Transcriptome Analysis and Identification of Differentially Expressed Transcripts of Immune-Related Genes in Spleen of Gosling and Adult Goose
Abstract
:1. Introduction
2. Results and Discussion
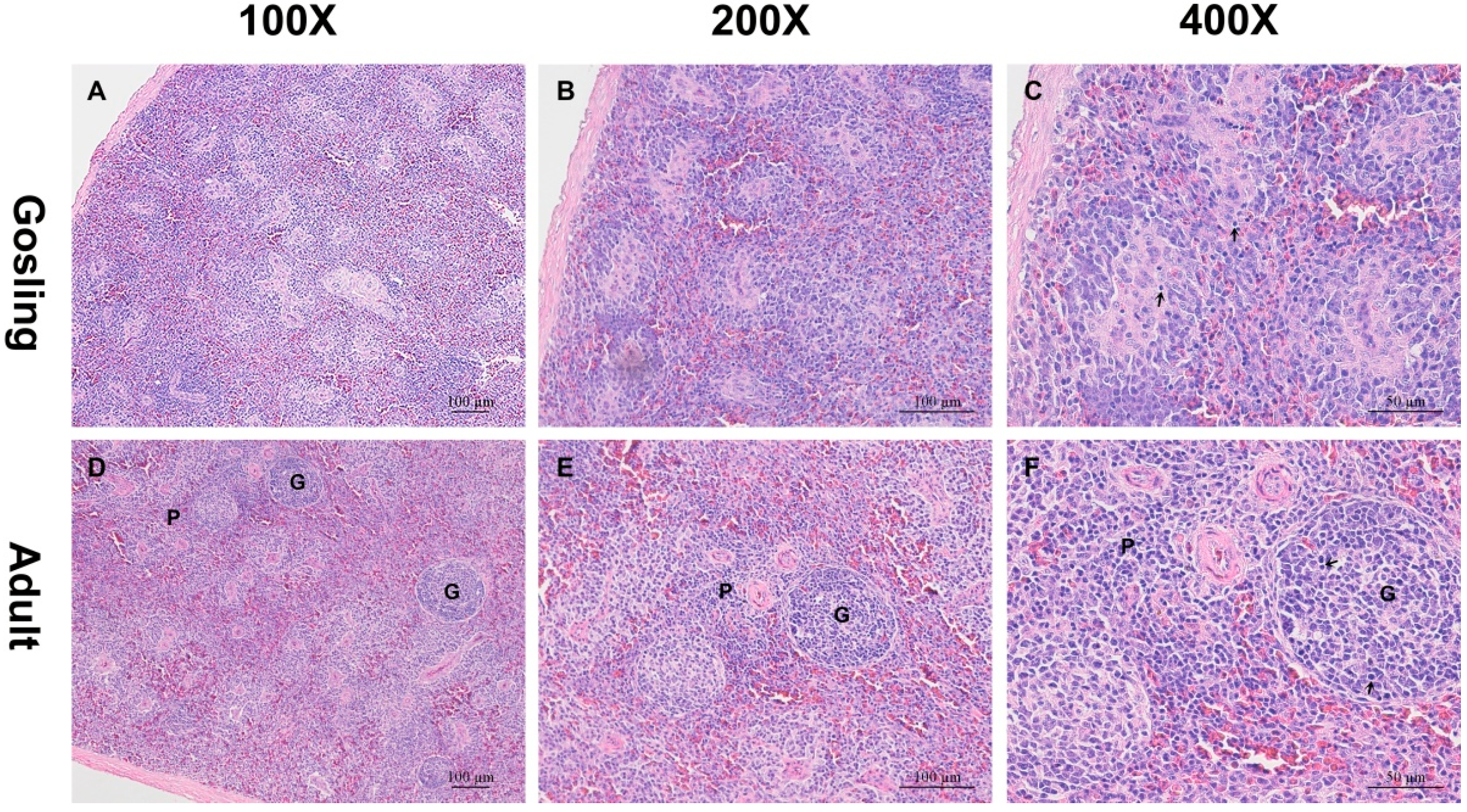
2.1. Histological Analysis
2.2. Illumina Sequencing and de Novo Assembly
| Description | 2 Week-Gosling | Adult Goose | Both |
|---|---|---|---|
| RIN (RNA integrity Number) | 10.0 | 10.0 | |
| Raw reads | 63,906,458 | 69,459,974 | |
| Clean reads | 63,121,772 | 68,655,944 | |
| Q30 (Q-score) | 93.18% | 93.25% | |
| Total number of unigenes | 50,390 | ||
| Total length of unigene | 79,599,367 | ||
| Max length of unigene | 31,052 | ||
| Min length of unigene | 301 | ||
| Average length of unigene | 1579.67 | ||
| N50 | 2585 |
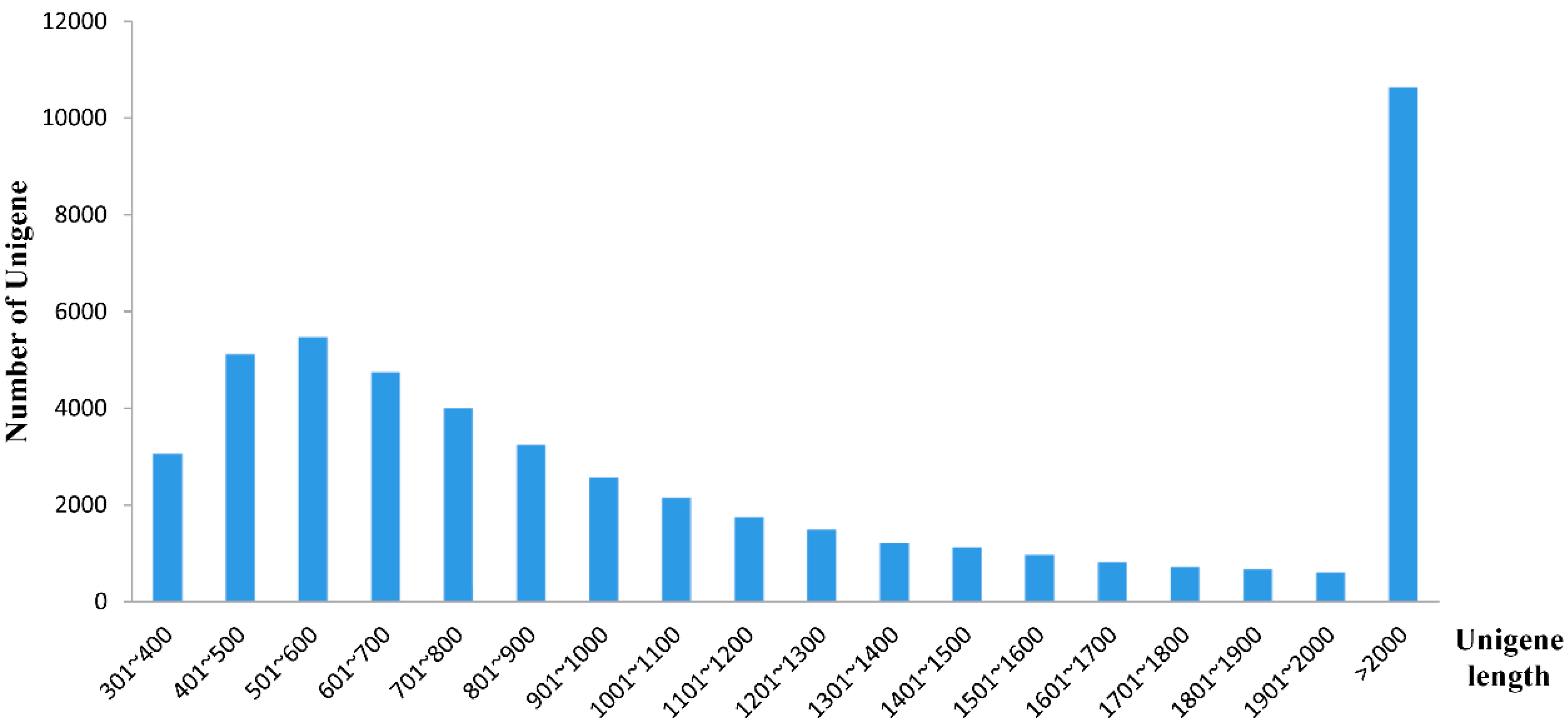
2.3. Assembly Evaluation and Annotation
| Database | Number of Annotated Unigenes | Annotated Unigene Ratio (%) |
|---|---|---|
| Nr | 13,841 | 27.47% |
| Swiss-Prot | 11,742 | 23.30% |
| GO | 9830 | 19.51% |
| KOG | 9534 | 18.92% |
| KEGG | 3607 | 7.16% |
| unknown | 36,448 | 72.33% |
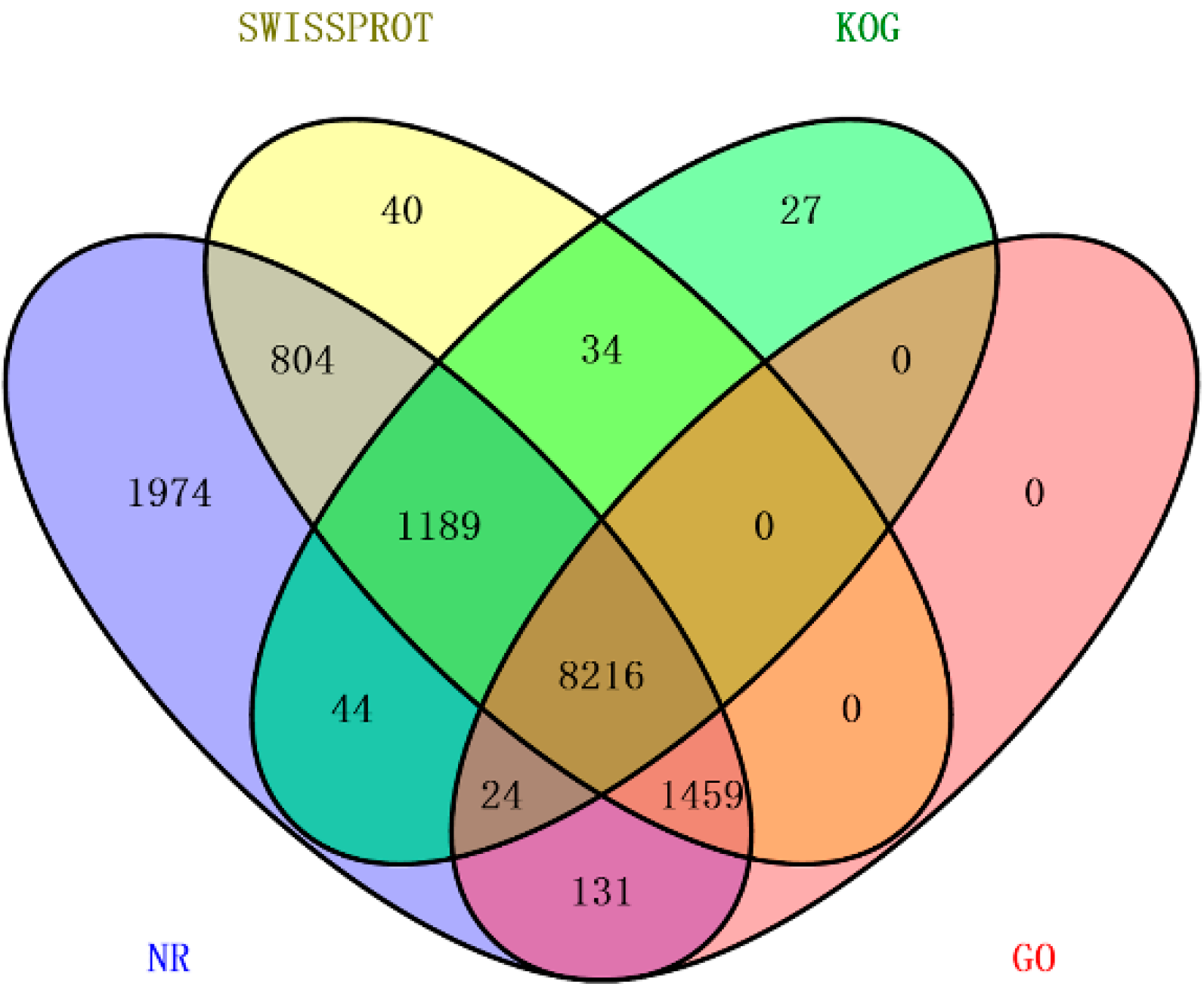
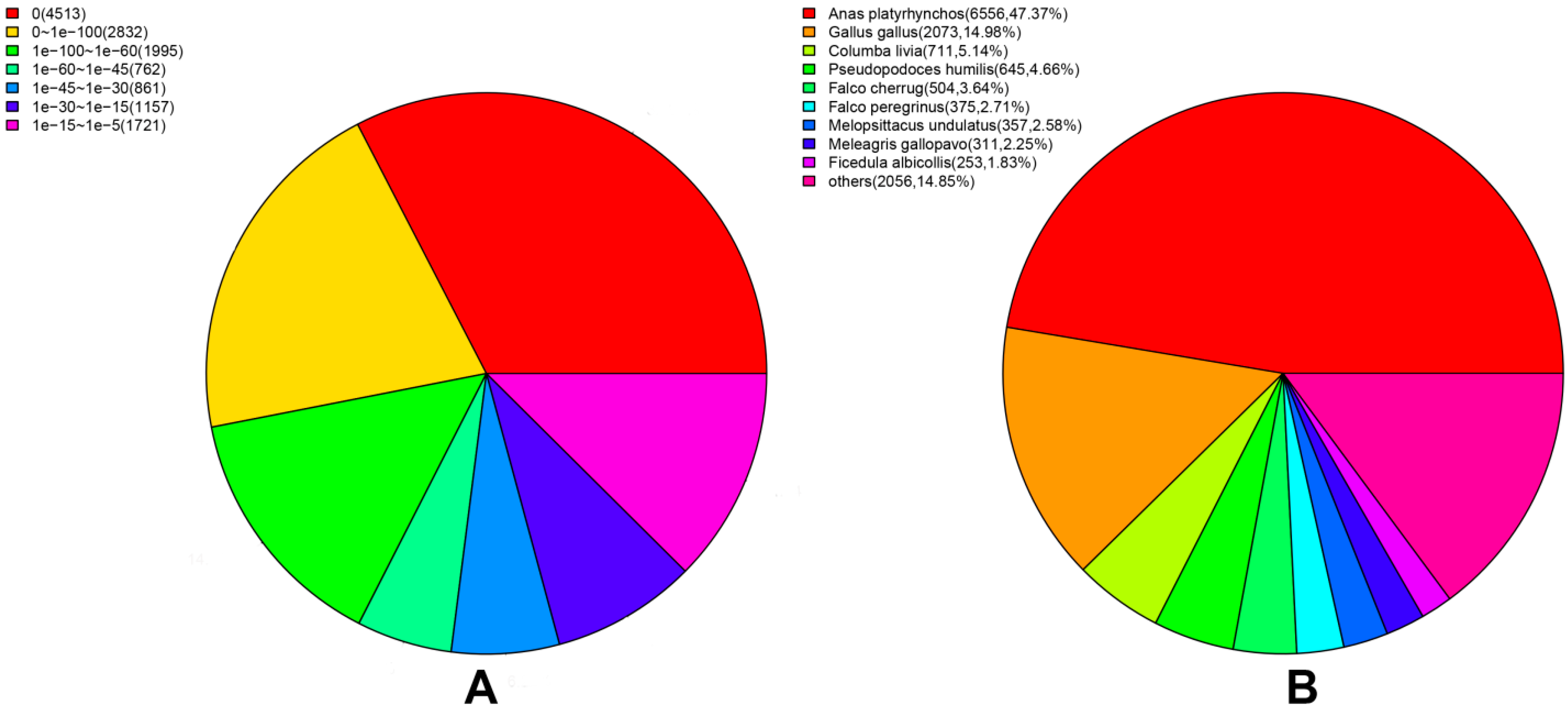
2.4. Function Classification and Pathway Analysis
2.4.1. KOG Analysis
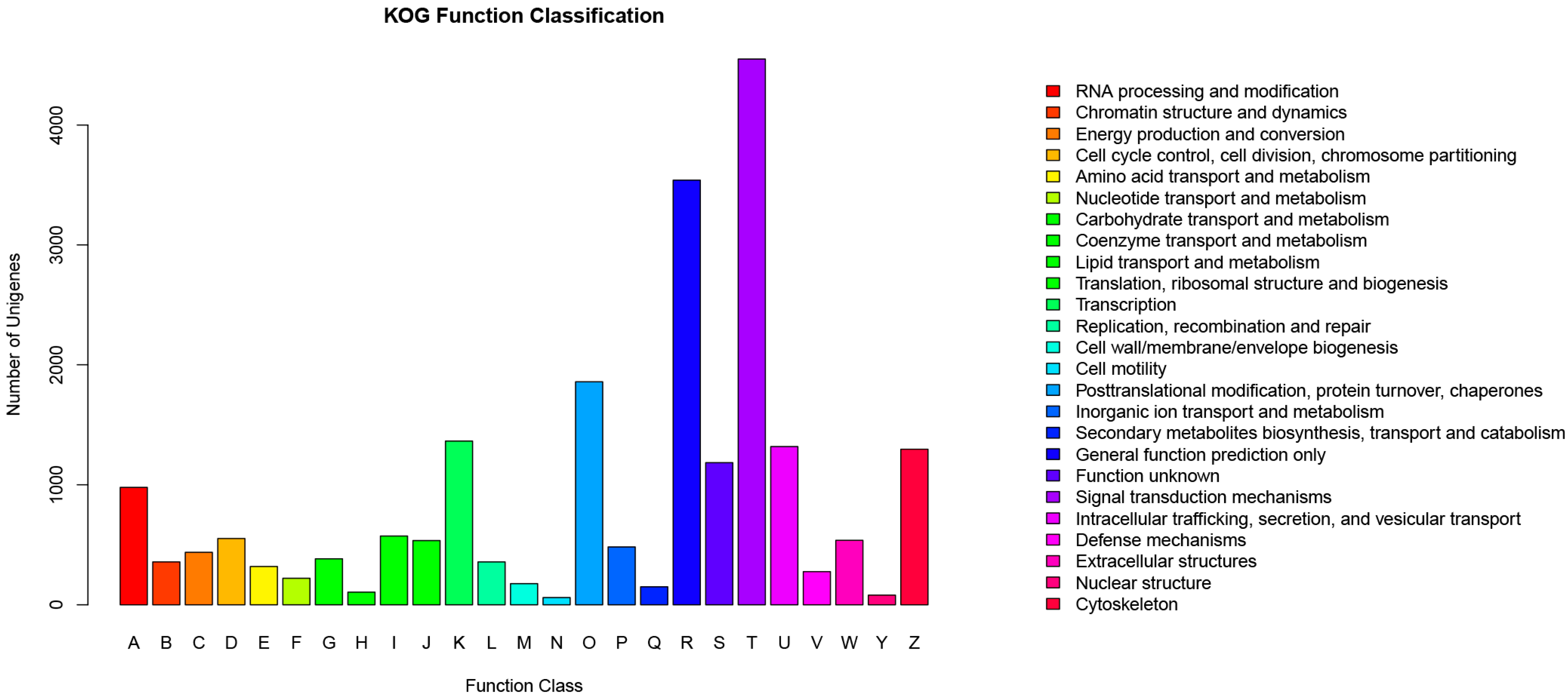
2.4.2. GO Analysis
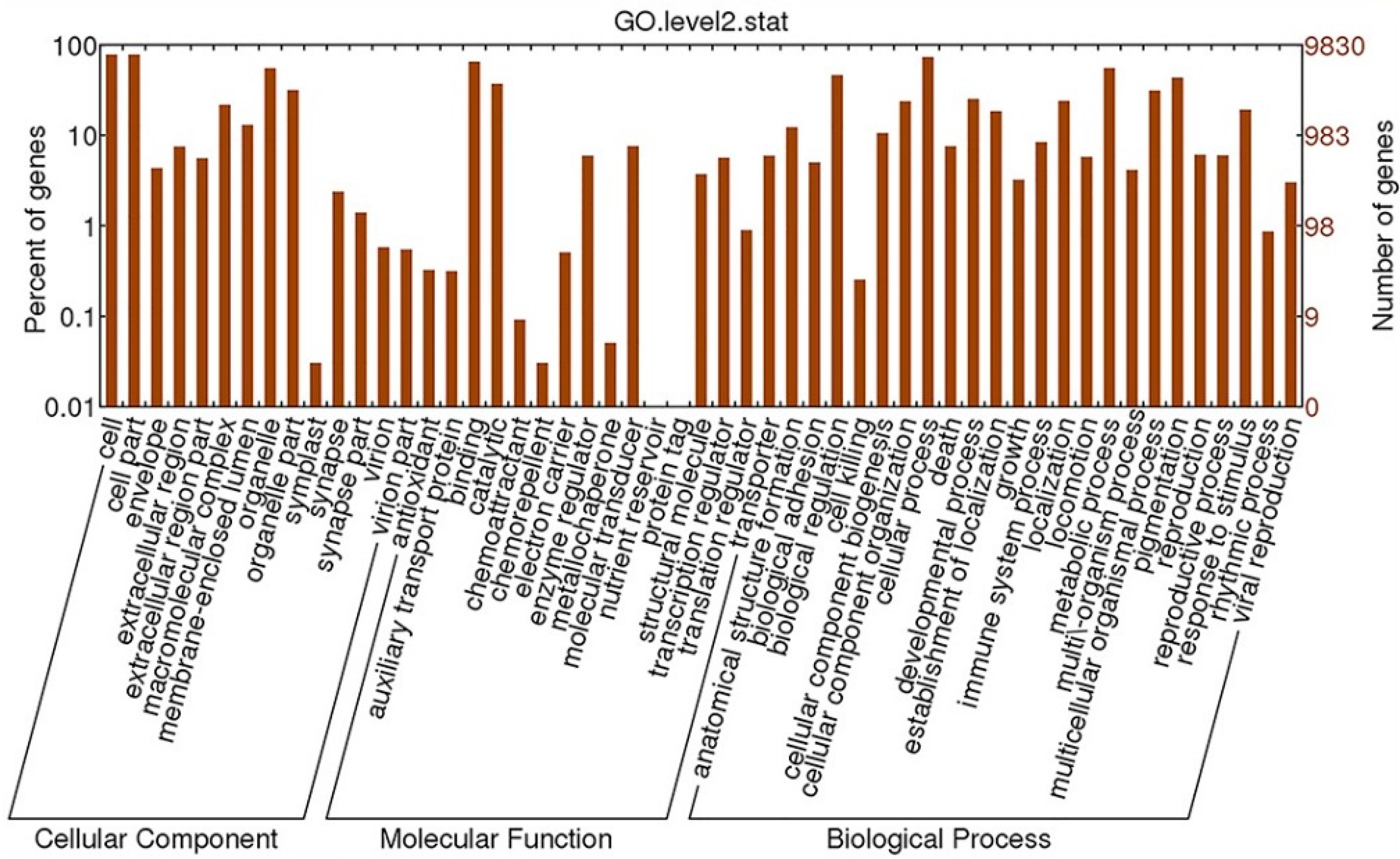
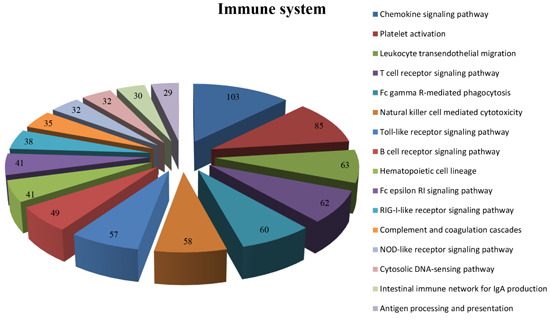
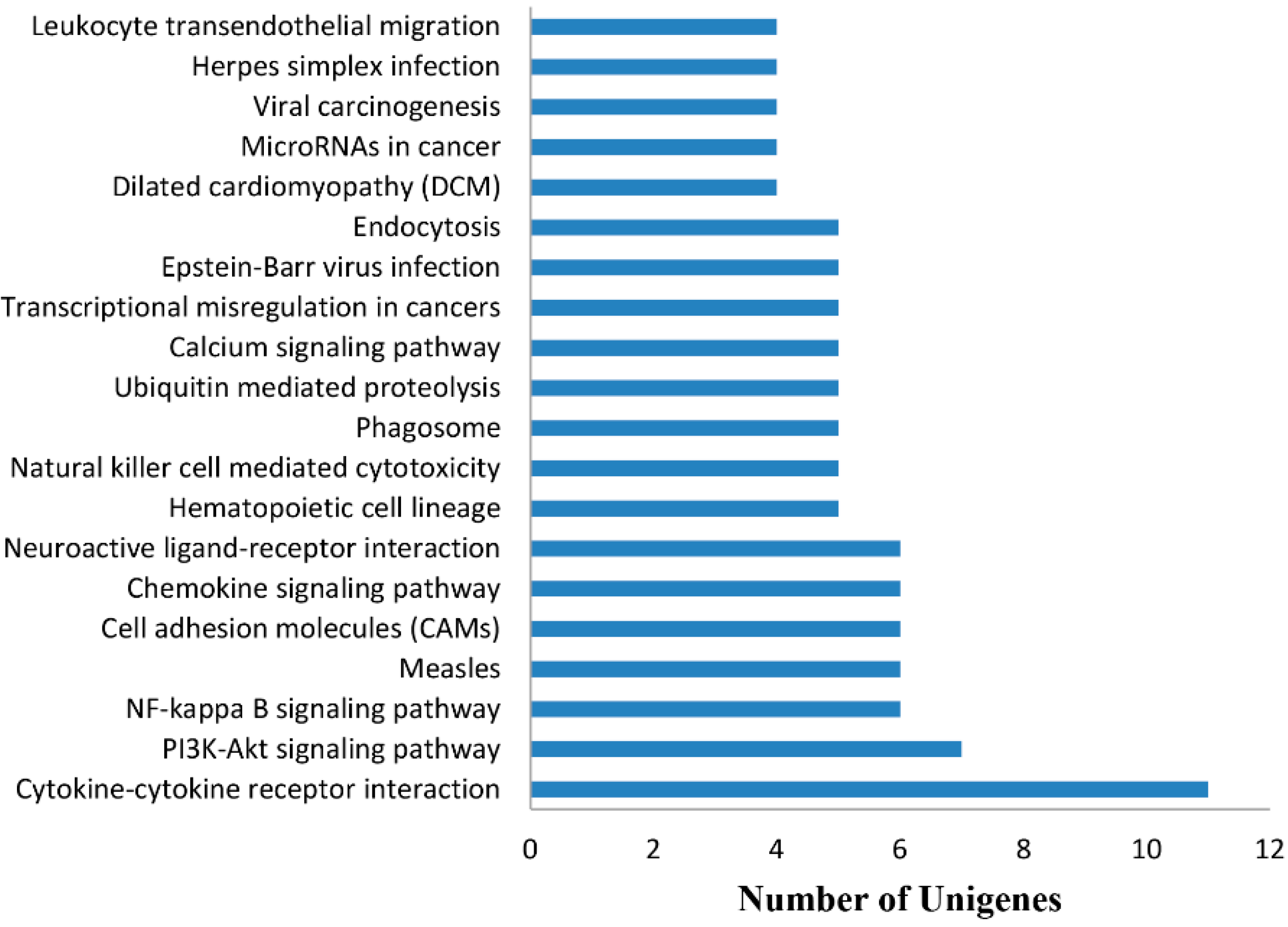
2.4.3. KEGG Analysis
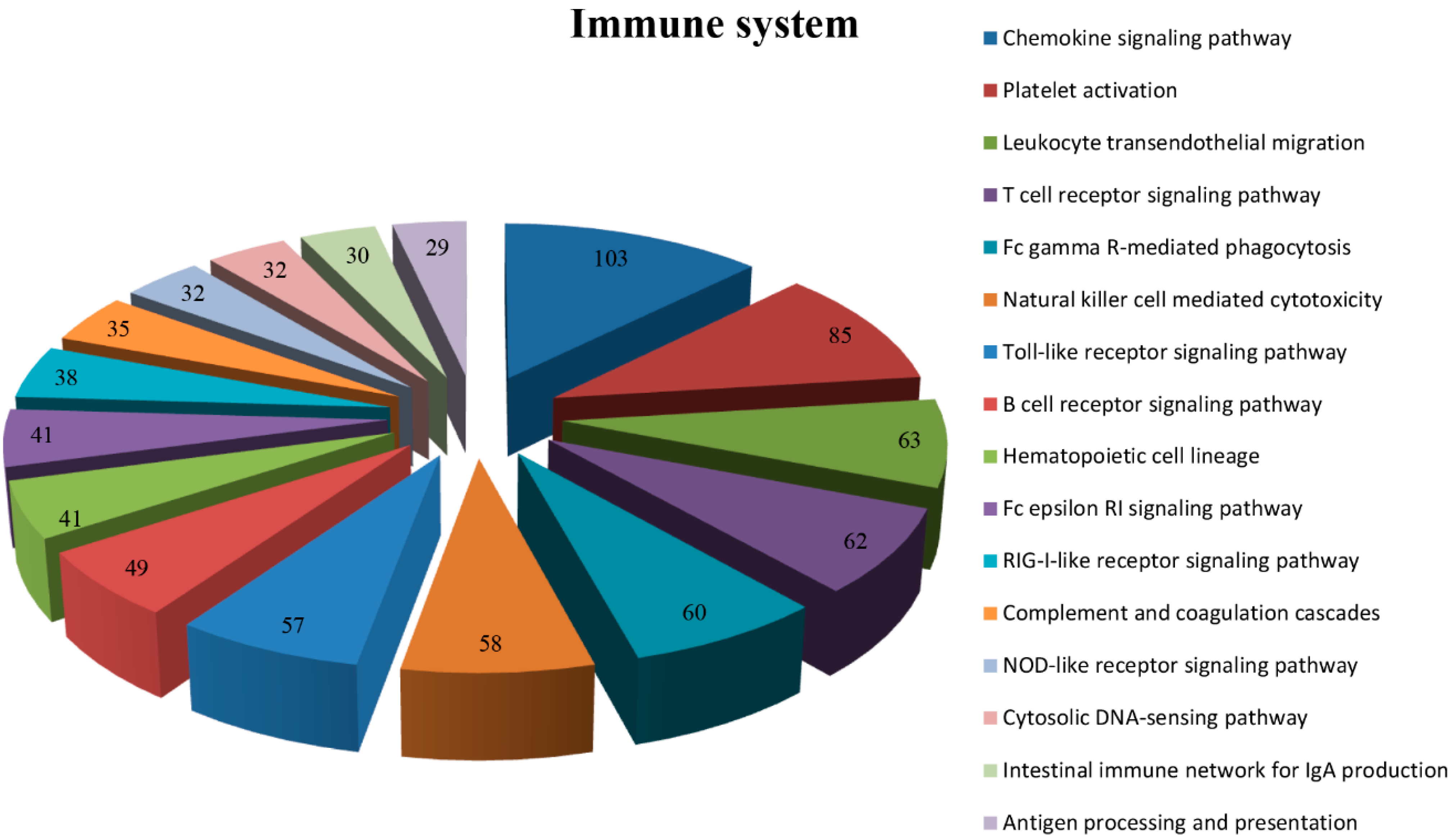
| Group Name | Unigene ID | Cytokine | Gene Description | Human | Chicken | Duck | Goose | Identity |
|---|---|---|---|---|---|---|---|---|
| Interleukin | CL32153Contig1 | IL6 | interleukin 6 | M54894.1 | NM_204628.1 | XM_005020542.1 | JF437643.1 | 73% |
| CL35348Contig1 | IL1B | interleukin 1 beta | M15330.1 | NM_204524.1 | Absent | JF505290.1 | 93% | |
| CL30075Contig1 | IL18, IL1F4 | interleukin 18 | AY044641.1 | NM_204608.1 | DQ522948.1 | JF505289.1 | 99% | |
| Interferon | CL30945Contig1 | IFNA | interferon alpha | M54886.1 | EU367971.1 | KF731866.1 | HQ1155831 | 88% |
| Tumor necrosis factor | CL13660Contig1 | TNFSF4, OX40L | tumor necrosis factor ligand | NM_003326.4 | XM_430147.3 | Absent | Absent | 94% |
| superfamily member 4 | NM_001297562.1 | |||||||
| CL33231Contig1 | TNFSF5, CD40L | tumor necrosis factor ligand | AY197739.1 | AJ293700.1 | DQ267671.2 | Absent | 99% | |
| superfamily member 5 | ||||||||
| CL36855Contig1 | TNFSF6, FASL | tumor necrosis factor ligand | AY225406.1 | AJ890143.1 | Absent | Absent | 99% | |
| superfamily member 6 | ||||||||
| CL36714Contig1 | TNFSF10, TRAIL | tumor necrosis factor ligand | NM_003810.3 NM_001190942.1 NM_001190943.1 | NM_204379.2 | XM_005017465.1 | Absent | 99% | |
| superfamily member 10 | XM_005017466.1 | |||||||
| CL44515Contig1 | TNFSF11, RANKL | tumor necrosis factor ligand | NM_003701.3 NM_033012.3 | NM_001083361.1 | XM_005008959.1 | Absent | 85% | |
| superfamily member 11 | ||||||||
| CL14930Contig1 | TNFSF15, VEGI | tumor necrosis factor ligand | NM_005118.3 NM_001204344.1 | NM_001024578.1 | XM_005022362.1 | Absent | 92% | |
| superfamily member 15 | ||||||||
| CL448Contig2 | TNFSF13B, BAFF | tumor necrosis factor ligand | NM_006573.4 NM_001145645.2 | NM_204327.2 | XM_005021732.1 | DQ874394.1 | 99% | |
| XM_005021733.1 | ||||||||
| superfamily member 13B | XM_005021734.1 | |||||||
| XM_005021735.1 | ||||||||
| CL34135Contig1 | EDA | ectodysplasin-A | NM_001399.4 | AY885699.1 | XM_005019557.1 | Absent | 99% | |
| NM_001005609.1 | ||||||||
| NM_001005612.2 | ||||||||
| NM_001005610.3 | ||||||||
| NM_001005613.3 | ||||||||
| Transforming growth factor | CL37958Contig1 | TGFB1 | transforming growth factor beta-1 | NM_000660.5 | JQ423909.1 | Absent | Absent | 83% |
| CL35089Contig1 | TGFB2 | transforming growth factor beta-2 | NM_001135599.2 NM_003238.3 | NM_001031045.3 | EU737316.1 | EF541127.1 | 99% | |
| Unigene ID | Cytokine | Gene Description | Human | Chicken | Duck | Goose | Identity | |
| Transforming growth factor | CL30050Contig1 | TGFB3 | transforming growth factor beta-3 | NM_003239.3 | NM_205454.1 | XM_005013912.1 | Absent | 99% |
| Colony-stimulating factor | CL39251Contig1 | CSF1, MCSF | macrophage colony-stimulating factor 1 | NM_000757.5 NM_172210.2 NM_172211.3 NM_172212.2 | GQ249403.1 | XM_005015071.1 | Absent | 94% |
| Chemokine | CL34468Contig1 | CCL4 | C-C motif chemokine 4 | NM_002984.3 | NM_204720.1 | Absent | Absent | 99% |
| CL38070Contig1 | CCL5 | C-C motif chemokine 5 | NM_002985.2 NM_001278736.1 | NM_001045832.1 | Absent | Absent | 99% | |
| CL35006Contig1 | CCL19, ELC | C-C motif chemokine 19 | NM_006274.2 | NM_001302168.1 | AY682098.1 | Absent | 100% | |
| CL39853Contig1 | CX3CL1, NTT | C-X3-C motif chemokine 1 | NM_002996.4 NM_001304392.1 | NM_001077232.1 | Absent | Absent | 99% | |
| CL29319Contig1 | CXCL12 | C–X–C motif chemokine 12 | NM_199168.3 NM_000609.6 | NM_204510.1 | XM_005029409.1XM_005029410.1 | Absent | 88% | |
| NM_001033886.2 NM_001178134.1 | ||||||||
| NM_001277990.11 | ||||||||
| CL19694Contig1 | CXCL13 | C–X–C motif chemokine 13 | NM_006419.2 | FR874037.1 | Absent | Absent | 99% | |
| FR874038.1 | ||||||||
| CL12315Contig1 | IL8, CXCL8 | interleukin 8 | M28130.1 | HM179639.1 |
2.5. Analysis of Differentially Expressed Genes
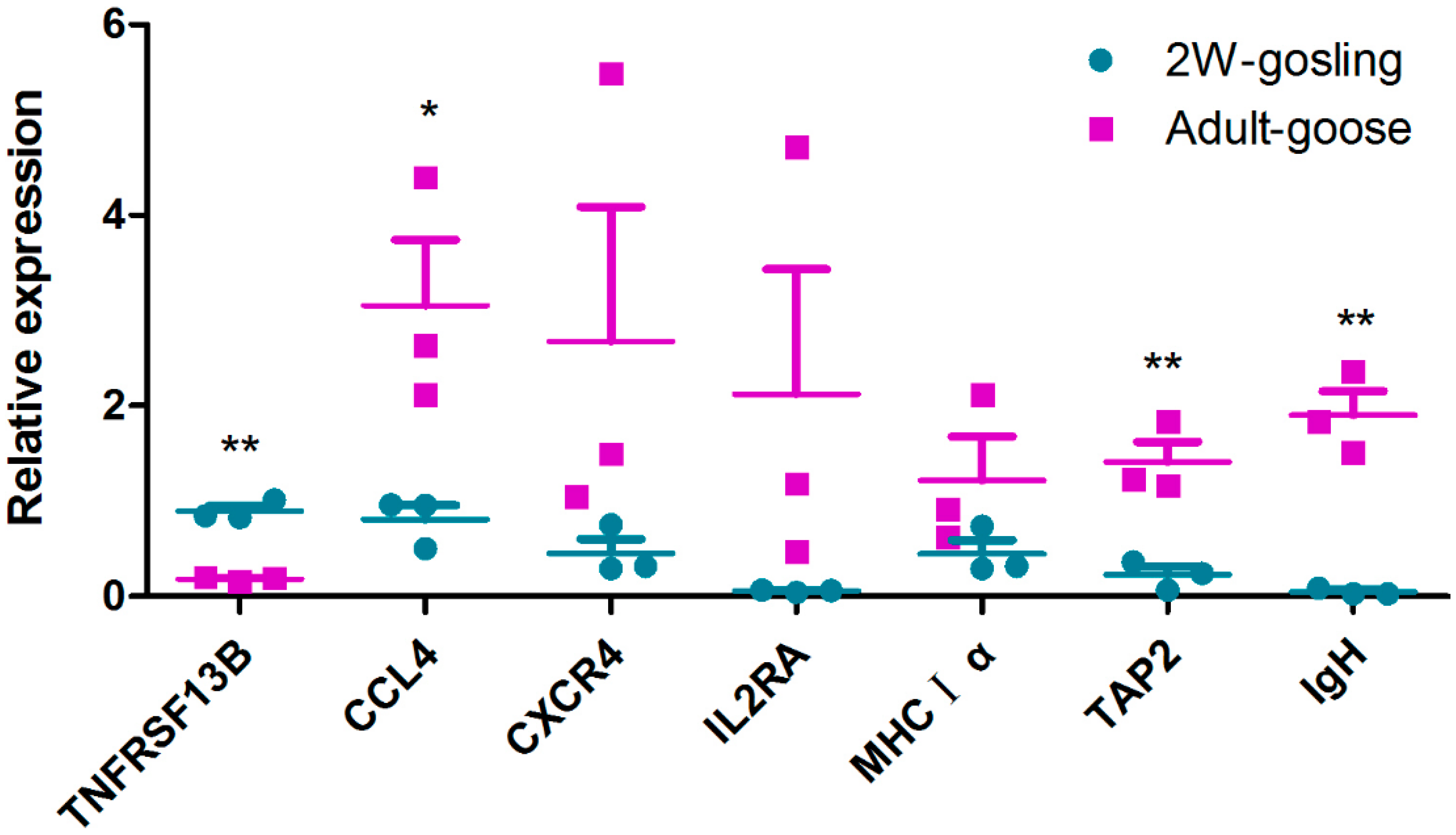
2.6. Mining for Immunity Relevant Genes and Validation by Quantitative Real-Time PCR (qRT-PCR)
| Unigene ID | Gene Symbol | Gene Description | Organism | log2 Fold Change | Pathway |
|---|---|---|---|---|---|
| CL22500Contig1 | IgH | immunoglobulin alpha heavy chain | Anser anser domesticus | 8.01485715881499 | HCL, NKC, BCR, FcεRI, FcγR, IIN |
| CL36733Contig1 | CXCR4 | C–X–C chemokine receptor type 4 | Melopsittacus undulatus | 2.82461947963658 | CSP, LTM, IIN |
| CL28326Contig1 | CXCR6 | C–X–C chemokine receptor type 6 | Anas platyrhynchos | 2.26986828363545 | CSP |
| CL36371Contig1 | XCR1 | chemokine XC receptor 1 | Anas platyrhynchos | 1.9920108201249 | CSP, CCR |
| CL35006Contig1 | CCL19 | chemokine C–C motif ligand 19 precursor | Anas platyrhynchos | 1.87882406908735 | CSP |
| CL34468Contig1 | CCL4 | c-C motif chemokine 4-like | Anas platyrhynchos | 1.87191382409167 | CSP, TLR, CDP, CCR |
| CL45402Contig1 | IgH | immunoglobulin alpha heavy chain | Anser anser domesticus | 1.86841583899162 | HCL, NKC, BCR, FcεRI, FcγR, IIN |
| CL46049Contig1 | CCL26 | c-C motif chemokine 26-like | Meleagris gallopavo | 1.80369233288299 | CSP |
| CL13415Contig1 | PLCG2 | 1-phosphatidylinositol 4,5-bisphosphate phosphodiesterase gamma-2 isoform X4 | Gallus gallus | 1.72847057141819 | NKC, BCR, FcεRI, FcγR, LTM |
| CL36217Contig1 | OPN | osteopontin | Anas platyrhynchos | 1.723521984199 | TLR |
| comp93267_c1_seq1 | MHCIα | MHC class I heavy chain | Anser anser | 1.55884992861107 | APP |
| CL45933Contig1 | TAP2 | transporter of antigen processing 2 | Anas platyrhynchos | 1.18018757654374 | APP |
| CL6275Contig1 | IL2RA | interleukin 2 receptor alpha | Anser cygnoides | 1.17080106279218 | HCL, CCR |
| CL15887Contig1 | IL21R | interleukin-21 receptor isoform X1 | Anas platyrhynchos | 1.09522512352901 | CCR |
| CL7617Contig1 | M3K8 | mitogen-activated protein kinase kinase kinase 8 isoform X2 | Anas platyrhynchos | 1.04305025312537 | TLR, TCR |
| CL47027Contig1 | MASP1 | mannan-binding lectin serine protease 1 isoform X1 | Columba livia | – | CCC |
| 3.10860084300754 | |||||
| CL12315Contig1 | IL8 | interleukin-8-like | Anas platyrhynchos | – | CSP, TLR, NODLR, RIGLR, CCR |
| 2.12963662747173 | |||||
| CL36341Contig1 | CSF3R | colony stimulating factor 3 receptor (granulocyte) | Anas platyrhynchos | – | HCL |
| 1.92500564529962 | |||||
| CL1203Contig2 | TNFRSF13B | tumor necrosis factor receptor superfamily member 13B | Anas platyrhynchos | – | IIN |
| 1.36191602157385 | |||||
| CL20554Contig1 | ITB3 | integrin beta-3-like | Anas platyrhynchos | – | HCL |
| 1.34147190439098 | |||||
| CL519Contig2 | TPA | tissue-type plasminogen activator | Anas platyrhynchos | – | CCC |
| 1.27377380687262 | |||||
| CL45872Contig1 | MP2K3 | dual specificity mitogen-activated protein kinase kinase 3 | Columba livia | – | TLR, FcεRI |
| 1.02391521347926 |
2.6.1. Cytokines and Cytokine Receptors
2.6.2. Antigen Processing and Presentation
2.6.3. Humoral Immune Effectors
3. Materials and Methods
3.1. Ethics Statement
3.2. Sample, H&E Staining and cDNA Library Preparation
3.3. Quality Assessment and de Novo Assembly
3.4. Functional Annotation and Classification
3.5. Differential Expression Analysis and Enrichment Analysis
3.6. Identification of Candidate Genes and qRT-PCR Validation
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Wang, Z.; Gerstein, M.; Snyder, M. RNA-Seq: A revolutionary tool for transcriptomics. Nat. Rev. Genet. 2009, 10, 57–63. [Google Scholar] [CrossRef] [PubMed]
- El-Metwally, S.; Ouda, O.M.; Helmy, M. Novel next-generation sequencing applications. In Next Generation Sequencing Technologies and Challenges in Sequence Assembly; Springer: New York, NY, USA, 2014; pp. 61–70. [Google Scholar]
- Dheilly, N.M.; Adema, C.; Raftos, D.A.; Gourbal, B.; Grunau, C.; du Pasquier, L. No more non-model species: The promise of next generation sequencing for comparative immunology. Dev. Comp. Immunol. 2014, 45, 56–66. [Google Scholar] [CrossRef] [PubMed]
- Xu, Q.; Zhao, W.; Chen, Y.; Tong, Y.; Rong, G.; Huang, Z.; Zhang, Y.; Chang, G.; Wu, X.; Chen, G. Transcriptome profiling of the goose (anser cygnoides) ovaries identify laying and broodiness phenotypes. PLoS ONE 2013, 8, e55496. [Google Scholar] [CrossRef] [PubMed]
- Luan, X.; Liu, D.; Cao, Z.; Luo, L.; Liu, M.; Gao, M.; Zhang, X. Transcriptome profiling identifies differentially expressed genes in huoyan goose ovaries between the laying period and ceased period. PLoS ONE 2014, 9, e13211. [Google Scholar] [CrossRef] [PubMed]
- Ding, N.; Han, Q.; Li, Q.; Zhao, X.; Li, J.; Su, J.; Wang, Q. Comprehensive analysis of sichuan white geese (anser cygnoides) transcriptome. Anim. Sci. J. 2014, 85, 650–659. [Google Scholar] [CrossRef] [PubMed]
- Gao, G.; Li, Q.; Zhao, X.; Ding, N.; Han, Q.; Su, J.; Wang, Q. Transcriptome profiling of the hypothalamus during prelaying and laying periods in sichuan white geese (anser cygnoides). Anim. Sci. J. 2015, 86, 800–805. [Google Scholar] [CrossRef] [PubMed]
- Yen, C.; Lin, E.; Wang, Y.; Wang, P.; Lin, H.; Hsu, J.; Wu, L.; Jiang, Y.; Ding, S. Abundantly expressed hepatic genes and their differential expression in liver of prelaying and laying geese. Poult. Sci. 2009, 88, 1955–1962. [Google Scholar] [CrossRef] [PubMed]
- Tariq, M.; Chen, R.; Yuan, H.; Liu, Y.; Wu, Y.; Wang, J.; Xia, C. De novo transcriptomic analysis of peripheral blood lymphocytes from the chinese goose: Gene discovery and immune system pathway description. PLoS ONE 2015, 10, e0121015. [Google Scholar] [CrossRef] [PubMed]
- Mebius, R.E.; Kraal, G. Structure and function of the spleen. Nat. Rev. Immunol. 2005, 5, 606–616. [Google Scholar] [CrossRef] [PubMed]
- Oláh, I.; Vervelde, L. Structure of the avian lymphoid system. In Avian Immunology; Elsevier: London, UK, 2008; pp. 13–50. [Google Scholar]
- Grabherr, M.G.; Haas, B.J.; Yassour, M.; Levin, J.Z.; Thompson, D.A.; Amit, I.; Adiconis, X.; Fan, L.; Raychowdhury, R.; Zeng, Q. Trinity: Reconstructing a full-length transcriptome without a genome from RNA-Seq data. Nat. Biotechnol. 2013, 29, 644–652. [Google Scholar] [CrossRef] [PubMed]
- Pertea, G.; Huang, X.; Liang, F.; Antonescu, V.; Sultana, R.; Karamycheva, S.; Lee, Y.; White, J.; Cheung, F.; Parvizi, B.; et al. TIGR gene indices clustering tools (TGICL): A software system for fast clustering of large est datasets. Bioinformatics 2003, 19, 651–652. [Google Scholar] [CrossRef] [PubMed]
- Venny. An interactive tool for comparing lists with venn diagrams. Available online: http://bioinfogp.cnb.csic.es/tools/venny/index.html (accessed on 17 July 2015).
- Tang, C.; Lan, D.; Zhang, H.; Ma, J.; Yue, H. Transcriptome analysis of duck liver and identification of differentially expressed transcripts in response to duck hepatitis A virus genotype C infection. PLoS ONE 2013, 8, e71051. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.W.; Luan, J.B.; Li, J.M.; Bao, Y.Y.; Zhang, C.X.; Liu, S.S. De novo characterization of a whitefly transcriptome and analysis of its gene expression during development. BMC Genom. 2010, 11, 400. [Google Scholar] [CrossRef] [PubMed]
- Martini, M.; de Santis, M.C.; Braccini, L.; Gulluni, F.; Hirsch, E. Pi3k/akt signaling pathway and cancer: An updated review. Ann. Med. 2014, 46, 372–383. [Google Scholar] [CrossRef] [PubMed]
- Vanhaesebroeck, B.; Guillermet-Guibert, J.; Graupera, M.; Bilanges, B. The emerging mechanisms of isoform-specific PI3K signalling. Nat. Rev. Mol. Cell Biol. 2010, 11, 329–341. [Google Scholar] [CrossRef] [PubMed]
- Isaacs, A.; Lindenmann, J. Virus interference. I. The interferon. Proc. R. Soc. Lond. B 1957, 147, 258–267. [Google Scholar] [CrossRef] [PubMed]
- Staeheli, P.; Puehler, F.; Schneider, K.; Göbel, T.W.; Kaspers, B. Cytokines of birds: Conserved functions-a largely different look. J. Interferon Cytokine Res. 2001, 21, 993–1010. [Google Scholar] [CrossRef] [PubMed]
- Asif, M.; Jenkins, K.A.; Hilton, L.S.; Kimpton, W.G.; Bean, A.G.; Lowenthal, J.W. Cytokines as adjuvants for avian vaccines. Immunol. Cell Biol. 2004, 82, 638–643. [Google Scholar] [CrossRef] [PubMed]
- Rahman, M.M.; Eo, S.K. Prospects and challenges of using chicken cytokines in disease prevention. Vaccine 2012, 30, 7165–7173. [Google Scholar] [CrossRef] [PubMed]
- Deng, R.; Lu, M.; Korteweg, C.; Gao, Z.; McNutt, M.A.; Ye, J.; Zhang, T.; Gu, J. Distinctly different expression of cytokines and chemokines in the lungs of two H5N1 avian influenza patients. J. Pathol. 2008, 216, 328–336. [Google Scholar] [CrossRef] [PubMed]
- Hirano, T. Interleukin 6 and its receptor: Ten years later. Int. Rev. Immunol. 1998, 16, 249–284. [Google Scholar] [CrossRef] [PubMed]
- Nakae, S.; Asano, M.; Horai, R.; Iwakura, Y. Interleukin-1β, but not interleukin-1α, is required for T-cell-dependent antibody production. Immunology 2001, 104, 402–409. [Google Scholar] [CrossRef] [PubMed]
- Dinarello, C.A. Immunological and inflammatory functions of the interleukin-1 family. Annu. Rev. Immunol. 2009, 27, 519–550. [Google Scholar] [CrossRef] [PubMed]
- Rahman, M.M.; Uyangaa, E.; Han, Y.W.; Kim, S.B.; Kim, J.H.; Choi, J.Y.; Eo, S.K. Enhancement of Th1-biased protective immunity against avian influenza H9N2 virus via oral co-administration of attenuated salmonella enterica serovar typhimurium expressing chicken interferon-alpha and interleukin-18 along with an inactivated vaccine. BMC Vet. Res. 2012, 8, 105. [Google Scholar] [CrossRef] [PubMed]
- Yao, Q.; Fischer, K.P.; Li, L.; Agrawal, B.; Berhane, Y.; Tyrrell, D.L.; Gutfreund, K.S.; Pasick, J. Immunogenicity and protective efficacy of a DNA vaccine encoding a chimeric protein of avian influenza hemagglutinin subtype H5 fused to CD154 (CD40L) in Pekin ducks. Vaccine 2010, 28, 8147–8156. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Ran, M.J.; Shan, X.X.; Cao, M.; Cao, P.; Yang, X.M.; Zhang, S.Q. Baff enhances B-cell-mediated immune response and vaccine-protection against a very virulent ibdv in chickens. Vaccine 2009, 27, 1393–1399. [Google Scholar] [CrossRef] [PubMed]
- Ansel, K.M.; Ngo, V.N.; Hyman, P.L.; Luther, S.A.; Förster, R.; Sedgwick, J.D.; Browning, J.L.; Lipp, M.; Cyster, J.G. A chemokine-driven positive feedback loop organizes lymphoid follicles. Nature 2000, 406, 309–314. [Google Scholar] [PubMed]
- Gunn, M.D.; Kyuwa, S.; Tam, C.; Kakiuchi, T.; Matsuzawa, A.; Williams, L.T.; Nakano, H. Mice lacking expression of secondary lymphoid organ chemokine have defects in lymphocyte homing and dendritic cell localization. J. Exp. Med. 1999, 189, 451–460. [Google Scholar] [CrossRef] [PubMed]
- Langmead, B.; Salzberg, S.L. Fast gapped-read alignment with bowtie 2. Nat. Methods 2012, 9, 357–359. [Google Scholar] [CrossRef] [PubMed]
- Benjamini, Y.; Hochberg, Y. Controlling the false discovery rate: A practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B 1995, 57, 289–300. [Google Scholar]
- Roberts, A.; Pachter, L. Streaming fragment assignment for real-time analysis of sequencing experiments. Nat. Methods 2013, 10, 71–73. [Google Scholar] [CrossRef] [PubMed]
- Akkoyunlu, M. Taci expression is low both in human and mouse newborns. Scand. J. Immunol. 2012, 75, 368–368. [Google Scholar] [CrossRef] [PubMed]
- Seshasayee, D.; Valdez, P.; Yan, M.; Dixit, V.M.; Tumas, D.; Grewal, I.S. Loss of taci causes fatal lymphoproliferation and autoimmunity, establishing taci as an inhibitory blys receptor. Immunity 2003, 18, 279–288. [Google Scholar] [CrossRef]
- Rossi, D.; Zlotnik, A. The biology of chemokines and their receptors. Annu. Rev. Immunol. 2000, 18, 217–242. [Google Scholar] [CrossRef] [PubMed]
- Hargreaves, D.C.; Hyman, P.L.; Lu, T.T.; Ngo, V.N.; Bidgol, A.; Suzuki, G.; Zou, Y.R.; Littman, D.R.; Cyster, J.G. A coordinated change in chemokine responsiveness guides plasma cell movements. J. Exp. Med. 2001, 194, 45–56. [Google Scholar] [CrossRef] [PubMed]
- Vanderven, H.A.; Petkau, K.; Ryan-Jean, K.E.; Aldridge, J.R.; Webster, R.G.; Magor, K.E. Avian influenza rapidly induces antiviral genes in duck lung and intestine. Mol. Immunol. 2012, 51, 316–324. [Google Scholar] [CrossRef] [PubMed]
- Teng, Q.Y.; Zhou, J.Y.; Wu, J.J.; Guo, J.Q.; Shen, H.G. Characterization of chicken interleukin 2 receptor alpha chain, a homolog to mammalian CD25. FEBS Lett. 2006, 580, 4274–4281. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Fang, J.; Guo, J.; Teng, Q.; Huang, Z.; Gu, J.; Shen, H.; Zhou, J. Molecular cloning and characterization of duck CD25. Vet. Immunol. Immunopathol. 2007, 117, 266–274. [Google Scholar] [CrossRef] [PubMed]
- Sakaguchi, S.; Sakaguchi, N.; Asano, M.; Itoh, M.; Toda, M. Immunologic self-tolerance maintained by activated T cells expressing IL-2 receptor alpha-chains (CD25). Breakdown of a single mechanism of self-tolerance causes various autoimmune diseases. J. Immunol. 1995, 155, 1151–1164. [Google Scholar] [PubMed]
- Suh, W.K.; Cohen-Doyle, M.F.; Fruh, K.; Wang, K.; Peterson, P.A.; Williams, D.B. Interaction of MHC class I molecules with the transporter associated with antigen processing. Science 1994, 264, 1322–1326. [Google Scholar] [CrossRef] [PubMed]
- Patel, R.K.; Jain, M. NGS QC Toolkit: A toolkit for quality control of next generation sequencing data. PLoS ONE 2012, 7, e30619. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Tatusov, R.L.; Fedorova, N.D.; Jackson, J.D.; Jacobs, A.R.; Kiryutin, B.; Koonin, E.V.; Krylov, D.M.; Mazumder, R.; Mekhedov, S.L.; Nikolskaya, A.N. The COG database: An updated version includes eukaryotes. BMC Bioinform. 2003, 4, 41. [Google Scholar] [CrossRef] [PubMed]
- Conesa, A.; Gotz, S.; Garcia-Gomez, J.M.; Terol, J.; Talon, M.; Robles, M. Blast2GO: A universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 2005, 21, 3674–3676. [Google Scholar] [CrossRef] [PubMed]
- Kanehisa, M.; Goto, S.; Kawashima, S.; Okuno, Y.; Hattori, M. The KEGG resource for deciphering the genome. Nucleic Acids Res. 2004, 32, 277–280. [Google Scholar] [CrossRef] [PubMed]
- Anders, S.; Huber, W. Differential Expression of RNA-Seq Data at the Gene Level—The DESeq Package; European Molecular Biology Laboratory (EMBL): Heidelberg, Germany, 2012. [Google Scholar]
- Mortazavi, A.; Williams, B.A.; McCue, K.; Schaeffer, L.; Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat. Methods 2008, 5, 621–628. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, A.; Liu, F.; Chen, S.; Wang, M.; Jia, R.; Zhu, D.; Liu, M.; Sun, K.; Wu, Y.; Chen, X.; et al. Transcriptome Analysis and Identification of Differentially Expressed Transcripts of Immune-Related Genes in Spleen of Gosling and Adult Goose. Int. J. Mol. Sci. 2015, 16, 22904-22926. https://doi.org/10.3390/ijms160922904
Wang A, Liu F, Chen S, Wang M, Jia R, Zhu D, Liu M, Sun K, Wu Y, Chen X, et al. Transcriptome Analysis and Identification of Differentially Expressed Transcripts of Immune-Related Genes in Spleen of Gosling and Adult Goose. International Journal of Molecular Sciences. 2015; 16(9):22904-22926. https://doi.org/10.3390/ijms160922904
Chicago/Turabian StyleWang, Anqi, Fei Liu, Shun Chen, Mingshu Wang, Renyong Jia, Dekang Zhu, Mafeng Liu, Kunfeng Sun, Ying Wu, Xiaoyue Chen, and et al. 2015. "Transcriptome Analysis and Identification of Differentially Expressed Transcripts of Immune-Related Genes in Spleen of Gosling and Adult Goose" International Journal of Molecular Sciences 16, no. 9: 22904-22926. https://doi.org/10.3390/ijms160922904
APA StyleWang, A., Liu, F., Chen, S., Wang, M., Jia, R., Zhu, D., Liu, M., Sun, K., Wu, Y., Chen, X., & Cheng, A. (2015). Transcriptome Analysis and Identification of Differentially Expressed Transcripts of Immune-Related Genes in Spleen of Gosling and Adult Goose. International Journal of Molecular Sciences, 16(9), 22904-22926. https://doi.org/10.3390/ijms160922904