Spatial Temporal Dynamics and Molecular Evolution of Re-Emerging Rabies Virus in Taiwan
Abstract
:1. Introduction
2. Results
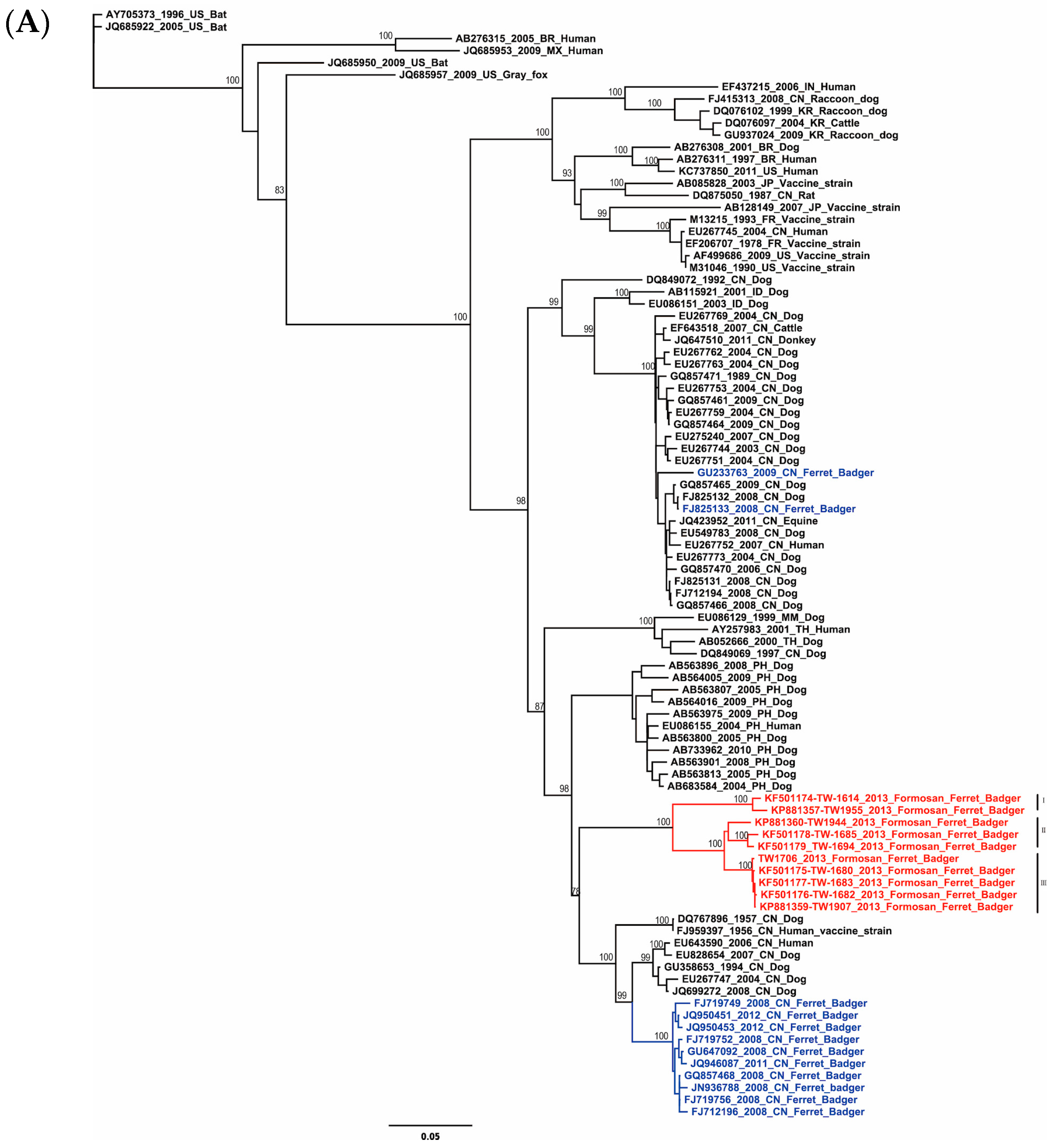
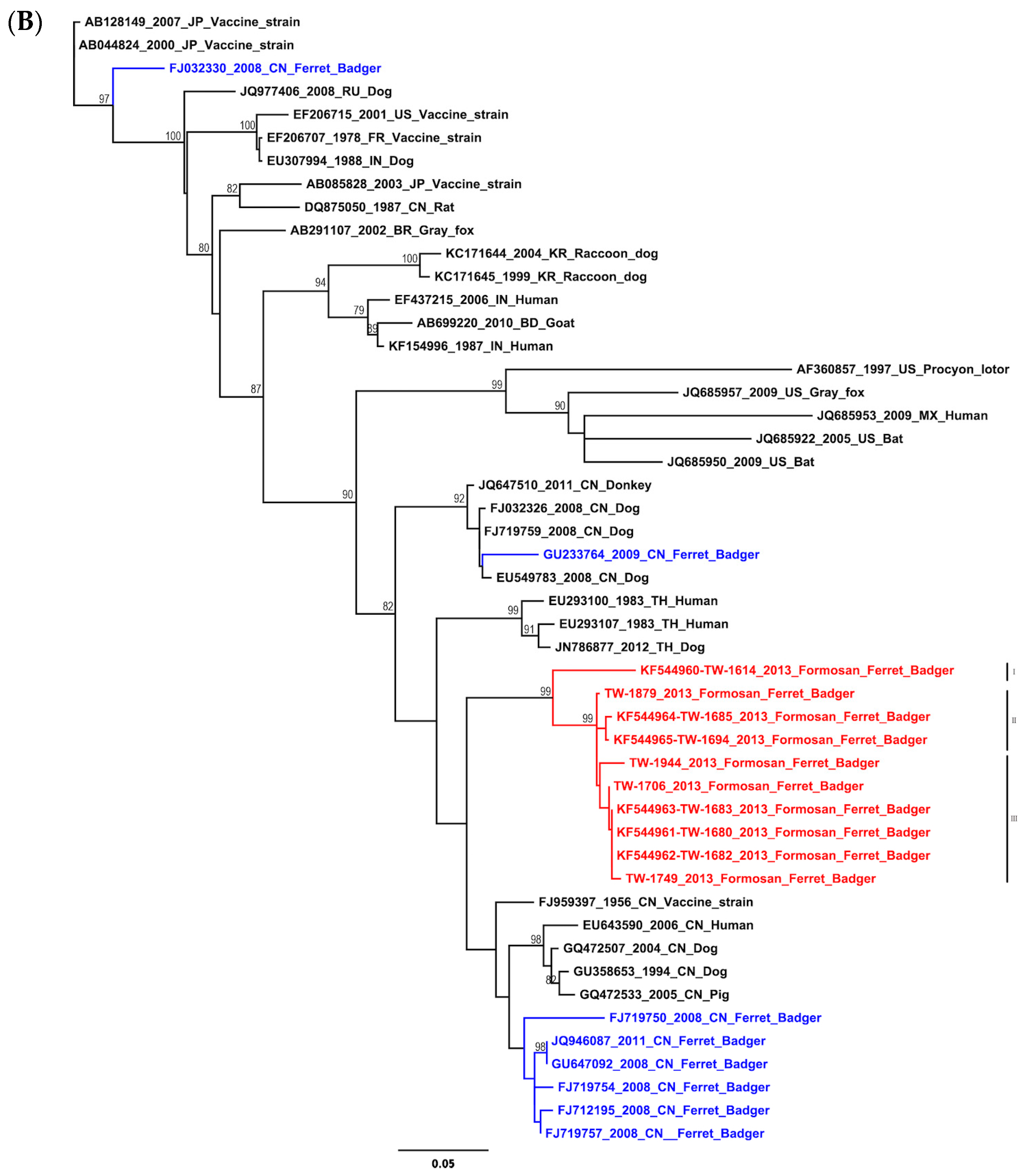
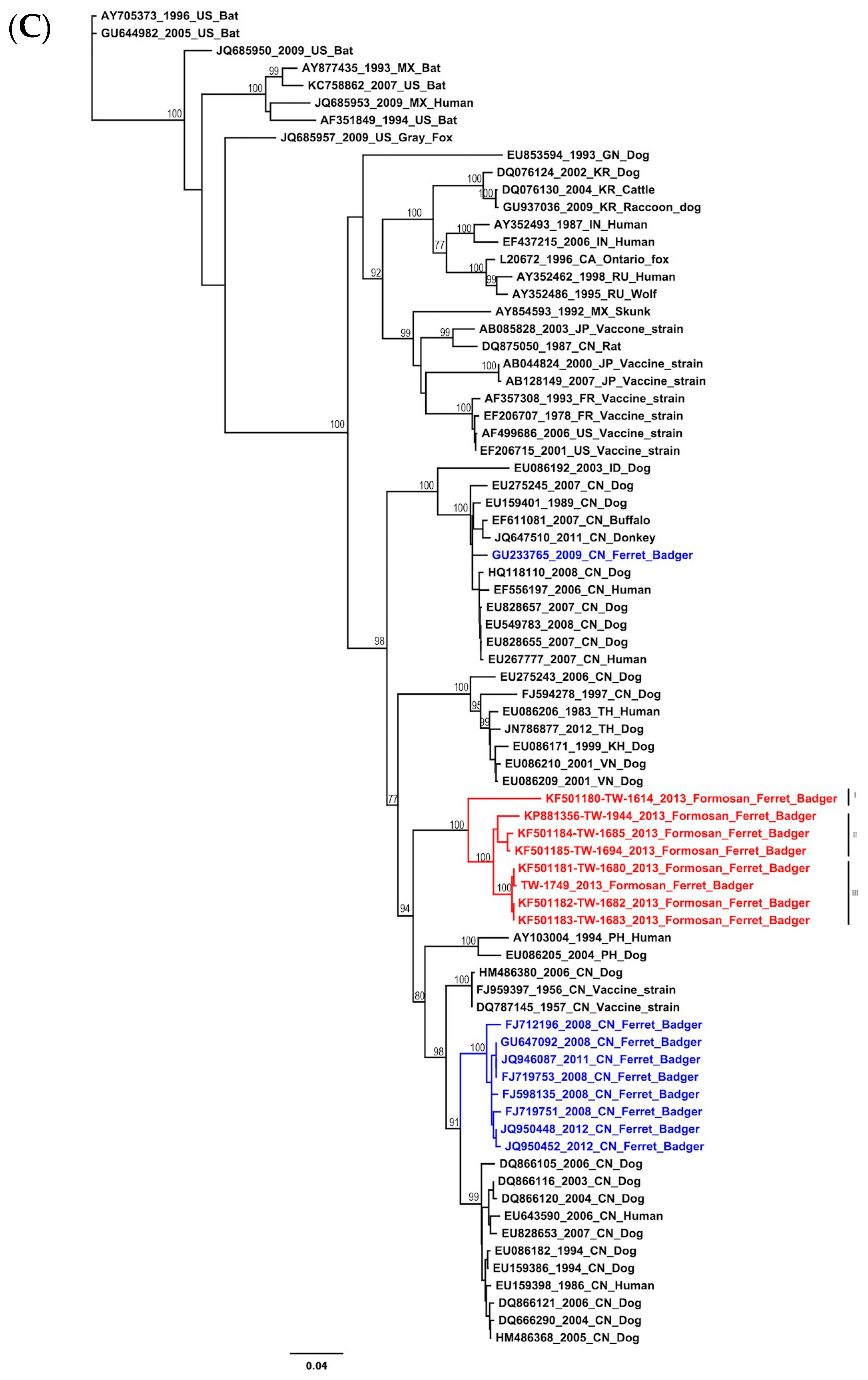
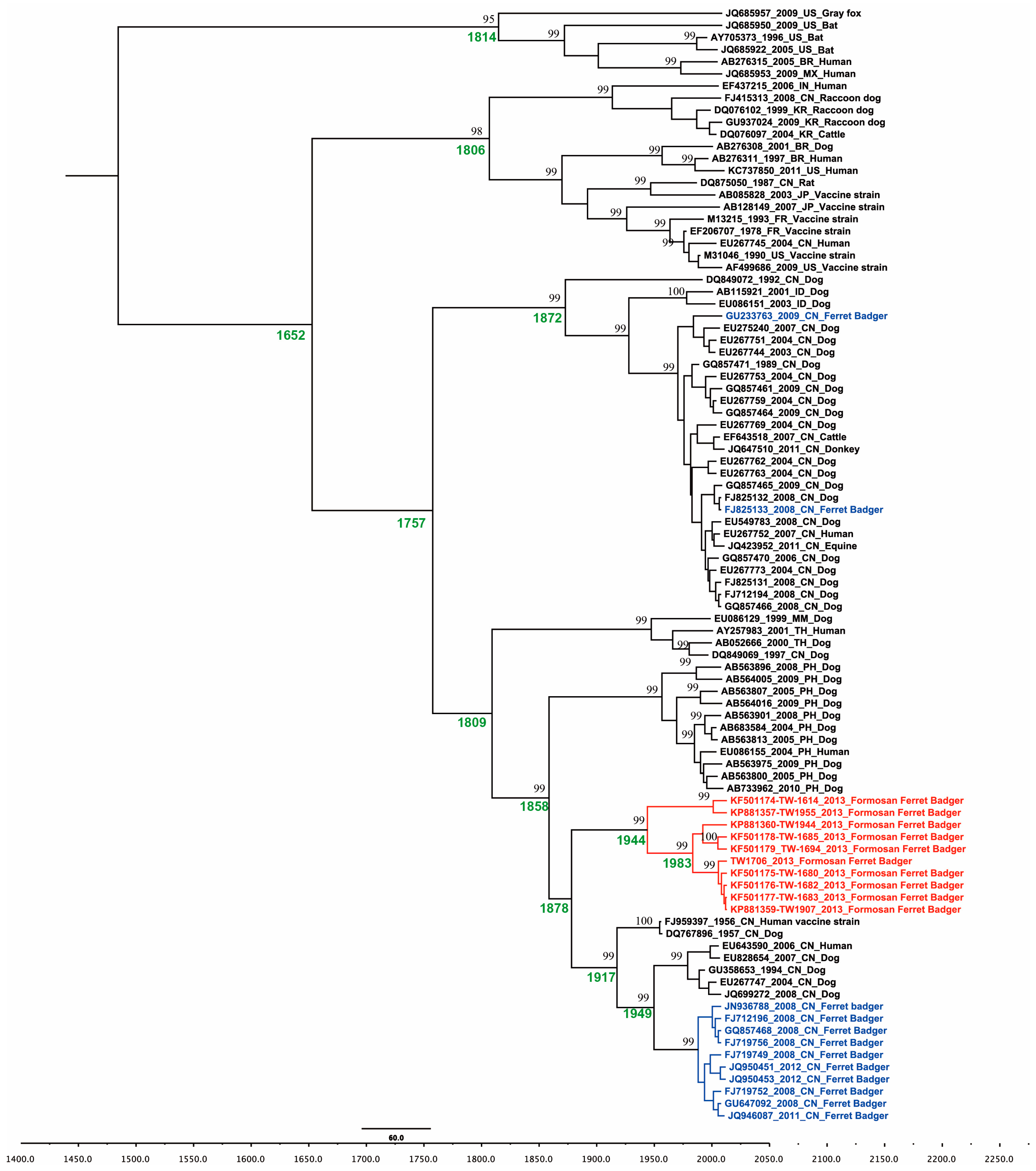
2.1. Phylogenetic and Molecular Evolution of the Rabies Virus
2.2. Selection Pressures and Co-Evolution RABV in the Formosan Ferret Badger
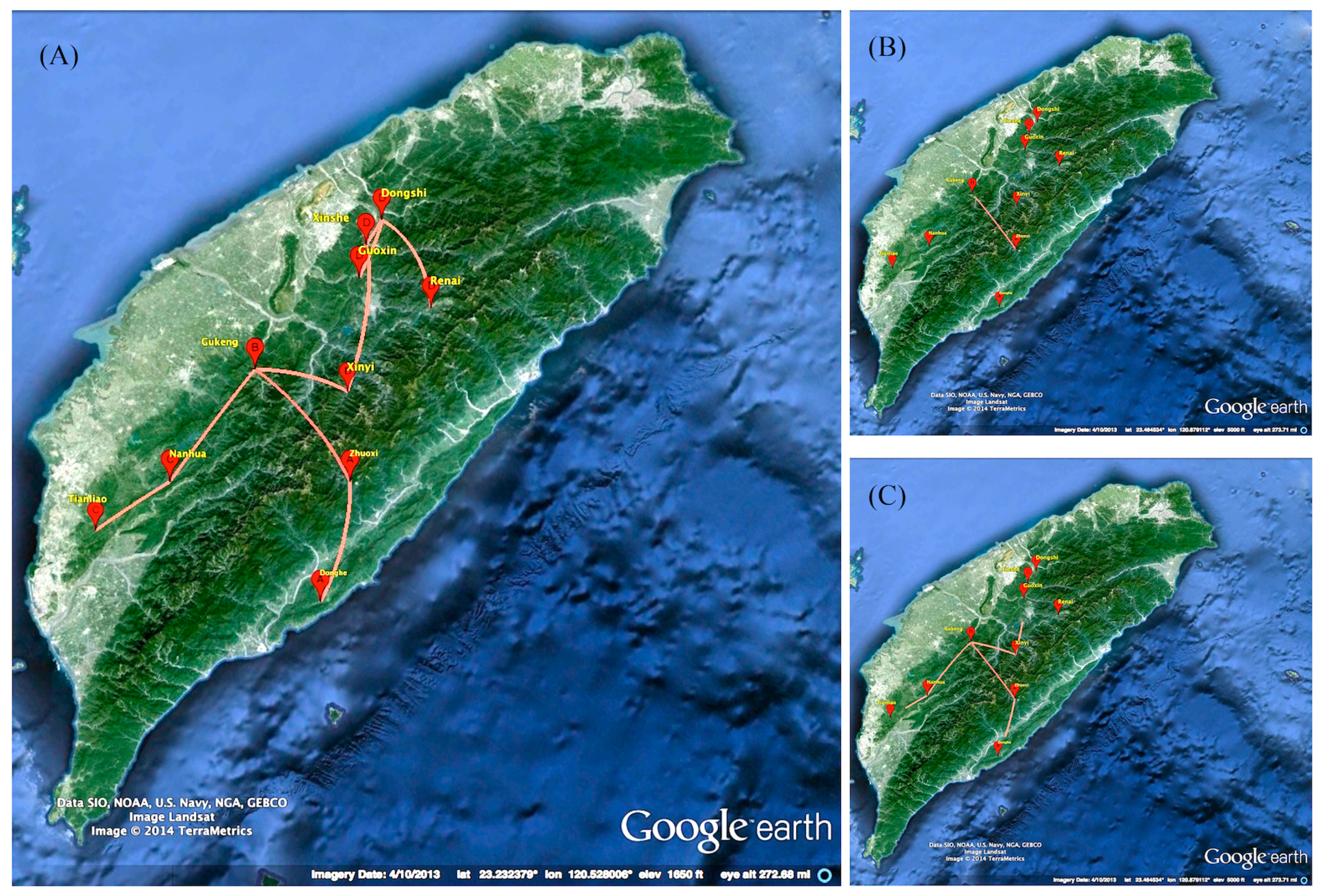
2.3. Phylogeographic of RABV in the Formosan Ferret Badger
3. Discussion
4. Materials and Methods
4.1. Source of Sequences
4.2. Phylogenetic Analysis
4.3. Phylodynamic and Phylogeography Analysis
4.4. Selection Pressure and Co-Evolution Analysis
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Knobel, D.L.; Cleaveland, S.; Coleman, P.G.; Fevre, E.M.; Meltzer, M.I.; Miranda, M.E.; Shaw, A.; Zinsstag, J.; Meslin, F.X. Re-evaluating the burden of rabies in Africa and Asia. Bull. World Health Organ. 2005, 83, 360–368. [Google Scholar] [PubMed]
- Warrell, M.J.; Warrell, D.A. Rabies and other lyssavirus diseases. Lancet 2004, 363, 959–969. [Google Scholar] [CrossRef]
- Carnieli, P., Jr.; Ruthner Batista, H.B.; de Novaes Oliveira, R.; Castilho, J.G.; Vieira, L.F. Phylogeographic dispersion and diversification of rabies virus lineages associated with dogs and crab-eating foxes (cerdocyon thous) in Brazil. Arch. Virol. 2013, 158, 2307–2313. [Google Scholar] [CrossRef] [PubMed]
- Carnieli, P., Jr.; de Novaes Oliveira, R.; Macedo, C.I.; Castilho, J.G. Phylogeography of rabies virus isolated from dogs in Brazil between 1985 and 2006. Arch. Virol. 2011, 156, 1007–1012. [Google Scholar] [CrossRef] [PubMed]
- Bourhy, H.; Reynes, J.M.; Dunham, E.J.; Dacheux, L.; Larrous, F.; Huong, V.T.; Xu, G.; Yan, J.; Miranda, M.E.; Holmes, E.C. The origin and phylogeography of dog rabies virus. J. Gen. Virol. 2008, 89, 2673–2681. [Google Scholar] [CrossRef] [PubMed]
- Delmas, O.; Holmes, E.C.; Talbi, C.; Larrous, F.; Dacheux, L.; Bouchier, C.; Bourhy, H. Genomic diversity and evolution of the lyssaviruses. PLoS ONE 2008, 3, e2057. [Google Scholar] [CrossRef] [PubMed]
- Rupprecht, C.E. Current Laboratory Techniques in Rabies Diagnosis, Research and Prevention; Elsevier: Amsterdam, The Netherlands, 2014; Volume 1, p. 350. [Google Scholar]
- Arechiga Ceballos, N.; Vazquez Moron, S.; Berciano, J.M.; Nicolas, O.; Aznar Lopez, C.; Juste, J.; Rodriguez Nevado, C.; Aguilar Setien, A.; Echevarria, J.E. Novel lyssavirus in bat, spain. Emerg. Infect. Dis. 2013, 19, 793–795. [Google Scholar] [CrossRef] [PubMed]
- Rupprecht, C.E.; Turmelle, A.; Kuzmin, I.V. A perspective on lyssavirus emergence and perpetuation. Curr. Opin. Virol. 2011, 1, 662–670. [Google Scholar] [CrossRef] [PubMed]
- Chiou, H.Y.; Hsieh, C.H.; Jeng, C.R.; Chan, F.T.; Wang, H.Y.; Pang, V.F. Molecular characterization of cryptically circulating rabies virus from ferret badgers, taiwan. Emerg. Infect. Dis. 2014, 20, 790–798. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.-H. History of rabies control in Taiwan and China. Taiwan Epidemiol. Bull. 2013, 29, 44–52. [Google Scholar]
- Poon, A.F.; Frost, S.D.; Pond, S.L. Detecting signatures of selection from DNA sequences using datamonkey. Methods Mol. Biol. 2009, 537, 163–183. [Google Scholar] [PubMed]
- Yang, Z.; Nielsen, R. Estimating synonymous and nonsynonymous substitution rates under realistic evolutionary models. Mol. Biol. Evol. 2000, 17, 32–43. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Zhang, S.; Wu, X.; Zhao, J.; Hou, Y.; Zhang, F.; Velasco-Villa, A.; Rupprecht, C.E.; Hu, R. Ferret badger rabies origin and its revisited importance as potential source of rabies transmission in southeast China. BMC Infect. Dis. 2010, 10. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Tang, Q.; Wu, X.; Liu, Y.; Zhang, F.; Rupprecht, C.E.; Hu, R. Rabies in ferret badgers, southeastern China. Emerg. Infect. Dis. 2009, 15, 946–949. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.; Wang, L.; Tao, X.; Li, H.; Rayner, S.; Liang, G.; Tang, Q. Genetic diversity and molecular evolution of the rabies virus matrix protein gene in China. Infect. Genet. Evol. 2013, 16, 248–253. [Google Scholar] [CrossRef] [PubMed]
- Kuzmina, N.A.; Kuzmin, I.V.; Ellison, J.A.; Taylor, S.T.; Bergman, D.L.; Dew, B.; Rupprecht, C.E. A reassessment of the evolutionary timescale of bat rabies viruses based upon glycoprotein gene sequences. Virus Genes 2013, 47, 305–310. [Google Scholar] [CrossRef] [PubMed]
- Meng, S.; Sun, Y.; Wu, X.; Tang, J.; Xu, G.; Lei, Y.; Wu, J.; Yan, J.; Yang, X.; Rupprecht, C.E. Evolutionary dynamics of rabies viruses highlights the importance of China rabies transmission in Asia. Virology 2011, 410, 403–409. [Google Scholar] [CrossRef] [PubMed]
- Talbi, C.; Holmes, E.C.; de Benedictis, P.; Faye, O.; Nakoune, E.; Gamatie, D.; Diarra, A.; Elmamy, B.O.; Sow, A.; Adjogoua, E.V.; et al. Evolutionary history and dynamics of dog rabies virus in western and central Africa. J. Gen. Virol. 2009, 90, 783–791. [Google Scholar] [CrossRef] [PubMed]
- Hughes, G.J.; Orciari, L.A.; Rupprecht, C.E. Evolutionary timescale of rabies virus adaptation to north American bats inferred from the substitution rate of the nucleoprotein gene. J. Gen. Virol. 2005, 86, 1467–1474. [Google Scholar] [CrossRef] [PubMed]
- Notredame, C.; Higgins, D.G.; Heringa, J. T-coffee: A novel method for fast and accurate multiple sequence alignment. J. Mol. Biol. 2000, 302, 205–217. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, H.A.; Strimmer, K.; Vingron, M.; von Haeseler, A. Tree-puzzle: Maximum likelihood phylogenetic analysis using quartets and parallel computing. Bioinformatics 2002, 18, 502–504. [Google Scholar] [CrossRef] [PubMed]
- Guindon, S.; Dufayard, J.F.; Lefort, V.; Anisimova, M.; Hordijk, W.; Gascuel, O. New algorithms and methods to estimate maximum-likelihood phylogenies: Assessing the performance of phyml 3.0. Syst. Biol. 2010, 59, 307–321. [Google Scholar] [CrossRef] [PubMed]
- Drummond, A.J.; Rambaut, A. Beast: Bayesian evolutionary analysis by sampling trees. BMC Evolut. Biol. 2007, 7. [Google Scholar] [CrossRef] [PubMed]
- Ayres, D.L.; Darling, A.; Zwickl, D.J.; Beerli, P.; Holder, M.T.; Lewis, P.O.; Huelsenbeck, J.P.; Ronquist, F.; Swofford, D.L.; Cummings, M.P.; et al. Beagle: An application programming interface and high-performance computing library for statistical phylogenetics. Syst. Biol. 2012, 61, 170–173. [Google Scholar] [CrossRef] [PubMed]
- Shapiro, B.; Rambaut, A.; Drummond, A.J. Choosing appropriate substitution models for the phylogenetic analysis of protein-coding sequences. Mol. Biol. Evol. 2006, 23, 7–9. [Google Scholar] [CrossRef] [PubMed]
- Drummond, A.J.; Suchard, M.A.; Xie, D.; Rambaut, A. Bayesian phylogenetics with beauti and the beast 1.7. Mol. Biol. Evol. 2012, 29, 1969–1973. [Google Scholar] [CrossRef] [PubMed]
- Baele, G.; Lemey, P.; Bedford, T.; Rambaut, A.; Suchard, M.A.; Alekseyenko, A.V. Improving the accuracy of demographic and molecular clock model comparison while accommodating phylogenetic uncertainty. Mol. Biol. Evol. 2012, 29, 2157–2167. [Google Scholar] [CrossRef] [PubMed]
- Bielejec, F.; Rambaut, A.; Suchard, M.A.; Lemey, P. Spread: Spatial phylogenetic reconstruction of evolutionary dynamics. Bioinformatics 2011, 27, 2910–2912. [Google Scholar] [CrossRef] [PubMed]
- Delport, W.; Poon, A.F.; Frost, S.D.; Kosakovsky Pond, S.L. Datamonkey 2010: A suite of phylogenetic analysis tools for evolutionary biology. Bioinformatics 2010, 26, 2455–2457. [Google Scholar] [CrossRef] [PubMed]
- Poon, A.F.; Lewis, F.I.; Frost, S.D.; Kosakovsky Pond, S.L. Spidermonkey: Rapid detection of co-evolving sites using bayesian graphical models. Bioinformatics 2008, 24, 1949–1950. [Google Scholar] [CrossRef] [PubMed]
| Gene | tMRCA | Substitution Rates Sub/Site/Year (10−4) | Mean Relative Substitution Rate | Standard Error of Mean |
|---|---|---|---|---|
| Nucleoprotein | 721.62 (406.06–1096.16) | 2.49 (1.40–3.63) | - | - |
| Ferret badger (Taiwan) | 170.37 (83.22–270.70) | - | - | - |
| Ferret badger (China) | 428.67 (243.95–644.24) | - | - | - |
| 1st + 2nd codon position | - | - | 0.22 (0.19–0.26) | 1.26 × 10−4 |
| 3rd codon position | - | - | 2.55 (2.48–2.62) | 2.52 × 10−4 |
| Matrix protein | 523.03 (218.04–931.53) | 4.03 (1.63–6.55) | - | - |
| Ferret badger (Taiwan) | 89.55 (32.37–166.28) | - | - | - |
| Ferret badger (China) | 391.62 (173.13–692.32) | - | - | - |
| 1st + 2nd codon position | - | - | 0.30 (0.24–0.36) | 2.05 × 10−4 |
| 3rd codon position | - | - | 2.395 (2.28–2.51) | 4.09 × 10−4 |
| Glycoprotein | 630.69 (206.44–1309.38) | 4.75 (2.05–7.32) | - | - |
| Ferret badger (Taiwan) | 75.68 (26.03–136.00) | - | - | - |
| Ferret badger (China) | 306.95 (105.09–578.58) | - | - | - |
| 1st + 2nd codon position codon | - | - | 0.37 (0.33–0.40) | 1.37 × 10−4 |
| 3rd codon position | - | - | 2.26 (2.19–2.34) | 2.75 × 10−4 |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lin, Y.-C.; Chu, P.-Y.; Chang, M.-Y.; Hsiao, K.-L.; Lin, J.-H.; Liu, H.-F. Spatial Temporal Dynamics and Molecular Evolution of Re-Emerging Rabies Virus in Taiwan. Int. J. Mol. Sci. 2016, 17, 392. https://doi.org/10.3390/ijms17030392
Lin Y-C, Chu P-Y, Chang M-Y, Hsiao K-L, Lin J-H, Liu H-F. Spatial Temporal Dynamics and Molecular Evolution of Re-Emerging Rabies Virus in Taiwan. International Journal of Molecular Sciences. 2016; 17(3):392. https://doi.org/10.3390/ijms17030392
Chicago/Turabian StyleLin, Yung-Cheng, Pei-Yu Chu, Mei-Yin Chang, Kuang-Liang Hsiao, Jih-Hui Lin, and Hsin-Fu Liu. 2016. "Spatial Temporal Dynamics and Molecular Evolution of Re-Emerging Rabies Virus in Taiwan" International Journal of Molecular Sciences 17, no. 3: 392. https://doi.org/10.3390/ijms17030392
APA StyleLin, Y.-C., Chu, P.-Y., Chang, M.-Y., Hsiao, K.-L., Lin, J.-H., & Liu, H.-F. (2016). Spatial Temporal Dynamics and Molecular Evolution of Re-Emerging Rabies Virus in Taiwan. International Journal of Molecular Sciences, 17(3), 392. https://doi.org/10.3390/ijms17030392








