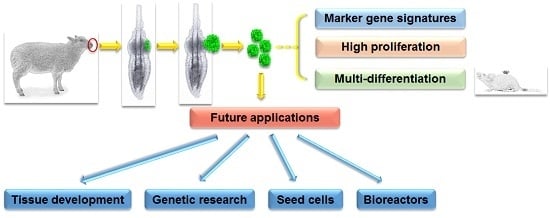
Ovine Hair Follicle Stem Cells Derived from Single Vibrissae Reconstitute Haired Skin
Abstract
:1. Introduction
2. Results
2.1. A Feasible Method to Obtain Abundant Ovine Keratinocytes from Single Vibrissa Hair Follicles
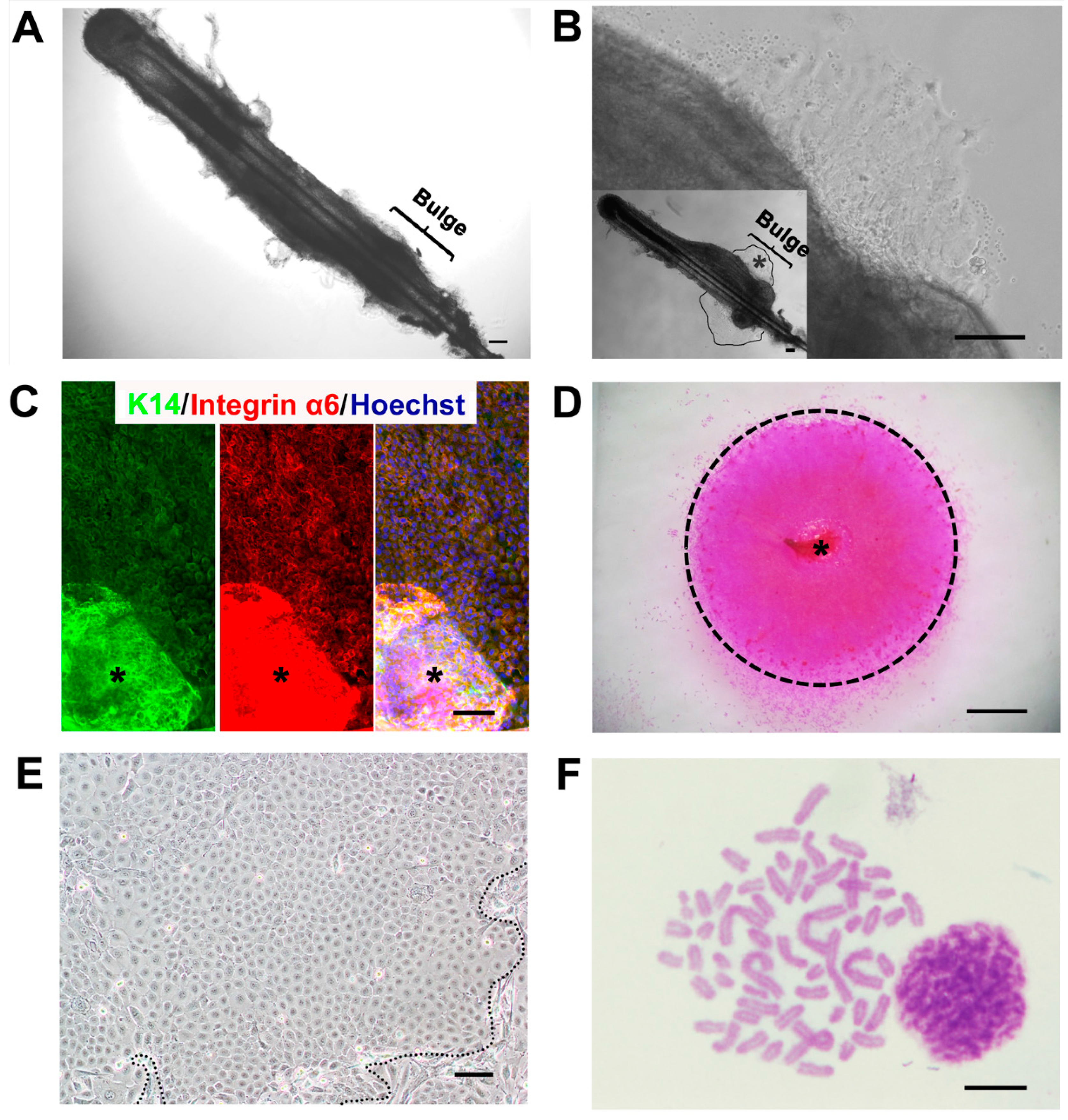
| Group | No. of Hair Follicles Cultured | No. of Hair Follicles that Formed a Colony | Percentage of Yield Ratio |
|---|---|---|---|
| 1 | 12 | 12 | 100% |
| 2 | 15 | 14 | 93.33% |
| 3 | 12 | 11 | 91.66% |
| Average Percentage | 94.99% | ||
2.2. Expression Profiles of Ovine Bulge-Derived Keratinocytes
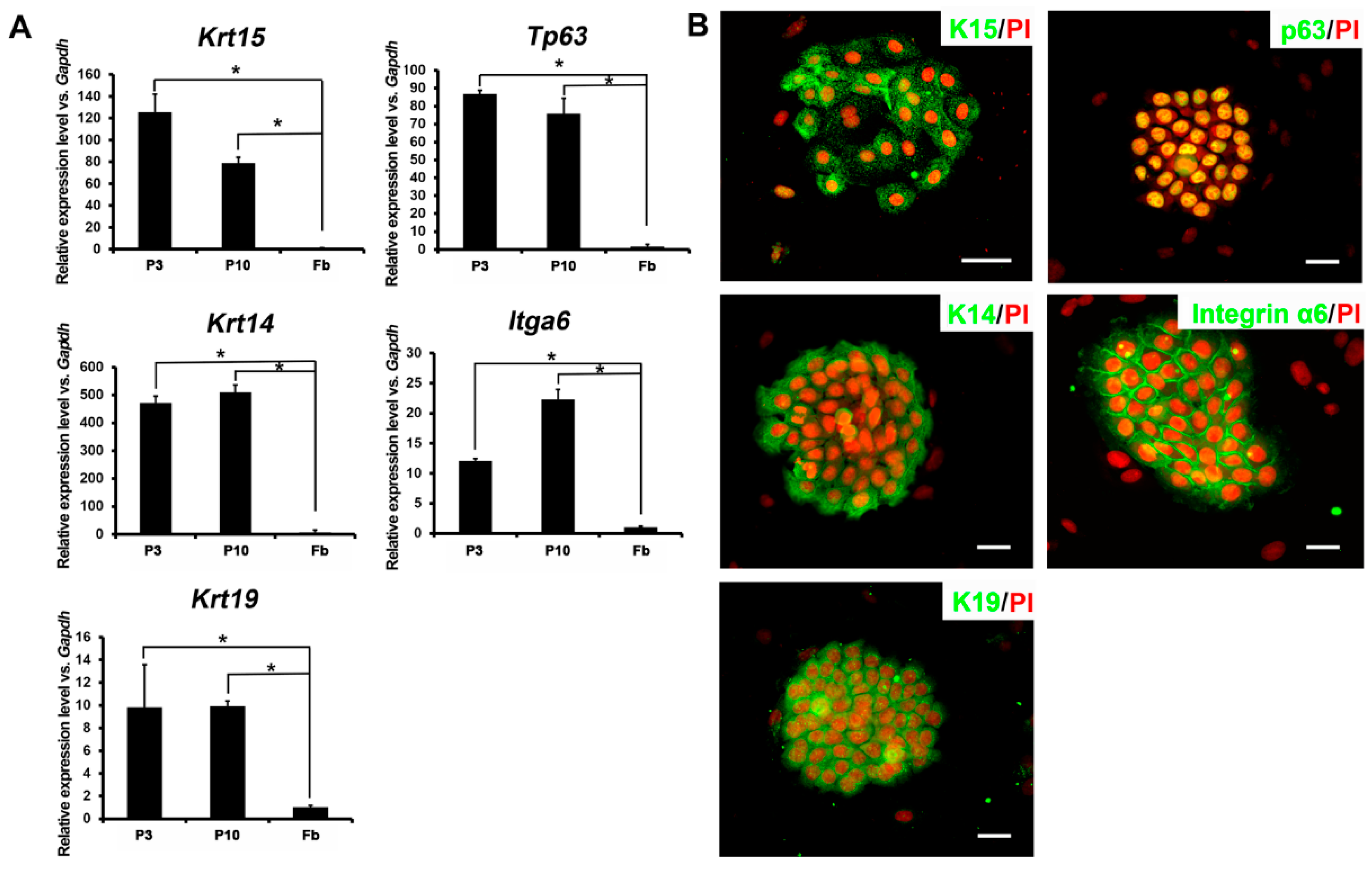
2.3. The Proliferative Capacity of Ovine Bulge-Derived Keratinocytes in Culture
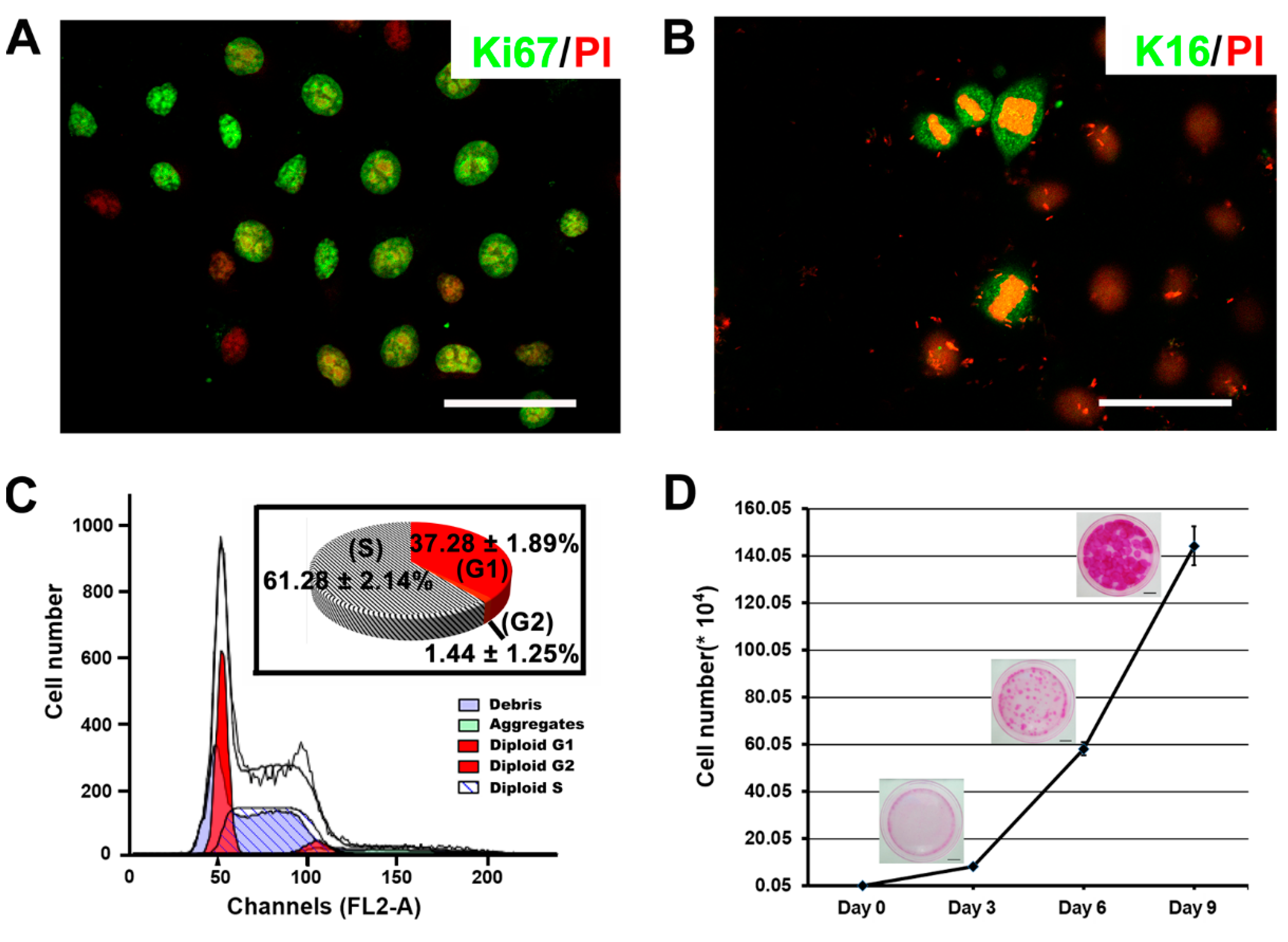
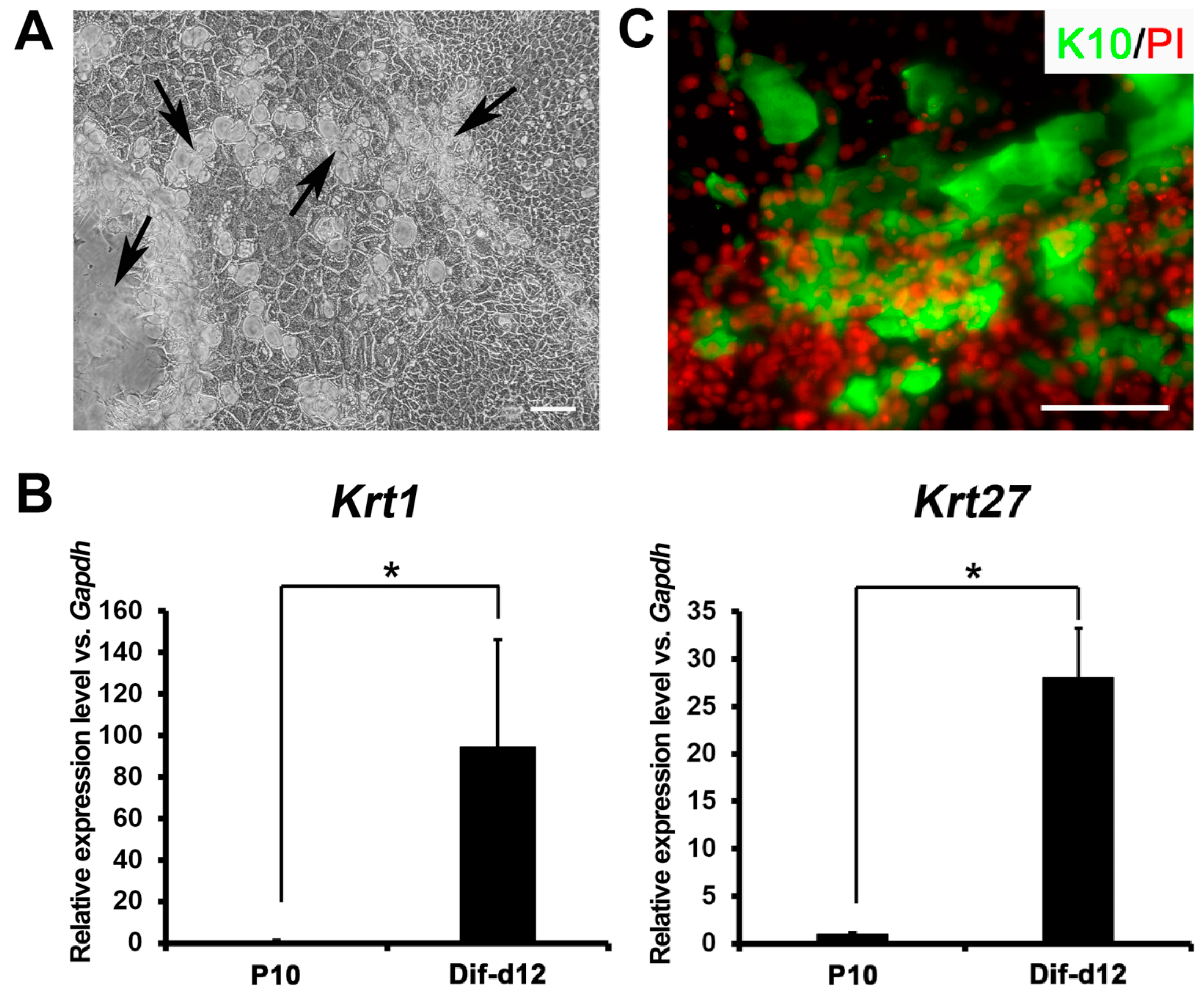
2.4. In Vitro Differentiation Capacity of Ovine Bulge-Derived Keratinocytes
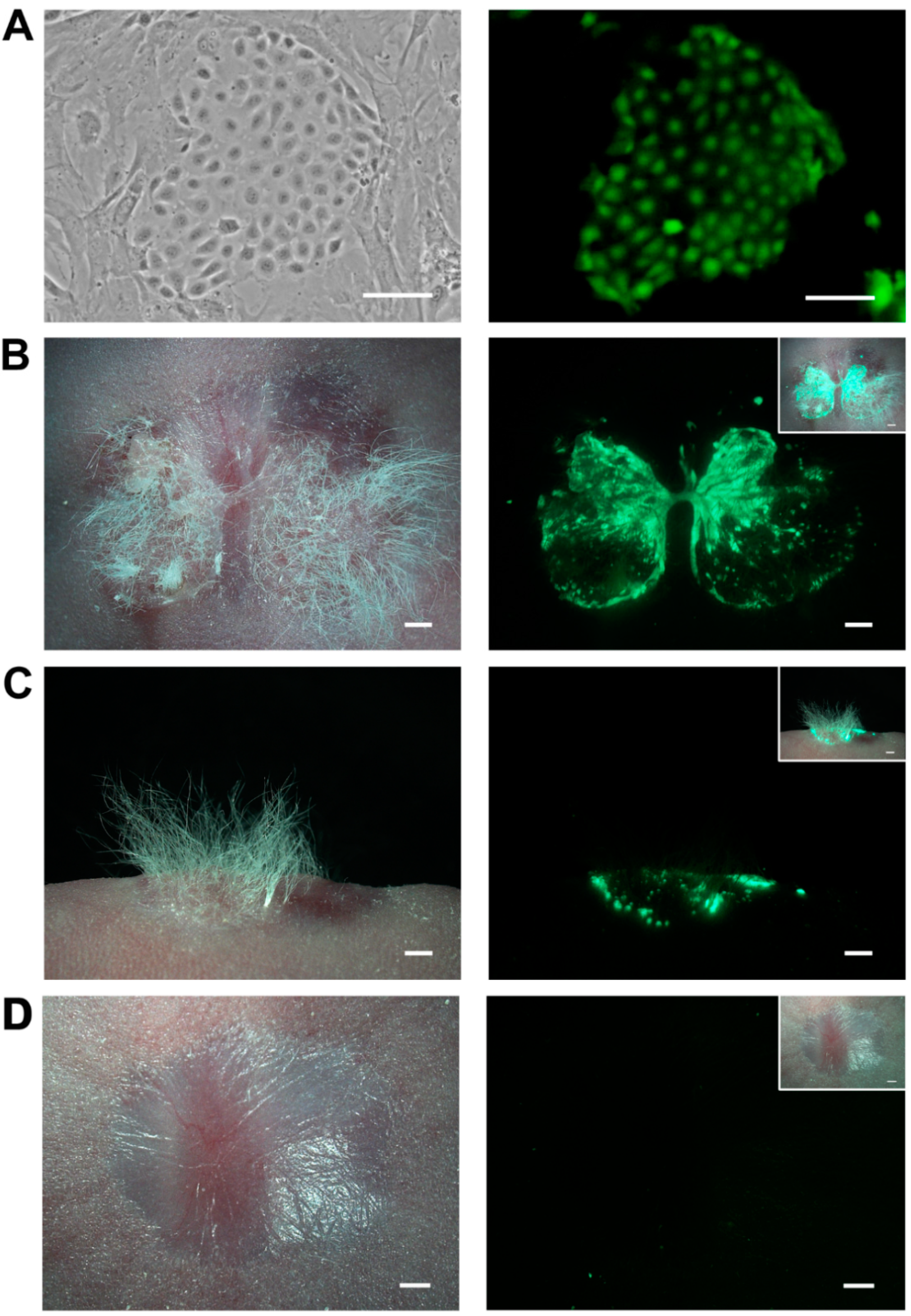
2.5. Skin Reconstitution with Ovine Bulge-Derived Keratinocytes and Neonatal Dermal Cells
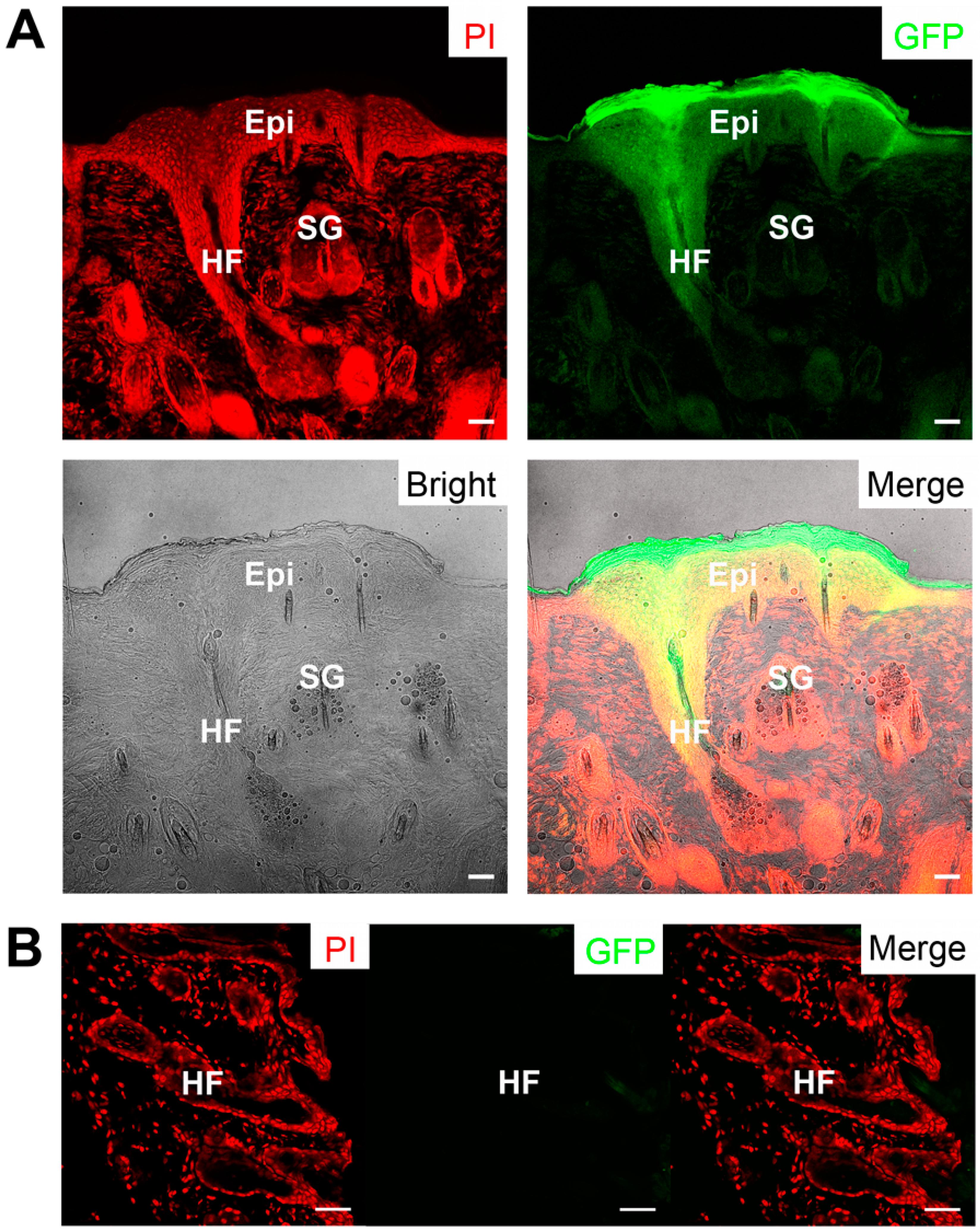
3. Discussion
4. Experimental Section
4.1. Animals
4.2. Microdissection
4.3. Cell Culture and Passaging
4.4. Fluorescence-Activated Cell Sorting (FACS)
4.5. Karyotype Analysis
4.6. RNA Purification, Reverse Transcription and qRT-PCR
4.7. Immunofluorescence
4.8. GFP Lentivirus Production and Keratinocyte Infection and Sorting
4.9. Skin Reconstitution Assay
4.10. Statistical Analysis
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Liu, S.; Zhang, H.; Duan, E. Epidermal development in mammals: Key regulators, signals from beneath, and stem cells. Int. J. Mol. Sci. 2013, 14, 10869–10895. [Google Scholar] [CrossRef] [PubMed]
- Muller-Rover, S.; Handjiski, B.; van der Veen, C.; Eichmuller, S.; Foitzik, K.; McKay, I.A.; Stenn, K.S.; Paus, R. A comprehensive guide for the accurate classification of murine hair follicles in distinct hair cycle stages. J. Investig. Dermatol. 2001, 117, 3–15. [Google Scholar] [CrossRef] [PubMed]
- Alonso, L.; Fuchs, E. The hair cycle. J. Cell Sci. 2006, 119, 391–393. [Google Scholar] [CrossRef] [PubMed]
- Blanpain, C.; Fuchs, E. Epidermal stem cells of the skin. Annu. Rev. Cell Dev. Biol. 2006, 22, 339–373. [Google Scholar] [CrossRef] [PubMed]
- Deng, Z.; Lei, X.; Zhang, X.; Zhang, H.; Liu, S.; Chen, Q.; Hu, H.; Wang, X.; Ning, L.; Cao, Y.; et al. mTOR signaling promotes stem cell activation via counterbalancing BMP-mediated suppression during hair regeneration. J. Mol. Cell Biol. 2015, 7, 62–72. [Google Scholar] [CrossRef] [PubMed]
- Blanpain, C.; Lowry, W.E.; Geoghegan, A.; Polak, L.; Fuchs, E. Self-renewal, multipotency, and the existence of two cell populations within an epithelial stem cell niche. Cell 2004, 118, 635–648. [Google Scholar] [CrossRef] [PubMed]
- Jaks, V.; Kasper, M.; Toftgard, R. The hair follicle-a stem cell zoo. Exp. Cell Res. 2010, 316, 1422–1428. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, K.; Rochat, A.; Barrandon, Y. Segregation of keratinocyte colony-forming cells in the bulge of the rat vibrissa. Proc. Natl. Acad. Sci. USA 1993, 90, 7391–7395. [Google Scholar] [CrossRef] [PubMed]
- Morris, R.J.; Liu, Y.; Marles, L.; Yang, Z.; Trempus, C.; Li, S.; Lin, J.S.; Sawicki, J.A.; Cotsarelis, G. Capturing and profiling adult hair follicle stem cells. Nat. Biotechnol. 2004, 22, 411–417. [Google Scholar] [CrossRef] [PubMed]
- Tumbar, T.; Guasch, G.; Greco, V.; Blanpain, C.; Lowry, W.E.; Rendl, M.; Fuchs, E. Defining the epithelial stem cell niche in skin. Science 2004, 303, 359–363. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Hu, H.; Zhang, H.; Liu, S.; Liu, S.; Zhang, Y.; Lei, X.; Ning, L.; Cao, Y.; Duan, E. Hair follicle stem cells derived from single rat vibrissa via organ culture reconstitute hair follicles in vivo. Cell Transplant. 2012, 21, 1075–1085. [Google Scholar] [CrossRef] [PubMed]
- Reynolds, A.J.; Jahoda, C.A. Hair follicle stem cells? A distinct germinative epidermal cell population is activated in vitro by the presence of hair dermal papilla cells. J. Cell Sci. 1991, 99 Pt 2, 373–385. [Google Scholar] [PubMed]
- Amici, A.W.; Yamato, M.; Okano, T.; Kobayashi, K. The multipotency of adult vibrissa follicle stem cells. Differentiation 2009, 77, 317–323. [Google Scholar] [CrossRef] [PubMed]
- Roh, C.; Tao, Q.; Photopoulos, C.; Lyle, S. In vitro differences between keratinocyte stem cells and transit-amplifying cells of the human hair follicle. J. Investig. Dermatol. 2005, 125, 1099–1105. [Google Scholar] [CrossRef] [PubMed]
- Ohyama, M.; Terunuma, A.; Tock, C.L.; Radonovich, M.F.; Pise-Masison, C.A.; Hopping, S.B.; Brady, J.N.; Udey, M.C.; Vogel, J.C. Characterization and isolation of stem cell-enriched human hair follicle bulge cells. J. Clin. Investig. 2006, 116, 249–260. [Google Scholar] [CrossRef] [PubMed]
- Ehama, R.; Ishimatsu-Tsuji, Y.; Iriyama, S.; Ideta, R.; Soma, T.; Yano, K.; Kawasaki, C.; Suzuki, S.; Shirakata, Y.; Hashimoto, K.; et al. Hair follicle regeneration using grafted rodent and human cells. J. Investig. Dermatol. 2007, 127, 2106–2115. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, T.; Shimizu, A.; Nishifuji, K.; Amagai, M.; Iwasaki, T.; Ohyama, M. Canine hair-follicle keratinocytes enriched with bulge cells have the highly proliferative characteristic of stem cells. Vet. Dermatol. 2009, 20, 338–346. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, T.; Iwasaki, T.; Amagai, M.; Ohyama, M. Canine follicle stem cell candidates reside in the bulge and share characteristic features with human bulge cells. J. Investig. Dermatol. 2010, 130, 1988–1995. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, T.; Enomoto, K.; Wang, Y.H.; Yoon, J.S.; Okamura, R.; Ide, K.; Ohyama, M.; Nishiyama, T.; Iwasaki, T.; Nishifuji, K. Epidermal structure created by canine hair follicle keratinocytes enriched with bulge cells in a three-dimensional skin equivalent model in vitro: Implications for regenerative therapy of canine epidermis. Vet. Dermatol. 2013, 24, 77–83, e19–e20. [Google Scholar] [CrossRef] [PubMed]
- Weinberg, W.C.; Goodman, L.V.; George, C.; Morgan, D.L.; Ledbetter, S.; Yuspa, S.H.; Lichti, U. Reconstitution of hair follicle development in vivo: Determination of follicle formation, hair growth, and hair quality by dermal cells. J. Investig. Dermatol. 1993, 100, 229–236. [Google Scholar] [CrossRef] [PubMed]
- Blanpain, C.; Fuchs, E. Stem cell plasticity. Plasticity of epithelial stem cells in tissue regeneration. Science 2014, 344, 1242281. [Google Scholar] [CrossRef] [PubMed]
- Trempus, C.S.; Morris, R.J.; Bortner, C.D.; Cotsarelis, G.; Faircloth, R.S.; Reece, J.M.; Tennant, R.W. Enrichment for living murine keratinocytes from the hair follicle bulge with the cell surface marker CD34. J. Investig. Dermatol. 2003, 120, 501–511. [Google Scholar] [CrossRef] [PubMed]
- Oh, J.H.; Mohebi, P.; Farkas, D.L.; Tajbakhsh, J. Towards expansion of human hair follicle stem cells in vitro. Cell Prolif. 2011, 44, 244–253. [Google Scholar] [CrossRef] [PubMed]
- Oshima, H.; Rochat, A.; Kedzia, C.; Kobayashi, K.; Barrandon, Y. Morphogenesis and renewal of hair follicles from adult multipotent stem cells. Cell 2001, 104, 233–245. [Google Scholar] [CrossRef]
- Ito, M.; Kizawa, K.; Toyoda, M.; Morohashi, M. Label-retaining cells in the bulge region are directed to cell death after plucking, followed by healing from the surviving hair germ. J. Investig. Dermatol. 2002, 119, 1310–1316. [Google Scholar] [CrossRef] [PubMed]
- Coulombe, P.A.; Kopan, R.; Fuchs, E. Expression of keratin K14 in the epidermis and hair follicle: Insights into complex programs of differentiation. J. Cell Biol. 1989, 109, 2295–2312. [Google Scholar] [CrossRef] [PubMed]
- Mills, A.A.; Zheng, B.; Wang, X.J.; Vogel, H.; Roop, D.R.; Bradley, A. p63 is a p53 homologue required for limb and epidermal morphogenesis. Nature 1999, 398, 708–713. [Google Scholar] [PubMed]
- Pellegrini, G.; Dellambra, E.; Golisano, O.; Martinelli, E.; Fantozzi, I.; Bondanza, S.; Ponzin, D.; McKeon, F.; de Luca, M. p63 identifies keratinocyte stem cells. Proc. Natl. Acad. Sci. USA 2001, 98, 3156–3161. [Google Scholar] [CrossRef] [PubMed]
- Tani, H.; Morris, R.J.; Kaur, P. Enrichment for murine keratinocyte stem cells based on cell surface phenotype. Proc. Natl. Acad. Sci. USA 2000, 97, 10960–10965. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Lyle, S.; Yang, Z.; Cotsarelis, G. Keratin 15 promoter targets putative epithelial stem cells in the hair follicle bulge. J. Investig. Dermatol. 2003, 121, 963–968. [Google Scholar] [CrossRef] [PubMed]
- Youssef, K.K.; van Keymeulen, A.; Lapouge, G.; Beck, B.; Michaux, C.; Achouri, Y.; Sotiropoulou, P.A.; Blanpain, C. Identification of the cell lineage at the origin of basal cell carcinoma. Nat. Cell Biol. 2010, 12, 299–305. [Google Scholar] [CrossRef] [PubMed]
- Jaks, V.; Barker, N.; Kasper, M.; van Es, J.H.; Snippert, H.J.; Clevers, H.; Toftgard, R. Lgr5 marks cycling, yet long-lived, hair follicle stem cells. Nat. Genet. 2008, 40, 1291–1299. [Google Scholar] [CrossRef] [PubMed]
- Snippert, H.J.; Haegebarth, A.; Kasper, M.; Jaks, V.; van Es, J.H.; Barker, N.; van de Wetering, M.; van den Born, M.; Begthel, H.; Vries, R.G.; et al. Lgr6 marks stem cells in the hair follicle that generate all cell lineages of the skin. Science 2010, 327, 1385–1389. [Google Scholar] [CrossRef] [PubMed]
- Waseem, A.; Dogan, B.; Tidman, N.; Alam, Y.; Purkis, P.; Jackson, S.; Lalli, A.; Machesney, M.; Leigh, I.M. Keratin 15 expression in stratified epithelia: Downregulation in activated keratinocytes. J. Investig. Dermatol. 1999, 112, 362–369. [Google Scholar] [CrossRef] [PubMed]
- Abbas, O.; Mahalingam, M. Epidermal stem cells: Practical perspectives and potential uses. Br. J. Dermatol. 2009, 161, 228–236. [Google Scholar] [CrossRef] [PubMed]
- Bose, A.; Teh, M.T.; Hutchison, I.L.; Wan, H.; Leigh, I.M.; Waseem, A. Two mechanisms regulate keratin K15 expression in keratinocytes: Role of PKC/AP-1 and FOXM1 mediated signalling. PLoS ONE 2012, 7, e38599. [Google Scholar] [CrossRef] [PubMed]
- Bose, A.; Teh, M.T.; Mackenzie, I.C.; Waseem, A. Keratin K15 as a biomarker of epidermal stem cells. Int. J. Mol. Sci. 2013, 14, 19385–19398. [Google Scholar] [CrossRef] [PubMed]
- Whitbread, L.A.; Powell, B.C. Expression of the intermediate filament keratin gene, K15, in the basal cell layers of epithelia and the hair follicle. Exp. Cell Res. 1998, 244, 448–459. [Google Scholar] [CrossRef] [PubMed]
- Barrandon, Y.; Grasset, N.; Zaffalon, A.; Gorostidi, F.; Claudinot, S.; Droz-Georget, S.L.; Nanba, D.; Rochat, A. Capturing epidermal stemness for regenerative medicine. Semin. Cell Dev. Biol. 2012, 23, 937–944. [Google Scholar] [CrossRef] [PubMed]
- Lichti, U.; Anders, J.; Yuspa, S.H. Isolation and short-term culture of primary keratinocytes, hair follicle populations and dermal cells from newborn mice and keratinocytes from adult mice for in vitro analysis and for grafting to immunodeficient mice. Nat. Protoc. 2008, 3, 799–810. [Google Scholar] [CrossRef] [PubMed]
- Lee, L.F.; Jiang, T.X.; Garner, W.; Chuong, C.M. A simplified procedure to reconstitute hair-producing skin. Tissue Eng. Part C Methods 2011, 17, 391–400. [Google Scholar] [CrossRef] [PubMed]
- Nowak, J.A.; Polak, L.; Pasolli, H.A.; Fuchs, E. Hair follicle stem cells are specified and function in early skin morphogenesis. Cell Stem Cell 2008, 3, 33–43. [Google Scholar] [CrossRef] [PubMed]
- Kretzschmar, K.; Watt, F.M. Lineage tracing. Cell 2012, 148, 33–45. [Google Scholar] [CrossRef] [PubMed]
- Goodell, M.A.; Nguyen, H.; Shroyer, N. Somatic stem cell heterogeneity: Diversity in the blood, skin and intestinal stem cell compartments. Nat. Rev. Mol. Cell Biol. 2015, 16, 299–309. [Google Scholar] [CrossRef] [PubMed]
- Gordon, W.; Andersen, B. A nervous hedgehog rolls into the hair follicle stem cell scene. Cell Stem Cell 2011, 8, 459–460. [Google Scholar] [CrossRef] [PubMed]
- Whitelaw, C.B. Toward ovine iPSCs. Cell Reprogr. 2013, 15, 385–388. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.Y.; Li, W.; Lv, Z.; Liu, L.; Tong, M.; Hai, T.; Hao, J.; Guo, C.L.; Ma, Q.W.; Wang, L.; et al. iPS cells produce viable mice through tetraploid complementation. Nature 2009, 461, 86–90. [Google Scholar] [CrossRef] [PubMed]
- Pfaffl, M.W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 2001, 29, e45. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Su, P.; Wang, L.; Chen, J.; Zimmermann, M.; Genbacev, O.; Afonja, O.; Horne, M.C.; Tanaka, T.; Duan, E.; et al. mTOR supports long-term self-renewal and suppresses mesoderm and endoderm activities of human embryonic stem cells. Proc. Natl. Acad. Sci. USA 2009, 106, 7840–7845. [Google Scholar] [CrossRef] [PubMed]
- Qiu, Z.; Miao, C.; Li, J.; Lei, X.; Liu, S.; Guo, W.; Cao, Y.; Duan, E.K. Skeletal myogenic potential of mouse skin-derived precursors. Stem Cells Dev. 2010, 19, 259–268. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, H.; Zhang, S.; Zhao, H.; Qiao, J.; Liu, S.; Deng, Z.; Lei, X.; Ning, L.; Cao, Y.; Zhao, Y.; et al. Ovine Hair Follicle Stem Cells Derived from Single Vibrissae Reconstitute Haired Skin. Int. J. Mol. Sci. 2015, 16, 17779-17797. https://doi.org/10.3390/ijms160817779
Zhang H, Zhang S, Zhao H, Qiao J, Liu S, Deng Z, Lei X, Ning L, Cao Y, Zhao Y, et al. Ovine Hair Follicle Stem Cells Derived from Single Vibrissae Reconstitute Haired Skin. International Journal of Molecular Sciences. 2015; 16(8):17779-17797. https://doi.org/10.3390/ijms160817779
Chicago/Turabian StyleZhang, Huishan, Shoubing Zhang, Huashan Zhao, Jingqiao Qiao, Shuang Liu, Zhili Deng, Xiaohua Lei, Lina Ning, Yujing Cao, Yong Zhao, and et al. 2015. "Ovine Hair Follicle Stem Cells Derived from Single Vibrissae Reconstitute Haired Skin" International Journal of Molecular Sciences 16, no. 8: 17779-17797. https://doi.org/10.3390/ijms160817779
APA StyleZhang, H., Zhang, S., Zhao, H., Qiao, J., Liu, S., Deng, Z., Lei, X., Ning, L., Cao, Y., Zhao, Y., & Duan, E. (2015). Ovine Hair Follicle Stem Cells Derived from Single Vibrissae Reconstitute Haired Skin. International Journal of Molecular Sciences, 16(8), 17779-17797. https://doi.org/10.3390/ijms160817779