Development of Cell-SELEX Technology and Its Application in Cancer Diagnosis and Therapy
Abstract
:1. Introduction
2. Advantages and Limitations of Cell-SELEX
2.1. Advantages of Cell-SELEX
2.2. Limitations of Cell-SELEX
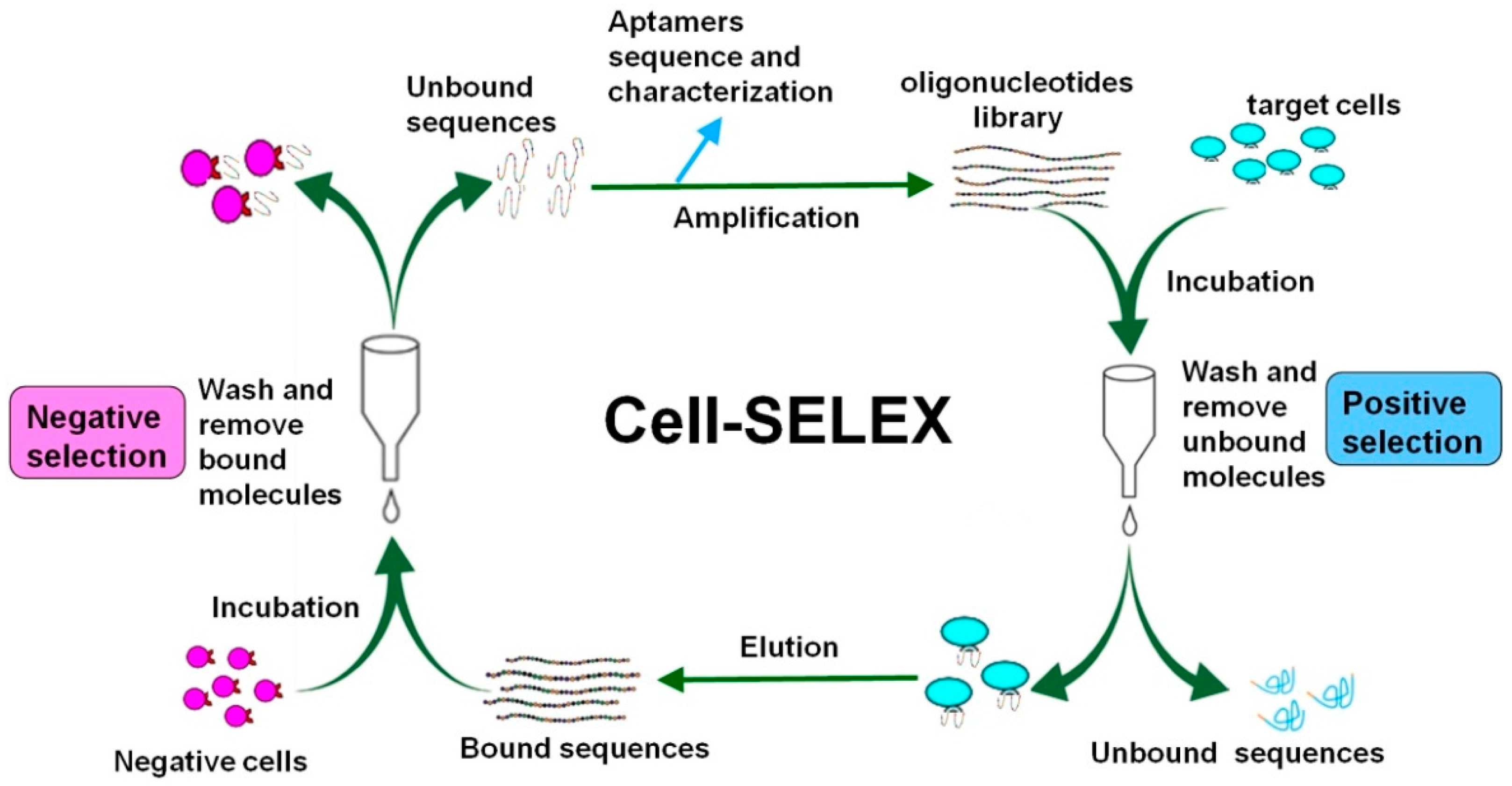
3. Cell-SELEX Strategy and Procedure
4. New Methods Derived from Cell-SELEX
4.1. TECS-SELEX
4.2. FACS-SELEX
4.3. 3D Cell-SELEX
4.4. Hybrid-SELEX
4.5. Cell-Internalization SELEX
5. Applications of Cell-SELEX in Cancer Diagnosis and Therapy
5.1. Application of Cell-SELEX Aptamers in Cancer Diagnosis
5.2. Application of Cell-SELEX Aptamers in Cancer Therapy
6. Future Perspectives
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Ma, H.; Liu, J. Nucleic acid aptamers in cancer research, diagnosis and therapy. Chem. Soc. Rev. 2015, 44, 1240–1256. [Google Scholar] [CrossRef] [PubMed]
- Song, K.M.; Lee, S. Aptamers and their biological applications. Sensors 2012, 12, 612–631. [Google Scholar] [CrossRef] [PubMed]
- Tuerk, C.; Gold, L. Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science 1990, 249, 505–510. [Google Scholar] [CrossRef] [PubMed]
- Ellington, A.D.; Szostak, J.W. In vitro selection of RNA molecules that bind specific ligands. Nature 1990, 346, 818–822. [Google Scholar] [CrossRef] [PubMed]
- Sun, H.; Zhu, X. Oligonucleotide aptamers: New tools for targeted cancer therapy. Mol. Ther. Nucleic Acids 2014, 3, 182. [Google Scholar] [CrossRef] [PubMed]
- Gopinath, S.C.B. Methods developed for SELEX. Anal. Bioanal. Chem. 2007, 387, 171–182. [Google Scholar] [CrossRef] [PubMed]
- Morris, K.N.; Jensen, K.B.; Julin, C.M.; Weil, M.; Gold, L. High affinity ligands from in vitro selection: Complex targets. Proc. Natl. Acad. Sci. USA 1998, 95, 2902–2907. [Google Scholar] [CrossRef] [PubMed]
- Sefah, K.; Shangguan, D.; Xiong, X.; O’Donoghue, M.B.; Tan, W. Development of DNA aptamers using Cell-SELEX. Nat. Protoc. 2010, 5, 1169–1185. [Google Scholar] [CrossRef] [PubMed]
- Fang, X.H.; Tan, W.H. Aptamers generated from cell-SELEX for molecular medicine: A chemical biology approach. Acc. Chem. Res. 2010, 43, 48–57. [Google Scholar] [CrossRef] [PubMed]
- Shangguan, D.; Cao, Z.; Meng, L. Cell-specific aptamer probes for membrane protein elucidation in cancer cells. J. Proteome Res. 2008, 7, 2133–2139. [Google Scholar] [CrossRef] [PubMed]
- Guo, K.T.; Ziemer, G.; Paul, A.; Wendel, H.P. CELL-SELEX: Novel perspectives of aptamer-based therapeutics. Int. J. Mol. Sci. 2008, 9, 668–678. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.C.; Yates, J.R. The application of mass spectrometry to membrane proteomics. Nat. Biotechnol. 2003, 21, 262–267. [Google Scholar] [CrossRef] [PubMed]
- Mallikaratchy, P.; Tang, Z.; Kwame, S.; Meng, L.; Shangguan, D.; Tan, W. Aptamer directly evolved from live cells recognizes membrane bound immunoglobin heavy mu chain in Burkitt’s lymphoma cells. Mol. Cell. Proteom. 2008, 6, 2230–2238. [Google Scholar] [CrossRef] [PubMed]
- Tang, Z.; Shangguan, D. Selection of aptamers for molecular recognition and characterization of cancer cells. Anal Chem. 2007, 79, 4900–4907. [Google Scholar] [CrossRef] [PubMed]
- Raddatz, M.S.; Dolf, A. Enrichment of cell-targeting and population-specific aptamers by fluorescence-activated cell sorting. Angew. Chem. Int. Ed. Engl. 2008, 47, 5190–5193. [Google Scholar] [CrossRef] [PubMed]
- Avci-Adali, M.; Metzger, M.; Perle, N.; Ziemer, G.; Wendel, H.P. Pitfalls of cell-systematic evolution of ligands by exponential enrichment (SELEX): Existing dead cells during in vitro selection anticipate the enrichment of specific aptamers. Oligonucleotides 2010, 20, 317–323. [Google Scholar] [CrossRef] [PubMed]
- Cox, J.C.; Rajendran, M.; Riedel, T. Automated acquisition of aptamer sequences. Comb. Chem. High Throughput Screen. 2002, 5, 289–299. [Google Scholar] [PubMed]
- Cox, J.C.; Ellington, A.D. Automated selection of anti-Protein aptamers. Biorg. Med. Chem. 2001, 9, 2525–2531. [Google Scholar] [CrossRef]
- Zhou, J.; Rossi, J. Aptamers as targeted therapeutics: Current potential and challenges. Nat. Rev. Drug Discov. 2016. [Google Scholar] [CrossRef] [PubMed]
- Darmostuk, M.; Rimpelova, S. Current approaches in SELEX: An update to aptamer selection technology. Biotechnol. Adv. 2015, 33, 1141–1161. [Google Scholar] [CrossRef] [PubMed]
- Mayer, G.; Ahmed, M.S. Fluorescence-activated cell sorting for aptamer SELEX with cell mixtures. Nat. Protoc. 2010, 5, 1993–2004. [Google Scholar] [CrossRef] [PubMed]
- Souza, A.G.; Marangoni, K. 3D Cell-SELEX: Development of RNA aptamers as molecular probes for PC-3 tumor cell line. Exp. Cell Res. 2016, 341, 147–156. [Google Scholar] [CrossRef] [PubMed]
- Bowser, M.T. SELEX: Just another separation. Analyst 2005, 130, 128–130. [Google Scholar] [CrossRef] [PubMed]
- Kuwahara, M.; Obika, S. In vitro selection of BNA (LNA) aptamers. Artif. DNA 2013, 4, 39–48. [Google Scholar] [CrossRef] [PubMed]
- Ninomiya, K.; Kaneda, K.; Kawashima, S.; Miyachi, Y.; Ogino, C.; Shimizu, N. Cell-SELEX based selection and characterization of DNA aptamer recognizing human hepatocarcinoma. Bioorg. Med. Chem. Lett. 2013, 23, 1797–1802. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Luo, Y.; Bing, T. DNA aptamer evolved by cell-SELEX for recognition of prostate cancer. PLoS ONE 2014, 9, e100243. [Google Scholar] [CrossRef] [PubMed]
- Shigdar, S.; Qiao, L.; Zhou, S.F. RNA aptamers targeting cancer stem cell marker CD133. Cancer Lett. 2013, 330, 84–95. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.K.; Kuo, T.L. Subtractive SELEX against two heterogeneous target samples: Numerical simulations and analysis. Comput. Biol. Med. 2007, 37, 750–759. [Google Scholar] [CrossRef] [PubMed]
- Gyllensten, U.B.; Erlich, H.A. Generation of single-stranded DNA by the polymerase chain reaction and its application to direct sequencing of the HLA-DQA locus. Proc. Natl. Acad. Sci. USA 1988, 85, 7652–7656. [Google Scholar] [CrossRef] [PubMed]
- Dickman, M.; Hornby, D.P. Isolation of single-stranded DNA using denaturing DNA chromatography. Anal. Biochem. 2000, 284, 164–167. [Google Scholar] [CrossRef] [PubMed]
- Higuchi, R.G.; Ochman, H. Production of single-stranded DNA templates by exonuclease digestion following the polymerase chain reaction. Nucleic Acids Res. 1989, 17, 5865. [Google Scholar] [CrossRef] [PubMed]
- Liang, C.; Li, D. Comparison of the methods for generating single-stranded DNA in SELEX. Analyst 2015, 140, 3439–3444. [Google Scholar] [CrossRef] [PubMed]
- Huafang, G.; Shengce, T.; Dong, W. Comparison of different methods for preparing single stranded DNA for oligonucleotide microarray. Anal. Lett. 2003, 13, 2849–2863. [Google Scholar]
- Ohuchi, S. Cell-SELEX technology. Biores Open Access 2012, 1, 265–272. [Google Scholar] [CrossRef] [PubMed]
- Cerchia, L.; Ducongé, F. Neutralizing aptamers from whole-cell SELEX inhibit the RET receptor tyrosine kinase. PLoS Biol. 2005, 3, 123. [Google Scholar] [CrossRef] [PubMed]
- Ohuchi, S.P.; Ohtsu, T.P.; Nakamura, Y. Selection of RNA aptamers against recombinant transforming growth factor-beta type III receptor displayed on cell surface. Biochimie 2006, 88, 897–904. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.W.; Kim, E.Y. Identification of DNA aptamers toward epithelial cell adhesion molecule via cell-SELEX. Mol. Cells 2014, 37, 742–746. [Google Scholar] [CrossRef] [PubMed]
- Hicke, B.J.; Marion, C. Tenascin-C aptamers are generated using tumor cells and purified protein. J. Biol. Chem. 2001, 276, 48644–48654. [Google Scholar] [CrossRef] [PubMed]
- Boltz, A.; Piater, B. Bi-specific aptamers mediating tumor cell lysis. J. Biol. Chem. 2011, 286, 21896–21905. [Google Scholar] [CrossRef] [PubMed]
- Thiel, W.H.; Thiel, K.W. Cell-internalization SELEX: Method for identifying cell-internalizing RNA aptamers for delivering siRNAs to target cells. Method Mol. Biol. 2015, 1218, 187–199. [Google Scholar]
- Thiel, K.W.; Hernandez, L.I. Delivery of chemo-sensitizing siRNAs to her2+-breast cancer cells using RNA aptamers. Nucleic Acids Res. 2012, 40, 6319–6337. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.Z.; Hernandez, F.J. RNA aptamer-based functional ligands of the neurotrophin receptor, TrkB. Mol. Pharmacol. 2012, 82, 623–635. [Google Scholar] [CrossRef] [PubMed]
- Shangguan, D.; Li, Y. Aptamers evolved from live cells as effective molecular probes for cancer study. Proc. Natl. Acad. Sci. USA 2006, 103, 11838–11843. [Google Scholar] [CrossRef] [PubMed]
- Sefah, K.; Meng, L.; Lopezcolon, D. DNA aptamers as molecular probes for colorectal cancer study. PLoS ONE 2010, 5, e14269. [Google Scholar] [CrossRef] [PubMed]
- Bamrungsap, S.; Chen, T.; Shukoor, M.I. Pattern recognition of cancer cells using aptamer-conjugated magnetic nanoparticles. ACS Nano 2012, 6, 3974–3981. [Google Scholar] [CrossRef] [PubMed]
- Hicke, B.J.; Stephens, A.W.T. Tumor targeting by an aptamer. Nucl. Med. 2006, 47, 668–678. [Google Scholar]
- Girvan, A.C.; Teng, Y.; Casson, L.K. AGRO100 inhibits activation of nuclear factor-κB (NF-κB) by forming a complex with NF-κB essential modulator (NEMO) and nucleolin. Mol. Cancer Ther. 2006, 5, 1790–1799. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, C.S.M.; Matthews, C.S.; Missailidis, S. DNA aptamers that bind to MUC1 tumour marker: Design and characterization of MUC1-binding single-stranded DNA aptamers. Tumour Biol. 2006, 7, 289–301. [Google Scholar] [CrossRef] [PubMed]
- Kang, W.J.; Chae, J.R.; Cho, Y.L.; Lee, J.; Kim, S. Multiplex imaging of single tumor cells using quantum-dot-conjugated aptamers. Small 2009, 22, 2519–2522. [Google Scholar] [CrossRef] [PubMed]
- Hung, M.C.; Matin, A.; Zhang, Y. HER-2/NEU-targeting gene therapy—A review. Gene 1995, 159, 65–71. [Google Scholar] [CrossRef]
- Niazi, J.H.; Verma, S.K. In vitro HER2 protein-induced affinity dissociation of carbon nanotube-wrapped anti-HER2 aptamers for HER2 protein detection. Analyst 2015, 140, 243–249. [Google Scholar] [CrossRef] [PubMed]
- Rudge, J.S.; Holash, J. VEGF Trap complex formation measures production rates of VEGF, providing a biomarker for predicting efficacious angiogenic blockade. Proc. Natl. Acad. Sci. USA 2007, 104, 18363–18370. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Schluesener, H.J. Pigpen is a cellular binding protein of therapeutic oligonucleotides. Cytotherapy 2005, 7, 186–194. [Google Scholar] [CrossRef] [PubMed]
- Mhawech-Fauceglia, P.; Zhang, S.; Terracciano, L. Prostate-specific membrane antigen (PSMA) protein expression in normal and neoplastic tissues and its sensitivity and specificity in prostate adenocarcinoma: an immunohistochemical study using multiple tumour tissue microarray technique. Histopathology 2007, 50, 472–483. [Google Scholar] [CrossRef] [PubMed]
- Zwick, E.; Bange, J. Receptor tyrosine kinase signalling as a target for cancer intervention strategies. Endocr. Relat. Cancer 2001, 8, 161–173. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Chen, Y.; Han, D.; Ocsoy, I.; Tan, W. Aptamers selected by cell-SELEX for application in cancer studies. Bioanalysis 2010, 2, 907–918. [Google Scholar] [CrossRef] [PubMed]
- Lupold, S.E.; Hicke, B.J. Identification and characterization of nuclease-stabilized RNA molecules that bind human prostate cancer cells via the prostate-specific membrane antigen. Cancer Res. 2002, 62, 4029–4033. [Google Scholar] [PubMed]
- Farokhzad, O.C.; Jon, S. Nanoparticle-aptamer bioconjugates: A new approach for targeting prostate cancer cell. Cancer Res. 2004, 64, 7668–7672. [Google Scholar] [CrossRef] [PubMed]
- Farokhzad, O.C.; Cheng, J. Targeted nanoparticle-aptamer bioconjugates for cancer chemotherapy in vivo. Proc. Natl. Acad. Sci. USA 2006, 103, 6315–6320. [Google Scholar] [CrossRef] [PubMed]
- Taghdisi, S.M.; Danesh, N.M. Targeted delivery of Epirubicin to cancer cells by PEGylated A10 aptamer. J. Drug Target 2013, 21, 739–744. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.; Chen, J.; Wu, M. Aptamers: active targeting ligands for cancer diagnosis and therapy. Theranostics. 2015, 5, 322–344. [Google Scholar] [CrossRef] [PubMed]
- Bates, P.J.; Laber, D.A.; Miller, D.M.; Thomas, S.D.; Trent, J.O. Discovery and development of the G-rich oligonucleotide AS1411 as a novel treatment for cancer. Exp. Mol. Pathol. 2009, 86, 151–164. [Google Scholar] [CrossRef] [PubMed]
- Rosenberg, J.E.; Bambury, R.M. A phase II trial of AS1411 (a novel nucleolin-targeted DNA aptamer) in metastatic renal cell carcinoma. Investig. New Drug 2014, 32, 178–187. [Google Scholar] [CrossRef] [PubMed]
- Lu, M.; Zhou, L.; Zheng, X. A novel molecular marker of breast cancer stem cells identified by cell-SELEX method. Cancer Biomark. 2015, 15, 163–170. [Google Scholar] [PubMed]
- Mallikaratchy, P.; Tang, Z.; Kwame, S. Aptamer directly evolved from live cells recognizes membrane bound immunoglobin heavy Mu chain in Burkitt′s lymphoma cells. Mol. Cell. Proteom. 2007, 6, 2230–2238. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, C.S.M.; Cheung, M.C. Phototoxic aptamers selectively enter and kill epithelial cancer cells. Nucleic Acids Res. 2009, 37, 866–876. [Google Scholar] [CrossRef] [PubMed]
- Cerchia, L.; Esposito, C.L.; Jacobs, A.H. Differential SELEX in human glioma cell lines. PLoS ONE 2009, 4, 7971. [Google Scholar] [CrossRef] [PubMed]
- Cerchia, L.; Esposito, C.L. Targeting Axl with a high-affinity inhibitory aptamer. Mol. Ther. 2012, 20, 2291–2303. [Google Scholar] [CrossRef] [PubMed]
- Dastjerdi, K.; Tabar, G.H. Generation of an enriched pool of DNA aptamers for an HER2-overexpressing cell line selected by Cell SELEX. Biotechnol. Appl. Biochem. 2011, 58, 226–230. [Google Scholar] [CrossRef] [PubMed]
- Shigdar, S.; Lin, J. RNA aptamer against a cancer stem cell marker epithelial cell adhesion molecule. Cancer Sci. 2011, 102, 991–998. [Google Scholar] [CrossRef] [PubMed]
- Subramanian, N.; Kanwar, J.R. EpCAM aptamer mediated cancer cell specific delivery of EpCAM siRNA using polymeric nanocomplex. J. Biomed. Sci. 2015, 22, 4. [Google Scholar] [CrossRef] [PubMed]
- Dua, P.; Kang, H.S.; Hong, S.M.; Tsao, M.S.; Kim, S.; Lee, D.K. Alkaline phosphatase ALPPL-2 is a novel pancreatic carcinoma-associated protein. Cancer Res. 2013, 73, 1934–1945. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Satheesan, S. Cell-Specific RNA aptamer against human CCR5 specifically targets HIV-1 susceptible cells and inhibits HIV-1 infectivity. Chem. Biol. 2015, 22, 379–390. [Google Scholar] [CrossRef] [PubMed]
- Rong, Y.; Chen, H.; Zhou, X.F. Identification of an aptamer through whole cell-SELEX for targeting high metastatic liver cancers. Oncotarget 2016, 7, 8282–8294. [Google Scholar] [PubMed]
- Soldevilla, M.M.; Villanueva, H. MRP1-CD28 bi-specific oligonucleotide aptamers: Target costimulation to drug-resistant melanoma cancer stem cells. Oncotarget 2016, 7, 23182–23196. [Google Scholar] [CrossRef] [PubMed]
- Zhu, H.; Li, J. Nucleic acid aptamer-mediated drug delivery for targeted cancer therapy. Chem. Med. Chem. 2015, 10, 39–45. [Google Scholar] [CrossRef] [PubMed]
- Mayer, G. The chemical biology of aptamers. Angew. Chem. Int. Ed. 2009, 48, 2672–2689. [Google Scholar] [CrossRef] [PubMed]
- Oney, S.; Lam, R.T.S. Development of universal antidotes to control aptamer activity. Nat. Med. 2009, 15, 1224–1228. [Google Scholar] [CrossRef] [PubMed]
- Ye, M.; Hu, J.; Peng, M. Generating aptamers by cell-SELEX for applications in molecular medicine. Int. J. Mol. Sci. 2012, 13, 3341–3353. [Google Scholar] [CrossRef] [PubMed]
- Hwang, D.W.; Ko, H.Y. A nucleolin-targeted multimodal nanoparticle imaging probe for tracking cancer cells using an aptamer. J. Nucl. Med. 2010, 51, 98–105. [Google Scholar] [CrossRef] [PubMed]
| Target | Method | Aptamer | Applications | Year | Reference |
|---|---|---|---|---|---|
| RET | cell-SELEX | RNA | Metastatic breast cancer, gastrointestinal stromal tumors and non-small cell lung cancer therapy | 2005 | [35] |
| PTK7 | cell-SELEX | DNA | Acute lymphoblastic leukemia therapy and diagnosis | 2006 | [10,43] |
| Immunoglobin Heavy Mu Chain (IGHM) | cell-SELEX | DNA | Burkitt lymphoma diagnosis and therapy | 2007 | [14,65] |
| O-Glycan-Peptide Signatures | cell-SELEX | DNA | Targeted delivery by conjugation with chlorine e6 (Ce6) for epithelial cells cancer therapy | 2009 | [66] |
| AXL | cell-SELEX | RNA | human glioma cell cancer therapy | 2009 | [67,68] |
| CD16α (FcγRIIIα) | hybrid-SELEX | DNA | Cancer immunotherapy via mediating cellular cytotoxicity | 2011 | [39] |
| HER2 | cell-SELEX | DNA | HER2 positive breast cancer therapy and diagnosis | 2011 | [69] |
| Epithelial Cell Adhesion Molecule (EpCAM) | Cell-SELEX | RNA | Novel targeted nanomedicine and molecular imaging agents for cancer theranostics | 2011 | [70,71] |
| CD133 | cell-SELEX | RNA | Cancer stem cell targeted therapeutics and molecular imaging. | 2013 | [27] |
| Alkaline Phosphatase Placental-Like 2 (ALPPL-2) | cell-SELEX | RNA | Pancreatic carcinoma diagnosis and therapy | 2013 | [72] |
| CD44/CD24 | cell-SELEX | DNA | Breast cancer diagnosis and therapy | 2014 | [64] |
| The C-C Chemokine Receptor Type 5 (CCR5) | cell-SELEX | RNA | HIV therapy | 2015 | [73] |
| Cytokeratin 19 | cell-SELEX | DNA | Metastatic hepatocellular carcinoma diagnosis and chemotherapy | 2016 | [74] |
| MRP1 | hybrid-SELEX | RNA | Binding MRP1-expressing tumors and delivering the CD28 costimulatory signal to tumor-infiltrating lymphocytes. | 2016 | [75] |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chen, M.; Yu, Y.; Jiang, F.; Zhou, J.; Li, Y.; Liang, C.; Dang, L.; Lu, A.; Zhang, G. Development of Cell-SELEX Technology and Its Application in Cancer Diagnosis and Therapy. Int. J. Mol. Sci. 2016, 17, 2079. https://doi.org/10.3390/ijms17122079
Chen M, Yu Y, Jiang F, Zhou J, Li Y, Liang C, Dang L, Lu A, Zhang G. Development of Cell-SELEX Technology and Its Application in Cancer Diagnosis and Therapy. International Journal of Molecular Sciences. 2016; 17(12):2079. https://doi.org/10.3390/ijms17122079
Chicago/Turabian StyleChen, Man, Yuanyuan Yu, Feng Jiang, Junwei Zhou, Yongshu Li, Chao Liang, Lei Dang, Aiping Lu, and Ge Zhang. 2016. "Development of Cell-SELEX Technology and Its Application in Cancer Diagnosis and Therapy" International Journal of Molecular Sciences 17, no. 12: 2079. https://doi.org/10.3390/ijms17122079
APA StyleChen, M., Yu, Y., Jiang, F., Zhou, J., Li, Y., Liang, C., Dang, L., Lu, A., & Zhang, G. (2016). Development of Cell-SELEX Technology and Its Application in Cancer Diagnosis and Therapy. International Journal of Molecular Sciences, 17(12), 2079. https://doi.org/10.3390/ijms17122079






