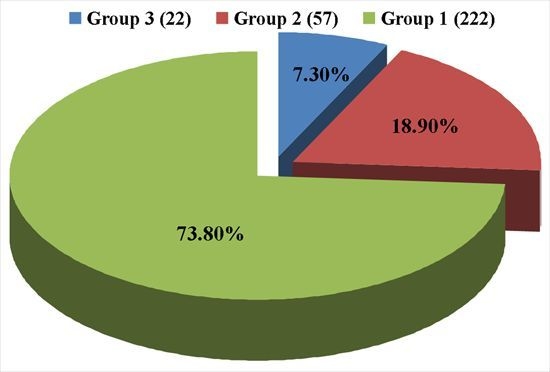
Identification of Pathogenicity-Related Genes in Biofilm-Defective Acidovorax citrulli by Transposon Tn5 Mutagenesis
Abstract
:1. Introduction
2. Results and Discussion
2.1. Construction of Virulence-Related Mutants
| Primers Name | Sequence (5’-3’) | Tm (°C) | GC (%) | Characteristics of Primers |
|---|---|---|---|---|
| AC-F | GACCAGCCACACTGGGAC | 55 | 67 | identification of A. citrulli strains |
| AC-R | CTGCCGTACTCCAGCGAT | 53 | 61 | |
| Kan-F | TTGTCAGCTTCGGTCAGTTG | 52 | 50 | identification of the Tn5 mutants |
| Kan-R | GCCTGAGCGAGACGAAATAC | 54 | 55 | |
| AD-1 | ACGATGGACTCCAGAGCANANNNGGAA | 57 | 52 | amplification of unknown sequence flanking transposon Tn5 in pre-amplification step |
| SP-1 (get 3’ sequence) | TTGCGCCTGAGCGAGACGAAATAC | 59 | 54 | |
| SP-4 (get 5’ sequence) | ATCAGATCACGCATCTTCCC | 52 | 50 | |
| SP-2 (get 5’ sequence) | ACGATGGACTCCAGATTGATGGTCGGAAGAGGC | 67 | 55 | amplification of unknown sequence flanking transposon Tn5 in the primary step |
| SP-5 (get 3’ sequence) | ACCTACAACAAAGCTCTCATCAACC | 56 | 44 | |
| AD-2 | ACGATGGACTCCAGAG | 46 | 56 | amplification of unknown sequence flanking transposon Tn5 in the secondary step |
| SP-3 (get 3’ sequence) | TCGCACCTGATTGCCCGACATTAT | 57 | 50 | |
| SP-6 (get 5’ sequence) | AGATGTGTATAAGAGACAG | 45 | 37 |
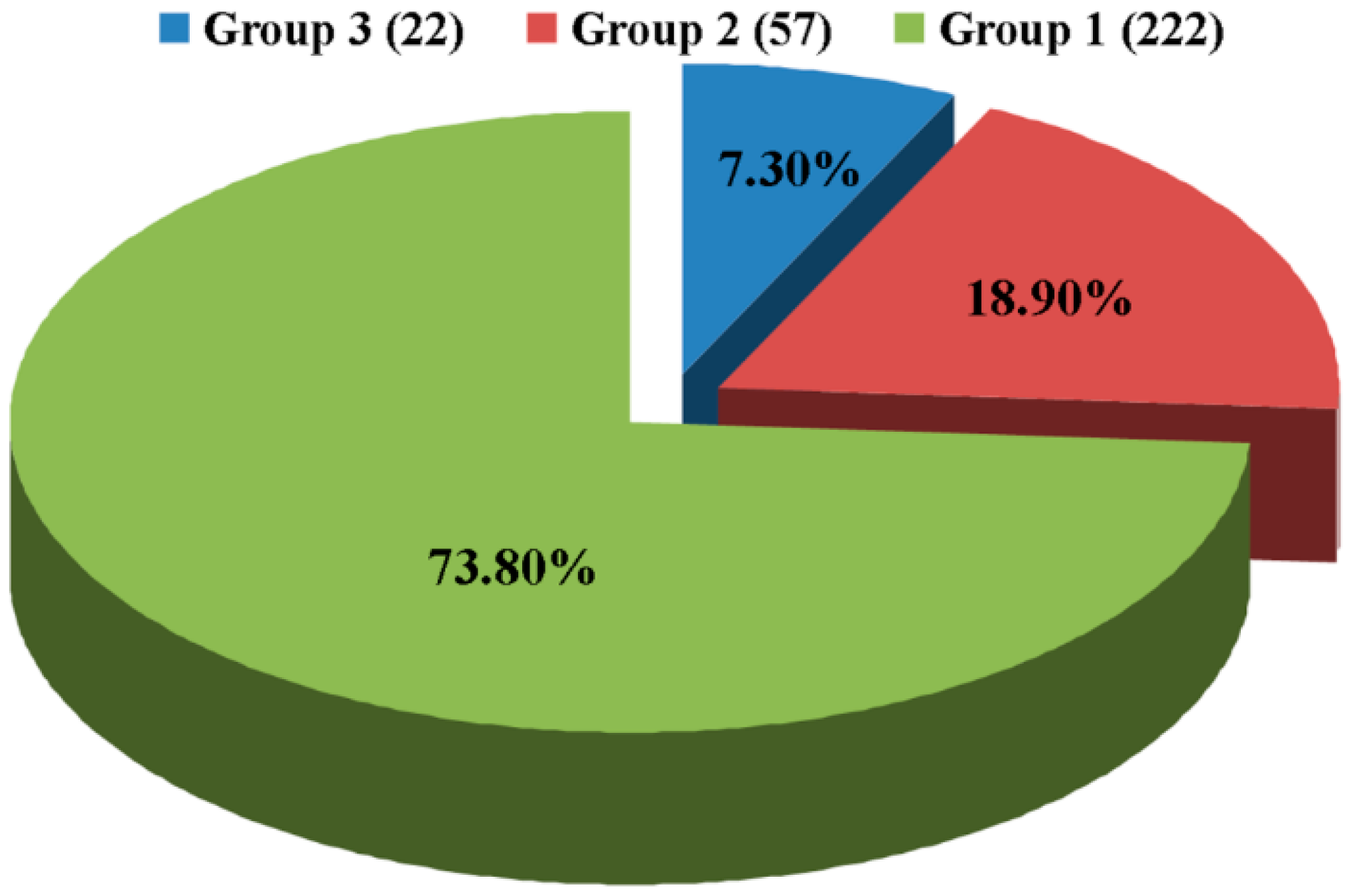
2.2. Location of Tn5 Inserted Sites
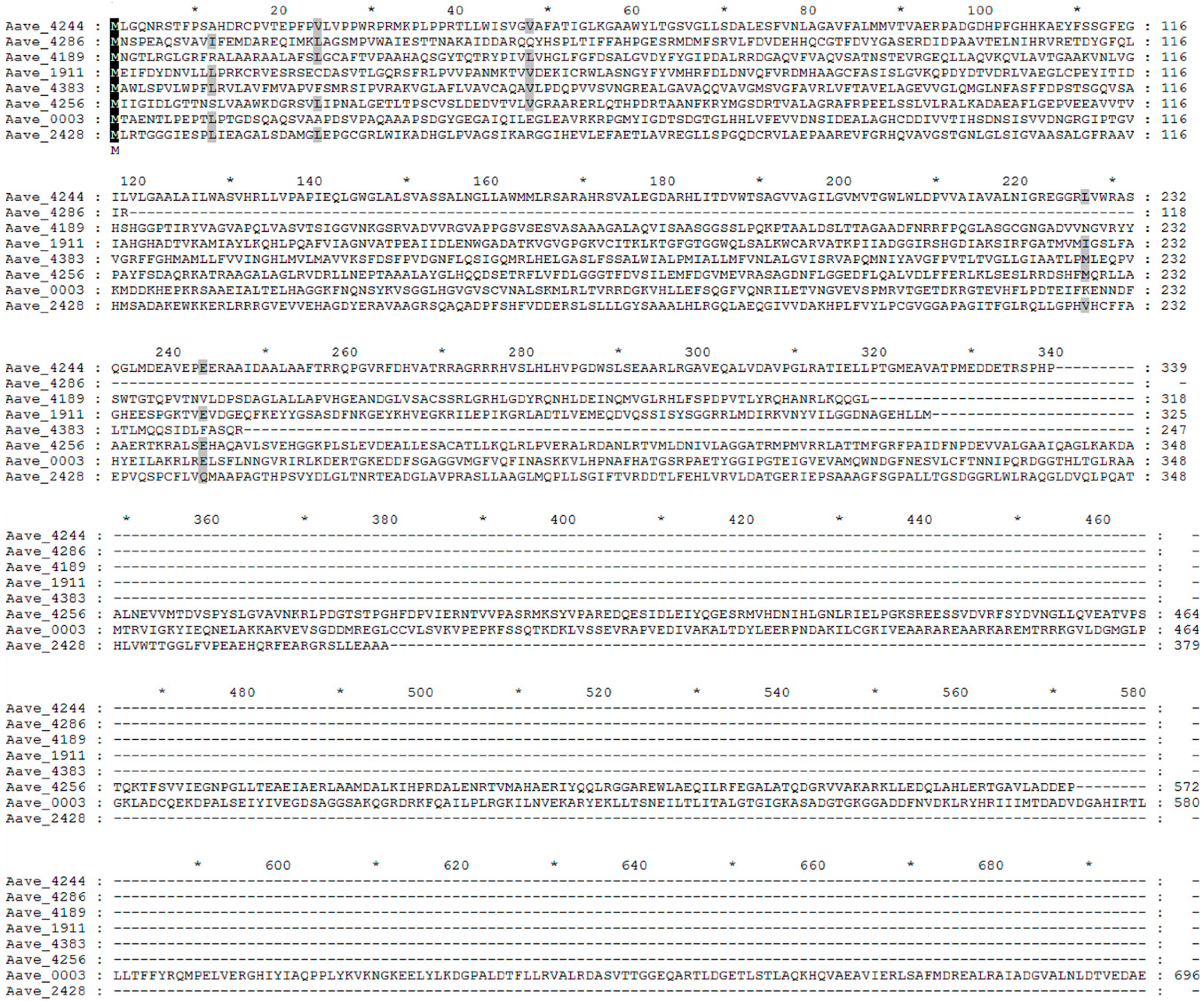
2.3. Functional Prediction of Virulence-Related Genes
| Locus_Tag | Gene Name | Pfam No. | Pfam E-Value | Mutant Nos. | Function Prediction |
|---|---|---|---|---|---|
| Aave 4244 | Cation_efflux | PF01545 | 1 × 1039 | 3 | Cation diffusion facilitator family transporter |
| Aave 4286 | Not found | Not found | – | 4 | Hypothetical protein |
| Aave 4189 | Abhydrolase_1 | PF00561 | 4.2 × 1016 | 2 | Alpha/beta hydrolase fold |
| Aave 1911 | IMPDH | PF00478 | 2.1 × 1062 | 1 | IMPdehydrogenase/GMP reductase domain |
| Aave 4383 | Bac_export_1 | PF01311 | 1.1 × 1066 | 5 | Bacterial export proteins, family 1 |
| Aave 4256 | HSP70 | PF00012 | 5.8 × 1017 | 2 | Hsp70 protein |
| Aave 0003 | HATPase_c | PF02518 | 2.2 × 1018 | 4 | Histidine kinase, DNA gyrase B, and HSP90-like ATPase |
| Aave 2428 | PALP | PF00291 | 1.7 × 1042 | 1 | Pyridoxal-phosphate dependent enzyme |
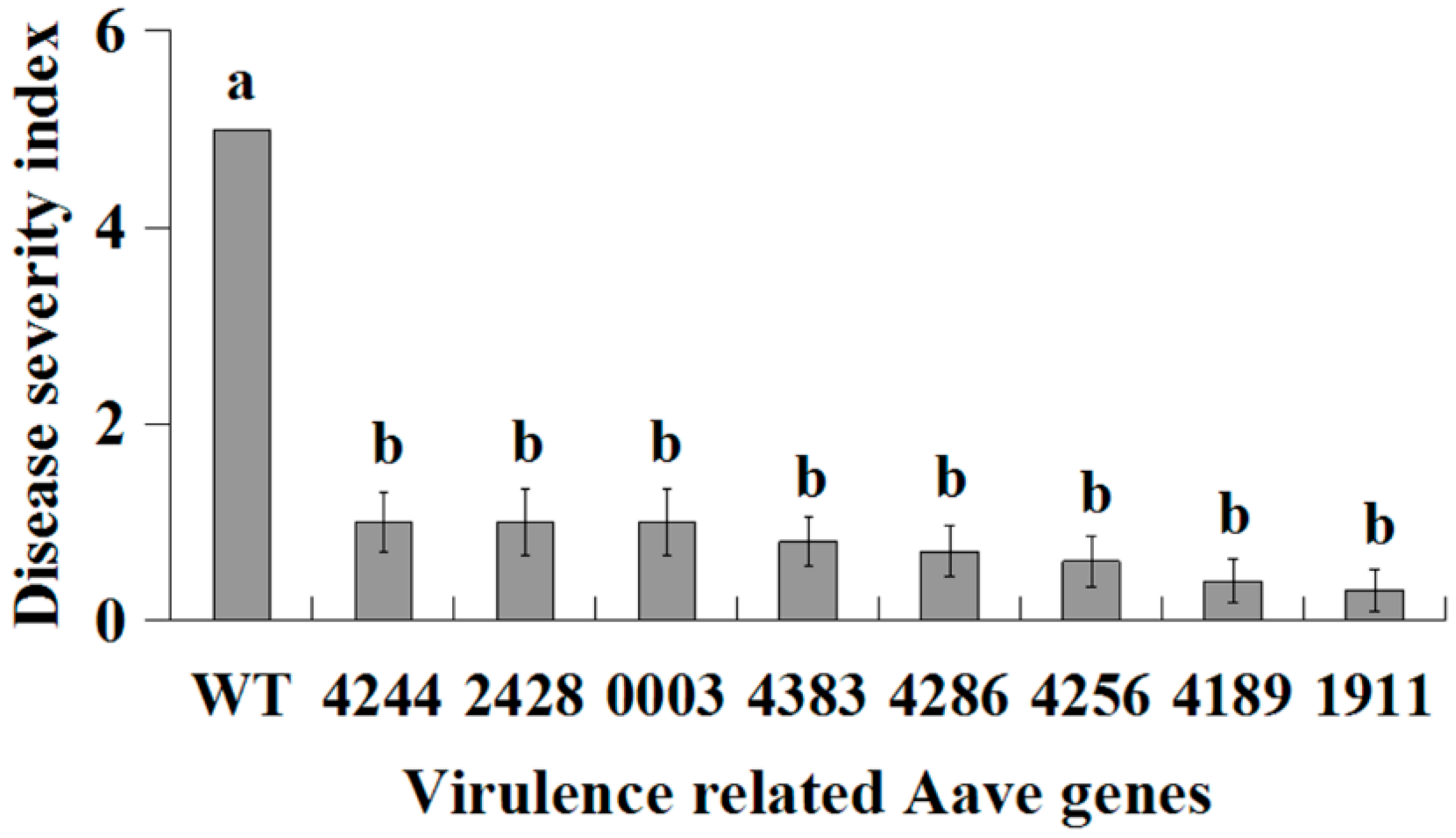
2.4. Growth of Virulence-Related Mutants
| Bacterial Strains | Incubation Time (h) | |
|---|---|---|
| 12 | 24 | |
| Wild type | 0.82 ± 0.02 a | 1.27 ± 0.02 b |
| Aave 2428 | 0.95 ± 0.01 a | 1.47 ± 0.01 a |
| Aave 4256 | 0.45 ± 0.05 b | 0.80 ± 0.02 c |
| Aave 0003 | 0.41 ± 0.05 b | 0.51 ± 0.04 d |
| Aave 4189 | 0.37 ± 0.04 b | 0.46 ± 0.04 d |
| Aave 4383 | 0.40 ± 0.05 b | 0.45 ± 0.04 d |
| Aave 1911 | 0.16 ± 0.01 c | 0.35 ± 0.01 e |
| Aave 4244 | 0.17 ± 0.01 c | 0.30 ± 0.01 e |
| Aave 4286 | 0.17 ± 0.00 c | 0.29 ± 0.01 e |
2.5. Stress Response of Virulence-Related Mutants
| Bacterial Strains | Temperature (°C) | ||
|---|---|---|---|
| 15 | 30 | 45 | |
| WT | 0.38 ± 0.04 a | 1.27 ± 0.02 b | 0.35 ± 0.02 b |
| Aave 4189 | 0.38 ± 0.01 a | 0.46 ± 0.04 d | 0.30 ± 0.03 b |
| Aave 4383 | 0.37 ± 0.01 a | 0.45 ± 0.04 d | 0.25 ± 0.05 b |
| Aave 0003 | 0.36 ± 0.02 a | 0.51 ± 0.04 d | 0.24 ± 0.04 b |
| Aave 2428 | 0.34 ± 0.01 a | 1.47 ± 0.01 a | 0.51 ± 0.03 a |
| Aave 4256 | 0.33 ± 0.02 a | 0.80 ± 0.02 c | 0.20 ± 0.04 b |
| Aave 4244 | 0.07 ± 0.00 b | 0.30 ± 0.01 e | 0.03 ± 0.00 c |
| Aave 4286 | 0.07 ± 0.00 b | 0.29 ± 0.01 e | 0.07 ± 0.05 c |
| Aave 1911 | 0.06 ± 0.00 b | 0.35 ± 0.01 e | 0.07 ± 0.04 c |
| Bacterial Strains | NaCl | |
|---|---|---|
| 170 mM | 340 mM | |
| WT | 1.27 ± 0.02 b | 0.96 ± 0.02 a |
| Aave 2428 | 1.47 ± 0.01 a | 1.01 ± 0.02 a |
| Aave 4256 | 0.80 ± 0.02 c | 0.46 ± 0.05 b |
| Aave 0003 | 0.51 ± 0.04 d | 0.32 ± 0.03 c |
| Aave 4189 | 0.46 ± 0.04 d | 0.32 ± 0.03 c |
| Aave 4383 | 0.45 ± 0.04 d | 0.35 ± 0.03 bc |
| Aave 1911 | 0.35 ± 0.01 e | 0.15 ± 0.01 d |
| Aave 4244 | 0.30 ± 0.01 e | 0.14 ± 0.02 d |
| Aave 4286 | 0.29 ± 0.01 e | 0.10 ± 0.01 d |
| Bacterial Strains | pH | ||
|---|---|---|---|
| 5.0 | 7.0 | 9.0 | |
| WT | 0.94 ± 0.19 b | 1.27 ± 0.02 b | 1.10 ± 0.02 a |
| Aave 2428 | 1.28 ± 0.02 a | 1.47 ± 0.01 a | 1.21 ± 0.07 a |
| Aave 4256 | 0.31 ± 0.03 c | 0.80 ± 0.02 c | 0.58 ± 0.06 b |
| Aave 0003 | 0.26 ± 0.06 c | 0.51 ± 0.04 d | 0.34 ± 0.03 cd |
| Aave 4189 | 0.15 ± 0.06 c | 0.46 ± 0.04 d | 0.32 ± 0.05 cd |
| Aave 4383 | 0.19 ± 0.04 c | 0.45 ± 0.04 d | 0.40 ± 0.05 bc |
| Aave 1911 | 0.35 ± 0.04 c | 0.35 ± 0.01 e | 0.26 ± 0.04 de |
| Aave 4244 | 1.00 ± 0.03 ab | 0.30 ± 0.01 e | 0.10 ± 0.02 e |
| Aave 4286 | 0.70 ± 0.02 b | 0.29 ± 0.01 e | 0.24 ± 0.05 de |
3. Experimental Section
3.1. Bacterial Strains and Growth Conditions
3.2. Transposon Insertion of Tn5
3.3. Virulence of Mutants to Watermelon
3.4. Cloning of Tn5-Tagged Regions by hiTAIL-PCR
3.5. Identification of Virulence-Related Genes
3.6. Growth Measurement of the Wild Type and Mutants
3.7. Growth of Mutants under Different Conditions
3.8. Statistical Analyses
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Burdman, S.; Walcott, R. Acidovorax citrulli: Generating basic and applied knowledge to tackle a global threat to the cucurbit industry. Mol. Plant Pathol. 2012, 13, 805–815. [Google Scholar] [CrossRef] [PubMed]
- Johnson, K.; Minsavage, G.; Le, T.; Jones, J.; Walcott, R. Efficacy of a nonpathogenic Acidovorax citrulli strain as a biocontrol seed treatment for bacterial fruit blotch of cucurbits. Plant Dis. 2011, 95, 697–704. [Google Scholar] [CrossRef]
- Wall, G.; Santos, V.; Cruz, F.; Nelson, D.; Cabrera, I. Outbreak of watermelon fruit blotch in the Mariana islands. Plant Dis. 1990, 74, 80. [Google Scholar] [CrossRef]
- Chalupowicz, L.; Dror, O.; Reuven, M.; Burdman, S.; Manulis-Sasson, S. Cotyledons are the main source of secondary spread of Acidovorax citrulli in melon nurseries. Plant Pathol. 2015, 64, 528–536. [Google Scholar] [CrossRef]
- Dutta, B.; Ha, Y.; Lessl, J.; Avci, U.; Sparks, A.; Johnson, K.; Walcott, R. Pathways of bacterial invasion and watermelon seed infection by Acidovorax citrulli. Plant Pathol. 2015, 64, 537–544. [Google Scholar] [CrossRef]
- Dutta, B.; Sanders, H.; Langston, D.; Booth, C.; Smith, S.; Gitaitis, R. Long-term survival of Acidovorax citrulli in citron melon (Citrullus lanatus var. citroides) seeds. Plant Pathol. 2014, 63, 1130–1137. [Google Scholar] [CrossRef]
- Ren, Z.; Jiang, W.; Ni, X.; Lin, M.; Zhang, W.; Tian, G.; Zhang, L. Multiplication of Acidovorax citrulli in planta during infection of melon seedlings requires the ability to synthesize leucine. Plant Pathol. 2014, 63, 784–791. [Google Scholar] [CrossRef]
- Shrestha, R.K.; Rosenberg, T.; Makarovsky, D.; Eckshtain-Levi, N.; Zelinger, E.; Kopelowitz, J.; Sikorski, J.; Burdman, S. Phenotypic variation in the plant pathogenic bacterium Acidovorax citrulli. PLoS ONE 2013, 8, e73189. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Johnson, K.L.; Walcott, R.R. Quorum Sensing Contributes to Seed-to-Seedling Transmission of Acidovorax citrulli on Watermelon. J. Phytopathol. 2013, 161, 562–573. [Google Scholar] [CrossRef]
- Li, B.; Shi, Y.; Shan, C.; Zhou, Q.; Ibrahim, M.; Wang, Y.; Wu, G.; Li, H.; Xie, G.; Sun, G. Effect of chitosan solution on the inhibition of Acidovorax citrulli causing bacterial fruit blotch of watermelon. J. Sci. Food Agric. 2013, 93, 1010–1015. [Google Scholar] [CrossRef] [PubMed]
- Horuz, S.; Cetinkaya-Yildiz, R.; Mirik, M.; Aysan, Y. Occurrence, Isolation, and Identification of Acidovorax citrulli from Melon in Turkey. Plant Prot. Sci. 2014, 50, 179–183. [Google Scholar]
- Cho, M.; Park, D.; Ahn, T.; Park, D. Rapid and specific detection of Acidovorax avenae subsp. citrulli by SYBR Green real-time PCR using YD repeat protein gene. J. Microbiol. Biotechnol. 2015, 25, 1401–1409. [Google Scholar] [PubMed]
- Himananto, O.; Thummabenjapone, P.; Luxananil, P.; Kumpoosiri, M.; Hongprayoon, R.; Kositratana, W.; Gajanandana, O. Novel and highly specific monoclonal antibody to Acidovorax citrulli and development of ELISA-based detection in cucurbit leaves and seed. Plant Dis. 2011, 95, 1172–1178. [Google Scholar] [CrossRef]
- Wang, Y.; Zhou, Q.; Li, B.; Liu, B.; Wu, G.; Ibrahim, M.; Xie, G.; Li, H.; Sun, G. Differentiation in MALDI-TOF MS and FTIR spectra between two closely related species Acidovorax oryzae and Acidovorax citrulli. BMC Microbiol. 2012, 12, 182. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Sun, B.; Yang, Y.; Zhao, T. Genome sequence of Acidovorax citrulli group 1 strain pslb65 causing bacterial fruit blotch of melons. Genome Announc. 2015, 3, pii:e00327–e00315. [Google Scholar]
- Wang, T.; Yang, Y.; Zhao, T. Genome sequence of a copper-resistant strain of Acidovorax citrulli causing bacterial fruit blotch of melons. Genome Announc. 2015, 3, pii:e00310–e00315. [Google Scholar]
- Liu, J.; Luo, S.; Zhang, Q.; Wang, Q.; Chen, J.; Guo, A.; Shan, W. Tn5 transposon mutagenesis in Acidovorax citrulli for identification of genes required for pathogenicity on cucumber. Plant Pathol. 2012, 61, 364–374. [Google Scholar] [CrossRef]
- Bahar, O.; Levi, N.; Burdman, S. The cucurbit pathogenic bacterium Acidovorax citrulli requires a polar flagellum for full virulence before and after host-tissue penetration. Mol. Plant. Microbe Interact. 2011, 24, 1040–1050. [Google Scholar] [CrossRef] [PubMed]
- Bahar, O.; Goffer, T.; Burdman, S. Type IV pili are required for virulence, twitching motility, and biofilm formation of Acidovorax avenae subsp. citrulli. Mol. Plant. Microbe Interact. 2009, 22, 909–920. [Google Scholar] [CrossRef] [PubMed]
- Tian, Y.; Zhao, Y.; Wu, X.; Liu, F.; Hu, B.; Walcott, R.R. The type VI protein secretion system contributes to biofilm formation and seed-to-seedling transmission of Acidovorax citrulli on melon. Mol. Plant Pathol. 2015, 16, 38–47. [Google Scholar] [CrossRef] [PubMed]
- Eckshtain-Levi, N.; Munitz, T.; Zivanovic, M.; Traore, S.M.; Spröer, C.; Zhao, B.; Welbaum, G.; Walcott, R.; Sikorski, J.; Burdman, S. Comparative analysis of type III secreted effector genes reflects divergence of Acidovorax citrulli strains into three distinct lineages. Phytopathology 2014, 104, 1152–1162. [Google Scholar] [CrossRef] [PubMed]
- Kristich, C.J.; Nguyen, V.T.; Le, T.; Barnes, A.M.T.; Grindle, S.; Dunny, G.M. Development and use of an efficient system for random mariner transposon mutagenesis to identify novel genetic determinants of biofilm formation in the core Enterococcus faecalis genome. Appl. Environ. Microbiol. 2008, 74, 3377–3386. [Google Scholar] [CrossRef] [PubMed]
- Filloux, A.; Vallet, I. Biofilm: Positioning and organisation of bacterial communities. Med. Sci. 2003, 19, 77–83. [Google Scholar]
- Nies, D.H. Efflux-mediated heavy metal resistance in prokaryotes. FEMS Microbiol. Rev. 2003, 27, 313–339. [Google Scholar] [CrossRef]
- Ollis, D.L.; Cheah, E.; Cygler, M.; Dijkstra, B.; Frolow, F.; Franken, S.M.; Harel, M.; Remington, S.J.; Silman, I.; Schrag, J.; et al. The alpha/beta hydrolase fold. Protein Eng. 1992, 5, 197–211. [Google Scholar] [CrossRef] [PubMed]
- Whitby, F.G.; Luecke, H.; Kuhn, P.; Somoza, J.R.; Huete-Perez, J.A.; Phillips, J.D.; Hill, C.P.; Fletterick, R.J.; Wang, C.C. Crystal structure of Tritrichomonas foetus inosine-5’-monophosphate dehydrogenase and the enzyme-product complex. Biochemistry 1997, 36, 10666–10674. [Google Scholar] [CrossRef] [PubMed]
- Zhang, R.-G.; Evans, G.; Rotella, F.J.; Westbrook, E.M.; Beno, D.; Huberman, E.; Joachimiak, A.; Collart, F.R. Characteristics and crystal structure of bacterial inosine-5'-monophosphate dehydrogenase. Biochemistry 1999, 38, 4691–4700. [Google Scholar] [CrossRef] [PubMed]
- Zhao, H.; Chiaro, C.R.; Zhang, L.; Smith, P.B.; Chan, C.Y.; Pedley, A.M.; Pugh, R.J.; French, J.B.; Patterson, A.D.; Benkovic, S.J. Quantitative analysis of purine nucleotides indicates that purinosomes increase de novo purine biosynthesis. J. Biol. Chem. 2015, 290, 6705–6713. [Google Scholar] [CrossRef] [PubMed]
- Zhuang, W.Y.; Shapiro, L. Caulobacter FliQ and FliR membrane proteins, required for flagellar biogenesis and cell division, belong to a family of virulence factor export proteins. J. Bacteriol. 1995, 177, 343–356. [Google Scholar] [PubMed]
- Tavaria, M.; Gabriele, T.; Kola, I.; Anderson, R.L. A hitchhiker's guide to the human Hsp70 family. Cell Stress Chaperones 1996, 1, 23. [Google Scholar] [CrossRef]
- Morano, K.A. New Tricks for an Old Dog. Ann. N. Y. Acad. Sci. 2007, 1113, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Beere, H.M.; Wolf, B.B.; Cain, K.; Mosser, D.D.; Mahboubi, A.; Kuwana, T.; Tailor, P.; Morimoto, R.I.; Cohen, G.M.; Green, D.R. Heat-shock protein 70 inhibits apoptosis by preventing recruitment of procaspase-9 to the Apaf-1 apoptosome. Nat. Cell. Biol. 2000, 2, 469–475. [Google Scholar] [PubMed]
- Dutta, R.; Inouye, M. GHKL, an emergent ATPase/kinase superfamily. Trends Biochem. Sci. 2000, 25, 24–28. [Google Scholar] [CrossRef]
- Phillips, A.; Wood, W. The mechanism of action of 5’-adenylic acid-activated threonine dehydrase. J. Biol. Chem. 1965, 240, 4703–4709. [Google Scholar] [PubMed]
- Parsot, C. Evolution of biosynthetic pathways: A common ancestor for threonine synthase, threonine dehydratase and D-serine dehydratase. EMBO J. 1986, 5, 3013. [Google Scholar] [PubMed]
- Soutourina, J.; Blanquet, S.; Plateau, P. Role of d-cysteine desulfhydrase in the adaptation of Escherichia coli to d-cysteine. J. Biol. Chem. 2001, 276, 40864–40872. [Google Scholar] [CrossRef] [PubMed]
- Xiong, A.; Jayaswal, R.K. Molecular characterization of a chromosomal determinant conferring resistance to zinc and cobalt ions in Staphylococcus aureus. J. Bacteriol. 1998, 180, 4024–4029. [Google Scholar] [PubMed]
- Li, B.; Ibrahim, M.; Ge, M.; Cui, Z.; Sun, G.; Xu, F.; Kube, M. Transcriptome analysis of Acidovorax avenae subsp. avenae cultivated in vivo and co-culture with Burkholderia seminalis. Sci. Rep. 2014, 4, 5698. [Google Scholar] [PubMed]
- Wang, Y.; Li, L.; Li, B.; Wu, G.; Tang, Q.; Ibrahim, M.; Li, H.; Xie, G.; Sun, G. Action of chitosan against Xanthomonas pathogenic bacteria isolated from Euphorbia pulcherrima. Molecules 2012, 17, 7028–7041. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Mueller, A.P.; Leong, C.R.; Matsumoto, K.i.; Taguchi, S.; Nomura, C.T. Quick and efficient method for genetic transformation of biopolymer-producing bacteria. J. Chem. Technol. Biotechnol. 2010, 85, 775–778. [Google Scholar] [CrossRef]
- Fan, J.; Qian, G.; Chen, T.; Zhao, Y.; Liu, F.; Walcott, R.R.; Hu, B. The acyl-homoserine lactone (AHL)-type quorum sensing system affects growth rate, swimming motility and virulence in Acidovorax avenae subsp. citrulli. World J. Microbiol. Biotechnol. 2011, 27, 1155–1166. [Google Scholar] [CrossRef]
- Jiang, C.H.; Wu, F.; Yu, Z.Y.; Xie, P.; Ke, H.J.; Li, H.W.; Yu, Y.Y.; Guo, J.H. Study on screening and antagonistic mechanisms of Bacillus amyloliquefaciens 54 against bacterial fruit blotch (BFB) caused by Acidovorax avenae subsp. citrulli. Microbiol. Res. 2015, 170, 95–104. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.G.; Whittier, R.F. Thermal asymmetric interlaced PCR: Automatable amplification and sequencing of insert end fragments from P1 and YAC clones for chromosome walking. Genomics 1995, 25, 674–681. [Google Scholar] [CrossRef]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Finn, R.D.; Bateman, A.; Clements, J.; Coggill, P.; Eberhardt, R.Y.; Eddy, S.R.; Heger, A.; Hetherington, K.; Holm, L.; Mistry, J.; et al. Pfam: The protein families database. Nucleic Acids Res. 2014, 42, 222–230. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Wang, X.; Chen, R.; Huangfu, W.; Xie, G. Antibacterial activity of chitosan solution against Xanthomonas pathogenic bacteria isolated from Euphorbia pulcherrima. Carbohydr. Polym. 2008, 72, 287–292. [Google Scholar] [CrossRef]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Luo, J.; Qiu, W.; Chen, L.; Anjum, S.I.; Yu, M.; Shan, C.; Ilyas, M.; Li, B.; Wang, Y.; Sun, G. Identification of Pathogenicity-Related Genes in Biofilm-Defective Acidovorax citrulli by Transposon Tn5 Mutagenesis. Int. J. Mol. Sci. 2015, 16, 28050-28062. https://doi.org/10.3390/ijms161226076
Luo J, Qiu W, Chen L, Anjum SI, Yu M, Shan C, Ilyas M, Li B, Wang Y, Sun G. Identification of Pathogenicity-Related Genes in Biofilm-Defective Acidovorax citrulli by Transposon Tn5 Mutagenesis. International Journal of Molecular Sciences. 2015; 16(12):28050-28062. https://doi.org/10.3390/ijms161226076
Chicago/Turabian StyleLuo, Jinyan, Wen Qiu, Lei Chen, Syed Ishtiaq Anjum, Menghao Yu, Changlin Shan, Mehmoona Ilyas, Bin Li, Yanli Wang, and Guochang Sun. 2015. "Identification of Pathogenicity-Related Genes in Biofilm-Defective Acidovorax citrulli by Transposon Tn5 Mutagenesis" International Journal of Molecular Sciences 16, no. 12: 28050-28062. https://doi.org/10.3390/ijms161226076
APA StyleLuo, J., Qiu, W., Chen, L., Anjum, S. I., Yu, M., Shan, C., Ilyas, M., Li, B., Wang, Y., & Sun, G. (2015). Identification of Pathogenicity-Related Genes in Biofilm-Defective Acidovorax citrulli by Transposon Tn5 Mutagenesis. International Journal of Molecular Sciences, 16(12), 28050-28062. https://doi.org/10.3390/ijms161226076