Cloning, Characterization and Expression Pattern Analysis of a Cytosolic Copper/Zinc Superoxide Dismutase (SaCSD1) in a Highly Salt Tolerant Mangrove (Sonneratia alba)
Abstract
:1. Introduction
2. Results
2.1. Cloning of SaCSD1 and Sequence Analysis

| Species | GenBank Accession Number | Identity |
|---|---|---|
| Tetradium ruticarpum | AFF57842.1 | 90% |
| Bruguiera gymnorhiza | BAB78597.1 | 85% |
| Litchi chinensis | ABY65355.1 | 89% |
| Avicennia marina | ACA50531.1 | 84% |
| Arabidopsis thaliana | P24704.2 | 86% |
| Oryza sativa | AAA33917.1 | 85% |
| Cenchrus americanus | ABP65325.1 | 87% |
| Zea mays | NP001105704.1 | 85% |
| Bambusa oldhamii | ACX94084.1 | 87% |
2.2. Expression, Purification, and Western Blot Analysis of SaCSD1
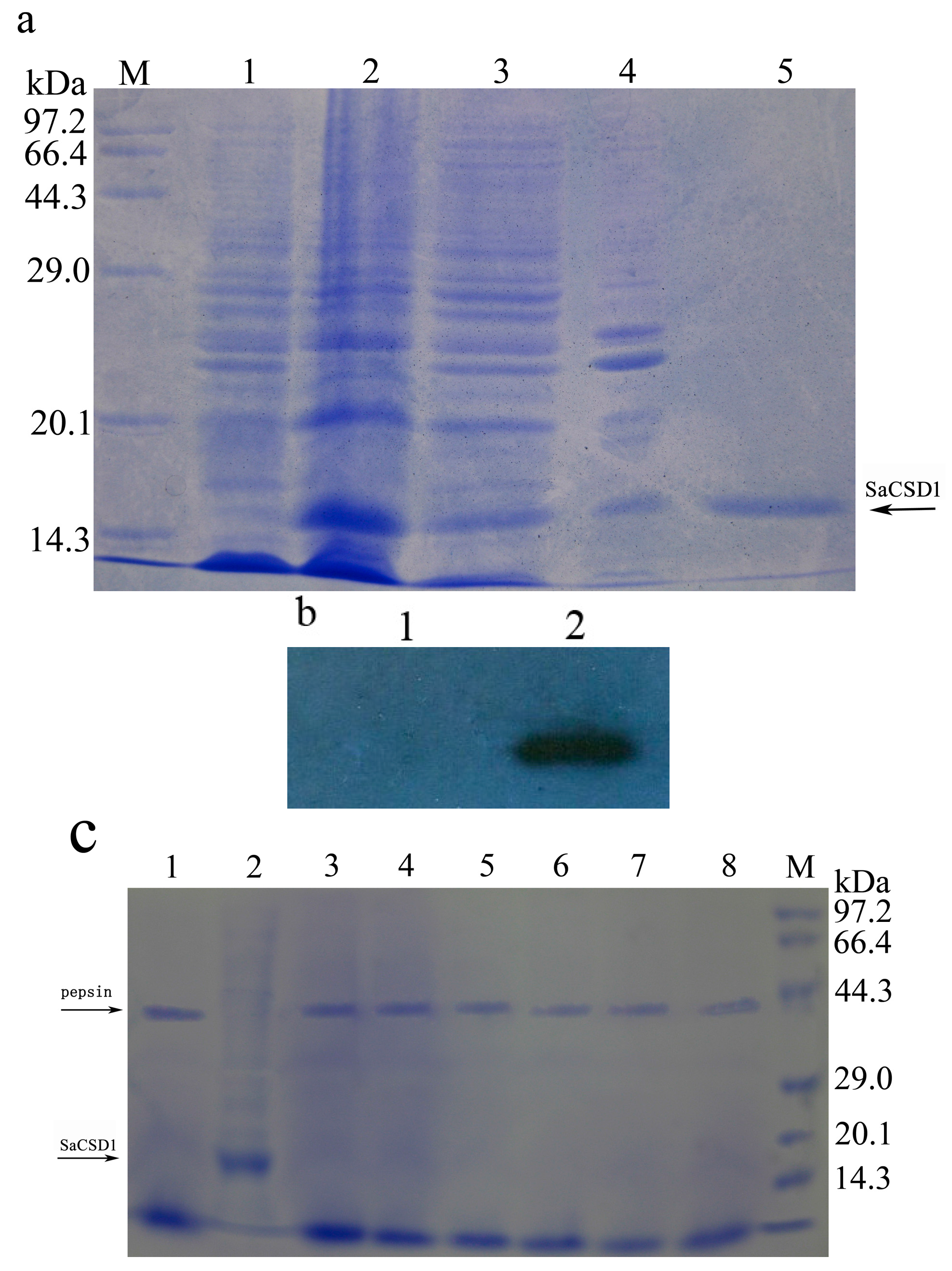
2.3. Effects of pH, Temperature, Chemicals, and Salt on Activity of SaCSD1
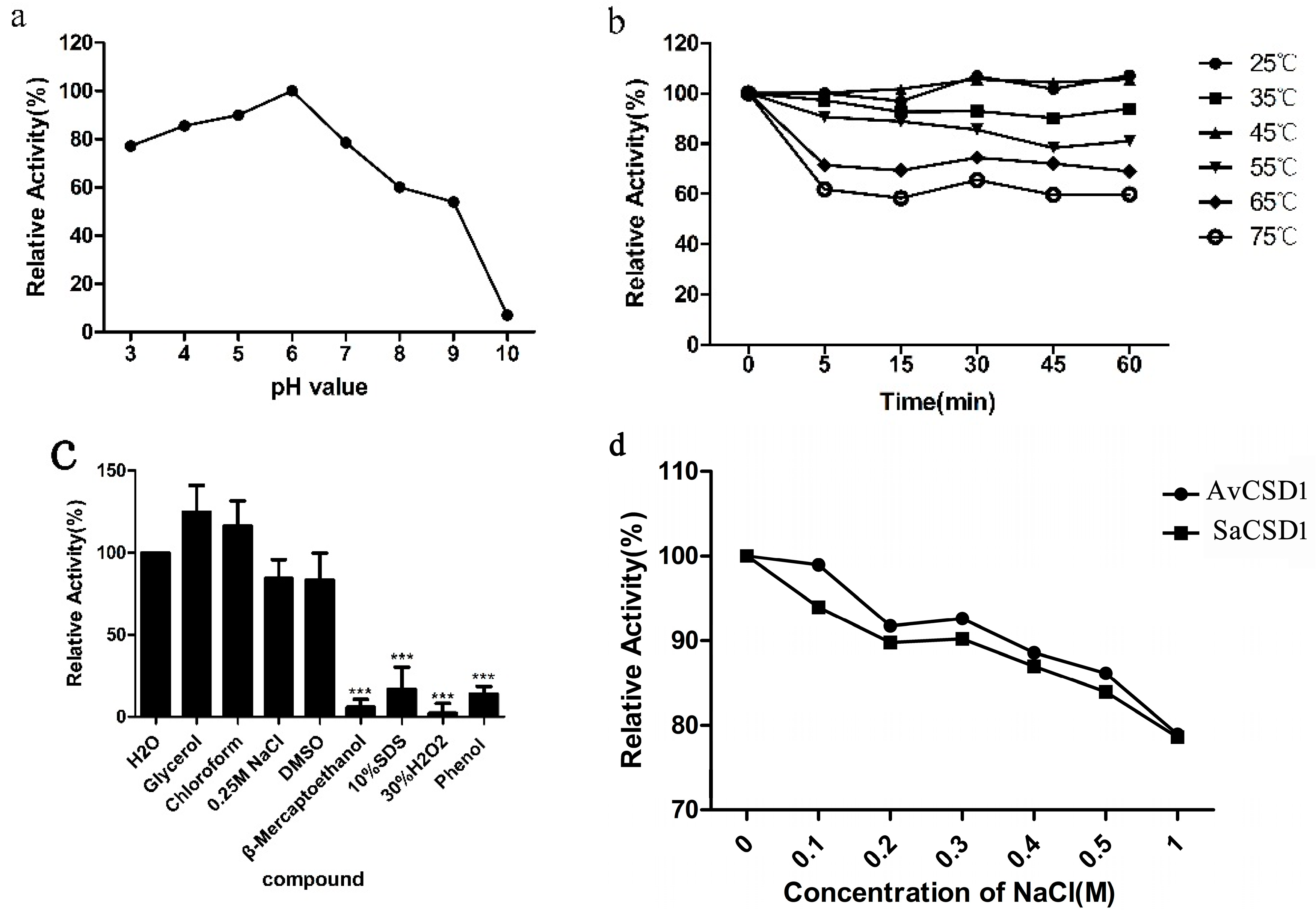
2.4. Pepsin Digestion Characteristics of SaCSD1
2.5. Quantification of SaCSD1 Gene Expression in Different Organs and under Salt Stress
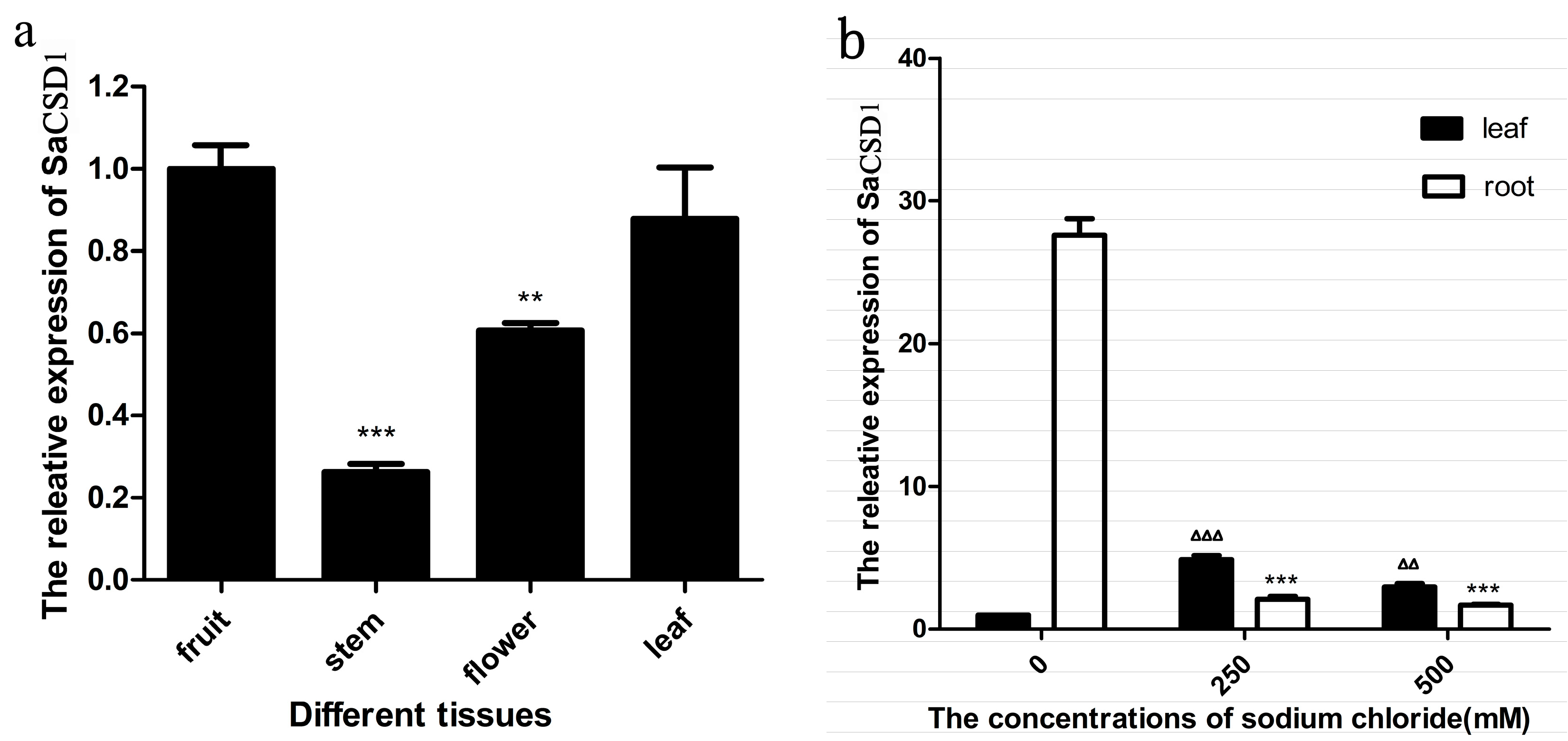
3. Discussion
4. Materials and Methods
4.1. Materials
4.2. RNA Preparation, cDNA Synthesis and Cloning
4.3. Nucleotide and Amino Acid Sequence Analysis of SaCSD1
4.4. Cloning into the Expression Vector, Expression and Purification of SaCSD1
4.5. Western Blot Analysis
4.6. Determination of Protein Concentration and Activity Assay of SaCSD1
4.7. Pepsin Digestion Assay Conditions
4.8. Quantification of SaCSD1 Gene Expression by Real-Time PCR
4.9. Statistical Analysis
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
References
- Donato, D.C.; Kauffman, J.B.; Murdiyarso, D.; Kurnianto, S.; Stidham, M.; Kanninen, M. Mangroves among the most carbon-rich forests in the tropics. Nat. Geosci. 2011, 4, 293–297. [Google Scholar] [CrossRef]
- Liang, S.; Zhou, R.; Dong, S.; Shi, S. Adaptation to salinity in mangroves: Implication on the evolution of salt-tolerance. Chin. Sci. Bull. 2008, 53, 1708–1715. [Google Scholar] [CrossRef]
- Berlyn, G.P. Coastal trees: The botany of mangroves. Science 1986, 234, 373. [Google Scholar] [CrossRef] [PubMed]
- Ball, M.; Pidsley, S. Growth responses to salinity in relation to distribution of two mangrove species, Sonneratia alba and s. Lanceolata, in northern australia. Funct. Ecol. 1995, 77–85. [Google Scholar] [CrossRef]
- Suzuki, M.; Yasumoto, E.; Baba, S.; Ashihara, H. Effect of salt stress on the metabolism of ethanolamine and choline in leaves of the betaine-producing Mangrove species Avicennia marina. Phytochemistry 2003, 64, 941–948. [Google Scholar] [CrossRef]
- Jithesh, M.N.; Prashanth, S.R.; Sivaprakash, K.R.; Parida, A. Monitoring expression profiles of antioxidant genes to salinity, iron, oxidative, light and hyperosmotic stresses in the highly salt tolerant grey mangrove, Avicennia marina (forsk.) vierh. by mRNA analysis. Plant Cell Rep. 2006, 25, 865–876. [Google Scholar] [CrossRef] [PubMed]
- Parida, A.K.; Das, A.B.; Mohanty, P. Investigations on the antioxidative defence responses to nacl stress in a mangrove, Bruguiera parviflora: Differential regulations of isoforms of some antioxidative enzymes. Plant Growth Regul. 2004, 42, 213–226. [Google Scholar] [CrossRef]
- Miller, R.A.; Britigan, B.E. Role of oxidants in microbial pathophysiology. Clin. Microbiol. Rev. 1997, 10, 1–18. [Google Scholar] [PubMed]
- Prashanth, S.R.; Sadhasivam, V.; Parida, A. Over expression of cytosolic copper/zinc superoxide dismutase from a mangrove plant Avicennia marina in indica rice var pusa basmati-1 confers abiotic stress tolerance. Transgenic Res. 2008, 17, 281–291. [Google Scholar] [CrossRef] [PubMed]
- Brouwer, M.; Brouwer, T.H.; Grater, W.; Brown-Peterson, N. Replacement of a cytosolic copper/zinc superoxide dismutase by a novel cytosolic manganese superoxide dismutase in crustaceans that use copper (haemocyanin) for oxygen transport. Biochem. J. 2003, 374, 219–228. [Google Scholar] [CrossRef] [PubMed]
- Kochhar, S.; Kochhar, V.K. Expression of Antioxidant enzymes and heat shock proteins in relation to combined stress of cadmium and heat in Vigna mungo seedlings. Plant Sci. 2005, 168, 921–929. [Google Scholar] [CrossRef]
- Cheeseman, J.M.; Herendeen, L.B.; Cheeseman, A.T.; Clough, B.F. Photosynthesis and photoprotection in mangroves under field conditions. Plant Cell Environ. 1997, 20, 579–588. [Google Scholar] [CrossRef]
- Parida, A.K.; Das, A.B.; Mohanty, P. Defense potentials to nacl in a mangrove, Bruguiera parviflora: Differential changes of isoforms of some antioxidative enzymes. J. Plant Physiol. 2004, 161, 531–542. [Google Scholar] [CrossRef] [PubMed]
- Lin, C.T.; Yeh, K.W.; Kao, M.C.; Shaw, J.F. Cloning and characterization of a cDNA encoding the cytosolic copper/zinc-superoxide dismutase from sweet potato tuberous root. Plant Mol. Biol. 1993, 23, 911–913. [Google Scholar] [CrossRef] [PubMed]
- Scioli, J.R.; Zilinskas, B.A. Cloning and characterization of a cDNA encoding the chloroplastic copper/zinc-superoxide dismutase from pea. Proc. Natl. Acad. Sci. USA 1988, 85, 7661–7665. [Google Scholar] [CrossRef] [PubMed]
- Kaminaka, H.; Morita, S.; Yokoi, H.; Masumura, T.; Tanaka, K. Molecular cloning and characterization of a cDNA for plastidic copper/zinc-superoxide dismutase in rice (Oryza sativa L.). Plant Cell Physiol. 1997, 38, 65–69. [Google Scholar] [CrossRef] [PubMed]
- Cui, H.; Zhang, T. Application of SOD in Food and Cosmetic and Its Production by rmentation. Pharm. Biotechnol. 2000, 7, 187–189. (In Chinese) [Google Scholar]
- Dutta, S.; Padhye, S.; Ahmed, F.; Sarkar, F. Pyridazolate-bridged dicopper (II) sod mimics with enhanced antiproliferative activities against estrogen and androgen independent cancer cell lines. Inorg. Chim. Acta 2005, 358, 3617–3624. [Google Scholar] [CrossRef]
- Madamanchi, N.R.; Vendrov, A.; Runge, M.S. Oxidative stress and vascular disease. Arterioscl. Throm. Vas. 2005, 25, 29–38. [Google Scholar] [CrossRef] [PubMed]
- Edeas, M.A.; Emerit, I.; Khalfoun, Y.; Lazizi, Y.; Cernjavski, L.; Levy, A.; Lindenbaum, A. Clastogenic factors in plasma of HIV-1 infected patients activate HIV-1 replication in vitro: Inhibition by superoxide dismutase. Free Radic. Biol. Med. 1997, 23, 571–578. [Google Scholar] [CrossRef]
- Bafana, A.; Dutt, S.; Kumar, A.; Kumar, S.; Ahuja, P.S. The basic and applied aspects of superoxide dismutase. J. Mol. Catal. B Enzym. 2011, 68, 129–138. [Google Scholar] [CrossRef]
- Lee, F.J.S.; Hassan, H.M. Biosynthesis of superoxide dismutase and catalase in chemostat culture of Saccharomyces cerevisiae. Appl. Microbiol. Biotechnol. 1987, 26, 531–536. [Google Scholar] [CrossRef]
- Xie, X.; Pan, X.; Gong, L.; Chen, H.; Lin, L. Effects of Some Organic Solvents on the Activity and Conformation of Superoxide Dismutase (SOD) from Porcine Blood. J. Quanzhou Norm. Univ. 2010, 28, 47–50. (In Chinese) [Google Scholar]
- Wang, F.; Li, X.Y.; Mo, X.M.; Zhang, G.; Sun, H.X. A biologically active vMIP-II-IgG3-TfN fusion protein, secreted from methylotrophic yeast pichia pastoris. Protein Express. Purif. 2013, 87, 47–54. [Google Scholar] [CrossRef] [PubMed]
- Thomas, K.; Aalbers, M.; Bannon, G.A.; Bartels, M.; Dearman, R.J.; Esdaile, D.J.; Fu, T.J.; Glatt, C.M.; Hadfield, N.; Hatzos, C.; et al. A multi-laboratory evaluation of a common in vitro pepsin digestion assay protocol used in assessing the safety of novel proteins. Regul. Toxicol. Pharm. 2004, 39, 87–98. [Google Scholar] [CrossRef] [PubMed]
- Grattan, S.R.; Grieve, C.M. Salinity–mineral nutrient relations in horticultural crops. Sci. Hortic. 1998, 78, 127–157. [Google Scholar] [CrossRef]
- Zhu, J.K. Regulation of ion homeostasis under salt stress. Curr. Opin. Plant Biol. 2003, 6, 441–445. [Google Scholar] [CrossRef]
- Lee, D.H.; Kim, Y.S.; Lee, C.B. The inductive responses of the antioxidant enzymes by salt stress in the rice (Oryza sativa L.). J. Plant Physiol. 2001, 158, 737–745. [Google Scholar] [CrossRef]
- Tsai, Y.C.; Hong, C.Y.; Liu, L.F.; Kao, C.H. Relative importance of na+ and cl- in nacl-induced antioxidant systems in roots of rice seedlings. Physiol. Plant. 2004, 122, 86–94. [Google Scholar] [CrossRef]
- Mittler, R. Oxidative stress, antioxidants and stress tolerance. Trends Plant Sci. 2002, 7, 405–410. [Google Scholar] [CrossRef]
- Van Breusegem, F.; Vranova, E.; Dat, J.F.; Inze, D. The role of active oxygen species in plant signal transduction. Plant Sci. 2001, 161, 405–414. [Google Scholar] [CrossRef]
- Sen Raychaudhuri, S.; Deng, X.W. The role of superoxide dismutase in combating oxidative stress in higher plants. Bot. Rev. 2000, 66, 89–98. [Google Scholar] [CrossRef]
- Bandeoglu, E.; Eyidogan, F.; Yucel, M.; Oktem, H.A. Antioxidant responses of shoots and roots of lentil to nacl-salinity stress. Plant Growth Regul. 2004, 42, 69–77. [Google Scholar] [CrossRef]
- Gomez, J.M.; Jimenez, A.; Olmos, E.; Sevilla, F. Location and effects of long-term nacl stress on superoxide dismutase and ascorbate peroxidase isoenzymes of pea (Pisum sativum cv. Puget) chloroplasts. J. Exp. Bot. 2004, 55, 119–130. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.F.; Zhou, R.C.; Huang, Y.L.; Zhang, M.; Yang, G.L.; Zhong, C.R.; Shi, S.H. Transcriptome sequencing of a highly salt tolerant mangrove species Sonneratia alba using illumina platform. Mar. Genom. 2011, 4, 129–136. [Google Scholar] [CrossRef] [PubMed]
- Bao, Y.; Li, L.; Wu, Q.; Zhang, G. Cloning, characterization, and expression analysis of extracellular copper/zinc superoxide dismutase gene from bay scallop argopecten irradians. Fish Shellfish Immunol. 2009, 27, 17–25. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.F.; Li, Y.Y.; Pang, C.H.; Lu, C.M.; Wang, B.S. Nacl enhances thylakoid-bound sod activity in the leaves of c-3 halophyte Suaeda salsa L. Plant Sci. 2005, 168, 423–430. [Google Scholar]
- Laemmli, U.K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 1970, 227, 680–685. [Google Scholar] [CrossRef] [PubMed]
- Li, D.C.; Gao, J.; Li, Y.L.; Lu, J. A thermostable manganese-containing superoxide dismutase from the thermophilic fungus thermomyces lanuginosus. Extrem. Life Extreme Cond. 2005, 9, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCt method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yang, E.; Yi, S.; Bai, F.; Niu, D.; Zhong, J.; Wu, Q.; Chen, S.; Zhou, R.; Wang, F. Cloning, Characterization and Expression Pattern Analysis of a Cytosolic Copper/Zinc Superoxide Dismutase (SaCSD1) in a Highly Salt Tolerant Mangrove (Sonneratia alba). Int. J. Mol. Sci. 2016, 17, 4. https://doi.org/10.3390/ijms17010004
Yang E, Yi S, Bai F, Niu D, Zhong J, Wu Q, Chen S, Zhou R, Wang F. Cloning, Characterization and Expression Pattern Analysis of a Cytosolic Copper/Zinc Superoxide Dismutase (SaCSD1) in a Highly Salt Tolerant Mangrove (Sonneratia alba). International Journal of Molecular Sciences. 2016; 17(1):4. https://doi.org/10.3390/ijms17010004
Chicago/Turabian StyleYang, Enze, Shanze Yi, Fang Bai, Dewei Niu, Junjie Zhong, Qiuhong Wu, Shufang Chen, Renchao Zhou, and Feng Wang. 2016. "Cloning, Characterization and Expression Pattern Analysis of a Cytosolic Copper/Zinc Superoxide Dismutase (SaCSD1) in a Highly Salt Tolerant Mangrove (Sonneratia alba)" International Journal of Molecular Sciences 17, no. 1: 4. https://doi.org/10.3390/ijms17010004
APA StyleYang, E., Yi, S., Bai, F., Niu, D., Zhong, J., Wu, Q., Chen, S., Zhou, R., & Wang, F. (2016). Cloning, Characterization and Expression Pattern Analysis of a Cytosolic Copper/Zinc Superoxide Dismutase (SaCSD1) in a Highly Salt Tolerant Mangrove (Sonneratia alba). International Journal of Molecular Sciences, 17(1), 4. https://doi.org/10.3390/ijms17010004






