The Influence of Fatty Acids on the GpA Dimer Interface by Coarse-Grained Molecular Dynamics Simulation
Abstract
:1. Introduction
2. Results
2.1. Self-Assembly of the GpA-Transmembrane Region Reconstructs the NMR Interface with Low Frequency
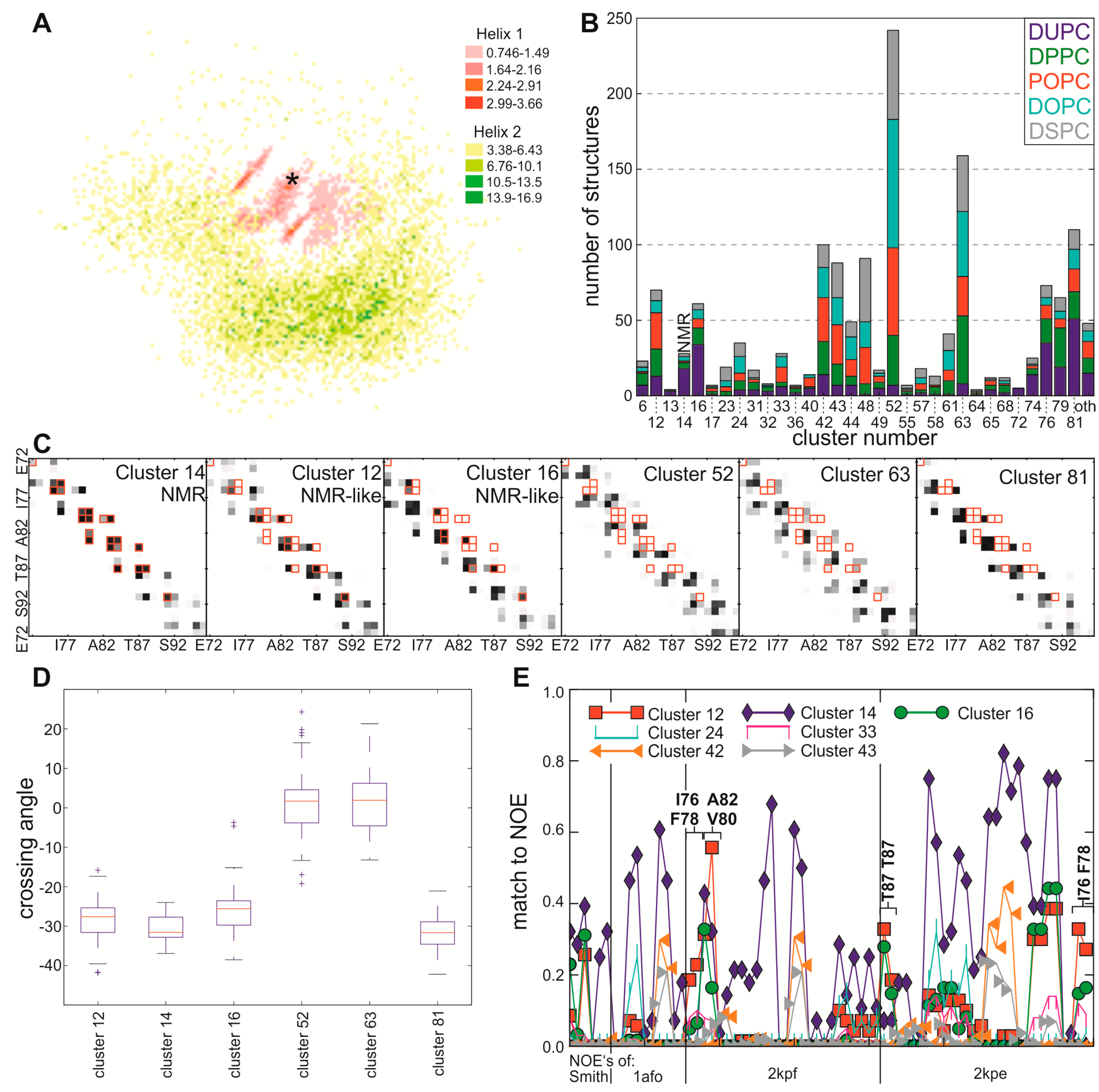
2.2. Interface Frequencies Are Depend on Different Fatty Acid Chain Environments
| Lipid | a | b | c | d | Lim |
|---|---|---|---|---|---|
| DUPC-DUPC | 0.51 | 0.10 | 0.42 | 0.02 | 0.93 |
| DUPC-DPPC | 0.37 | 0.07 | 0.18 | 0.01 | 0.56 |
| DUPC-POPC | 0.24 | 0.09 | 0.12 | 0.01 | 0.36 |
| DUPC-DOPC | 0.18 | 0.11 | 0.07 | 0.02 | 0.25 |
| DUPC-DSCP | 0.18 | 0.10 | 0.09 | 0.02 | 0.27 |
| DPPC-DPPC | 0.55 | 0.07 | 0.38 | 0.01 | 0.94 |
| DPPC-POPC | 0.58 | 0.07 | 0.24 | 0.01 | 0.82 |
| DPPC-DOPC | 0.58 | 0.08 | 0.22 | 0.01 | 0.80 |
| DPPC-DSPC | 0.57 | 0.07 | 0.22 | 0.01 | 0.79 |
| POPC-POPC | 0.59 | 0.09 | 0.36 | 0.02 | 0.95 |
| POPC-DOPC | 0.73 | 0.08 | 0.20 | 0.01 | 0.93 |
| POCP-DSPC | 0.67 | 0.08 | 0.25 | 0.01 | 0.92 |
| DOPC-DOPC | 0.65 | 0.13 | 0.31 | 0.02 | 0.96 |
| DOPC-DSPC | 0.73 | 0.09 | 0.19 | 0.01 | 0.93 |
| DSPC-DSPC | 0.59 | 0.11 | 0.35 | 0.02 | 0.95 |
2.3. Transitions between Dimer Conformations Are Visited by Long Self-Assembly Simulations

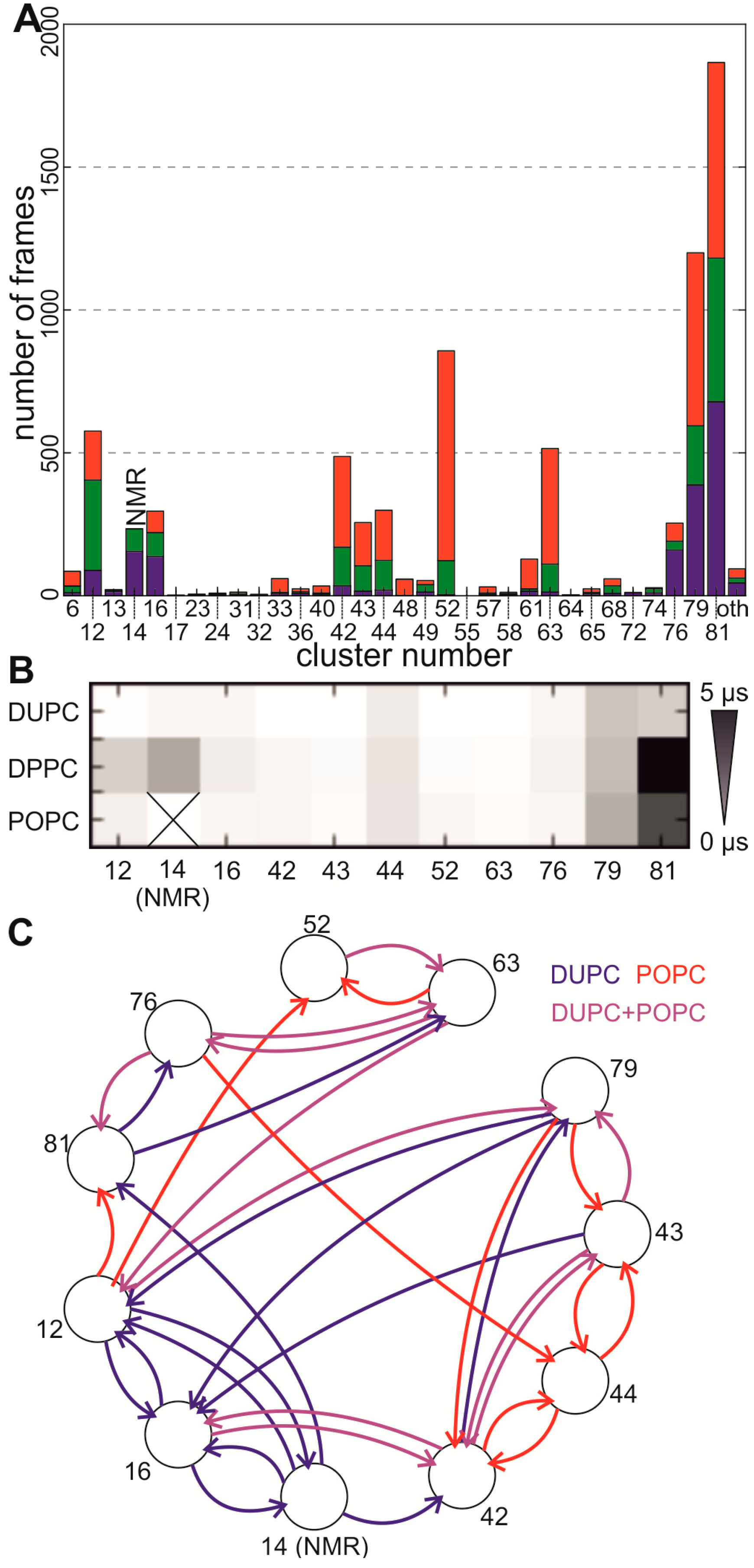
2.4. Stability of the NMR Interface Depends on Helix Length and Helix Conformation
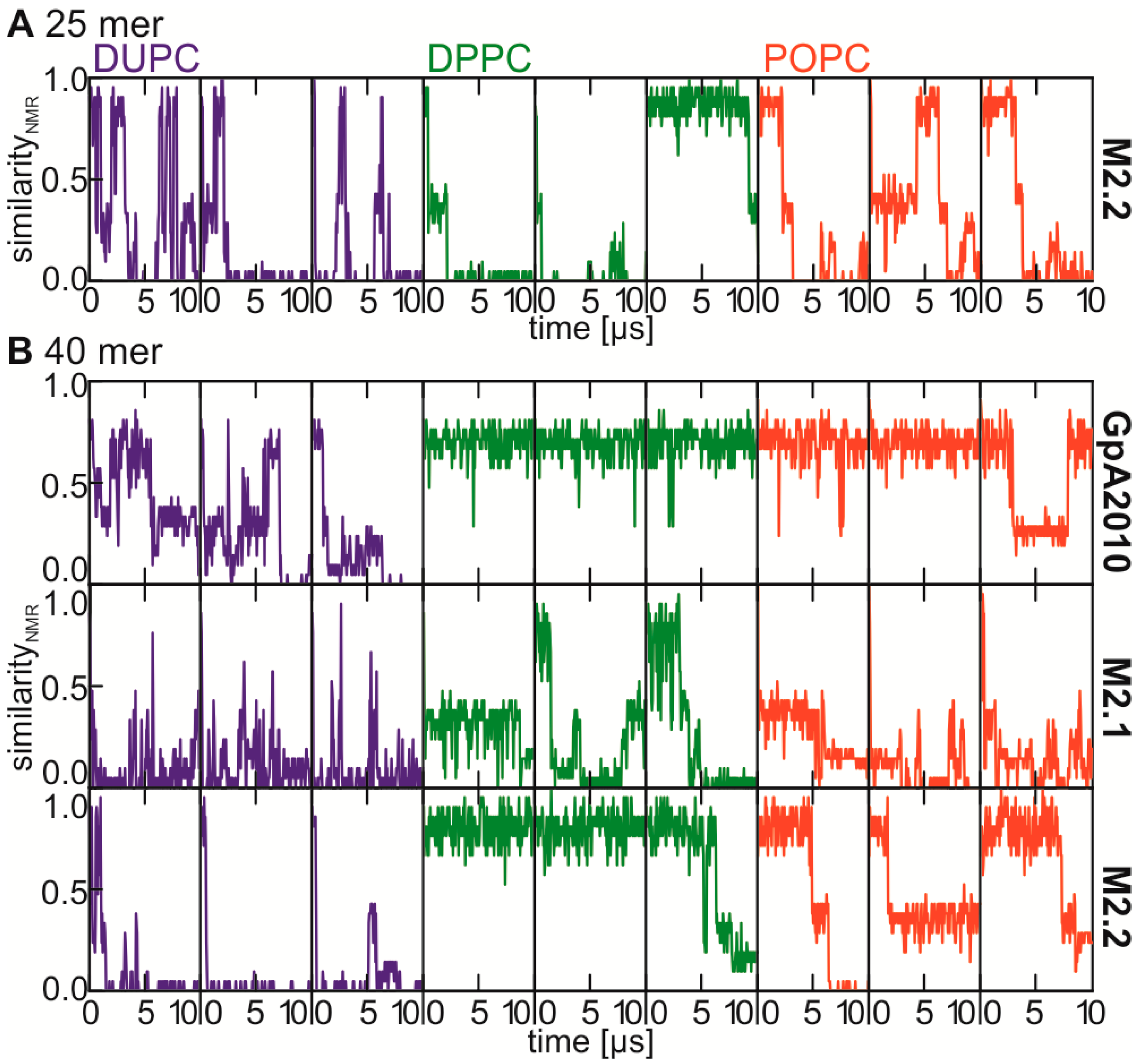
2.5. Correction of the Helix Conformation Leads to Increased NMR Interface Frequency
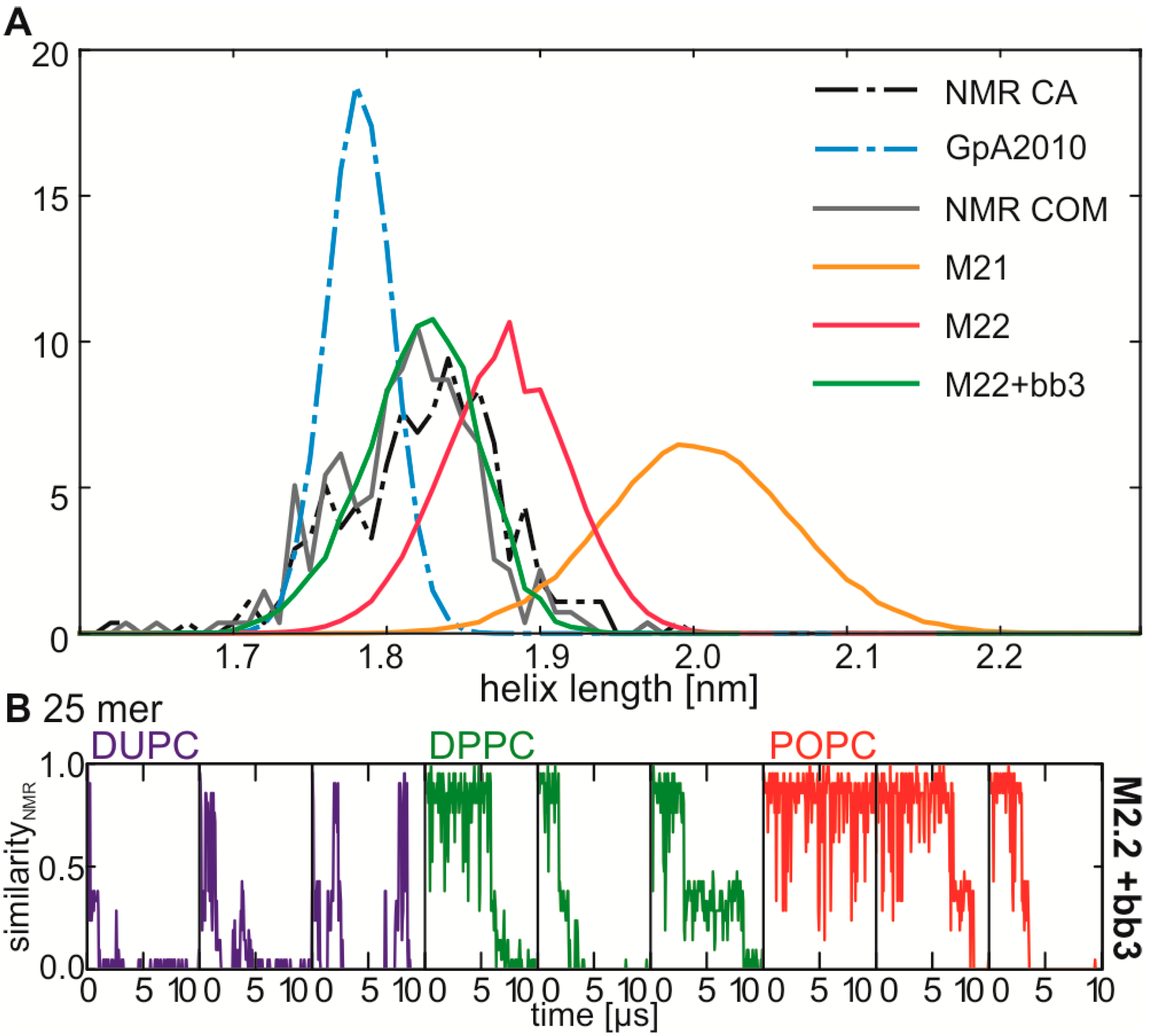
| bb5 | None | 500 | 1000 | 5000 | 10000 | bb3 | |
|---|---|---|---|---|---|---|---|
| bb4 | |||||||
| None | 133.07 | 115.7 | 100.2 | 60.34 | 57.02 | None | |
| 500 | 137.10 | 123.6 | 114.4 | 72.56 | 62.92 | ||
| 1000 | 121.99 | 114.0 | 104.0 | 70.09 | 62.57 | ||
| 5000 | 83.59 | 84.61 | 82.69 | 73.35 | 71.52 | ||
| 10,000 | 75.11 | 77.43 | 76.97 | 76.83 | 79.96 | ||
| None | 71.72 | 61.61 | 51.89 | 40.38 | 44.57 | 500 | |
| 500 | 92.88 | 83.73 | 70.28 | 51.18 | 50.20 | ||
| 1000 | 81.57 | 78.54 | 69.55 | 51.53 | 52.24 | ||
| 5000 | 58.17 | 60.20 | 60.39 | 61.08 | 63.49 | ||
| 10,000 | 64.70 | 66.86 | 67.18 | 73.00 | 76.51 | ||
| None | 54.63 | 56.27 | 45.74 | 39.17 | 47.65 | 1000 | |
| 500 | 87.97 | 79.35 | 69.90 | 50.15 | 50.48 | ||
| 1000 | 75.77 | 72.60 | 65.51 | 51.49 | 52.83 | ||
| 5000 | 56.98 | 60.49 | 59.77 | 60.89 | 63.18 | ||
| 10,000 | 64.17 | 67.19 | 66.17 | 72.05 | 76.92 | ||
| None | 27.38 | 32.15 | 34.26 | 46.53 | 57.14 | 5000 | |
| 500 | 48.72 | 50.96 | 49.47 | 50.73 | 56.61 | ||
| 1000 | 49.67 | 49.56 | 49.15 | 51.87 | 59.93 | ||
| 5000 | 53.08 | 56.04 | 55.63 | 63.25 | 71.04 | ||
| 10,000 | 67.34 | 68.75 | 68.40 | 75.04 | 80.79 | ||
| None | 32.34 | 33.56 | 37.52 | 55.03 | 64.71 | 10,000 | |
| 500 | 38.60 | 41.83 | 45.41 | 53.83 | 65.94 | ||
| 1000 | 40.73 | 44.60 | 46.12 | 57.61 | 66.66 | ||
| 5000 | 59.00 | 60.18 | 59.61 | 69.77 | 75.88 | ||
| 10,000 | 71.35 | 71.95 | 73.49 | 79.16 | 84.68 | ||
| Place | DUPC | DPPC | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| m22 | m22 + bb3 | GpA2010 | m22 | m22 + bb3 | GpA2010 | |||||||
| nr | freq | nr | freq | nr | freq | nr | freq | nr | freq | nr | freq | |
| 1 | 81 | 17.0% | 81 | 13.3% | 2 | 31.3% | 63 | 15.0% | 52 | 19.7% | 2 | 15.7% |
| 2 | 76 | 11.7% | NMR | 10.3% | 0 | 19.3% | 52 | 11.0% | 63 | 12.7% | 0 | 11.7% |
| 3 | 16 | 11.3% | 76 | 9.3% | 12 | 12.3% | 79 | 8.7% | 42 | 8.0% | 7 | 10.0% |
| 4 | 79 | 6.3% | 33 | 6.7% | 16 | 4.7% | 42 | 7.3% | 76/79 | 5.0% | 6 | 6.7% |
| 5 | NMR | 6.0% | 16/79 | 6.0% | 6/9 | 3.0% | 81/12 | 6.0% | 12 | 4.3% | 12 | 6.0% |
3. Discussion
4. Methods
4.1. Simulation Details
4.2. System Composition
4.3. Details for the Short Self-Assembly Simulations
4.4. Description of the Similarity to the NMR Structure and Clustering of Results
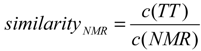
4.5. Autocorrelation, Correlation and Fitting
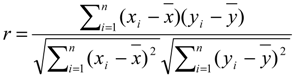
4.6. Construction of the Transition Net
4.7. Backmapping and Comparison to NOE’s
5. Conclusions
Supplementary Files
Supplementary File 1Acknowledgments
Author Contributions
Conflicts of Interest
References
- Ohvo-Rekilä, H.; Ramstedt, B.; Leppimäki, P.; Slotte, J.P. Cholesterol interactions with phospholipids in membranes. Prog. Lipid Res. 2002, 41, 66–97. [Google Scholar] [CrossRef]
- Ikonen, E. Cellular cholesterol trafficking and compartmentalization. Nat. Rev. Mol. Cell Biol. 2008, 9, 125–138. [Google Scholar] [CrossRef]
- Van Meer, G.; de Kroon, A.I.P.M. Lipid map of the mammalian cell. J. Cell Sci. 2010, 124, 5–8. [Google Scholar] [CrossRef]
- Risselada, H.J.; Marrink, S.J. The molecular face of lipid rafts in model membranes. Proc. Natl. Acad. Sci. USA 2008, 105, 17367–17372. [Google Scholar] [CrossRef]
- Davis, R.S.; Sunil Kumar, P.B.; Sperotto, M.M.; Laradji, M. Predictions of phase separation in three-component lipid membranes by the MARTINI force field. J. Phys. Chem. B 2013, 117, 4072–4080. [Google Scholar]
- Domański, J.; Marrink, S.J.; Schäfer, L.V. Transmembrane helices can induce domain formation in crowded model membranes. Biochim. Biophys. Acta 2012, 1818, 984–994. [Google Scholar] [CrossRef]
- Lee, A. Membrane structure. Curr. Biol. 2001, 11, R811–R814. [Google Scholar] [CrossRef]
- Yeagle, P.L. Lipid regulation of cell membrane structure and function. FASEB J. 1989, 3, 1833–1842. [Google Scholar]
- Engelman, D.M. Membranes are more mosaic than fluid. Nature 2005, 438, 578–580. [Google Scholar] [CrossRef]
- Lee, A.G. Lipid-protein interactions in biological membranes: A structural perspective. Biochim. Biophys. Acta 2003, 1612, 1–40. [Google Scholar] [CrossRef]
- Schäfer, L.V.; de Jong, D.H.; Holt, A.; Rzepiela, A.J.; de Vries, A.H.; Poolman, B.; Killian, J.A.; Marrink, S.J. Lipid packing drives the segregation of transmembrane helices into disordered lipid domains in model membranes. Proc. Natl. Acad. Sci. USA 2011, 108, 1343–1348. [Google Scholar] [CrossRef]
- De Jong, D.H.; Lopez, C.A.; Marrink, S.J. Molecular view on protein sorting into liquid-ordered membrane domains mediated by gangliosides and lipid anchors. Faraday Discuss. 2013, 161 discussion 419-459, 347–363. [Google Scholar] [CrossRef]
- Phillips, R.; Ursell, T.; Wiggins, P.; Sens, P. Emerging roles for lipids in shaping membrane-protein function. Nature 2009, 459, 379–385. [Google Scholar] [CrossRef]
- Holt, A.; Killian, J.A. Orientation and dynamics of transmembrane peptides: The power of simple models. Eur. Biophys. J. 2010, 39, 609–621. [Google Scholar] [CrossRef]
- Strandberg, E.; Esteban-Martín, S.; Ulrich, A.S.; Salgado, J. Hydrophobic mismatch of mobile transmembrane helices: Merging theory and experiments. Biochim. Biophys. Acta 2012, 1818, 1242–1249. [Google Scholar] [CrossRef]
- Parton, D.L.; Klingelhoefer, J.W.; Sansom, M.S.P. Aggregation of model membrane proteins, modulated by hydrophobic mismatch, membrane curvature, and protein class. Biophys. J. 2011, 101, 691–699. [Google Scholar] [CrossRef]
- Leidl, K.; Liebisch, G.; Richter, D.; Schmitz, G. Mass spectrometric analysis of lipid species of human circulating blood cells. Biochim. Biophys. Acta 2008, 1781, 655–664. [Google Scholar]
- De Oliveira, S.; Saldanha, C. An overview about erythrocyte membrane. Clin. Hemorheol. Microcirc. 2010, 44, 63–74. [Google Scholar]
- Reid, M.E. MNS blood group system: A review. Immunohematology 2009, 25, 95–101. [Google Scholar]
- Mankelow, T.; Satchwell, T.; Burton, N. Refined views of multi-protein complexes in the erythrocyte membrane. Blood Cells Mol. Dis. 2012, 49, 1–10. [Google Scholar] [CrossRef]
- MacKenzie, K.R.; Prestegard, J.H.; Engelman, D.M. A transmembrane helix dimer: Structure and implications. Science 1997, 276, 131–133. [Google Scholar] [CrossRef]
- Mineev, K.S.; Bocharov, E.V.; Volynsky, P.E.; Goncharuk, M.V.; Tkach, E.N.; Ermolyuk, Y.S.; Schulga, A.A.; Chupin, V.V.; Maslennikov, I.V.; Efremov, R.G.; et al. Dimeric structure of the transmembrane domain of glycophorin a in lipidic and detergent environments. Acta Naturae 2011, 3, 90–98. [Google Scholar]
- Russ, W.P.; Engelman, D.M. The GxxxG motif: A framework for transmembrane helix-helix association. J. Mol. Biol. 2000, 296, 911–919. [Google Scholar] [CrossRef]
- Bond, P.J.; Sansom, M.S.P. Insertion and assembly of membrane proteins via simulation. J. Am. Chem. Soc. 2006, 128, 2697–2704. [Google Scholar] [CrossRef]
- Psachoulia, E.; Fowler, P.W.; Bond, P.J.; Sansom, M.S.P. Helix-helix interactions in membrane proteins: Coarse-grained simulations of glycophorin a helix dimerization. Biochemistry 2008, 47, 10503–10512. [Google Scholar] [CrossRef]
- Sengupta, D.; Marrink, S.J. Lipid-mediated interactions tune the association of glycophorin A helix and its disruptive mutants in membranes. Phys. Chem. Chem. Phys. 2010, 12, 12987–12996. [Google Scholar] [CrossRef]
- Marrink, S.J.; de Vries, A.H.; Mark, A.E. Coarse grained model for semiquantitative lipid simulations. J. Phys. Chem. B 2004, 108, 750–760. [Google Scholar]
- Monticelli, L.; Kandasamy, S.K.; Periole, X.; Larson, R.G.; Tieleman, D.P.; Marrink, S.-J. The MARTINI coarse-grained force field: Extension to proteins. J. Chem. Theory Comput. 2008, 4, 819–834. [Google Scholar]
- De Jong, D.H.; Singh, G.; Bennett, W.F.D.; Arnarez, C.; Wassenaar, T.A.; Schäfer, L.V.; Periole, X.; Tieleman, D.P.; Marrink, S.J. Improved parameters for the martini coarse-grained protein force field. J. Chem. Theory Comput. 2013, 9, 687–697. [Google Scholar]
- Long, F.; Du, L.-C.; Mei, D.-C. Asymmetric effects on the associated relaxation time and the correlation function of a bistable system with correlated noises. Phys. Scr. 2009, 79, 45007. [Google Scholar] [CrossRef]
- Gaspard, P. The correlation time of mesoscopic chemical clocks. J. Chem. Phys. 2002, 117, 8905. [Google Scholar] [CrossRef]
- Periole, X.; Cavalli, M.; Marrink, S.-J.; Ceruso, M.A. Combining an elastic network with a coarse-grained molecular force field: Structure, dynamics, and intermolecular recognition. J. Chem. Theory Comput. 2009, 5, 2531–2543. [Google Scholar]
- Smith, S.O.; Eilers, M.; Song, D.; Crocker, E.; Ying, W.; Groesbeek, M.; Metz, G.; Ziliox, M.; Aimoto, S. Implications of threonine hydrogen bonding in the glycophorin A transmembrane helix dimer. Biophys. J. 2002, 82, 2476–2486. [Google Scholar] [CrossRef]
- Marrink, S.J.; Tieleman, D.P. Perspective on the Martini model. Chem. Soc. Rev. 2013, 42, 6801–6822. [Google Scholar] [CrossRef]
- Anbazhagan, V.; Schneider, D. The membrane environment modulates self-association of the human GpA TM domain—Implications for membrane protein folding and transmembrane signaling. Biochim. Biophys. Acta 2010, 1798, 1899–1907. [Google Scholar] [CrossRef]
- Dony, N.; Crowet, J.M.; Joris, B.; Brasseur, R.; Lins, L. SAHBNET, an accessible surface-based elastic network: An application to membrane protein. Int. J. Mol. Sci. 2013, 14, 11510–11526. [Google Scholar] [CrossRef]
- Globisch, C.; Krishnamani, V.; Deserno, M.; Peter, C. Optimization of an elastic network augmented coarse grained model to study CCMV capsid deformation. PLoS One 2013, 8, e60582. [Google Scholar]
- North, B.; Cristian, L.; Fu Stowell, X.; Lear, J.D.; Saven, J.G.; Degrado, W.F. Characterization of a membrane protein folding motif, the Ser zipper, using designed peptides. J. Mol. Biol. 2006, 359, 930–939. [Google Scholar] [CrossRef]
- Cuthbertson, J.M.; Bond, P.J.; Sansom, M.S.P. Transmembrane helix-helix interactions: Comparative simulations of the glycophorin a dimer. Biochemistry 2006, 45, 14298–14310. [Google Scholar] [CrossRef]
- Polyansky, A.A.; Volynsky, P.E.; Efremov, R.G. Multistate organization of transmembrane helical protein dimers governed by the host membrane. J. Am. Chem. Soc. 2012, 134, 14390–14400. [Google Scholar] [CrossRef]
- Doura, A.K.; Kobus, F.J.; Dubrovsky, L.; Hibbard, E.; Fleming, K.G. Sequence context modulates the stability of a GxxxG-mediated transmembrane helix-helix dimer. J Mol Biol. 2004, 341, 991–998. [Google Scholar] [CrossRef]
- Manni, S.; Mineev, K.S.; Usmanova, D.; Lyukmanova, E.N.; Shulepko, M.A.; Kirpichnikov, M.P.; Winter, J.; Matkovic, M.; Deupi, X.; Arseniev, A.S.; et al. Structural and Functional Characterization of Alternative Transmembrane Domain Conformations in VEGF Receptor 2 Activation. Structure 2014, 22, 1077–1089. [Google Scholar] [CrossRef]
- Bocharov, E.V.; Mayzel, M.L.; Volynsky, P.E.; Goncharuk, M.V.; Ermolyuk, Y.S.; Schulga, A.A.; Artemenko, E.O.; Efremov, R.G.; Arseniev, A.S. Spatial structure and pH-dependent conformational diversity of dimeric transmembrane domain of the receptor tyrosine kinase EphA1. J. Biol. Chem. 2008, 283, 29385–29395. [Google Scholar] [CrossRef]
- Bocharov, E.V.; Mayzel, M.L.; Volynsky, P.E.; Mineev, K.S.; Tkach, E.N.; Ermolyuk, Y.S.; Schulga, A.A.; Efremov, R.G.; Arseniev, A.S. Left-handed dimer of EphA2 transmembrane domain: Helix packing diversity among receptor tyrosine kinases. Biophys. J. 2010, 98, 881–889. [Google Scholar] [CrossRef]
- Tang, T.C.; Hu, Y.; Kienlen-Campard, P.; El Haylani, L.; Decock, M.; van Hees, J.; Fu, Z.; Octave, J.N.; Constantinescu, S.N.; Smith, S.O. Conformational changes induced by the A21G Flemish mutation in the amyloid precursor protein lead to increased Aβ production. Structure 2014, 22, 387–396. [Google Scholar] [CrossRef]
- Janosi, L.; Prakash, A.; Doxastakis, M. Lipid-modulated sequence-specific association of glycophorin A in membranes. Biophys. J. 2010, 99, 284–292. [Google Scholar] [CrossRef]
- Petrache, H.I.; Grossfield, A.; MacKenzie, K.R.; Engelman, D.M.; Woolf, T.B. Modulation of glycophorin A transmembrane helix interactions by lipid bilayers: Molecular dynamics calculations. J. Mol. Biol. 2000, 302, 727–746. [Google Scholar] [CrossRef]
- Orzáez, M.; Pérez-Payá, E.; Mingarro, I. Influence of the C-terminus of the glycophorin A transmembrane fragment on the dimerization process. Protein Sci. 2000, 9, 1246–1253. [Google Scholar] [CrossRef]
- Lemmon, M.A.; Flanagan, J.M.; Hunt, J.F.; Adair, B.D.; Bormann, B.J.; Dempsey, C.E.; Engelman, D.M. Glycophorin A dimerization is driven by specific interactions between transmembrane alpha-helices. J. Biol. Chem. 1992, 267, 7683–7689. [Google Scholar]
- Gerber, D.; Shai, Y. In vivo detection of hetero-association of glycophorin-A and its mutants within the membrane. J. Biol. Chem. 2001, 276, 31229–31232. [Google Scholar] [CrossRef]
- Brosig, B.; Langosch, D. The dimerization motif of the glycophorin A transmembrane segment in membranes: Importance of glycine residues. Protein Sci. 1998, 7, 1052–1056. [Google Scholar] [CrossRef]
- Lindberg, L.; Santos, A.X.; Riezman, H.; Olsson, L.; Bettiga, M. Lipidomic profiling of Saccharomyces cerevisiae and Zygosaccharomyces bailii reveals critical changes in lipid composition in response to acetic acid stress. PLoS One 2013, 8, e73936. [Google Scholar]
- Ghorbal, S.K.B.; Chatti, A.; Sethom, M.M.; Maalej, L.; Mihoub, M.; Kefacha, S.; Feki, M.; Landoulsi, A.; Hassen, A. Changes in membrane fatty acid composition of Pseudomonas aeruginosa in response to UV-C radiations. Curr. Microbiol. 2013, 67, 112–117. [Google Scholar] [CrossRef]
- Oberg, T.S.; Ward, R.E.; Steele, J.L.; Broadbent, J.R. Genetic and physiological responses of Bifidobacterium animalis subsp. lactis to hydrogen peroxide stress. J. Bacteriol. 2013, 195, 3743–3751. [Google Scholar] [CrossRef]
- Shimura, Y.; Shiraiwa, Y.; Suzuki, I. Characterization of the subdomains in the N-terminal region of histidine kinase Hik33 in the cyanobacterium Synechocystis sp. PCC 6803. Plant Cell Physiol. 2012, 53, 1255–1266. [Google Scholar]
- Mikami, K.; Kanesaki, Y.; Suzuki, I.; Murata, N. The histidine kinase Hik33 perceives osmotic stress and cold stress in Synechocystis sp PCC 6803. Mol. Microbiol. 2002, 46, 905–915. [Google Scholar] [CrossRef]
- Mironov, K.S.; Sidorov, R.A.; Trofimova, M.S.; Bedbenov, V.S.; Tsydendambaev, V.D.; Allakhverdiev, S.I.; Los, D.A. Light-dependent cold-induced fatty acid unsaturation, changes in membrane fluidity, and alterations in gene expression in Synechocystis. Biochim. Biophys. Acta 2012, 1817, 1352–1359. [Google Scholar] [CrossRef]
- Cybulski, L.E.; Martín, M.; Mansilla, M.C.; Fernández, A.; de Mendoza, D. Membrane thickness cue for cold sensing in a bacterium. Curr. Biol. 2010, 20, 1539–1544. [Google Scholar] [CrossRef]
- Andersen, O.S.; Koeppe, R.E. Bilayer thickness and membrane protein function: An energetic perspective. Annu. Rev. Biophys. Biomol. Struct. 2007, 36, 107–130. [Google Scholar] [CrossRef]
- Carruthers, A.; Melchior, D.L. Human erythrocyte hexose transporter activity is governed by bilayer lipid composition in reconstituted vesicles. Biochemistry 1984, 23, 6901–6911. [Google Scholar] [CrossRef]
- Hess, B.; Kutzner, C.; van der Spoel, D.; Lindahl, E. GROMACS 4: Algorithms for Highly Efficient, Load-Balanced, and Scalable Molecular Simulation. J. Chem. Theory Comput. 2008, 4, 435–447. [Google Scholar]
- Berendsen, H.J.C.; Postma, J.P.M.; van Gunsteren, W.F.; DiNola, A.; Haak, J.R. Molecular dynamics with coupling to an external bath. J. Chem. Phys. 1984, 81, 3684. [Google Scholar] [CrossRef]
- Marrink, S.J.; Risselada, H.J.; Yefimov, S.; Tieleman, D.P.; de Vries, A.H. The MARTINI force field: Coarse grained model for biomolecular simulations. J. Phys. Chem. B 2007, 111, 7812–7824. [Google Scholar]
- Krieger, E.; Vriend, G. Models@Home: Distributed computing in bioinformatics using a screensaver based approach. Bioinformatics 2002, 18, 315–318. [Google Scholar] [CrossRef]
- Jones, E.; Oliphant, T.; Peterson, P. SciPy: Open Source Scientific Tools for Python. 2001. Available online: http://www.scipy.org/ (access on 22 May 2014).
- Dice, L.R. Measures of the Amount of Ecologic Association between Species. Ecology 1945, 26, 297–302. [Google Scholar] [CrossRef]
- Wassenaar, T.A.; Pluhackova, K.; Böckmann, R.A.; Marrink, S.J.; Tieleman, D.P. Going Backward: A Flexible Geometric Approach to Reverse Transformation from Coarse Grained to Atomistic Models. J. Chem. Theory Comput. 2014, 10, 676–690. [Google Scholar]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Flinner, N.; Mirus, O.; Schleiff, E. The Influence of Fatty Acids on the GpA Dimer Interface by Coarse-Grained Molecular Dynamics Simulation. Int. J. Mol. Sci. 2014, 15, 14247-14268. https://doi.org/10.3390/ijms150814247
Flinner N, Mirus O, Schleiff E. The Influence of Fatty Acids on the GpA Dimer Interface by Coarse-Grained Molecular Dynamics Simulation. International Journal of Molecular Sciences. 2014; 15(8):14247-14268. https://doi.org/10.3390/ijms150814247
Chicago/Turabian StyleFlinner, Nadine, Oliver Mirus, and Enrico Schleiff. 2014. "The Influence of Fatty Acids on the GpA Dimer Interface by Coarse-Grained Molecular Dynamics Simulation" International Journal of Molecular Sciences 15, no. 8: 14247-14268. https://doi.org/10.3390/ijms150814247
APA StyleFlinner, N., Mirus, O., & Schleiff, E. (2014). The Influence of Fatty Acids on the GpA Dimer Interface by Coarse-Grained Molecular Dynamics Simulation. International Journal of Molecular Sciences, 15(8), 14247-14268. https://doi.org/10.3390/ijms150814247




