PAMP Activity of Cerato-Platanin during Plant Interaction: An -Omic Approach
Abstract
:1. Introduction
2. Results
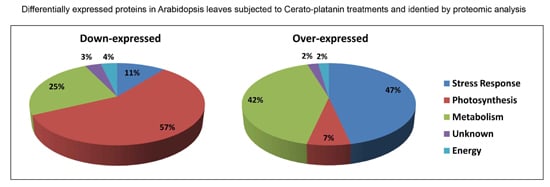
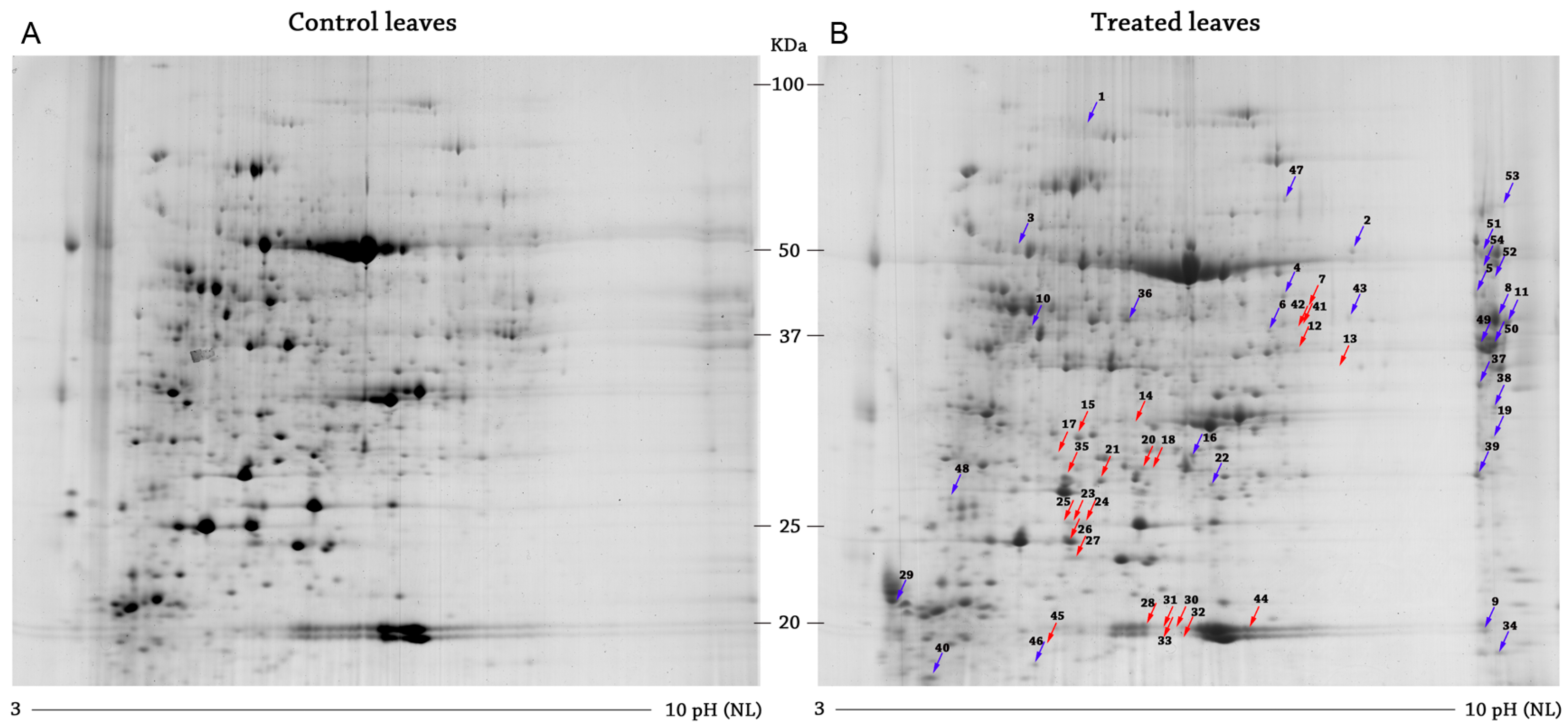
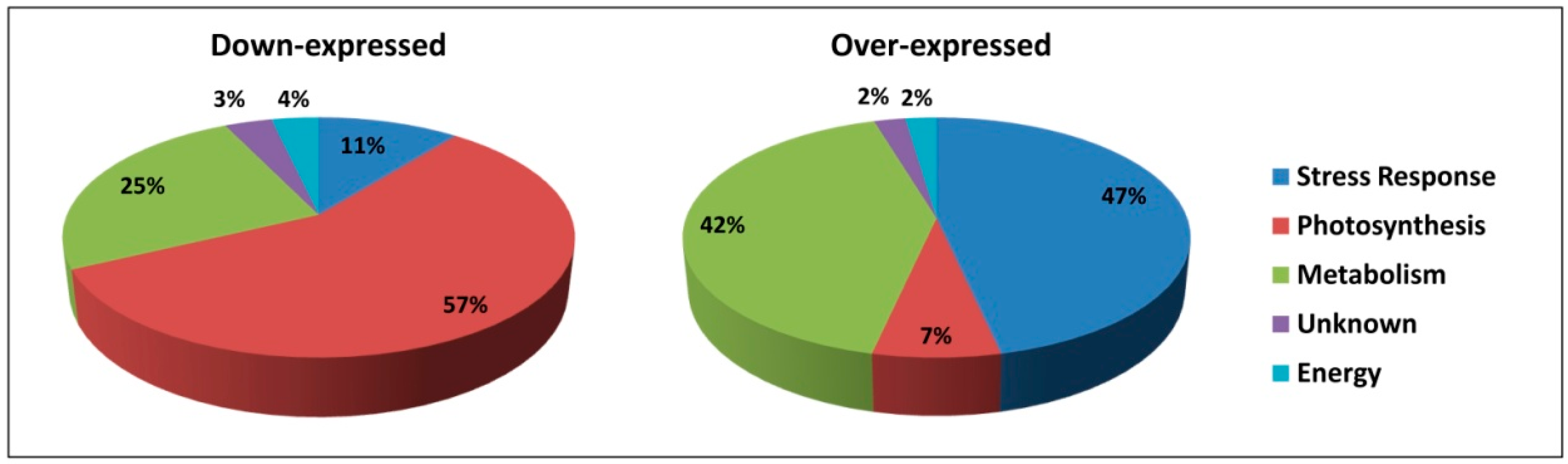
2.1. Differential Protein Expression
2.1.1. Downregulated Proteins
2.1.2. Upregulated Proteins
2.2. CO2 Assimilation and Transpiration Rate
2.3. VOC Accumulation
3. Discussion
3.1. CP Treatment Slows Metabolism and Negatively Affects Photosynthesis
3.2. CP Treatment Increases Expression of Defense-Related Proteins and VOC Emission
4. Materials and Methods
4.1. Plants and CP Treatment
4.2. Proteomic Experiments
4.2.1. Protein Extraction
4.2.2. 2D-Electrophoresis
4.2.3. Images Analysis
4.2.4. In-Gel Digestion and MALDI-ToF Analysis
4.3. Leaf Gas Exchange
4.4. Proton Transfer Reaction-Time-of-Flight-Mass Spectrometry and VOC Determination
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| 2DE-SDS | Bi-Dimensional Electrophoresis-Sodium Dodecyl-Sulfate |
| CP | Cerato-Platanin |
| CPF | Cerato-platanin Family |
| DTT | Dithiothreitol |
| GLs | Glucosinolates |
| GLVs | Green Leaves Volatiles |
| GSH | Glutathione |
| IEF | IsoElectric Focusing |
| IPG | Immobilized pH Gradient |
| LOX | Lipoxygenase |
| MALDI-TOF | Matrix Assisted Laser Desorption Ionization-Time of Flight |
| RP-HPLC | Reverse Phase-High Performance Liquid Chromatography |
| M/PAMPs | Microbe-/Pathogen-Associated Molecular Patterns |
| ppmv | Parts Per Million by Volume |
| PTR-ToF-MS | Proton-Transfer-Reaction-Time of Flight-Mass Spectrometry |
| ROS | Reactive Oxygen Species |
| RuBisCO | Ribulose-1,5-bisphosphate carboxylase/oxygenase |
| VOCs | Volatile Organic Compounds |
References
- Boller, T.; Felix, G. A renaissance of elicitors: Perception of microbeassociated molecular patterns and danger signals by pattern-recognition receptors. Annu. Rev. Plant Biol. 2009, 60, 379–406. [Google Scholar] [CrossRef] [PubMed]
- Jones, J.D.G.; Dangl, J.L. The plant immune system. Nature 2006, 444, 323–329. [Google Scholar] [CrossRef] [PubMed]
- Newman, M.A.; Sundelin, T.; Nielsen, J.T.; Erbs, G. MAMP (microbe-associated molecular pattern) triggered immunity in plants. Front. Plant Sci. 2013, 4, 139. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Kovalchuk, A.; Keriö, S.; Asiegbu, F.O. Distribution and bioinformatic analysis of the cerato-platanin protein family in Dikarya. Mycologia 2013, 105, 1479–1488. [Google Scholar] [CrossRef] [PubMed]
- Pazzagli, L.; Seidl-Seiboth, V.; Barsottini, M.; Vargas, W.A.; Scala, A.; Mukherjee, P.K. Cerato-platanins: Elicitors and effectors. Plant Sci. 2014, 228, 79–87. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Quintana, J.; Kovalchuk, A.; Ubhayasekera, W.; Asiegbu, F.O. A cerato-platanin-like protein HaCPL2 from Heterobasidion annosum sensu stricto induces cell death in Nicotiana tabacum and Pinus sylvestris. Fungal Genet. Biol. 2015, 84, 41–51. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Wu, J.; Kim, S.G.; Tsuda, K.; Gupta, R.; Park, S.Y.; Kim, S.T.; Kang, K.Y. Magnaporthe oryzae-secreted protein MSP1 Induces cell death and elicits defense responses in rice. Mol. Plant Microbe Interact. 2016, 29, 299–312. [Google Scholar] [CrossRef] [PubMed]
- Baccelli, I.; Luti, S.; Bernardi, R.; Scala, A.; Pazzagli, L. Cerato-platanin shows expansin-like activity on cellulosic materials. Appl. Microbiol. Biotechnol. 2014, 98, 175–184. [Google Scholar] [CrossRef] [PubMed]
- De Oliveira, A.L.; Gallo, M.; Pazzagli, L.; Benedetti, C.E.; Cappugi, G.; Scala, A.; Pantera, B.; Spisni, A.; Pertinhez, T.A.; Cicero, D.O. The structure of the elicitor Cerato-platanin (CP), the first member of the CP fungal protein family, reveals a double ψβ-barrel fold and carbohydrate binding. J. Biol. Chem. 2011, 286, 17560–17568. [Google Scholar] [CrossRef] [PubMed]
- Georgelis, N.; Nikolaidis, N.; Cosgrove, D.J. Bacterial expansins and related proteins from the world of microbes. Appl. Microbiol. Biotechnol. 2015, 99, 3807–3823. [Google Scholar] [CrossRef] [PubMed]
- Lombardi, L.; Faoro, F.; Luti, S.; Baccelli, I.; Martellini, F.; Bernardi, R.; Picciarelli, P.; Scala, A.; Pazzagli, L. Differential timing of defense-related responses induced by cerato-platanin and cerato-populin, two non-catalytic fungal elicitors. Physiol. Plant. 2013, 149, 408–421. [Google Scholar] [CrossRef] [PubMed]
- Baccelli, I.; Lombardi, L.; Luti, S.; Bernardi, R.; Picciarelli, P.; Scala, A.; Pazzagli, L. Cerato-platanin induces resistance in Arabidopsis leaves through stomatal perception, over-expression of salicylic acid- and ethylene-signalling genes and camalexin biosynthesis. PLoS ONE 2014, 9, e100959. [Google Scholar] [CrossRef] [PubMed]
- González-Fernández, R.; Valero-Galván, J.; Gómez-Gálvez, F.J.; Jorrín-Novo, J.V. Unraveling the in vitro secretome of the phytopathogen Botrytis cinerea to understand the interaction with its hosts. Front. Plant Sci. 2015, 6, 839. [Google Scholar] [CrossRef] [PubMed]
- Gaderer, R.; Lamdan, N.L.; Frischmann, A.; Sulyok, M.; Krska, R.; Horwitz, B.A.; Seidl-Seiboth, V. Sm2, a paralog of the Trichoderma cerato-platanin elicitor Sm1, is also highly important for plant protection conferred by the fungal-root interaction of Trichoderma with maize. BMC Microbiol. 2015, 15. [Google Scholar] [CrossRef] [PubMed]
- Kangasjarvi, S.; Neukermans, J.; Li, S.; Aro, E.M.; Noctor, G. Photosynthesis, photorespiration, and light signalling in defence responses. J. Exp. Bot. 2012, 63, 1619–1636. [Google Scholar] [CrossRef] [PubMed]
- Torres, M.A. ROS in biotic interactions. Physiol. Plant. 2010, 138, 414–429. [Google Scholar] [CrossRef] [PubMed]
- O’Brien, J.A.; Daudi, A.; Butt, V.S.; Bolwell, G.P. Reactive oxygen species and their role in plant defence and cell wall metabolism. Planta 2012, 236, 765–779. [Google Scholar] [CrossRef] [PubMed]
- Potters, G.; Horemans, N.; Jansen, M.A. The cellular redox state in plant stress biology—A charging concept. Plant Physiol. Biochem. 2010, 48, 292–300. [Google Scholar] [CrossRef] [PubMed]
- Mithöfer, A.; Schulze, B.; Boland, W. Biotic and heavy metal stress response in plants: Evidence for common signals. FEBS Lett. 2004, 566, 1–5. [Google Scholar] [CrossRef] [PubMed]
- Scala, A.; Allmann, S.; Mirabella, R.; Haring, M.A.; Schuurink, R.C. Green leaf volatiles: A plant’s multifunctional weapon against herbivores and pathogens. Int. J. Mol. Sci. 2013, 14, 17781–17811. [Google Scholar] [CrossRef] [PubMed]
- Denoux, C.; Galletti, R.; Mammarella, N.; Gopalan, S.; Werck, D.; De Lorenzo, G.; Ferrari, S.; Ausubel, F.M.; Dewdney, J. Activation of defense response pathways by OGs and Flg22 elicitors in Arabidopsis seedlings. Mol. Plant 2008, 1, 423–445. [Google Scholar] [CrossRef] [PubMed]
- Sánchez-Vallet, A.; Mesters, J.R.; Thomma, B.P. The battle for chitin recognition in plant-microbe interactions. FEMS Microbiol. Rev. 2015, 39, 171–183. [Google Scholar] [CrossRef] [PubMed]
- Iriti, M.; Varoni, E.M. Chitosan-induced antiviral activity and innate immunity in plants. Environ. Sci. Pollut. Res. Int. 2015, 22, 2935–2944. [Google Scholar] [CrossRef] [PubMed]
- Balmer, A.; Pastor, V.; Gamir, J.; Flors, V.; Mauch-Mani, B. The “prime-ome”: Towards a holistic approach to priming. Trends Plant Sci. 2015, 20, 443–452. [Google Scholar] [CrossRef] [PubMed]
- Dadakova, K.; Havelkova, M.; Kurkova, B.; Tlolkova, I.; Kasparovsky, T.; Zdrahal, Z.; Lochman, J. Proteome and transcript analysis of Vitis vinifera cell cultures subjected to Botrytis cinerea infection. J. Proteom. 2015, 119, 143–153. [Google Scholar] [CrossRef] [PubMed]
- Taiti, C.; Costa, C.; Menesatti, P.; Comparini, D.; Bazihizina, N.; Azzarello, E.; Masi, E.; Mancuso, S. Class-modeling approach to PTR-TOF-MS data: A peppers case study. J. Sci. Food Agric. 2014, 95, 1757–1763. [Google Scholar] [CrossRef] [PubMed]
- Brilli, F.; Ruuskanen, T.M.; Schnitzhofer, R.; Muller, M.; Breitenlechner, M.; Bittner, V.; Wohlfahrt, G.; Loreto, F.; Hansel, A. A detection of plant volatiles after leaf wounding and darkening by proton transfer reaction “time-of-flight” mass spectrometry (PTR-TOF). PLoS ONE 2011, 6, e20419. [Google Scholar] [CrossRef] [PubMed]
- Fall, R.; Hansel, A.; Jordan, A.; Lindinger, W. Volatile organic compounds emitted after leaf wounding: On-line analysis by proton-transfer-reaction mass spectrometry. J. Geophys. Res. 1999, 104, 15963–15974. [Google Scholar] [CrossRef]
- Sánchez-López, J.A.; Zimmermann, R.; Yeretzian, C. Insight into the time-resolved extraction of aroma compounds during espresso coffee preparation: Online monitoring by PTR-ToF-MS. Anal. Chem. 2014, 86, 11696–11704. [Google Scholar] [CrossRef] [PubMed]
- Brilli, F.; He Ortnagl, L.; Bamberger, I.; Schnitzhofer, R.; Ruuskanen, T.M.; Hansel, A.; Loreto, F.; Wohlfahrt, G. Qualitative and quantitative characterization of volatile organic compound emissions from cut grass. Environ. Sci. Technol. 2012, 46, 3859–3865. [Google Scholar] [CrossRef] [PubMed]
- Mancuso, S.; Taiti, C.; Bazihizina, N.; Costa, C.; Menesatti, P.; Giagnoni, L.; Arenella, M.; Nannipieri, P.; Renella, G. Soil volatile analysis by proton transfer reaction-time of flight mass spectrometry (PTR-TOF-MS). Appl. Soil Ecol. 2015, 86, 182–191. [Google Scholar] [CrossRef]
- Taiti, C.; Costa, C.; Menesatti, P.; Caparrotta, S.; Bazihizina, N.; Azzarello, E.; Petrucci, W.A.; Masi, E.; Giordani, E. Use of volatile organic compounds and physico-chemical parameters for monitoring the post-harvest ripening of imported tropical fruits. Eur. Food Res. Technol. 2015, 241, 91–102. [Google Scholar] [CrossRef]
- Berger, S.; Sinha, A.K.; Roitsch, T. Plant physiology meets phytopathology: Plant primary metabolism and plant–pathogen interactions. J. Exp. Bot. 2007, 58, 4019–4026. [Google Scholar] [CrossRef] [PubMed]
- Dong, X.; Yi, H.; Han, C.T.; Nou, I.S.; Hur, Y. GDSL esterase/lipase genes in Brassica rapa L.: Genome-wide identification and expression analysis. Mol. Genet. Genom. 2016, 291, 531–542. [Google Scholar] [CrossRef] [PubMed]
- Wingler, A.; Lea, P.J.; Quick, W.P.; Leegood, R.C. Photorespiration: Metabolic pathways and their role in stress protection. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2000, 355, 1517–1529. [Google Scholar] [CrossRef] [PubMed]
- Wang, N.; Zhao, J.; He, X.; Sun, H.; Zhang, G.; Wu, F. Comparative proteomic analysis of drought tolerance in the two contrasting Tibetan wild genotypes and cultivated genotype. BMC Genom. 2015, 16, 432. [Google Scholar] [CrossRef] [PubMed]
- Margaria, P.; Abbà, S.; Palmano, S. Novel aspects of grapevine response to phytoplasma infection investigated by a proteomic and phospho-proteomic approach with data integration into functional networks. BMC Genom. 2013, 14, 38. [Google Scholar] [CrossRef] [PubMed]
- Gollan, P.J.; Ziemann, M.; Bhave, M. PPIase activities and interaction partners of FK506-binding proteins in the wheat thylakoid. Physiol. Plant. 2011, 143, 385–395. [Google Scholar] [CrossRef] [PubMed]
- Mhamdi, A.; Queval, G.; Chaouch, S.; Vanderauwera, S.; van Breusegem, F.; Noctor, G. Catalase function in plants: A focus on Arabidopsis mutants as stress-mimic models. J. Exp. Bot. 2010, 61, 4197–4220. [Google Scholar] [CrossRef] [PubMed]
- Venugopal, S.C.; Chanda, B.; Vaillancourt, L.; Kachroo, A.; Kachroo, P. The common metabolite glycerol-3-phosphate is a novel regulator of plant defense signaling. Plant Signal. Behav. 2009, 4, 746–749. [Google Scholar] [CrossRef] [PubMed]
- Vescovi, M.; Zaffagnini, M.; Festa, M.; Trost, P.; Schiavo, F.L.; Costa, A. Nuclear accumulation of cytosolic glyceraldehyde-3-phosphate dehydrogenase in cadmium-stressed Arabidopsis roots. Plant Physiol. 2013, 162, 333–346. [Google Scholar] [CrossRef] [PubMed]
- Chepyshko, H.; Lai, C.-P.; Huang, L.-M.; Liu, J.-H.; Shaw, J.-F. Multifunctionality and diversity of GDSL esterase/lipase gene family in rice (Oryza. sativa L. japonica) genome: New insights from bioinformatics analysis. BMC Genom. 2012, 13, 309. [Google Scholar] [CrossRef] [PubMed]
- Barkan, A.; Small, I. Pentatricopeptide repeat proteins in plants. Annu. Rev. Plant Biol. 2014, 65, 415–442. [Google Scholar] [CrossRef] [PubMed]
- Casasoli, M.; Spadoni, S.; Lilley, K.S.; Cervone, F.; de Lorenzo, G.; Mattei, B. Identification by 2-D DIGE of apoplastic proteins regulated by oligogalacturonides in Arabidopsis thaliana. Proteomics 2008, 8, 1042–1054. [Google Scholar] [CrossRef] [PubMed]
- Angelino, D.; Dosz, E.B.; Sun, J.; Hoeflinger, J.H.; van Tassell, M.L.; Chen, P.; Harnly, J.M.; Miller, M.J.; Jeffery, E.H. Myrosinase-dependent and independent formation and control of isothiocyanate products of glucosinolate hydrolysis. Front. Plant Sci. 2015, 6, 831. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Liu, H.; Li, S.; Zhai, G.; Shao, J.; Tao, Y. Characterization and molecular cloning of a serine hydroxymethyltransferase 1 (OsSHM1) in rice. J. Integr. Plant Biol. 2015, 57, 745–756. [Google Scholar] [CrossRef] [PubMed]
- Dubreuil-Maurizi, C.; Poinssot, B. Role of glutathione in plant signaling under biotic stress. Plant Signal. Behav. 2012, 7, 210–212. [Google Scholar] [CrossRef] [PubMed]
- Moreno, J.I.; Martin, R.; Castresana, C. Arabidopsis SHMT1, a serine hydroxymethyltransferase that functions in the photorespiratory pathway influences resistance to biotic and abiotic stress. Plant J. 2005, 41, 451–463. [Google Scholar] [CrossRef] [PubMed]
- Giordano, D.; Provenzano, S.; Ferrandino, A.; Vitali, M.; Pagliarani, C.; Roman, F.; Cardinale, F.; Castellarin, S.D.; Schubert, A. Characterization of a multifunctional caffeoyl-CoA O-methyltransferase activated in grape berries upon drought stress. Plant Physiol. Biochem. 2016, 101, 23–32. [Google Scholar] [CrossRef] [PubMed]
- Jeong, Y.J.; An, C.H.; Woo, S.G.; Jeong, H.J.; Kim, Y.M.; Park, S.J.; Yoon, B.D.; Kim, C.Y. Production of pinostilbene compounds by the expression of resveratrol O-methyltransferase genes in Escherichia coli. Enzym. Microb. Technol. 2014, 54, 8–14. [Google Scholar] [CrossRef] [PubMed]
- Chen, I.C.; Huang, I.C.; Liu, M.J.; Wang, Z.G.; Chung, S.S.; Hsieh, H.L. Glutathione S-transferase interacting with far-red insensitive 219 is involved in phytochrome A-mediated signaling in Arabidopsis. Plant Physiol. 2007, 143, 1189–1202. [Google Scholar] [CrossRef] [PubMed]
- Dixon, D.P.; Edwards, R. Selective binding of glutathione conjugates of fatty acid derivatives by plant glutathione transferases. J. Biol. Chem. 2009, 284, 21249–21256. [Google Scholar] [CrossRef] [PubMed]
- Sasaki-Sekimoto, Y.; Taki, N.; Obayashi, T.; Aono, M.; Matsumoto, F.; Sakurai, N.; Suzuki, H.; Hirai, M.Y.; Noji, M.; Saito, K.; et al. Coordinated activation of metabolic pathways for antioxidants and defence compounds by jasmonates and their roles in stress tolerance in Arabidopsis. Plant J. 2005, 44, 653–668. [Google Scholar] [CrossRef] [PubMed]
- Hirschmann, F.; Krause, F.; Papenbrock, J. The multi-protein family of sulfotransferases in plants: Composition, occurrence, substrate specificity, and functions. Front. Plant Sci. 2014, 5, 556. [Google Scholar] [CrossRef] [PubMed]
- Taiti, C.; Giorni, E.; Colzi, I.; Pignattelli, S.; Bazihizina, N.; Buccianti, A.; Luti, S.; Pazzagli, L.; Mancuso, S.; Gonnelli, C. Under fungal attack on a metalliferous soil: ROS or not ROS? Insights from Silene paradoxa L. growing under copper stress. Environ. Pollut. 2016, 210, 282–292. [Google Scholar] [CrossRef] [PubMed]
- Bentley, R.; Chasteen, T.G. Environmental VOCs formation and degradation of dimethyl sulfide, methanethiol and related materials. Chemosphere 2004, 55, 291–317. [Google Scholar] [CrossRef] [PubMed]
- Van Dam, N.M.; Samudrala, D.; Harren, F.J.; Cristescu, S.M. Real-time analysis of sulfur-containing volatiles in Brassica plants infested with root-feeding Delia radicum larvae using proton-transfer reaction mass spectrometry. AoB Plants 2012, 2012, pls021. [Google Scholar] [CrossRef] [PubMed]
- Niinemets, U.; Kännaste, A.; Copolovici, L. Quantitative patterns between plant volatile emissions induced by biotic stresses and the degree of damage. Front. Plant Sci. 2013, 4, 262. [Google Scholar] [CrossRef] [PubMed]
- Jardine, K.; Karl, T.; Lerdau, M.; Harley, P.; Guenther, A.; Mak, J.E. Carbon isotope analysis of acetaldehyde emitted from leaves following mechanical stress and anoxia. Plant Biol. 2009, 11, 591–597. [Google Scholar] [CrossRef] [PubMed]
- Monson, R.K. Metabolic and gene expression controls on the production of biogenic volatile organic compounds. In Biology, Controls and Models of Tree Volatile Organic Compound Emissions; Springer: Berlin, Germany, 2013; Volume 5, pp. 153–179. [Google Scholar]
- Copolovici, L.; Väärtnõu, F.; Portillo Estrada, M.; Niinemets, Ü. Oak powdery mildew (Erysiphe alphitoides)-induced volatile emissions scale with the degree of infection in Quercus robur. Tree Physiol. 2014, 12, 1399–1410. [Google Scholar] [CrossRef] [PubMed]
- Neilson, E.H.; Goodger, J.Q.D.; Woodrow, I.E.; Møller, B.L. Plant chemical defense: At what cost? Trends Plant Sci. 2013, 18, 1360–1385. [Google Scholar] [CrossRef] [PubMed]
- Carresi, L.; Pantera, B.; Zoppi, C.; Cappugi, G.; Oliveira, A.L.; Pertinhez, T.A.; Spisni, A.; Scala, A.; Pazzagli, L. Cerato-platanin, a phytotoxic protein from Ceratocystis fimbriata: Expression in Pichia pastoris, purification and characterization. Protein Expr. Purif. 2006, 49, 159–167. [Google Scholar] [CrossRef] [PubMed]
- Pazzagli, L.; Zoppi, C.; Carresi, L.; Tiribilli, B.; Sbrana, F.; Schiff, S.; Pertinhez, T.A.; Scala, A.; Cappugi, G. Characterization of ordered aggregates of ceratoplatanin and their involvement in fungus–host interactions. Biochim. Biophys. Acta 2009, 1790, 1334–1344. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.J.; Lee, H.M.; Wang, Y.; Wu, J.; Kim, S.G.; Kang, K.Y.; Park, K.H.; Kim, Y.C.; Choi, I.S.; Agrawal, G.K.; et al. Depletion of abundant plant RuBisCO protein using the protamine sulfate precipitation method. Proteomics 2013, 13, 2176–2179. [Google Scholar] [CrossRef] [PubMed]
- Monici, M.; Cialdai, F.; Ranaldi, F.; Paoli, P.; Boscaro, F.; Moneti, G.; Caselli, A. Effect of IR laser on myoblasts: A proteomic study. Mol. Biosyst. 2013, 6, 1147–1161. [Google Scholar] [CrossRef] [PubMed]
- Bazihizina, N.; Colzi, I.; Giorni, E.; Mancuso, S.; Gonnelli, C. Photosynthesizing on metal excess: Copper differently induced changes in various photosynthetic parameters in copper tolerant and sensitive Silene paradoxa L. populations. Plant Sci. 2015, 232, 67–76. [Google Scholar] [CrossRef] [PubMed]
- Easlon, H.M.; Bloom, A.J. Easy Leaf Area: Automated digital image analysis for rapid and accurate measurement of leaf area. Appl. Plant Sci. 2014, 2. [Google Scholar] [CrossRef] [PubMed]
- Cappellin, L.; Biasioli, F.; Fabris, A.; Schuhfried, E.; Soukoulis, C.; Meark, T.D. Improved mass accuracy in PTR-TOF-MS: Another step towards better compound identification in PTR-MS. Int. J. Mass Spectr. 2010, 209, 60–63. [Google Scholar] [CrossRef]


| Spot No. | ID | Protein Name | MASCOT Search Results * | Theoretic pI/Mr (kDa) | Function | Localization | GO-Biological Process | |||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 2 | 3 | 4 | |||||||
| 7 | Q9LFA3 | Monodehydro-ascorbate reductase | 16 | 52 | 214 | 6.4/46.6 | Metabolism | Peroxisome | Oxidation-reduction process flavin adenine dinucleotide binding | |
| 12 | Q9SA52 | 15 | 54 | 169 | 8.2/42.7 | Metabolism | Chloroplast | Polysaccharide metabolic process Response to bacterium and abiotic stress | ||
| 13 | O03042 | RuBisCO large chain | 7 | 14 | 78 | 5.9/53.4 | Photosynthesis | Chloroplast | Photorespiration | |
| 14 | O03042 | RuBisCO large chain | 9 | 31 | 111 | 5.9/53.4 | Photosynthesis | Chloroplast | Photorespiration | |
| 15 | O80934 | Uncharacterized protein At2g37660 | 15 | 48 | 204 | 8.4/34.9 | Unknown | Chloroplast | Defense response to bacterium | |
| 17 | P48491 | Triosephosphate isomerase | 9 | 47 | 126 | 5.4/27.3 | Metabolism | Cytoplasm | Carbohydrate metabolic process | |
| 18 | O03042 | RuBisCO large chain (fragment 147–479) | 14 | 30 (75.9) | 209 | 5.9/53.4 (6.4/37.3) | Photosynthesis | Chloroplast | Photorespiration | |
| 20 | P27140 | Beta carbonic anhydrase 1 | 15 | 53 | 186 | 5.7/37.8 | Photosynthesis | Chloroplast | Carbon utilization Response to bacterium Regulation of stomatal complex | |
| 21 | Q9FWR4 | Glutathione S-transferase DHAR1 | 15 | 87 | 225 | 353 | 5.6/23.7 | Defense | mitochondrial | Response to fungus, response to Jasmonic acid |
| Q8L7C9 | Glutathione S-transferase U20 | 12 | 48 | 143 | 5.6/25.1 | Metabolism/Defense | Cytoplasm/Nucleus | Regulation of growth and flowering | ||
| 23 | O03042 | RuBisCO large chain | 12 | 34 | 121 | 5.9/53.4 | Photosynthesis | Chloroplast | Photorespiration | |
| 24 | O03042 | RuBisCO large chain | 10 | 26 | 97 | 5.9/53.4 | Photosynthesis | Chloroplast | Photorespiration | |
| 25 | O03042 | RuBisCO large chain (fragment 1–260) | 9 | 15 (61) | 84 | 5.9/53.4 (6.0/29.2) | Photosynthesis | Chloroplast | Photorespiration | |
| 26 | Q9SSS9 | ATP Synthase subunit delta | 12 | 52 | 134 | 9.0/25.6 | Energy | Chloroplast | ATP biosynthetic process photosynthetic electron transport | |
| 27 | P34791 | Peptidyl-prolyl cis-trans isomerase | 15 | 66 | 216 | 8.8/28.5 | Metabolism | Chloroplast | Protein folding | |
| 28 | P10797 | RuBisCO small chain 2B | 8 | 35 | 111 | 7.6/20.6 | Photosynthesis | Chloroplast | Photorespiration | |
| 30 | P10797 | RuBisCO small chain 2B | 8 | 35 | 111 | 7.6/20.6 | Photosynthesis | Chloroplast | Photorespiration | |
| 31 | P10795 | RuBisCO-small chain 1A | 6 | 32 | 80 | 7.6/20.4 | Photosynthesis | Chloroplast | Photorespiration | |
| 32 | P10795 | RuBisCO- small chain 1A | 11 | 51 | 183 | 7.6/20.4 | Photosynthesis | Chloroplast | Photorespiration | |
| 33 | P10795 | RuBisCO small chain 1A | 15 | 77 | 245 | 7.6/20.4 | Photosynthesis | Chloroplast | Photorespiration | |
| 35 | Q9ZRW8 | Glutathione S-transferase U19 | 7 | 34 | 95 | 5.8/25.6 | Metabolism/Defense | Cytoplasm/Chloroplast | Response to oxidative stress; response to Jasmonic Acid | |
| 41 | O65396 | Aminomethyl-transferase | 21 | 61 | 249 | 8.5/44.7 | Metabolism | Mitochondrion | Glycine catabolic process | |
| 42 | Q93X94 | GDS-Lesterase/lipase EXL6 | 10 | 42 | 147 | 9.5/38.9 | Metabolism | Secreted | Lipid catabolic process | |
| 44 | P10795 | RuBisCO-small chain 1A | 10 | 42 | 148 | 7.6/20.4 | Photosynthesis | Chloroplast | Photorespiration | |
| 45 | P10797 | RuBisCO small chain 2B | 5 | 28 | 101 | 7.6/20.6 | Photosynthesis | Chloroplast | Photorespiration | |
| Spot No. | ID | Protein Name | MASCOT Search Results * | Theoretical pI/Mr (kDa) | Function | Localization | GO Biological Process | |||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 2 | 3 | 4 | |||||||
| 1 | Q9FXH6 | cyclic nucleotide-gated ion channel 8 | 11 | 17 | 81 | 9.15/86.7 | Metabolism | Plasma membrane | Ion transport Trasmembrane potential | |
| 2 | Q9M5K3 | Dhydrolipoyl Dehydrogenase | 16 | 48 | 230 | 7.0/54.2 | Defense | Mitochondrion | Cell redox homeostasis Response to cadmium | |
| 3 | P56757 | ATP synthase subunit alpha | 13 | 29 | 143 | 5.2/55.3 | Energy | Chloroplast | ATP hydrolysis and synthesis | |
| 4 | P25819 | Catalase-2 | 20 | 50 | 275 | 6.6/57.2 | Defense | Mitochondrion/Peroxisome | Cell redox homeostasis Response to oxidative stress | |
| 5 | Q9FLS9 | Pentatricopeptide repeat-At5g61800 | 16 | 35 | 155 | 8.7/56.7 | Defense | Mitochondrion | Defense responses to oligogalatturonides | |
| 6 | Q9S7E9 | Glutamate—glyoxylate aminotransfe-rase 2 | 15 | 41 | 190 | 6.27/53.9 | Metabolism | Peroxisome | Biosynthetic process l-alanine catabolic process | |
| 8 | Q9SCX9 | Glycerol-3-phosphate dehydrogenase | 10 | 28 | 108 | 8.2/44.3 | Metabolism/Defense | Chloroplast | Carbohydrate metabolism Glycerol 3-phosphate catabolism | |
| 9 | F4JDF3 | Dirigent protein 8 | 7 | 59 | 104 | 9.76/18.9 | Metabolism/Defense | Apoplast | Phenylpropanoid biosynthetic process | |
| 10 | P25697 | Phosphoribulokinase | 7 | 31 | 104 | 5.7/44.7 | Metabolism/Defense | Chloroplast | Response to bacterium Pyrimidine salvage Pentose phosphate cycle | |
| 11 | O65396 | Aminomethyltransferase | 14 | 41 | 174 | 8.5/44.7 | Metabolism/Defense | Mitochondrion | Glycine catabolic process Response to cadmium | |
| 16 | Q05431 | l-ascorbate peroxidase 1 | 13 | 61 | 167 | 5.7/27.8 | Defense | Cytoplasm | Response to oxidative stress | |
| 19 | Q9SIT7 | Pentatricopepti de repeat-At2g13600 | 5 | 10 | 61 | 5.6/79.1 | Unknown | Mitochondrion | Mitochondrial mRNA modification | |
| 22 | P42760 | Glutathione S-transferase F6 | 13 | 85 | 205 | 5.8/23.4 | Defense | Cell wall/Cytoplasm | Glutathione catabolic process Response to bacterium Response to abiotic stress | |
| 29 | Q9C9C9 | Cytosolic sulfotransferase 18 | 9 | 29 | 92 | 5.5/40.2 | Metabolism/Defense | Cytoplasm | Glucosinolate biosynthetic process | |
| 34 | O80929 | 60S ribosomal protein L36-1 | 5 | 37 | 72 | 11.7/12.7 | Metabolism | Ribosome | Structural constituent of ribosome | |
| 36 | P35614 | Peptide chain release factor subunit 1–3 | 7 | 24 | 87 | 5.4/49.1 | Metabolism | Cytoplasm | Protein biosynthesis | |
| 37 | Q9AV97 | 2-dehydro-3-deoxyphosphooctonate aldolase | 12 | 55 | 176 | 6.3/31.9 | Metabolism | Cytoplasm | Pollen tube development and growth | |
| 38 | Q9LK72 | Lectin-like protein At3g16530 | 12 | 55 | 151 | 7.0/30.5 | Defense | Apoplast | Defense response to fungus Response to chitin | |
| 39 | Q9S789 | Probable inactive cytidine deaminase 9 | 7 | 25 | 89 | 8.1/33.1 | Metabolism | Cytoplasm | Cytidine deamination Pyrimidine savage | |
| 40 | Q39239 | Thioredoxin H4 | 5 | 34 | 63 | 5.3/13.2 | Defense | Cytoplasm | Cell redox homeostasis Response to oxidative stress | |
| 43 | Q9LJG3 | GDSL esterase/lipase ESM1 | 12 | 47 | 158 | 7.6/44.4 | Defense | Peroxisome/secreted | Responses to bacterium glucosinolate catabolism response to cold | |
| 46 | Q9LUV2 | Probable protein Pop3 | 5 | 73 | 90 | 5.4/12.2 | Defense | Cytoplasm/Plasma membrane | Defense response to fungus Defense response to fungus | |
| 47 | Q9C5C2 | Myrosinase 2 | 25 | 51 | 284 | 7.1/63.3 | Metabolism/Defense | Apoplast | Defense response to insect glucosinolate catabolism response to abscisic acid | |
| 48 | Q9C5R8 | 2-Cys peroxiredoxin BAS1-like | 12 | 51 | 200 | 5.5/29.9 | Defense | Chloroplast | Cellular oxidant detoxification Responses to bacterium response to cold | |
| 49 | Q9SA52 | Chloroplast stem-loop binding protein of 41 kDa | 17 | 52 | 172 | 242 | 8.2/42.7 | Metabolism/Defense | Chloroplast | Polysaccharide metabolism Defense response to bacterium Response to abiotic stress |
| Q9LPW0 | Glyceraldehyde-3-phosphate dehydrogenase GAPA2 | 13 | 39 | 120 | 8.2/43.1 | Metabolism | Chloroplast | Glucose metabolic process Reductive pentose phosphate cycle | ||
| 50 | Q9SA52 | Chloroplast stem-loop binding protein | 12 | 44 | 124 | 8.2/42.7 | Metabolism | Chloroplast | Photosynthesis Polysaccharide metabolic process | |
| 51 | Q9SA52 | Chloroplast stem-loop binding protein | 11 | 32 | 113 | 140 | 8.2/42.7 | Metabolism | Chloroplast | Photosynthesis Polysaccharide metabolic process |
| P25858 | Glyceraldehyde-3-phosphate dehydrogenase GAPC1 | 7 | 27 | 65 | 6.6/37.0 | Metabolism/Defense | Cytoplasm | Carbohydrate metabolism Response to redox state Response to abiotic stress | ||
| 52 | Q9LJG3 | GDSL esterase/lipase ESM1 | 16 | 54 | 188 | 7.6/44.3 | Defense | Secreted | Defence responses to bacterium glucosinolate catabolic process response to cold | |
| 53 | Q9SZJ5 | Serine hydroxymethy-transferase | 18 | 46 | 187 | 8.1/57.5 | Metabolism/Defense | Mitochondrion | Gly and Ser metabolism Hypersensitive Response tetrahydrofolate interconversion response to abiotic stress | |
| 54 | Q9LJG3 | GDSL esterase/lipase ESM1 | 6 | 28 | 95 | 7.6/44.3 | Defense | Secreted | Responses to bacterium glucosinolate catabolism response to cold | |
| Protonated Masses m/z | Time after Treatment | |||||||
|---|---|---|---|---|---|---|---|---|
| Control Plants (ncps) | CP-Treated Plants (ncps) | |||||||
| 0.5 h | 2 h | 4 h | 24 h | 0.5 h | 2 h | 4 h | 24 h | |
| 27.022 | 6.5 ± 0.9 aA | 9.7 ± 3.4 abA | 13.9 ± 1.4 bB | 13.8 ± 2.7 bB | 8.1 ± 1.7 aA | 8.5 ± 2.2 aA | 8.3 ± 2.1 aA | 7.3 ± 3.5 aA |
| 33.033 | 90.4 ± 33.3 aA | 180.1 ± 45.4 bA | 157.9 ± 45.1 bA | 150.6 ± 43.1 bA | 93.0 ± 23.2 aA | 166.2 ± 50.4 bA | 150.7 ± 24.3 bA | 128.4 ± 33.1 abA |
| 45.033 | 83.8 ± 5.3 aA | 98.1 ± 13.2 abA | 93.2 ± 16.8 abA | 111.3 ± 11.4 bA | 100.7 ± 12.0 aB | 117.3 ± 39.2 abA | 114.9 ± 37.8 abA | 142.3 ± 14.6 bB |
| 49.000 | 4.3 ± 1.0 aA | 4.9 ± 0.7 aA | 5.3 ± 0.4 aA | 11.0 ± 1.6 bA | 4.5 ± 0.7 aA | 7.8 ± 2.3 bB | 8.3 ± 1.9 bB | 18.4 ± 4.1 cB |
| 55.055 | 3.6 ± 0.7 aA | 6.4 ± 0.3 bA | 8.8 ± 3.6 cbA | 11.4 ± 0.8 cA | 5.0 ± 1.1 aA | 7.6 ± 0.3 bB | 9.6 ± 3.6 bA | 15.5 ± 0.8 cB |
| 57.033 | 5.0 ± 0.8 aA | 7.9 ± 3.1 aA | 6.2 ± 1.7 aA | 7.5 ± 1.0 bA | 4.3 ± 1.8 aA | 4.9 ± 3.1 abA | 4.4 ± 1.6 aA | 7.7 ± 1.0 bA |
| 63.027 | 0.7 ± 0.4 aA | 0.4 ± 0.1 aA | 4.1 ± 0.1 bA | 4.1 ± 0.5 bA | 1.8 ± 0.8 aB | 0.9 ± 0.1 aB | 5.0 ± 0.2 bB | 3.4 ± 0.5 bA |
| 67.054 | 1.9 ± 0.4 aA | 4.1 ± 0.8 bB | 4.9 ± 1.2 bB | 4.8 ± 1.0 bB | 1.9 ± 0.4 aA | 2.8 ± 1.0 aA | 2.1 ± 0.3 aA | 2.2 ± 0.7 aA |
| 69.069 | 0.8 ± 0.4 aA | 2.2 ± 0.1 bA | 3.1 ± 0.4 cA | 4.8 ± 1.3 dA | 1.2 ± 0.5 aA | 2.8 ± 0.3 bB | 5.0 ± 0.4 cB | 10.0 ± 1.3 dB |
| 71.049 | 0.8 ± 0.4 aA | 2.1 ± 0.3 bA | 1.8 ± 0.5 bA | 2.4 ± 0.2 bA | 1.9 ± 0.4 aB | 2.8 ± 0.5 bA | 2.7 ± 0.3 bB | 4.8 ± 1.3 cB |
| 73.065 | 2.3 ± 1.2 aA | 3.3 ± 0.8 abA | 6.0 ± 2.3 bcA | 5.5 ± 1.6 bcA | 4.2 ± 2.0 aA | 5.2 ± 1.8 aA | 7.0 ± 2.0 abA | 9.5 ± 2.1 bB |
| 75.044 | 18.7 ± 4.7 aA | 27.1 ± 14.4 aA | 88.9 ± 20.7 cA | 59.9 ± 10.7 bA | 23.0 ± 9.3 aA | 42.0 ± 14.4 abA | 72.4 ± 20.7 cA | 62.3 ± 20.7 bA |
| 81.069 | 1.4 ± 0.4 aB | 5.8 ± 2.4 bA | 5.6 ± 1.4 bB | 4.6 ± 1.0 bA | 0.6 ± 0.3 aA | 3.5 ± 1.0 bA | 1.5 ± 0.9 aA | 3.7 ± 1.0 bA |
| 89.059 | 3.3 ± 1.8 aA | 3.3 ± 0.3 aA | 3.6 ± 1.4 aA | 3.7 ± 0.8 aA | 4.1 ± 1.0 aA | 13.0 ± 2.4 bB | 15.7 ± 3.5 bB | 14.6 ± 4.8 bB |
| 93.069 | 1.4 ± 0.4 aA | 6.7 ± 1.3 bB | 6.5 ± 0.9 bA | 5.7 ± 0.9 bB | 1.7 ± 0.3 aA | 3.6 ± 1.1 bA | 5.5 ± 1.6 bA | 4.1 ± 0.9 bA |
| 101.060 | 0.9 ± 0.4 aA | 1.4 ± 0.3 aA | 4.9 ± 0.6 bA | 5.1 ± 0.5 bA | 0.8 ± 0.1 aA | 1.8 ± 0.1 bB | 4.3 ± 0.9 cA | 7.6 ± 1.3 dB |
| Protonated Masses m/z | Tentative Identification | Chemical Formulae | Reference |
|---|---|---|---|
| 27.022 | Acetylene | C2H3+ | [26] |
| 33.033 | Methanol | CH5O+ | [26] |
| 45.033 | Acetaldehyde | C2H5O+ | [27] |
| 49.000 | methanethiol | CH5S+ | [28] |
| 55.055 | Alkyl fragment | C4H7+ | [29] |
| 57.033 | hexenal fragments | C3H5O+ | [30] |
| 63.027 | Dimethylsulfide (DMS) | C2H7S+ | [31] |
| 67.054 | Terpene fragment | C5H7+ | [32] |
| 69.069 | Isoprene | C5H9+ | [30] |
| 71.049 | 2-butenal | C4H7O+ | [29] |
| 73.065 | Isobutanal/Butanone | C4H9O+ | [29] |
| 75.044 | Methyl acetate/Propanoates | C3H7O2+ | [29] |
| 81.069 | Fragment/hexanal fragments | C6H9+ | [27] |
| 89.059 | Ethyl acetate/Methyl-propanoate | C4H9O2+ | [29] |
| 93.069 | terpenes alkyl fragment | C7H9+ | [32] |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Luti, S.; Caselli, A.; Taiti, C.; Bazihizina, N.; Gonnelli, C.; Mancuso, S.; Pazzagli, L. PAMP Activity of Cerato-Platanin during Plant Interaction: An -Omic Approach. Int. J. Mol. Sci. 2016, 17, 866. https://doi.org/10.3390/ijms17060866
Luti S, Caselli A, Taiti C, Bazihizina N, Gonnelli C, Mancuso S, Pazzagli L. PAMP Activity of Cerato-Platanin during Plant Interaction: An -Omic Approach. International Journal of Molecular Sciences. 2016; 17(6):866. https://doi.org/10.3390/ijms17060866
Chicago/Turabian StyleLuti, Simone, Anna Caselli, Cosimo Taiti, Nadia Bazihizina, Cristina Gonnelli, Stefano Mancuso, and Luigia Pazzagli. 2016. "PAMP Activity of Cerato-Platanin during Plant Interaction: An -Omic Approach" International Journal of Molecular Sciences 17, no. 6: 866. https://doi.org/10.3390/ijms17060866
APA StyleLuti, S., Caselli, A., Taiti, C., Bazihizina, N., Gonnelli, C., Mancuso, S., & Pazzagli, L. (2016). PAMP Activity of Cerato-Platanin during Plant Interaction: An -Omic Approach. International Journal of Molecular Sciences, 17(6), 866. https://doi.org/10.3390/ijms17060866