Gene Expression Profiles of Main Olfactory Epithelium in Adenylyl Cyclase 3 Knockout Mice
Abstract
:1. Introduction
2. Results
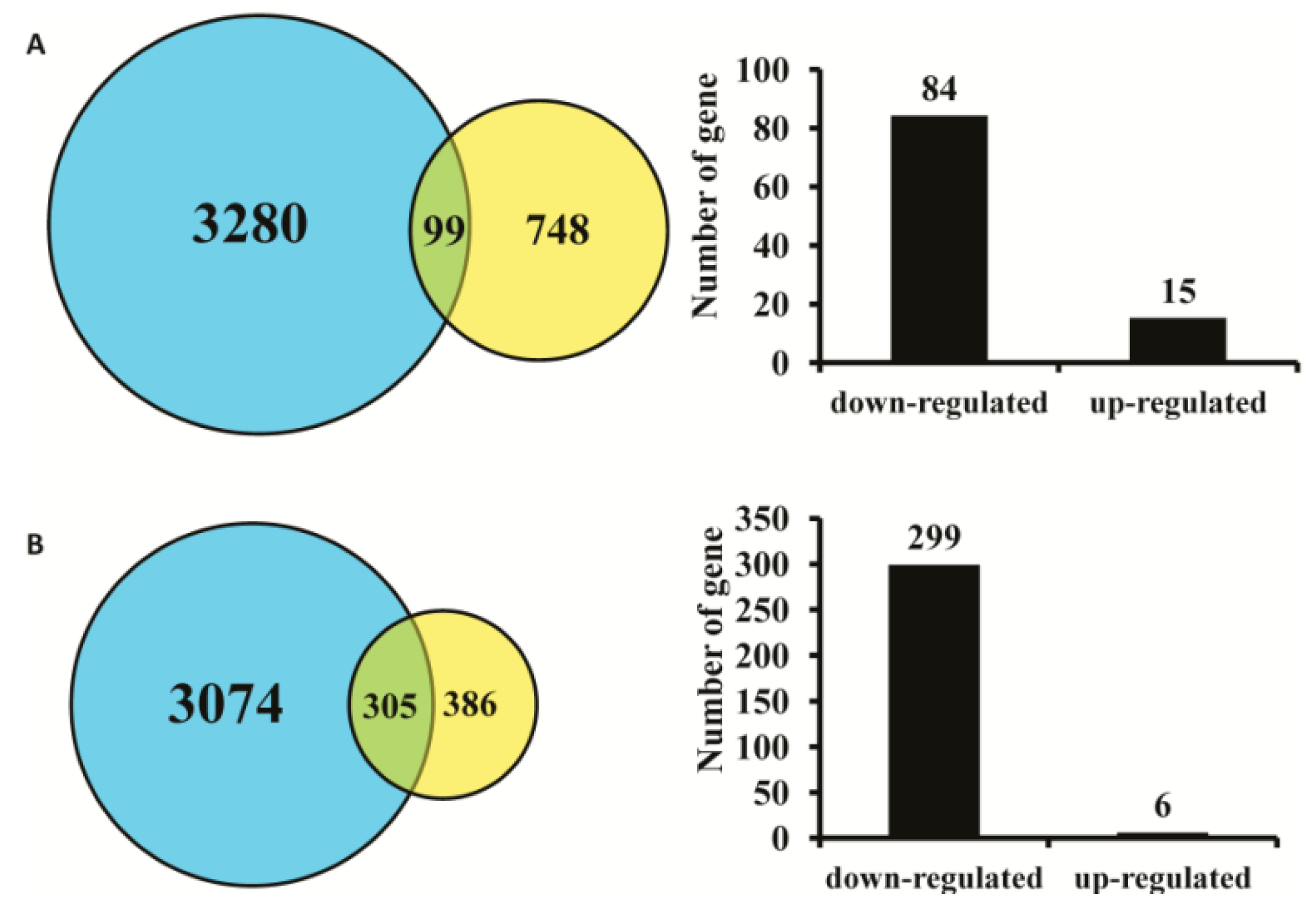
2.1. Effect of AC3 Knockout on Gene Expression in Mouse MOE
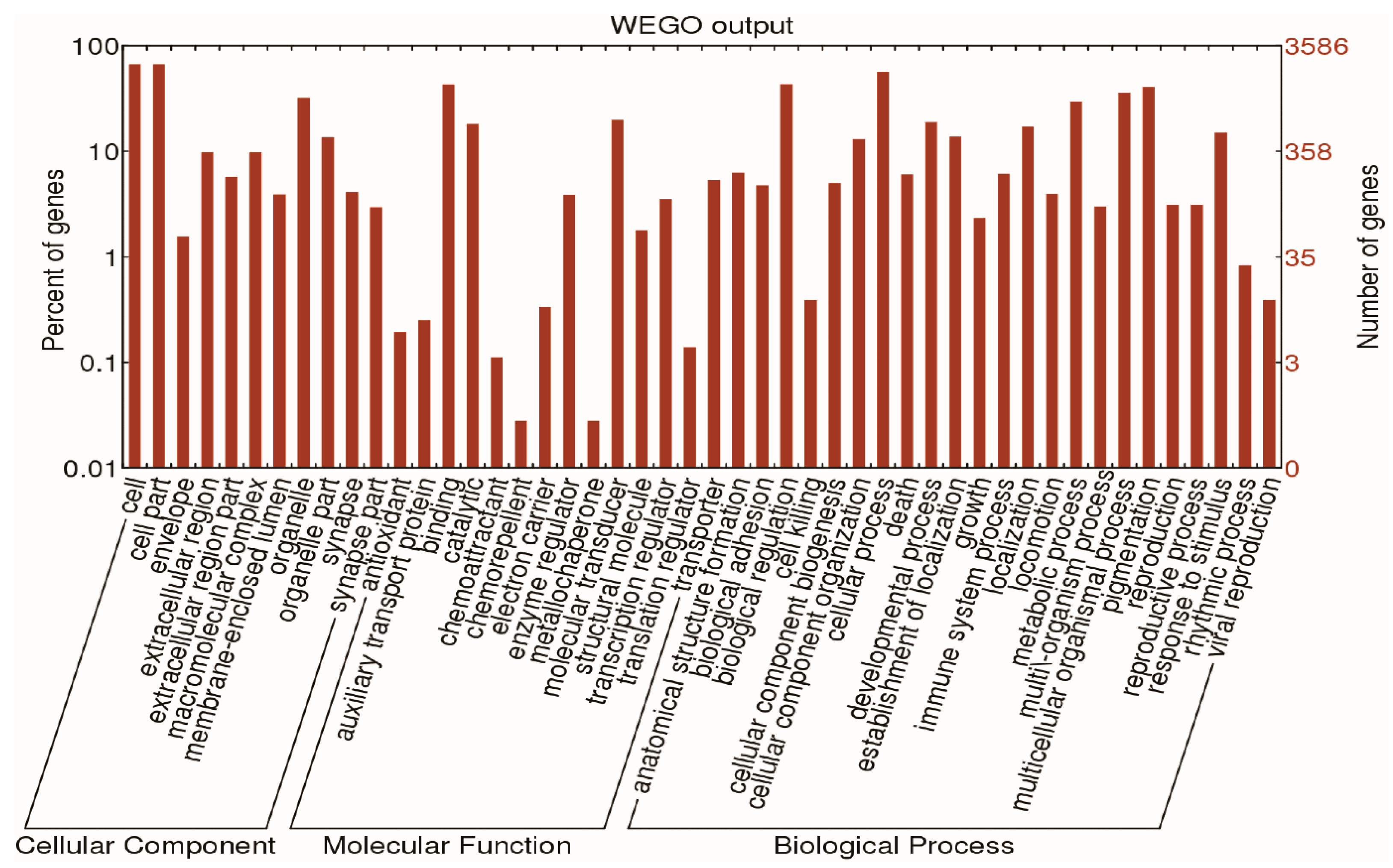
2.2. Functional Analysis of Differentially Expressed Genes
| Pathway Name | p-Value | Hit | Total | Percent |
|---|---|---|---|---|
| Metabolism correlation | ||||
| Alanine aspartate and glutamate metabolism * | 0.0101 | 6 | 34 | 17.65% |
| alpha-Linolenic acid metabolism | 0.132 | 4 | 20 | 20% |
| Amino sugar and nucleotide sugar metabolism * | 0.046 | 5 | 49 | 10.2% |
| Arachidonic acid metabolism | 0.081 | 12 | 56 | 21.43% |
| Arginine and proline metabolism ** | 0.0096 | 5 | 26 | 19.23% |
| Ascorbate and aldarate metabolism | 0.109 | 5 | 26 | 19.23% |
| Fatty acid metabolism * | 0.046 | 20 | 50 | 40% |
| Histidine metabolism | 0.134 | 5 | 28 | 17.86% |
| Inositol phosphate metabolism * | 0.027 | 17 | 57 | 29.8% |
| Fat digestion and absorption * | 0.024 | 16 | 45 | 35% |
| beta-Alanine metabolism ** | 0.0082 | 8 | 25 | 29% |
| Linoleic acid metabolism * | 0.023 | 8 | 46 | 17.39% |
| Disease association | ||||
| Type I diabetes mellitus ** | 0.000 | 21 | 63 | 33.3% |
| Olfactory transduction ** | 0.0 | 395 | 1041 | 35% |
| Graft-versus-host disease ** | 0.0 | 19 | 58 | 32.76% |
| Parkinson’s disease | 0.9 | 9 | 151 | 5.98% |
| Chagas disease * | 0.0324 | 16 | 101 | 15.84% |
| Alzheimer’s disease * | 0.037 | 19 | 75 | 25.3% |
| Autoimmune thyroid disease ** | 0.0004 | 18 | 72 | 25% |
| Allograft rejection ** | 0.0001 | 17 | 56 | 33.36% |
| Arrhythmogenic right ventricular cardiomyopathy (ARVC) ** | 0.0 | 22 | 74 | 29.73% |
| Dilated cardiomyopathy ** | 0.0 | 24 | 89 | 26.97% |
| Hypertrophic cardiomyopathy (HCM) ** | 0.0 | 24 | 83 | 28.92% |
| Cell growth and death | ||||
| Natural killer cell mediated cytotoxicity ** | 0.005 | 22 | 125 | 17.6% |
| Mitogen-activated protein kinase (MAPK) signaling pathway * | 0.0136 | 37 | 296 | 13.75% |
| Ras-Independent pathway in NK cell-mediated cytotoxicity * | 0.0313 | 5 | 17 | 29.41% |
| p53 signaling pathway * | 0.038 | 18 | 70 | 25.7% |
| Sexual maturity | ||||
| Maturity onset diabetes of the young * | 0.0453 | 6 | 26 | 23.08% |
| Oocyte meiosis * | 0.043 | 17 | 114 | 14.91% |
| GnRH signaling pathway * | 0.024 | 23 | 99 | 23.23% |
| Metabolism of xenobiotics by cytochrome P450 * | 0.011 | 11 | 78 | 15.1% |
| Neural development | ||||
| IL12 and Stat4 Dependent Signaling Pathway in Th1 Development ** | 0.0082 | 7 | 22 | 31.82% |
| Th1/Th2 Differentiation ** | 0.0034 | 7 | 18 | 38.89% |
| Neurotrophin signaling pathway * | 0.043 | 17 | 50 | 34% |
| Osteoclast differentiation * | 0.032 | 13 | 75 | 17.3% |
2.3. Effect of AC3 Knockout on cAMP Signaling Pathway-Related Genes
2.4. Effect of AC3 Knockout on the Expression of Genes Related to Cell Growth and Apoptosis, and on Their Signal Pathways

| Pathway Name | Hits | Total | p-Value | Percent |
|---|---|---|---|---|
| Apoptotic DNA fragmentation and tissue homeostasis | 1 | 9 | 0.000 | 11.1% |
| Apoptotic Signaling in Response to DNA Damage | 3 | 21 | 0.0 | 14.2% |
| Erk and PI-3 Kinase Are Necessary for Collagen Binding in Corneal Epithelia | 3 | 23 | 0.0 | 13% |
| Erk1Erk2 Mapk Signaling pathway | 4 | 28 | 0.9 | 14.2% |
| Erythropoietin mediated neuroprotection through NF-kB | 3 | 11 | 0.0324 | 27.3% |
| FAS signaling pathway (CD95) | 3 | 28 | 0.037 | 10.7% |
| MAPKinase Signaling Pathway | 5 | 77 | 0.0004 | 6% |
| p38 MAPK Signaling Pathway | 4 | 31 | 0.0001 | 13% |
| p53 Signaling Pathway | 4 | 16 | 0.0 | 25% |
| Th1/Th2 Differentiation | 7 | 18 | 0.0 | 39% |
| WNT Signaling Pathway | 2 | 25 | 0.0 | 8% |
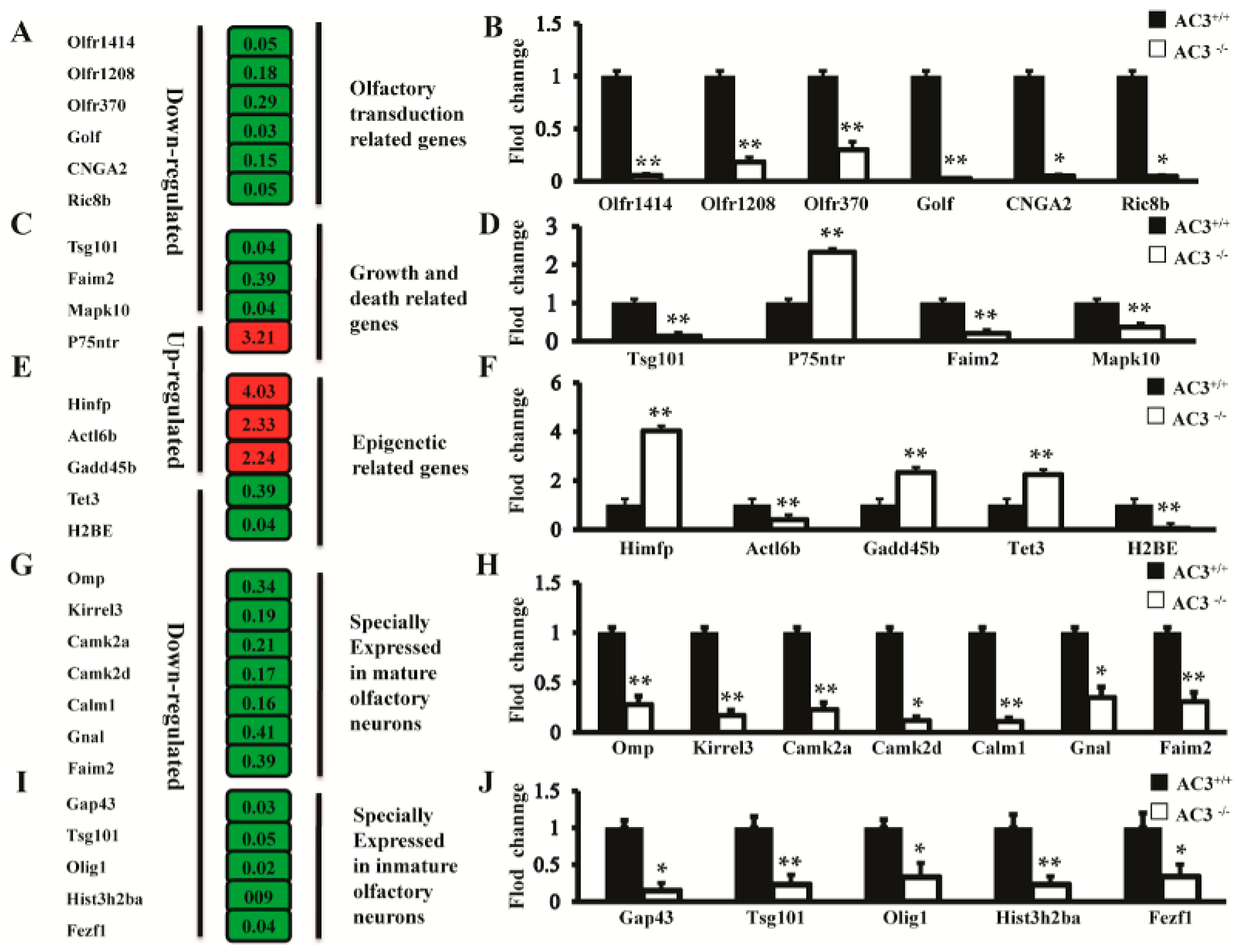
2.5. Effect of AC3 Knockout on the Expression of Genes Related to Olfactory Neuron Functions
2.6. Effect of AC3 Knockout on Epigenetic Regulation Genes
2.7. qRT-PCR Validation of the Differentially Expressed Genes
3. Discussion
4. Experimental Section
4.1. Animals and Sample Preparation
4.2. Total RNA Extraction and cDNA Synthesis
4.3. Microarray Hybridization and Screening of the Differentially Expressed Genes
4.4. Functional Annotation of the Differentially Expressed Genes
4.5. The Pathway Analysis for Differentially Expressed Genes
4.6. qRT-PCR
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Wang, Z.; Nudelman, A.; Storm, D.R. Are pheromones detected through the main olfactory epithelium? Mol. Neurobiol. 2007, 35, 317–323. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Rogers, M.; Tian, H.; Zhang, X.; Zou, D.J.; Liu, J.; Ma, M.; Shepherd, G.M.; Firestein, S.J. High-throughput microarray detection of olfactory receptor gene expression in the mouse. Proc. Natl. Acad. Sci. USA 2004, 101, 14168–14173. [Google Scholar] [CrossRef] [PubMed]
- Belluscio, L.; Gold, G.H.; Nemes, A.; Axel, R. Mice deficient in G(olf) are anosmic. Neuron 1998, 20, 69–81. [Google Scholar] [CrossRef]
- Wong, S.T.; Trinh, K.; Hacker, B.; Chan, G.C.; Lowe, G.; Gaggar, A.; Xia, Z.; Gold, G.H.; Storm, D.R. Disruption of the type III adenylyl cyclase gene leads to peripheral and behavioral anosmia in transgenic mice. Neuron 2000, 27, 487–497. [Google Scholar] [CrossRef]
- Brunet, L.J.; Gold, G.H.; Ngai, J. General anosmia caused by a targeted disruption of the mouse olfactory cyclic nucleotide-gated cation channel. Neuron 1996, 17, 681–693. [Google Scholar] [CrossRef]
- Zufall, F.; Munger, S.D. From odor and pheromone transduction to the organization of the sense of smell. Trends Neurosci. 2001, 24, 191–193. [Google Scholar] [CrossRef]
- Velez, Z.; Hubbard, P.C.; Barata, E.N.; Canario, A.V. Olfactory transduction pathways in the Senegalese sole Solea senegalensis. J. Fish Biol. 2013, 83, 501–514. [Google Scholar] [CrossRef] [PubMed]
- Rasche, S.; Toetter, B.; Adler, J.; Tschapek, A.; Doerner, J.F.; Kurtenbach, S.; Hatt, H.; Meyer, H.; Warscheid, B.; Neuhaus, E.M. Tmem16b is specifically expressed in the cilia of olfactory sensory neurons. Chem. Senses 2010, 35, 239–245. [Google Scholar] [CrossRef] [PubMed]
- Hengl, T.; Kaneko, H.; Dauner, K.; Vocke, K.; Frings, S.; Mohrlen, F. Molecular components of signal amplification in olfactory sensory cilia. Proc. Natl. Acad. Sci. USA 2010, 107, 6052–6057. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Li, V.; Chan, G.C.; Phan, T.; Nudelman, A.S.; Xia, Z.; Storm, D.R. Adult type 3 adenylyl cyclase-deficient mice are obese. PLoS ONE 2009, 4, 6979. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Balet Sindreu, C.; Li, V.; Nudelman, A.; Chan, G.C.; Storm, D.R. Pheromone detection in male mice depends on signaling through the type 3 adenylyl cyclase in the main olfactory epithelium. J. Neurosci. 2006, 26, 7375–7379. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Storm, D.R. Maternal behavior is impaired in female mice lacking type 3 adenylyl cyclase. Neuropsychopharmacology 2011, 36, 772–781. [Google Scholar] [CrossRef] [PubMed]
- Trinh, K.; Storm, D.R. Vomeronasal organ detects odorants in absence of signaling through main olfactory epithelium. Nat. Neurosci. 2003, 6, 519–525. [Google Scholar] [CrossRef] [PubMed]
- Zhenlong, C.; Jiangye, H.; Yanfen, Z.; Zhe, Z.; Zhihua, N.; Yuanxiang, H.; Weili, L.; Yongchao, L.; Daniel, R.S.; Runlin, Z.M.; et al. Differentially expressed genes identified in the main olfactory epithelium of mice with deficiency of adenylate cyclase 3 by using suppression subtractive hybridization approach. Yi Chuan 2014, 36, 574–583. [Google Scholar] [PubMed]
- Wang, S.Z.; Ou, J.; Zhu, L.J.; Green, M.R. Transcription factor ATF5 is required for terminal differentiation and survival of olfactory sensory neurons. Proc. Natl. Acad. Sci. USA 2012, 109, 18589–18594. [Google Scholar] [CrossRef] [PubMed]
- Santoro, S.W.; Dulac, C. The activity-dependent histone variant H2BE modulates the life span of olfactory neurons. Elife 2012, 1, e00070. [Google Scholar] [CrossRef] [PubMed]
- Bradley, J.; Reisert, J.; Frings, S. Regulation of cyclic nucleotide-gated channels. Curr. Opin. Neurobiol. 2005, 15, 343–349. [Google Scholar] [CrossRef] [PubMed]
- Lyons, D.B.; Allen, W.E.; Goh, T.; Tsai, L.; Barnea, G.; Lomvardas, S. An epigenetic trap stabilizes singular olfactory receptor expression. Cell 2013, 154, 325–336. [Google Scholar] [CrossRef] [PubMed]
- Rimbault, M.; Robin, S.; Vaysse, A.; Galibert, F. RNA profiles of rat olfactory epithelia: Individual and age related variations. BMC Genom. 2009, 10, 572. [Google Scholar] [CrossRef] [PubMed]
- Sammeta, N.; Yu, T.T.; Bose, S.C.; McClintock, T.S. Mouse olfactory sensory neurons express 10,000 genes. J. Comp. Neurol. 2007, 502, 1138–1156. [Google Scholar] [CrossRef] [PubMed]
- Nickell, M.D.; Breheny, P.; Stromberg, A.J.; McClintock, T.S. Genomics of mature and immature olfactory sensory neurons. J. Comp. Neurol. 2012, 520, 2608–2629. [Google Scholar] [CrossRef] [PubMed]
- Krolewski, R.C.; Packard, A.; Schwob, J.E. Global expression profiling of globose basal cells and neurogenic progression within the olfactory epithelium. J. Comp. Neurol. 2013, 521, 833–859. [Google Scholar] [CrossRef] [PubMed]
- Shetty, R.S.; Bose, S.C.; Nickell, M.D.; McIntyre, J.C.; Hardin, D.H.; Harris, A.M.; McClintock, T.S. Transcriptional changes during neuronal death and replacement in the olfactory epithelium. Mol. Cell. Neurosci. 2005, 30, 583–600. [Google Scholar] [CrossRef] [PubMed]
- McIntyre, J.C.; Bose, S.C.; Stromberg, A.J.; McClintock, T.S. Emx2 stimulates odorant receptor gene expression. Chem. Senses 2008, 33, 825–837. [Google Scholar] [CrossRef] [PubMed]
- Coppola, D.M.; Waggener, C.T. The effects of unilateral naris occlusion on gene expression profiles in mouse olfactory mucosa. J. Mol. Neurosci. 2012, 47, 604–618. [Google Scholar] [CrossRef] [PubMed]
- Fischl, A.M.; Heron, P.M.; Stromberg, A.J.; McClintock, T.S. Activity-dependent genes in mouse olfactory sensory neurons. Chem. Senses 2014, 39, 439–449. [Google Scholar] [CrossRef] [PubMed]
- Bennett, M.K.; Kulaga, H.M.; Reed, R.R. Odor-evoked gene regulation and visualization in olfactory receptor neurons. Mol. Cell. Neurosci. 2010, 43, 353–362. [Google Scholar] [CrossRef] [PubMed]
- Kajimura, D.; Dragomir, C.; Ramirez, F.; Laub, F. Identification of genes regulated by transcription factor KLF7 in differentiating olfactory sensory neurons. Gene 2007, 388, 34–42. [Google Scholar] [CrossRef] [PubMed]
- Sanabria, H.; Swulius, M.T.; Kolodziej, S.J.; Liu, J.; Waxham, M.N. CaMKII regulates actin assembly and structure. J. Biol. Chem. 2009, 284, 9770–9780. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Templeton, D.M. Cadmium activates CaMK-II and initiates CaMK-II-dependent apoptosis in mesangial cells. FEBS Lett. 2007, 581, 1481–1486. [Google Scholar] [CrossRef] [PubMed]
- Palomeque, J.; Rueda, O.V.; Sapia, L.; Valverde, C.A.; Salas, M.; Petroff, M.V.; Mattiazzi, A. Angiotensin II-induced oxidative stress resets the Ca2+ dependence of Ca2+-calmodulin protein kinase II and promotes a death pathway conserved across different species. Circ. Res. 2009, 105, 1204–1212. [Google Scholar] [CrossRef] [PubMed]
- Mercure, M.Z.; Ginnan, R.; Singer, H.A. CaM kinase II delta2-dependent regulation of vascular smooth muscle cell polarization and migration. Am. J. Physiol. Cell Physiol. 2008, 294, 1465–1475. [Google Scholar] [CrossRef] [PubMed]
- Piao, C.S.; Che, Y.; Han, P.L.; Lee, J.K. Delayed and differential induction of p38 MAPK isoforms in microglia and astrocytes in the brain after transient global ischemia. Brain Res. Mol. Brain Res. 2002, 107, 137–144. [Google Scholar] [CrossRef]
- Lee, S.M.; Tole, S.; Grove, E.; McMahon, A.P. A local Wnt-3a signal is required for development of the mammalian hippocampus. Development 2000, 127, 457–467. [Google Scholar] [PubMed]
- Megason, S.G.; McMahon, A.P. A mitogen gradient of dorsal midline Wnts organizes growth in the CNS. Development 2002, 129, 2087–2098. [Google Scholar] [PubMed]
- Cowan, C.M.; Roskams, A.J. Apoptosis in the mature and developing olfactory neuroepithelium. Microsc. Res. Tech. 2002, 58, 204–215. [Google Scholar] [CrossRef] [PubMed]
- Zheng, J.; Zagotta, W.N. Stoichiometry and assembly of olfactory cyclic nucleotide-gated channels. Neuron 2004, 42, 411–421. [Google Scholar] [CrossRef]
- Wang, Z.; Phan, T.; Storm, D.R. The type 3 adenylyl cyclase is required for novel object learning and extinction of contextual memory: Role of cAMP signaling in primary cilia. J. Neurosci. 2011, 31, 5557–5561. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Lane, A.P.; Bock, R.; Leinders-Zufall, T.; Zufall, F. Blocking adenylyl cyclase inhibits olfactory generator currents induced by “IP(3)-odors”. J. Neurophysiol. 2000, 84, 575–580. [Google Scholar] [PubMed]
- Rouquier, S.; Blancher, A.; Giorgi, D. The olfactory receptor gene repertoire in primates and mouse: Evidence for reduction of the functional fraction in primates. Proc. Natl. Acad. Sci. USA 2000, 97, 2870–2874. [Google Scholar] [CrossRef] [PubMed]
- Gilad, Y.; Przeworski, M.; Lancet, D. Loss of olfactory receptor genes coincides with the acquisition of full trichromatic vision in primates. PLoS Biol. 2004, 2, 5. [Google Scholar] [CrossRef] [PubMed]
- Laska, M.; Genzel, D.; Wieser, A. The number of functional olfactory receptor genes and the relative size of olfactory brain structures are poor predictors of olfactory discrimination performance with enantiomers. Chem. Senses 2005, 30, 171–175. [Google Scholar] [CrossRef] [PubMed]
- Niimura, Y.; Nei, M. Evolutionary dynamics of olfactory and other chemosensory receptor genes in vertebrates. J. Hum. Genet. 2006, 51, 505–517. [Google Scholar] [CrossRef] [PubMed]
- Mundy, N.I. Genetic basis of olfactory communication in primates. Am. J. Primatol. 2006, 68, 559–567. [Google Scholar] [CrossRef] [PubMed]
- Olender, T.; Feldmesser, E.; Atarot, T.; Eisenstein, M.; Lancet, D. The olfactory receptor universe—From whole genome analysis to structure and evolution. Genet. Mol. Res. 2004, 3, 545–553. [Google Scholar] [PubMed]
- Haering, C.; Kanageswaran, N.; Bouvain, P.; Scholz, P.; Altmuller, J.; Becker, C.; Gisselmann, G.; Waring-Bischof, J.; Hatt, H. Ion transporter NKCC1, modulator of neurogenesis in murine olfactory neurons. J. Biol. Chem. 2015, 290, 9767–9779. [Google Scholar] [CrossRef] [PubMed]
- Zhao, S.; Tian, H.; Ma, L.; Yuan, Y.; Yu, C.R.; Ma, M. Activity-dependent modulation of odorant receptor gene expression in the mouse olfactory epithelium. PLoS ONE 2013, 8, 69862. [Google Scholar] [CrossRef] [PubMed]
- Cheng, L.E.; Reed, R.R. Zfp423/OAZ participates in a developmental switch during olfactory neurogenesis. Neuron 2007, 54, 547–557. [Google Scholar] [CrossRef] [PubMed]
- Omura, M.; Mombaerts, P. Trpc2-expressing sensory neurons in the mouse main olfactory epithelium of type B express the soluble guanylate cyclase Gucy1b2. Mol. Cell. Neurosci. 2015, 65, 114–124. [Google Scholar] [CrossRef] [PubMed]
- Brunert, D.; Klasen, K.; Corey, E.A.; Ache, B.W. PI3Kγ-dependent signaling in mouse olfactory receptor neurons. Chem. Senses 2010, 35, 301–308. [Google Scholar] [CrossRef] [PubMed]
- Ukhanov, K.; Bruner, D.; Corey, EA.; Ache, B.W. Phosphoinositide 3-kinase-dependent antagonism in mammalian olfactory receptor neurons. J. Neurosci. 2011, 31, 273–280. [Google Scholar] [CrossRef] [PubMed]
- Kanageswaran, N.; Demond, M.; Nagel, M.; Schreiner, B.S.; Baumgart, S.; Scholz, P.; Altmuller, J.; Becker, C.; Doerner, J.F.; Conrad, H.; et al. Deep sequencing of the murine olfactory receptor neuron transcriptome. PLoS ONE 2015, 10, e0113170. [Google Scholar] [CrossRef] [PubMed]
- Ibarra-Soria, X.; Levitin, M.O.; Saraiva, L.R.; Logan, D.W. The olfactory transcriptomes of mice. PLoS Genet. 2014, 10, e1004593. [Google Scholar] [CrossRef] [PubMed]
- Von der Weid, B.; Rossier, D.; Lindup, M.; Tuberosa, J.; Widmer, A.; Col, J.D.; Kan, C.; Carleton, A.; Rodriguez, I. Large-scale transcriptional profiling of chemosensory neurons identifies receptor-ligand pairs in vivo. Nat. Neurosci. 2015, 18, 1455–1463. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Y.; Gong, N.N.; Hu, X.S.; Ni, M.J.; Pasi, R.; Matsunami, H. Molecular profiling of activated olfactory neurons identifies odorant receptors for odors in vivo. Nat. Neurosci. 2015, 18, 1446–1454. [Google Scholar] [CrossRef] [PubMed]
- Kolodziejczyk, A.A.; Kim, J.K.; Svensson, V.; Marioni, J.C.; Teichmann, S.A. The technology and biology of single-cell RNA sequencing. Mol. Cell 2015, 58, 610–620. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Storm, D.R. Extraction of DNA from mouse tails. BioTechniques 2006, 41, 410–412. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCt Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, Z.; Zhou, Y.; Luo, Y.; Zhang, J.; Zhai, Y.; Yang, D.; Zhang, Z.; Li, Y.; Storm, D.R.; Ma, R.Z. Gene Expression Profiles of Main Olfactory Epithelium in Adenylyl Cyclase 3 Knockout Mice. Int. J. Mol. Sci. 2015, 16, 28320-28333. https://doi.org/10.3390/ijms161226107
Wang Z, Zhou Y, Luo Y, Zhang J, Zhai Y, Yang D, Zhang Z, Li Y, Storm DR, Ma RZ. Gene Expression Profiles of Main Olfactory Epithelium in Adenylyl Cyclase 3 Knockout Mice. International Journal of Molecular Sciences. 2015; 16(12):28320-28333. https://doi.org/10.3390/ijms161226107
Chicago/Turabian StyleWang, Zhenshan, Yanfen Zhou, Yingtao Luo, Jing Zhang, Yunpeng Zhai, Dong Yang, Zhe Zhang, Yongchao Li, Daniel R. Storm, and Runlin Z. Ma. 2015. "Gene Expression Profiles of Main Olfactory Epithelium in Adenylyl Cyclase 3 Knockout Mice" International Journal of Molecular Sciences 16, no. 12: 28320-28333. https://doi.org/10.3390/ijms161226107
APA StyleWang, Z., Zhou, Y., Luo, Y., Zhang, J., Zhai, Y., Yang, D., Zhang, Z., Li, Y., Storm, D. R., & Ma, R. Z. (2015). Gene Expression Profiles of Main Olfactory Epithelium in Adenylyl Cyclase 3 Knockout Mice. International Journal of Molecular Sciences, 16(12), 28320-28333. https://doi.org/10.3390/ijms161226107





