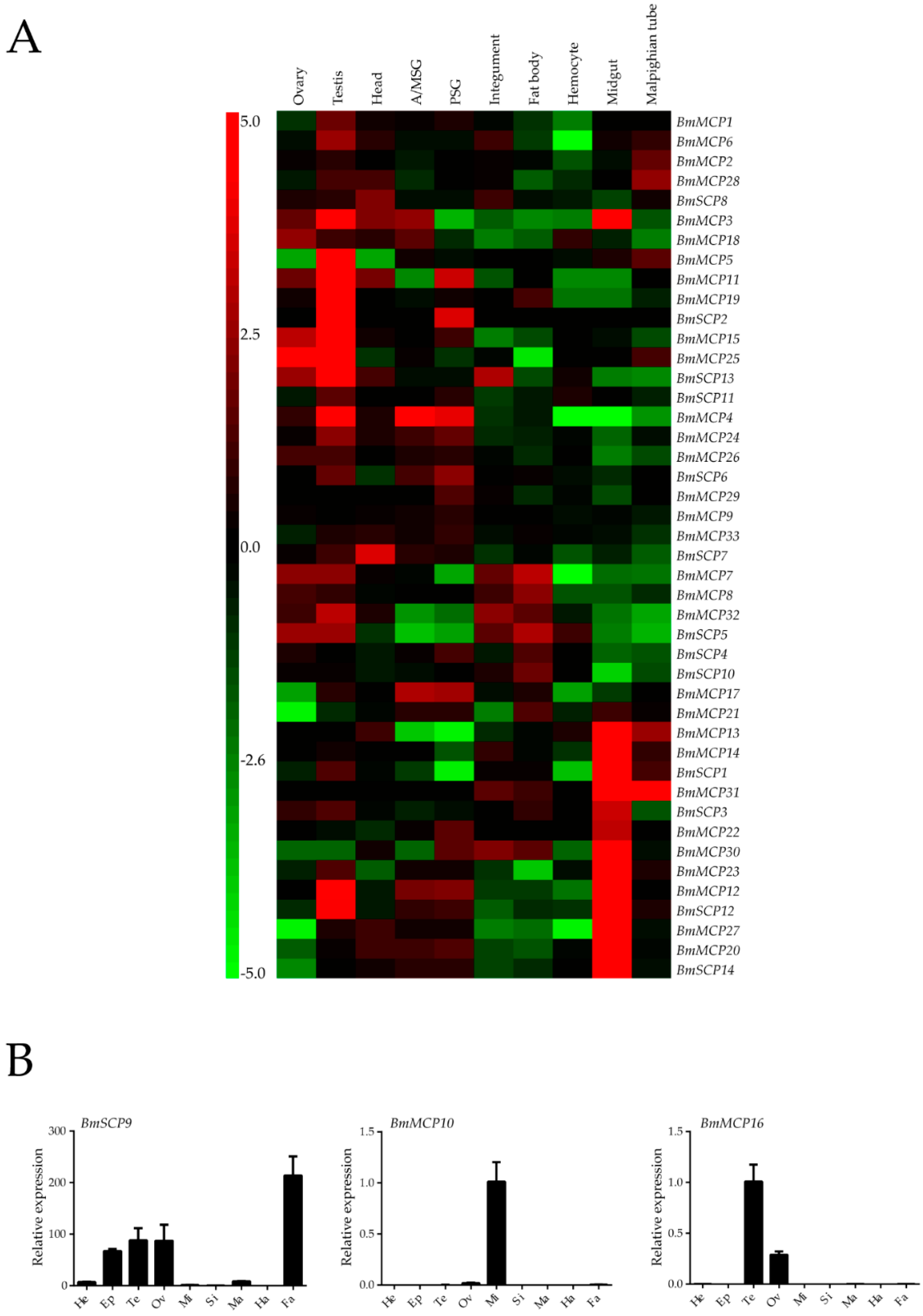
2.3. Expression Profiles
2.4. Carboxypeptidase Expression Profile after Starvation
3. Discussion
4. Materials and Methods
4.1. Biological Materials
4.2. Identification of the Carboxypeptidase Family in Silkworm
4.3. Bioinformatics and Phylogeny Analysis of the Silkworm Carboxypeptidase Family
4.4. Expression Profiles of Silkworm Carboxypeptidase Genes via Whole-Genome Microarrays
4.5. Silkworm Starvation Experiment
4.6. RNA Extraction
4.7. qRT-PCR
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Wang, P.; Li, G.; Kain, W. Characterization and cDNA cloning of midgut carboxypeptidases from Trichoplusia ni. Insect Biochem. Mol. Biol. 2004, 34, 831–843. [Google Scholar] [CrossRef] [PubMed]
- Vendrell, J.; Querol, E.; Avilés, F.X. Metallocarboxypeptidases and their protein inhibitors: Structure, function and biomedical properties. Biochim. Biophys. Acta 2000, 1477, 284–298. [Google Scholar] [CrossRef]
- Ferreira, C.; Rebola, K.G.; Cardoso, C.; Bragatto, I.; Ribeiro, A.F.; Terra, W.R. Insect midgut carboxypeptidases with emphasis on S10 hemipteran and M14 lepidopteran carboxypeptidases. Insect Mol. Biol. 2015, 24, 222–239. [Google Scholar] [CrossRef] [PubMed]
- Skidgel, R.A.; Erdös, E.G. Cellular carboxypeptidases. Immunol. Rev. 1998, 161, 129–141. [Google Scholar] [CrossRef] [PubMed]
- Fricker, L.D. Activation and membrane binding of carboxypeptidase E. J. Cell. Biochem. 1988, 38, 279–289. [Google Scholar] [CrossRef] [PubMed]
- Walter, M.F.; Zeineh, L.L.; Black, B.C.; Mcivor, W.E.; Wright, T.R.F.; Biessmann, H. Catecholamine metabolism and in vitro induction of premature cuticle melanization in wild type and pigmentation mutants of Drosophila melanogaster. Arch. Insect Biochem. Physiol. 1996, 31, 219–233. [Google Scholar] [CrossRef]
- Isoe, J.; Zamora, J.; Miesfeld, R.L. Molecular analysis of the Aedes aEgypti carboxypeptidase gene family. Insect Biochem. Mol. Biol. 2009, 39, 68–73. [Google Scholar] [CrossRef] [PubMed]
- Ramos, A.; Mahowald, A.; Jacobs-Lorena, M. Gut-specific genes from the black fly Simulium vittatum encoding trypsin-like and carboxypeptidase-like proteins. Insect Mol. Biol. 1993, 1, 149–163. [Google Scholar] [CrossRef] [PubMed]
- Yan, J.; Cheng, Q.; Li, C.B.; Aksoy, S. Molecular characterization of three gut genes from Glossina morsitans morsitans: Cathepsin B, zinc-metalloprotease and zinc-carboxypeptidase. Insect Mol. Biol. 2002, 11, 57–65. [Google Scholar] [CrossRef] [PubMed]
- Bown, D.P. Midgut carboxypeptidase from Helicoverpa armigera (Lepidoptera: Noctuidae) larvae: Enzyme characterisation, cDNA cloning and expression. Insect Biochem. Mol. Biol. 1998, 28, 739–749. [Google Scholar] [CrossRef]
- Hegedus, D.; Baldwin, D.; O’Grady, M.; Braun, L.; Gleddie, S.; Sharpe, A.; Lydiate, D.; Erlandson, M. Midgut proteases from Mamestra configurata (Lepidoptera: Noctuidae) larvae: Characterization, cDNA cloning, and expressed sequence tag analysis. Arch. Insect Biochem. Physiol. 2003, 53, 30–47. [Google Scholar] [CrossRef] [PubMed]
- Edwards, M.J.; Lemos, F.J.A.; Donnelly-DOman, M.; Jacobs-Lorena, M. Rapid induction by a blood meal of a carboxypeptidase gene in the gut of the mosQuito Anopheles gambiae. Insect Biochem. Mol. Biol. 1997, 27, 1063–1072. [Google Scholar] [CrossRef]
- Lavazec, C.; Bonnet, S.; Thiery, I.; Boisson, B.; Bourgouin, C. cpbAg1 encodes an active carboxypeptidase B expressed in the midgut of Anopheles gambiae. Insect Mol. Biol. 2005, 14, 163–174. [Google Scholar] [CrossRef] [PubMed]
- Broadway, R.M. Dietary regulation of serine proteinases that are resistant to serine proteinase inhibitors. J. Insect Physiol. 1997, 43, 855–874. [Google Scholar] [CrossRef]
- Waniek, P.J.; Araujo, C.A.; Momoli, M.M.; Azambuja, P.; Jansen, A.M.; Genta, F.A. Serine carboxypeptidases of Triatoma brasiliensis (Hemiptera, Reduviidae): Sequence characterization, expression pattern and activity localization. J. Insect Physiol. 2014, 63, 9–20. [Google Scholar] [CrossRef] [PubMed]
- Ward, C.W. Properties of the major carboxypeptidase in the larvae of the webbing clothes moth, Tineola bisselliella. Biochim. Biophys. Acta 1976, 429, 564–572. [Google Scholar] [CrossRef]
- Zhao, P.; Wang, G.H.; Dong, Z.M.; Duan, J.; Xu, P.Z.; Cheng, T.C.; Xiang, Z.H.; Xia, Q.Y. Genome-wide identification and expression analysis of serine proteases and homologs in the silkworm Bombyx mori. BMC Genom. 2010, 11, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Ote, M.; Mita, K.; Kawasaki, H.; Daimon, T.; Kobayashi, M.; Shimada, T. Identification of molting fluid carboxypeptidase A (MF-CPA) in Bombyx mori. Comp. Biochem. Physiol. 2005, 141, 314–322. [Google Scholar] [CrossRef] [PubMed]
- Li, J.Y.; Moghaddam, S.H.; Chen, J.E.; Chen, M.; Zhong, B.X. Shotgun proteomic analysis on the embryos of silkworm Bombyx mori at the end of organogenesis. Insect Biochem. Mol. Biol. 2010, 40, 293–302. [Google Scholar] [CrossRef] [PubMed]
- Xia, Q.; Zhou, Z.; Lu, C.; Cheng, D.; Dai, F.; Li, B.; Zhao, P.; Zha, X.; Cheng, T.; Chai, C. A draft sequence for the genome of the domesticated silkworm (Bombyx mori). Science 2004, 306, 1937–1940. [Google Scholar] [PubMed]
- Xia, Q.; Guo, Y.; Zhang, Z.; Li, D.; Xuan, Z.; Li, Z.; Dai, F.; Li, Y.; Cheng, D.; Li, R. Complete resequencing of 40 genomes reveals domestication events and genes in silkworm (Bombyx). Science 2009, 326, 433–436. [Google Scholar] [CrossRef] [PubMed]
- Xia, Q.; Li, S.; Feng, Q. Advances in silkworm studies accelerated by the genome sequencing of Bombyx mori. Annu. Rev. Entomol. 2013, 59, 513–536. [Google Scholar] [CrossRef] [PubMed]
- Stein, L.D.; Bao, Z.; Blasiar, D.; Blumenthal, T.; Brent, M.R.; Chen, N.; Chinwalla, A.; Clarke, L.; Clee, C.; Coghlan, A. The genome sequence of Caenorhabditis briggsae: A platform for comparative genomics. PLoS Biol. 2003, 1, e45. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sapio, M.R.; Fricker, L.D. Crboxypeptidases in disease: Insights from peptidomic studies. Proteom. Clin. Appl. 2014, 8, 327–337. [Google Scholar] [CrossRef] [PubMed]
- Zhang, K.X.; Tan, L.; Pellegrini, M.; Zipursky, S.L.; Mcewen, J.M. Rapid changes in the translatome during the conversion of growth cones to synaptic terminals. Cell Rep. 2016, 14, 1258–1271. [Google Scholar] [CrossRef] [PubMed]
- Houard, X.; Williams, T.A.; Michaud, A.; Dani, P.; Isaac, R.E.; Shirras, A.D.; Coates, D.; Corvol, P. The Drosophila melanogaster-related angiotensin-I-converting enzymes acer and ance–Distinct enzymic characteristics and alternative expression during pupal development. Eur. J. Biochem. 1998, 257, 599–606. [Google Scholar] [CrossRef] [PubMed]
- Burnham, S.; Smith, J.A.; Lee, A.J.; Isaac, R.E.; Shirras, A.D. The angiotensin-converting enzyme (ACE) gene family of Anopheles gambiae. BMC Genom. 2005, 6, 172. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fraga-Silva, R.A.; Costa-Fraga, F.P.; Murça, T.M.; Moraes, P.L.; Augusto, M.L.; Lautner, R.Q.; Castro, C.H.; Soares, C.M.A.; Borges, C.L.; Ana Paula, N. Angiotensin-converting enzyme 2 activation improves endothelial function. Hypertension 2013, 61, 1233–1238. [Google Scholar] [CrossRef] [PubMed]
- Kumar, A.; Sharma, A.; Sharma, R.; Gakhar, S.K. Identification, characterization and analysis of expression of gene encoding carboxypeptidase A in Anopheles culicifacies A (Diptera: Culicidae). Acta Trop. 2014, 139, 123–130. [Google Scholar] [CrossRef] [PubMed]
- Cool, D.R.; Loh, Y.P. Carboxypeptidase E is a sorting receptor for prohormones: Binding and kinetic studies. Mol. Cell. Endocrinol. 1998, 139, 7–13. [Google Scholar] [CrossRef]
- Hourdou, M.L.; Guinand, M.; Vacheron, M.J.; Michel, G.; Denoroy, L.; Duez, C.; Englebert, S.; Joris, B.; Weber, G.; Ghuysen, J.M. Characterization of the sporulation-related γ-d-glutamyl-(l)meso-diaminopimelic-acid-hydrolysing peptidase I of Bacillus sphaericus NCTC 9602 as a member of the metallo(zinc) carboxypeptidase A family. Modular design of the protein. Biochem. J. 1993, 292, 563–570. [Google Scholar] [CrossRef] [PubMed]
- Jung, G.; Ueno, H.; Hayashi, R. Proton-relay system of carboxypeptidase Y as a sole catalytic site: Studies on mutagenic replacement of his 397. J. Biochem. 1998, 124, 446–450. [Google Scholar] [CrossRef] [PubMed]
- Breddam, K. Serine carboxypeptidases. A review. Carlsberg Res. Commun. 1986, 51, 83–128. [Google Scholar] [CrossRef]
- Valls, L.A.; Hunter, C.P.; Rothman, J.H.; Stevens, T.H. Protein sorting in yeast: The localization determinant of yeast vacuolar carboxypeptidase Y resides in the propeptide. Cell 1987, 48, 887–897. [Google Scholar] [CrossRef]
- Jackman, H.L.; Morris, P.W.; Deddish, P.A.; Skidgel, R.A.; Erdös, E.G. Inactivation of endothelin I by deamidase (lysosomal protective protein). J. Biol. Chem. 1992, 267, 2872–2875. [Google Scholar] [PubMed]
- Breddam, K.; Sørensen, S.B.; Svendsen, I. Primary structure and enzymatic properties of carboxypeptidase II from wheat bran. Carlsberg Res. Commun. 1987, 52, 297–311. [Google Scholar] [CrossRef]
- Liao, D.I.; Breddam, K.; Sweet, R.M.; Bullock, T.; Remington, S.J. Refined atomic model of wheat serine carboxypeptidase II at 2.2-A resolution. Biochemistry 1992, 31, 9796–9812. [Google Scholar] [CrossRef] [PubMed]
- Barbara, L.; Pryor, K.A.D.; Wu, J.K.; Frank, M.; Patel, R.A.; Craik, C.S.; Ellman, J.A.; Cummings, R.T.; Thornberry, N.A. Catalytic properties and inhibition of proline-specific dipeptidyl peptidases II, IV and VII. Biochem. J. 2003, 371, 525–532. [Google Scholar]
- Shariat-Madar, Z.; Mahdi, F.; Schmaier, A.H. Recombinant prolylcarboxypeptidase activates plasma prekallikrein. Blood 2004, 103, 4554–4561. [Google Scholar] [CrossRef] [PubMed]
- Witt, H.; Beer, S.; Rosendahl, J.; Chen, J.M.; Chandak, G.R.; Masamune, A.; Bence, M.; Szmola, R.; Oracz, G.; Milan Macek, J. Variants in CPA1 are strongly associated with early-onset chronic pancreatitis. Nat. Genet. 2013, 45, 1216–1220. [Google Scholar] [CrossRef] [PubMed]
- Chun, Z.; Daohua, Z.; Sichun, Z.; Lin, L.; Shu, T.; Li, Y.; Songnian, H.; Qili, F. A chymotrypsin-like serine protease cDNA involved in food protein digestion in the common cutworm, Spodoptera litura: Cloning, characterization, developmental and induced expression patterns, and localization. J. Insect Physiol. 2010, 56, 788–799. [Google Scholar]
- Dmitriew, C.D.; Cooray, M.C.; Rowe, L.R. Effects of early resource-limiting conditions on patterns of growth, growth efficiency, and immune function at emergence in a damselfly (Odonata: Coenagrionidae). Can. J. Zool. 2007, 85, 310–318. [Google Scholar] [CrossRef]
- Larsson, A.; Lewander, K. Metabolic effects of starvation in the eel, Anguilla Anguilla L. Comp. Biochem. Physiol. Part A 1973, 44, 367–374. [Google Scholar] [CrossRef]
- Jobling, M. Effects of starvation on proximate chemical composition and energy utilizaton of plaice, Pleuronectes platessa L. J. Fish Biol. 2006, 17, 325–334. [Google Scholar] [CrossRef]
- MaChado, C.R.; Garofaloj, M. Effects of starvation, refeeding, and insulin on energy-linked metabolic processes in catfish (Rhamdia hilarii) adapted to a carbohydrate-rich diet. Gen. Comp. Endocrinol. 1988, 71, 3429–3437. [Google Scholar] [CrossRef]
- SilkDB. Available online: http://www.silkdb.org/silkdb/ (accessed on 9 January 2016).
- GenBank. Available online: http://www.ncbi.nlm.nih.gov/Genbank/ (accessed on 10 January 2016).
- The ftp site of the National Center for Biotechnology Information. Available online: ftp://ftp.ncbi.nih.gov/blast/ (accessed on 10 January 2016).
- Enright, A.J.; van Dongen, S.; Ouzounis, C.A. An efficient algorithm for large-scale detection of protein families. Nucleic Acids Res. 2002, 30, 1575–1584. [Google Scholar] [CrossRef] [PubMed]
- SMART. Available online: http://smart.embl-heidelberg.de/ (accessed on 14 January 2016).
- Pfam. Available online: http://pfam.sanger.ac.UK/ (accessed on 14 January 2016).
- ORF Finder. Available online: http://www.ncbi.nlm.nih.gov/gorf/gorf.html (accessed on 15 January 2016).
- SignaIP 4.1. Available online: http://www.cbs.dtu.dk/services/SignalP/ (accessed on 15 January 2016).
- ProtParam. Available online: http://web.expasy.org/protparam/ (accessed on 15 January 2016).
- Thompson, J.D.; Higgins, D.G.; Gibson, T.J. CLUSTAL W: Improving the sensitiv ity of progressive multiple sequence alignment through sequence weighting, position specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994, 22, 4673–4680. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis Version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Dudley, J.; Nei, M.; Kumar, S. MEGA4: Molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol. Biol. Evol. 2007, 24, 1596–1599. [Google Scholar] [CrossRef] [PubMed]
- Xia, Q.; Cheng, D.; Duan, J.; Wang, G.; Cheng, T.; Zha, X.; Liu, C.; Zhao, P.; Dai, F.; Zhang, Z. Microarray-based gene expression profiles in multiple tissues of the domesticated silkworm, Bombyx mori. Genome Biol. 2007. [Google Scholar] [CrossRef] [PubMed]
- Golub, T.R.; Slonim, D.K.; Tamayo, P.; Huard, C.; Gaasenbeek, M.; Mesirov, J.P.; Coller, H.; Loh, M.L.; Downing, J.R.; Caligiuri, M.A. Molecular classification of cancer: Class discovery and class prediction by gene expression monitoring. Science 1999, 286, 531–537. [Google Scholar] [CrossRef] [PubMed]
- Broehan, G.; Kemper, M.; Driemeier, D.; Vogelpohl, I.; Merzendorfer, H. Cloning and expression analysis of midgut chymotrypsin-like proteinases in the tobacco hornworm. J. Insect Physiol. 2008, 54, 1243–1252. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2 method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- GraphPad. Available online: http://www.graphpad.com/quickcalcs/ttest1.cfm (accessed on 25 February 2016).


| Gene | Name in SilkDB | Chromosome | Scaffold | Position | Strand | Domain | Size (AA) | Probe | MW/Da | pI | Signal Peptide |
|---|---|---|---|---|---|---|---|---|---|---|---|
| BmMCP1 | BGIBMGA002530-TA | 9 | nscaf2511 | 5048133–5052368 | + | Peptidase_M2 | 73 | sw03136 | 8984.4 | 9.3 | no |
| BmMCP2 | BGIBMGA006234-TA | 6 | nscaf2851 | 1576285–1581353 | − | Peptidase_M2 | 409 | sw03231 | 46,754.2 | 4.89 | no |
| BmMCP3 | BGIBMGA002527-TA | 9 | nscaf2511 | 4990965–5002524 | + | Peptidase_M2 | 649 | sw14172 | 74,229.8 | 5.11 | yes |
| BmMCP4 | BGIBMGA006539-TA | 6 | nscaf2853 | 6801913–6813068 | + | Peptidase_M2 | 647 | sw11710 | 74,185.1 | 5.39 | no |
| BmMCP5 | BGIBMGA002529-TA | 9 | nscaf2511 | 5030694–5035485 | + | Peptidase_M2 | 173 | sw20193 | – | – | no |
| BmMCP6 | BGIBMGA002531-TA | 9 | nscaf2511 | 5192136–5202359 | + | Peptidase_M2 | 153 | sw11006 | 17,724 | 7.01 | no |
| BmMCP7 | BGIBMGA003228-TA | 2 | nscaf2623 | 1029259–1049209 | − | Peptidase_M2 | 711 | sw17925 | 82,533.6 | 9.06 | no |
| BmMCP8 | BGIBMGA002526-TA | 9 | nscaf2511 | 4965783–4980141 | + | Peptidase_M2 | 648 | sw00643 | 74,913.1 | 5.34 | yes |
| BmMCP9 | BGIBMGA009693-TA | 2 | nscaf2964 | 820517–835616 | − | Peptidase_M2 | 535 | sw19815 | 62,129.8 | 7.32 | no |
| BmMCP10 | BGIBMGA009487-TA | 14 | nscaf2953 | 517470–526524 | + | Peptidase_M14 | 428 | – | 48,175.4 | 6.06 | yes |
| BmMCP11 | BGIBMGA004799-TA | 25 | nscaf2818 | 2043455–2064130 | − | Peptidase_M14 | 468 | sw03973 | 53,767.3 | 9.18 | no |
| BmMCP12 | BGIBMGA004800-TA | 25 | nscaf2818 | 2024210–2029825 | − | Peptidase_M14 | 344 | sw20598 | 38,598.6 | 6.55 | no |
| BmMCP13 | BGIBMGA013275-TA | 16 | nscaf3063 | 3241878–3245274 | + | Peptidase_M14 | 346 | sw18892 | 39,898 | 5.24 | yes |
| BmMCP14 | BGIBMGA013276-TA | 16 | nscaf3063 | 3256098–3269527 | + | Peptidase_M14 | 445 | sw05014 | 50,709.9 | 5.41 | yes |
| BmMCP15 | BGIBMGA004801-TA | 25 | nscaf2818 | 1996486–2007642 | − | Peptidase_M14 | 427 | sw18974 | 48,293.5 | 5.78 | no |
| BmMCP16 | BGIBMGA008973-TA | 3 | nscaf2930 | 2150368–2156343 | + | Peptidase_M14 | 365 | – | 41,363.1 | 9.22 | no |
| BmMCP17 | BGIBMGA001892-TA | 19 | nscaf2204 | 1727896–1729970 | − | Peptidase_M14 | 298 | sw09352 | 33,757.5 | 6.37 | no |
| BmMCP18 | BGIBMGA001891-TA | 19 | nscaf2204 | 1730646–1734010 | − | Peptidase_M14 | 418 | sw14852 | 48,425.2 | 9.24 | no |
| BmMCP19 | BGIBMGA008975-TA | 3 | nscaf2930 | 2171627–2175156 | + | Peptidase_M14 | 267 | sw07649 | 30,547.4 | 5.31 | no |
| BmMCP20 | BGIBMGA009477-TA | 14 | nscaf2953 | 493144–498033 | − | Peptidase_M14 | 429 | sw02069 | 48,772.3 | 6.08 | yes |
| BmMCP21 | BGIBMGA004797-TA | 25 | nscaf2818 | 2109792–2113829 | − | Peptidase_M14 | 383 | sw11839 | 43,402.5 | 5.51 | no |
| BmMCP22 | BGIBMGA009478-TA | 14 | nscaf2953 | 480242–485505 | − | Peptidase_M14 | 394 | sw00546 | 44,578.6 | 5.39 | no |
| BmMCP23 | BGIBMGA004798-TA | 25 | nscaf2818 | 2099617–2105069 | − | Peptidase_M14 | 395 | sw03321 | 44,319.1 | 4.8 | no |
| BmMCP24 | BGIBMGA006871-TA | 10 | nscaf2859 | 1283815–1295739 | + | Peptidase_M14 | 604 | sw12842 | 68,595.4 | 6.45 | yes |
| BmMCP25 | BGIBMGA008976-TA | 3 | nscaf2930 | 2176287–2190645 | + | Peptidase_M14 | 365 | sw03831 | 41,702.1 | 6.15 | no |
| BmMCP26 | BGIBMGA008910-TA | 3 | nscaf2930 | 1816975–1823616 | − | Peptidase_M14 | 479 | sw07559 | 53,979.4 | 5.75 | yes |
| BmMCP27 | BGIBMGA004830-TA | 25 | nscaf2818 | 155397–165470 | − | Peptidase_M14 | 670 | sw03365 | 75,788.8 | 8.55 | no |
| BmMCP28 | BGIBMGA006715-TA | 10 | nscaf2855 | 3350086–3361536 | + | Peptidase_M14 | 371 | sw20772 | 42,903 | 5.3 | no |
| BmMCP29 | BGIBMGA001890-TA | 19 | nscaf2204 | 1746161–1749490 | − | Peptidase_M14 | 353 | sw20818 | 40,328.8 | 6.09 | no |
| BmMCP30 | BGIBMGA009486-TA | 14 | nscaf2953 | 512395–515181 | + | Peptidase_M14 | 409 | sw10048 | 46,594.4 | 5.81 | no |
| BmMCP31 | BGIBMGA009476-TA | 14 | nscaf2953 | 546619–551421 | − | Peptidase_M14 | 397 | sw12317 | 45,444.3 | 6.06 | no |
| BmMCP32 | BGIBMGA000307-TA | 22 | nscaf1681 | 1661872–1673460 | − | Peptidase_M14 | 483 | sw11693 | 54,330.3 | 5.73 | yes |
| BmMCP33 | BGIBMGA012807-TA | 16 | nscaf3058 | 6657227–6684446 | − | Peptidase_M14 | 996 | sw17444 | 11,1592 | 5.73 | no |
| BmMCP34 | BGIBMGA012806-TA | 16 | nscaf3058 | 6684947–6690407 | − | Peptidase_M14 | 294 | – | 33,318.1 | 5.55 | yes |
| BmSCP1 | BGIBMGA012452-TA | 9 | nscaf3045 | 2246700–2252046 | − | Peptidase_S28 | 352 | sw12542 | 40,622 | 4.69 | yes |
| BmSCP2 | BGIBMGA003579-TA | 5 | nscaf2674 | 1284051–1285310 | − | Peptidase_S28 | 419 | sw13803 | 48,709.9 | 4.96 | no |
| BmSCP3 | BGIBMGA008167-TA | 24 | nscaf2891 | 256973–274367 | + | Peptidase_S28 | 282 | sw14786 | 31,433.3 | 4.83 | no |
| BmSCP4 | BGIBMGA000549-TA | 1 | nscaf1690 | 3748863–3757879 | − | Peptidase_S28 | 439 | sw20204 | 49,930.5 | 6.08 | yes |
| BmSCP5 | BGIBMGA013534-TA | 5 | nscaf3075 | 927568–931683 | + | Peptidase_S28 | 383 | sw14860 | 42,863.6 | 6 | no |
| BmSCP6 | BGIBMGA003110-TA | 4 | nscaf2589 | 2567114–2581357 | + | Peptidase_S10 | 374 | sw00555 | 43,079.3 | 6.05 | no |
| BmSCP7 | BGIBMGA012773-TA | 16 | nscaf3058 | 8186538–8187953 | − | Peptidase_S10 | 471 | sw04192 | 53,120.8 | 5.56 | yes |
| BmSCP8 | BGIBMGA003111-TA | 4 | nscaf2589 | 2593763–2602101 | + | Peptidase_S10 | 402 | sw17614 | 45,191.6 | 5.11 | no |
| BmSCP9 | BGIBMGA003109-TA | 4 | nscaf2589 | 2546587–2560291 | + | Peptidase_S10 | 285 | – | 32,834.9 | 5.7 | no |
| BmSCP10 | BGIBMGA013085-TA | 16 | nscaf3058 | 8193345–8194772 | + | Peptidase_S10 | 475 | sw14878 | 54,378 | 8.46 | yes |
| BmSCP11 | BGIBMGA003112-TA | 4 | nscaf2589 | 2615158–2633253 | + | Peptidase_S10 | 1051 | sw01972 | 11,9508 | 5.66 | no |
| BmSCP12 | BGIBMGA010348-TA | 12 | nscaf2990 | 809983–812392 | + | Peptidase_S10 | 403 | sw06212 | 46,108.2 | 4.99 | no |
| BmSCP13 | BGIBMGA006502-TA | 6 | nscaf2853 | 4266715–4273371 | + | Peptidase_S10 | 376 | sw15928 | 42,942.2 | 8.8 | no |
| BmSCP14 | BGIBMGA010349-TA | 12 | nscaf2990 | 814696–818250 | + | Peptidase_S10 | 365 | sw06213 | 40,585.3 | 4.95 | no |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ye, J.; Li, Y.; Liu, H.-W.; Li, J.; Dong, Z.; Xia, Q.; Zhao, P. Genome-Wide Identification and Characterization of Carboxypeptidase Genes in Silkworm (Bombyx mori). Int. J. Mol. Sci. 2016, 17, 1203. https://doi.org/10.3390/ijms17081203
Ye J, Li Y, Liu H-W, Li J, Dong Z, Xia Q, Zhao P. Genome-Wide Identification and Characterization of Carboxypeptidase Genes in Silkworm (Bombyx mori). International Journal of Molecular Sciences. 2016; 17(8):1203. https://doi.org/10.3390/ijms17081203
Chicago/Turabian StyleYe, Junhong, Yi Li, Hua-Wei Liu, Jifu Li, Zhaoming Dong, Qingyou Xia, and Ping Zhao. 2016. "Genome-Wide Identification and Characterization of Carboxypeptidase Genes in Silkworm (Bombyx mori)" International Journal of Molecular Sciences 17, no. 8: 1203. https://doi.org/10.3390/ijms17081203
APA StyleYe, J., Li, Y., Liu, H.-W., Li, J., Dong, Z., Xia, Q., & Zhao, P. (2016). Genome-Wide Identification and Characterization of Carboxypeptidase Genes in Silkworm (Bombyx mori). International Journal of Molecular Sciences, 17(8), 1203. https://doi.org/10.3390/ijms17081203







