Genome-Wide Analysis of the RNA Helicase Gene Family in Gossypium raimondii
Abstract
:1. Introduction
2. Results
2.1. Identification of RNA Helicase Family Genes in G. raimondii
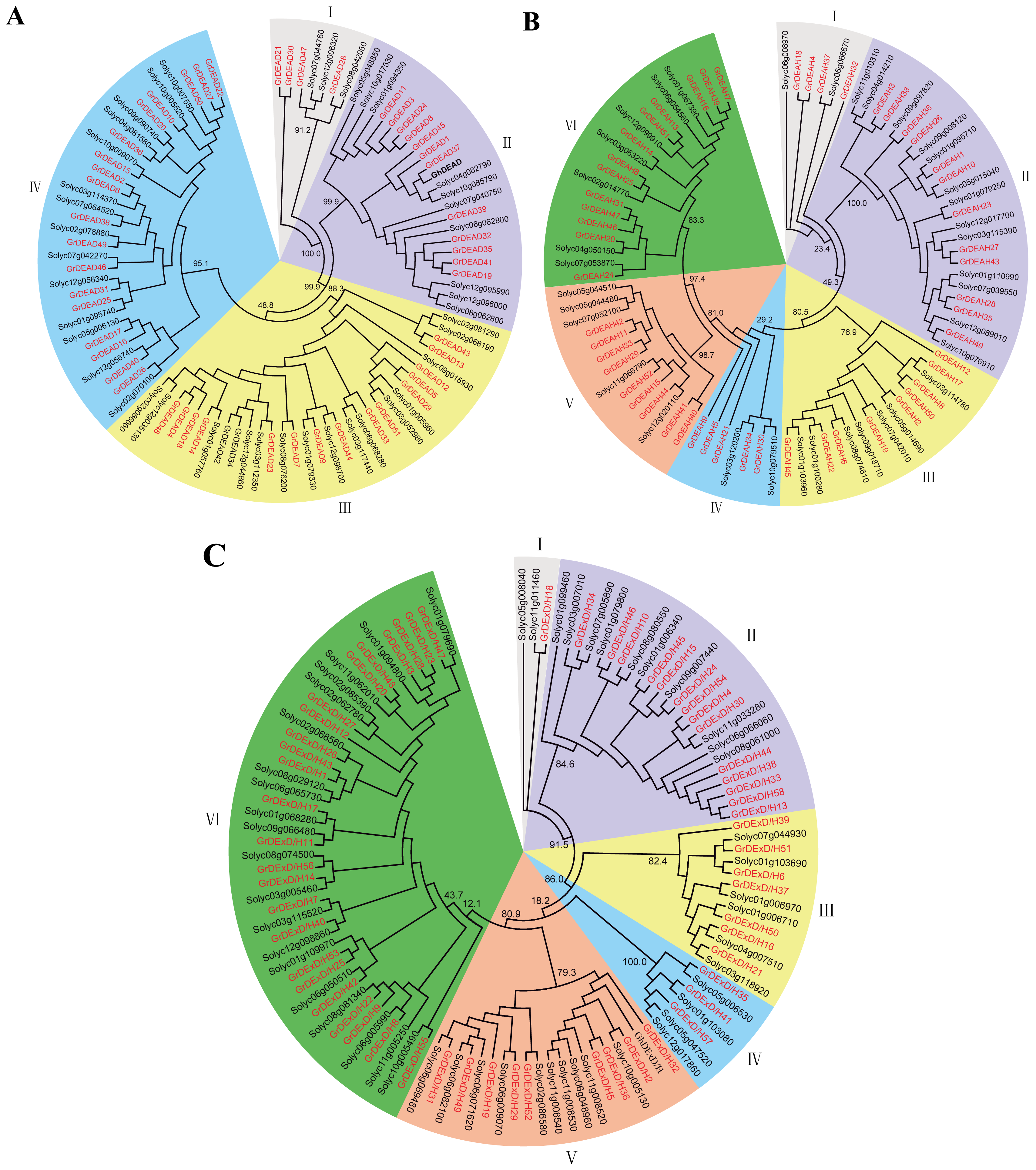
2.2. Phylogenetic Analysis of the RNA Helicase Family Genes in G. raimondii
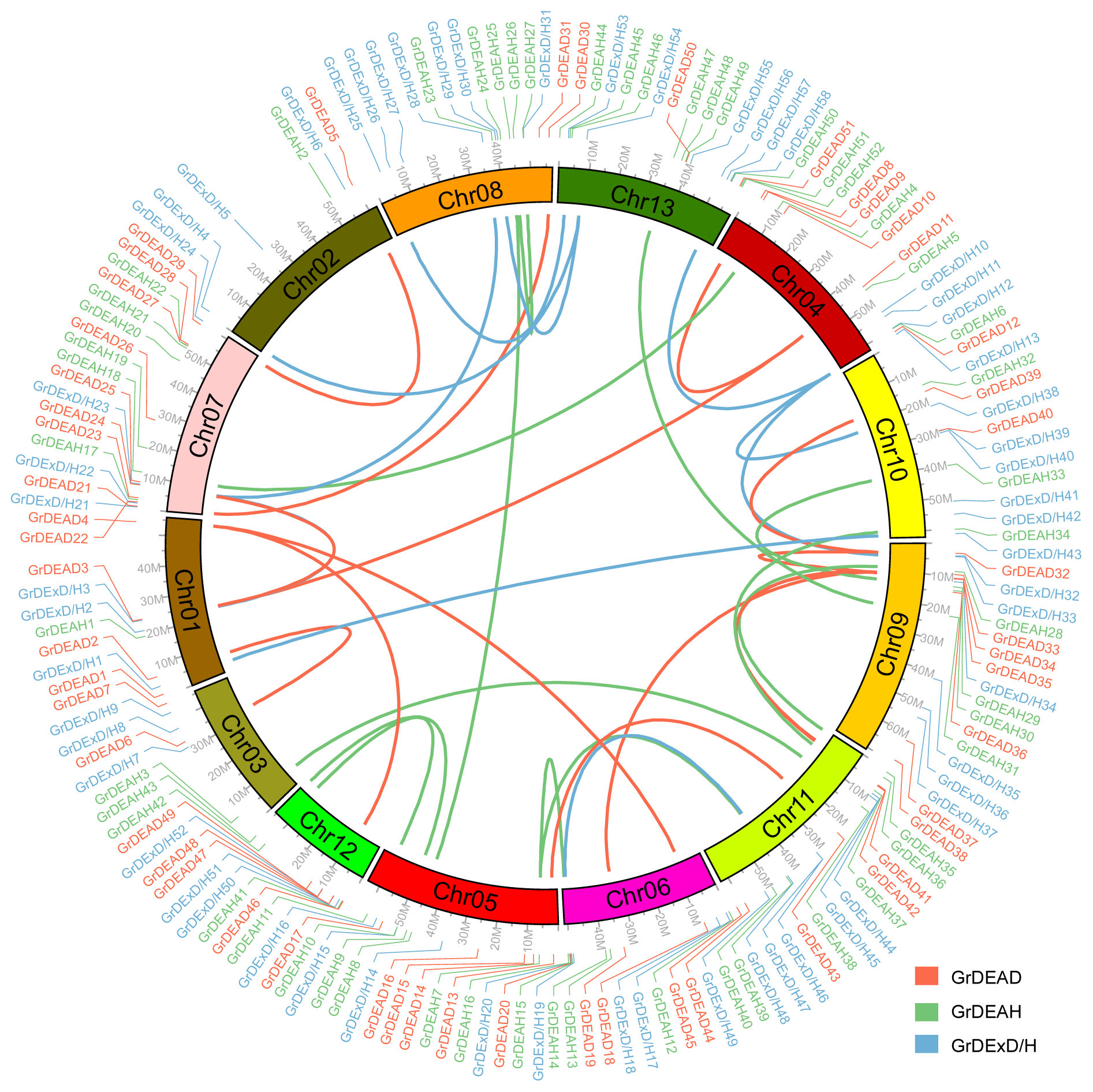
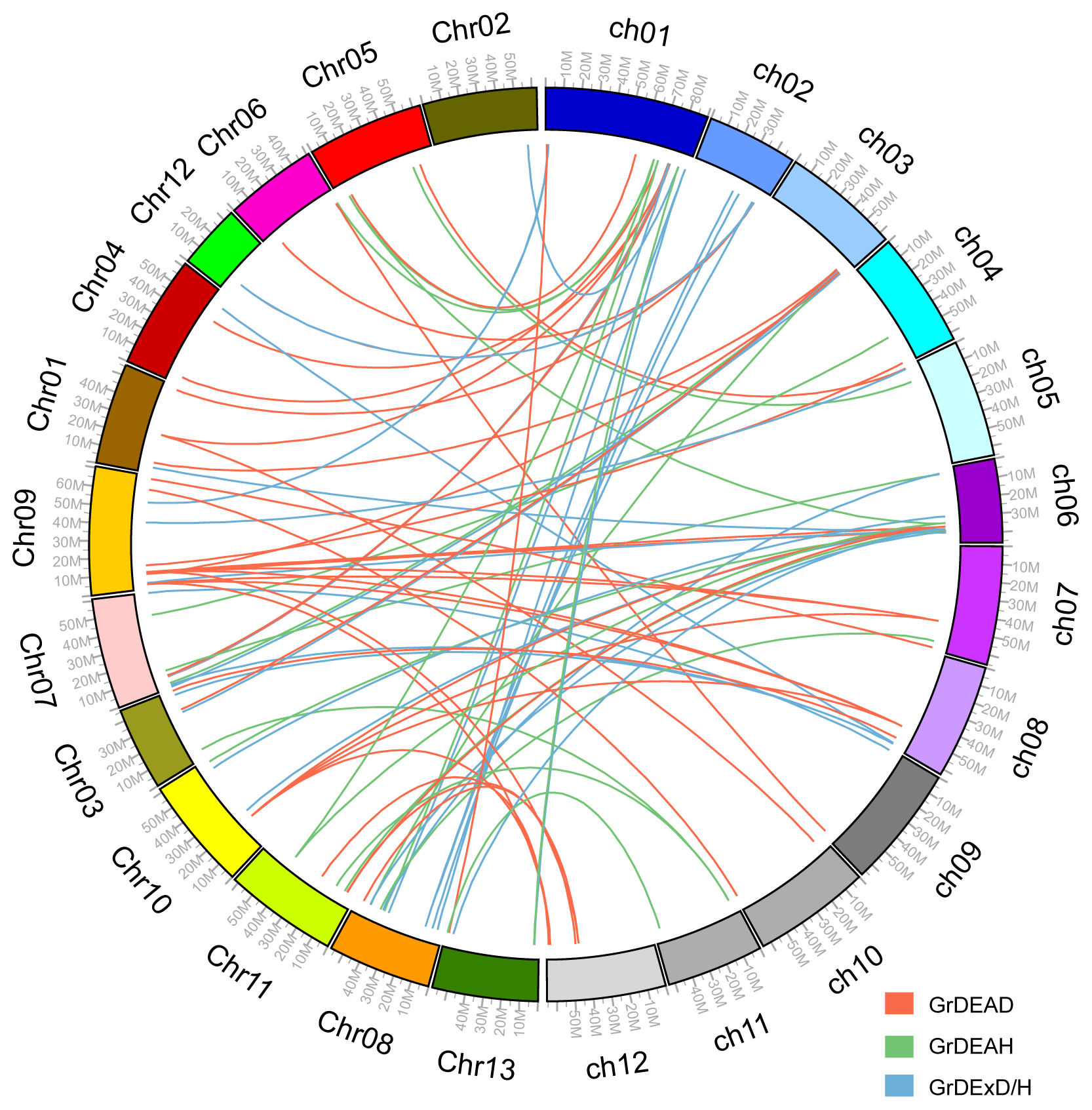
2.3. Chromosomal Position and Gene Duplications
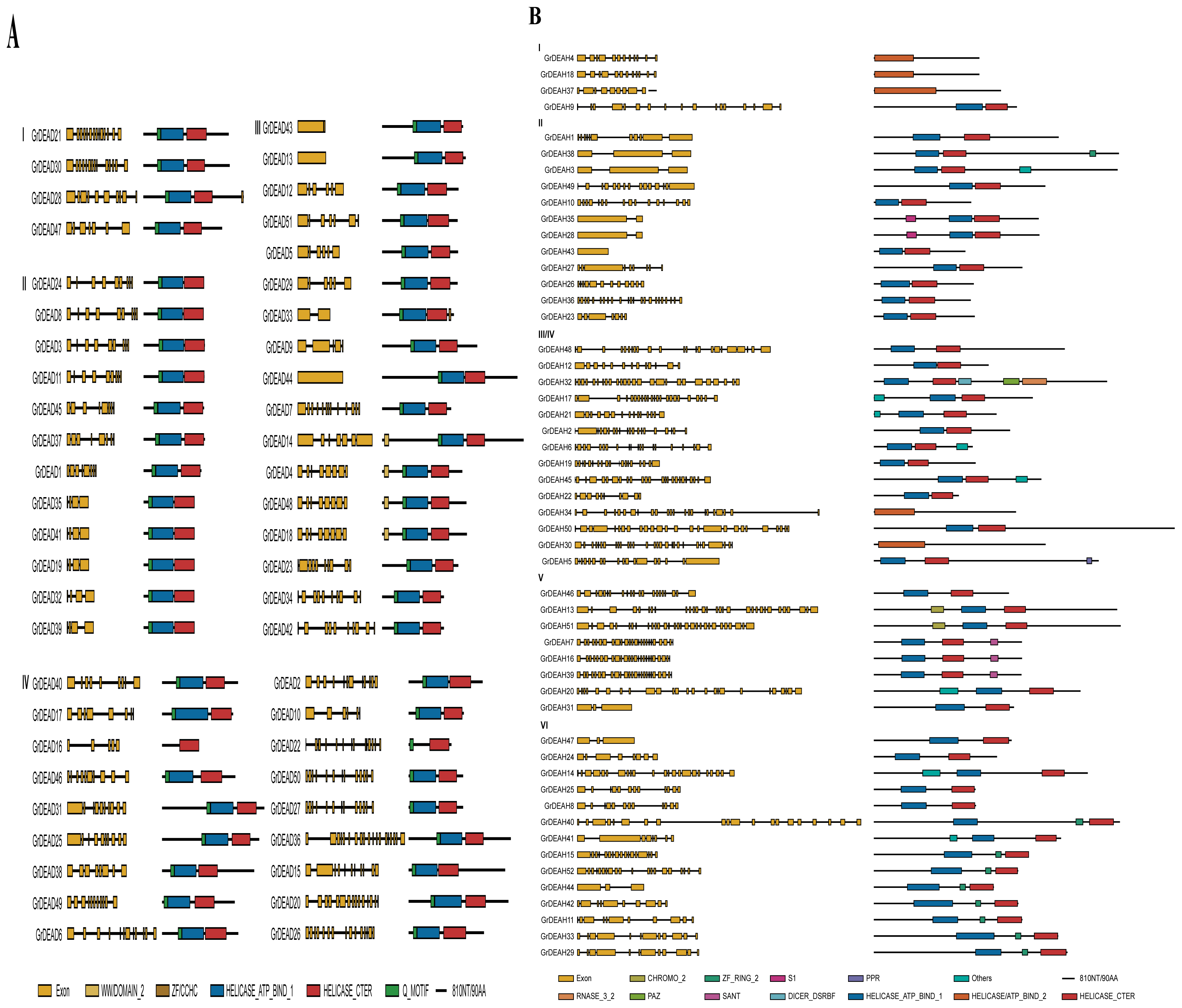
2.4. Structures and Domain Analysis of the Putative Helicase Genes
2.5. Expression of RNA Helicase Family Genes in G. raimondii
3. Discussion
3.1. RNA Helicases in G. raimondii
3.2. Expansion of the RNA Helicase Gene Family in G. raimondii
3.3. Phylogenetic Analysis and Gene Structural Organization
3.4. Expression Analysis Based on Transcriptome Sequencing Data
4. Experimental Section
4.1. Identification of RNA Helicase Genes
4.2. Phylogenetic Analysis
4.3. Chromosome Localization and Gene Duplications
4.4. Gene Structure and Domain Analysis
4.5. Gene Expression Analyses
5. Conclusions
Acknowledgments
Conflicts of Interest
References
- Umate, P.; Tuteja, R.; Tuteja, N. Genome-wide analysis of helicase gene family from rice and Arabidopsis: A comparison with yeast and human. Plant Mol. Biol. 2010, 73, 449–465. [Google Scholar]
- Lohman, T.M.; Bjornson, K.P. Mechanisms of helicase-catalyzed DNA unwinding. Annu. Rev. Biochem. 1996, 65, 169–214. [Google Scholar]
- Owttrim, G.W. RNA helicases and abiotic stress. Nucleic Acids Res. 2006, 34, 3220–3230. [Google Scholar]
- Schmid, S.; Linder, P. D-E-A-D protein family of putative RNA helicases. Mol. Microbiol. 1992, 6, 283–292. [Google Scholar]
- Xu, R.; Zhang, S.; Lu, L.; Cao, H.; Zheng, C. A genome-wide analysis of the RNA helicase gene family in Solanum lycopersicum. Gene 2013, 513, 128–140. [Google Scholar]
- Xu, R.; Zhang, S.; Huang, J.; Zheng, C. Genome-wide comparative in silico analysis of the RNA helicase gene family in Zea mays and Glycine max: A comparison with arabidopsis and Oryza sativa. PLoS One 2013, 8, e78982. [Google Scholar]
- Gong, Z.; Dong, C.H.; Lee, H.; Zhu, J.; Xiong, L.; Gong, D.; Stevenson, B.; Zhu, J.K. A DEAD box RNA helicase is essential for mrna export and important for development and stress responses in Arabidopsis. Plant Cell Online 2005, 17, 256–267. [Google Scholar]
- Gong, Z.; Lee, H.; Xiong, L.; Jagendorf, A.; Stevenson, B.; Zhu, J.K. RNA helicase-like protein as an early regulator of transcription factors for plant chilling and freezing tolerance. Proc. Natl. Acad. Sci. USA 2002, 99, 11507–11512. [Google Scholar]
- Kant, P.; Kant, S.; Gordon, M.; Shaked, R.; Barak, S. STRESS RESPONSE SUPPRESSOR1 and STRESS RESPONSE SUPPRESSOR2 two DEAD-Box RNA helicases that attenuate Arabidopsis responses to multiple abiotic stresses. Plant Physiol. 2007, 145, 814–830. [Google Scholar]
- Guan, Q.; Wu, J.; Zhang, Y.; Jiang, C.; Liu, R.; Chai, C.; Zhu, J. A DEAD vox RNA helicase is critical for pre-mrna splicing cold-responsive gene regulation and cold tolerance in Arabidopsis. Plant Cell Online 2013, 25, 342–356. [Google Scholar]
- Wang, Y.; Duby, G.; Purnelle, B.; Boutry, M. Tobacco VDL gene encodes a plastid DEAD box RNA Helicase and is involved in chloroplast differentiation and plant morphogenesis. Plant Cell Online 2000, 12, 2129–2142. [Google Scholar]
- Gendra, E.; Moreno, A.; Albà, M. Interaction of the plant glycine-rich RNA-binding protein MA16 with a novel nucleolar DEAD box RNA helicase protein from Zea mays. Plant J. 2004, 38, 875–886. [Google Scholar]
- Macovei, A.; Vaid, N.; Tula, S.; Tuteja, N. A new DEAD-box helicase ATP-binding protein (OsABP) from rice is responsive to abiotic stress. Plant Signal. Behav. 2012, 7, 1138–1143. [Google Scholar]
- Liu, H.; Liu, J.; Fan, S.; Song, M.; Han, X.; Liu, F.; Shen, F. Molecular cloning and characterization of a salinity stress-induced gene encoding DEAD-box helicase from the halophyte Apocynum venetum. J. Exp. Bot. 2008, 59, 633–644. [Google Scholar]
- Lange, H.; Sement, F.M.; Gagliardi, D. MTR4 a putative RNA helicase and exosome co-factor is required for proper rRNA biogenesis and development in Arabidopsis thaliana. Plant J. 2011, 68, 51–63. [Google Scholar]
- Kobayashi, K.; Otegui, M.S.; Krishnakumar, S.; Mindrinos, M.; Zambryski, P. INCREASED SIZE EXCLUSION LIMIT2 encodes a putative DEVH Box RNA helicase involved in plasmodesmata function during Arabidopsis embryogenesis. Plant Cell Online 2007, 19, 1885–1897. [Google Scholar]
- Xu, R.R.; Qi, S.D.; Lu, L.T.; Chen, C.T.; Wu, C.A.; Zheng, C.C. A DExD/H box RNA helicase is important for K+ deprivation responses and tolerance in Arabidopsis thaliana. FEBS J. 2011, 278, 2296–2306. [Google Scholar]
- Ohtani, M.; Demura, T.; Sugiyama, M. Arabidopsis ROOT INITIATION DEFECTIVE1 a DEAH-Box RNA helicase involved in pre-mrna splicing is essential for plant development. Plant Cell Online 2013. [Google Scholar] [CrossRef]
- Aubourg, S.; Kreis, M.; Lecharny, A. The DEAD box RNA helicase family in Arabidopsis thaliana. Nucleic Acids Res. 1999, 27, 628–636. [Google Scholar]
- Rathore, K.S. Cotton. In Genetic Modification of Plants; Springer: Berlin/Heidelberg, Germany, 2010; pp. 269–285. [Google Scholar]
- Wendel, J.F.; Brubaker, C.; Alvarez, I.; Cronn, R.; Stewart, J.M. Evolution and natural history of the cotton genus. In Genetics and Genomics of Cotton; Springer: New York, NY, USA, 2009; pp. 3–22. [Google Scholar]
- Hovav, R.; Udall, J.A.; Hovav, E.; Rapp, R.; Flagel, L.; Wendel, J.F. A majority of cotton genes are expressed in single-celled fiber. Planta 2008, 227, 319–329. [Google Scholar]
- Hovav, R.; Udall, J.A.; Chaudhary, B.; Hovav, E.; Flagel, L.; Hu, G.; Wendel, J.F. The evolution of spinnable cotton fiber entailed prolonged development and a novel metabolism. PLoS Genet. 2008, 4, e25. [Google Scholar]
- Yao, D.; Zhang, X.; Zhao, X.; Liu, C.; Wang, C.; Zhang, Z.; Zhang, C.; Wei, Q.; Wang, Q.; Yan, H. Transcriptome analysis reveals salt-stress-regulated biological processes and key pathways in roots of cotton (Gossypium hirsutum L). Genomics 2011, 98, 47–55. [Google Scholar]
- Wang, K.; Wang, Z.; Li, F.; Ye, W.; Wang, J.; Song, G.; Yue, Z.; Cong, L.; Shang, H.; Zhu, S.; et al. The draft genome of a diploid cotton Gossypium raimondii. Nat. Genet. 2012, 44, 1098–1103. [Google Scholar]
- NCBI. Available online: http://www.ncbi.nlm.nih.gov/ (accessed on 1 October 2013).
- Marchat, L.A.; Orozco, E.; Guillen, N.; Weber, C.; Lopez-Camarillo, C. Putative DEAD and DExH-box RNA helicases families in Entamoeba histolytica. Gene 2008, 424, 1–10. [Google Scholar]
- Rocak, S.; Linder, P. DEAD-box proteins: the driving forces behind RNA metabolism. Nat. Rev. Mol. Cell Biol. 2004, 5, 232–241. [Google Scholar]
- Zou, J.; Chang, M.; Nie, P.; Secombes, C. Origin and evolution of the RIG-I like RNA helicase gene family. BMC Evol. Biol. 2009, 9, 85. [Google Scholar]
- Edgar, R.C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004, 32, 1792–1797. [Google Scholar]
- Holub, E.B. The arms race is ancient history in Arabidopsis the wildflower. Nat. Rev. Genet. 2001, 2, 516–527. [Google Scholar]
- Zhang, C.; Zhang, H.; Zhao, Y.; Jiang, H.; Zhu, S.; Cheng, B.; Xiang, Y. Genome-wide analysis of the CCCH zinc finger gene family in Medicago truncatula. Plant Cell Rep. 2013, 1–13. [Google Scholar]
- Altenhoff, A.M.; Dessimoz, C. Phylogenetic and functional assessment of orthologs inference projects and methods. PLoS Comput. Biol. 2009, 5, e1000262. [Google Scholar]
- Linder, P.; Owttrim, G.W. Plant RNA helicases: Linking aberrant and silencing RNA. Trends Plant Sci. 2009, 14, 344–352. [Google Scholar]
- Asakura, Y.; Galarneau, E.; Watkins, K.P.; Barkan, A.; van Wijk, K.J. Chloroplast RH3 DEAD box RNA helicases in maize and Arabidopsis function in splicing of specific group II introns and affect chloroplast ribosome biogenesis. Plant Physiol. 2012, 159, 961–974. [Google Scholar]
- He, J.; Duan, Y.; Hua, D.; Fan, G.; Wang, L.; Liu, Y.; Chen, Z.; Han, L.; Qu, L.J.; Gong, Z. DEXH Box RNA helicase-mediated mitochondrial reactive oxygen species production in Arabidopsis mediates crosstalk between abscisic acid and auxin signaling. Plant Cell Online 2012, 24, 1815–1833. [Google Scholar]
- Shang, H.; Li, W.; Zou, C.; Yuan, Y. Analyses of the NAC transcription factor gene family in Gossypium raimondii Ulbr: Chromosomal location structure phylogeny and expression patterns. J. Integr. Plant Biol. 2013, 55, 663–676. [Google Scholar]
- Cannon, S.B.; Mitra, A.; Baumgarten, A.; Young, N.D.; May, G. The roles of segmental and tandem gene duplication in the evolution of large gene families in Arabidopsis thaliana. BMC Plant Biol. 2004, 4, 10. [Google Scholar]
- Du, D.; Hao, R.; Cheng, T.; Pan, H.; Yang, W.; Wang, J.; Zhang, Q. Genome-wide analysis of the AP2/ERF gene family in Prunus mume. Plant Mol. Biol. Rep. 2013, 31, 741–750. [Google Scholar]
- Tanner, N.K.; Cordin, O.; Banroques, J.; Doère, M.; Linder, P. The Q motif: A newly identified motif in DEAD Box helicases may regulate ATP binding and hydrolysis. Mol. Cell 2003, 11, 127–138. [Google Scholar]
- Ilsleya, J.L.; Sudolb, M.; Windera, S.J. The WW domain: Linking cell signalling to the membrane cytoskeleton. Cell. Signal. 2002, 14, 183–189. [Google Scholar]
- Klostermeier, D.; Rudolph, M.G. A novel dimerization motif in the C-terminal domain of the Thermus thermophilus DEAD box helicase Hera confers substantial flexibility. Nucleic Acids Res. 2009, 37, 421–430. [Google Scholar]
- Menges, M.; de Jager, S.M.; Gruissem, W.; Murray, J.A. Global analysis of the core cell cycle regulators of Arabidopsis identifies novel genes reveals multiple and highly specific profiles of expression and provides a coherent model for plant cell cycle control. Plant J. 2005, 41, 546–566. [Google Scholar]
- Lucas, W.J.; Ham, B.K.; Kim, J.Y. Plasmodesmata—Bridging the gap between neighboring plant cells. Trends Cell Biol. 2009, 19, 495–503. [Google Scholar]
- CottonGen. Available online: http://www.cottongen.org/ (accessed on 1 October 2013).
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar]
- The Arabidopsis Information Resource. Available online: http://www.arabidopsis.org/ (accessed on 6 October 2013).
- PlantGDB. Available online: http://www.plantgdb.org/ (accessed on 6 October 2013).
- PROSITE Program. Available online: http://www.expasy.ch/prosite/ (accessed on 25 October 2013).
- Compute pI/Mw Tool. ExPASy Server. Available online: http://web.expasy.org/compute_pi/ (accessed on 25 October 2013).
- CELLO v2.5. Available online: http://cello.life.nctu.edu.tw/ (accessed on 25 October 2013).
- Guindon, S.; Lethiec, F.; Duroux, P.; Gascuel, O. PHYML Online—A web server for fast maximum likelihood-based phylogenetic inference. Nucleic Acids Res. 2005, 33, W557–W559. [Google Scholar]
- Tang, H.; Bowers, J.E.; Wang, X.; Ming, R.; Alam, M.; Paterson, A.H. Synteny and collinearity in plant genomes. Science 2008, 320, 486–488. [Google Scholar]
- Wang, Y.; Tang, H.; DeBarry, J.D.; Tan, X.; Li, J.; Wang, X.; Lee, T.-H.; Jin, H.; Marler, B.; Guo, H. MCScanX: A toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 2012, 40, e49–e49. [Google Scholar]
- Krzywinski, M.; Schein, J.; Birol, İ; Connors, J.; Gascoyne, R.; Horsman, D.; Jones, S.J.; Marra, M.A. Circos: An information aesthetic for comparative genomics. Genome Res. 2009, 19, 1639–1645. [Google Scholar]
- Bairoch, A. PROSITE: A dictionary of sites and patterns in proteins. Nucleic Acids Res. 1991, 19, 2241. [Google Scholar]
- Gasteiger, E.; Gattiker, A.; Hoogland, C.; Ivanyi, I.; Appel, R.D.; Bairoch, A. ExPASy: The proteomics server for in-depth protein knowledge and analysis. Nucleic Acids Res. 2003, 31, 3784–3788. [Google Scholar]
- GSDS v2.0. Available online: http://gsds.cbi.pku.edu.cn/ (accessed on 25 October 2013).
- Li, H.; Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar]
- Mortazavi, A.; Williams, B.A.; McCue, K.; Schaeffer, L.; Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat. Methods 2008, 5, 621–628. [Google Scholar]
- Yin, Z.; Wang, J.; Wang, D.; Fan, W.; Wang, S.; Ye, W. The MAPKKK gene family in Gossypium raimondii: Genome-wide identification classification and expression analysis. Int. J. Mol. Sci. 2013, 14, 18740–18757. [Google Scholar]



| Gene name | Gene identifier | Genomic position | Size (AA) | Mw | pI | Subcellular localization |
|---|---|---|---|---|---|---|
| GrDEAD1 | Gorai.001G008400.1 | Chr01: 784950-787262 | 472 | 52.95 | 6.12 | Cytoplasmic |
| GrDEAD2 | Gorai.001G064600.1 | Chr01: 6445636-6451393 | 604 | 68.98 | 8.93 | Nuclear |
| GrDEAD3 | Gorai.001G169800.1 | Chr01: 24289833-24295275 | 499 | 57.13 | 8.66 | Nuclear |
| GrDEAD4 | Gorai.001G263500.1 | Chr01: 53921207-53926399 | 655 | 70.98 | 9.19 | Nuclear |
| GrDEAD5 | Gorai.002G232200.1 | Chr02: 59155342-59159290 | 620 | 67.94 | 8.4 | Nuclear |
| GrDEAD6 | Gorai.003G106600.1 | Chr03: 32786814-32793828 | 622 | 71.26 | 8.96 | Nuclear |
| GrDEAD7 | Gorai.003G186400.1 | Chr03: 45681420-45686317 | 564 | 62.89 | 8.97 | Nuclear |
| GrDEAD8 | Gorai.004G042200.1 | Chr04: 3636522-3642658 | 490 | 56.2 | 8.84 | Nuclear |
| GrDEAD9 | Gorai.004G050000.1 | Chr04: 4582045-4586206 | 776 | 85.09 | 5.72 | Nuclear |
| GrDEAD10 | Gorai.004G089100.1 | Chr04: 11968779-11974240 | 450 | 49.77 | 6.53 | Cytoplasmic |
| GrDEAD11 | Gorai.004G157400.1 | Chr04: 44565986-44571005 | 496 | 56.99 | 8.72 | Nuclear |
| GrDEAD12 | Gorai.004G277500.1 | Chr04: 61046063-61050155 | 624 | 68.33 | 6.93 | Chloroplast |
| GrDEAD13 | Gorai.005G021200.1 | Chr05: 1744957-1747872 | 683 | 78.2 | 8.94 | Nuclear |
| GrDEAD14 | Gorai.005G070600.1 | Chr05: 7665522-7673295 | 1154 | 126.09 | 10.07 | Nuclear |
| GrDEAD15 | Gorai.005G084500.1 | Chr05: 10403322-10409154 | 788 | 89.02 | 9.82 | Nuclear |
| GrDEAD16 | Gorai.005G118500.1 | Chr05: 24460627-24464773 | 300 | 33.4 | 9.26 | Cytoplasmic |
| GrDEAD17 | Gorai.005G186000.1 | Chr05: 54468800-54474652 | 580 | 63.66 | 7.68 | Chloroplast |
| GrDEAD18 | Gorai.006G031200.1 | Chr06: 8129011-8134006 | 692 | 76.22 | 9.66 | Nuclear |
| GrDEAD19 | Gorai.006G095400.1 | Chr06: 33433098-33436366 | 413 | 46.84 | 5.28 | Cytoplasmic |
| GrDEAD20 | Gorai.006G254000.1 | Chr06: 49690002-49695784 | 814 | 91.89 | 9.16 | Nuclear |
| GrDEAD21 | Gorai.007G009400.1 | Chr07: 733772-738313 | 697 | 75.73 | 8.99 | Nuclear |
| GrDEAD22 | Gorai.007G029000.1 | Chr07: 1983857-1989372 | 349 | 39.22 | 8.92 | Cytoplasmic |
| GrDEAD23 | Gorai.007G047000.1 | Chr07: 3268271-3272693 | 623 | 68.33 | 9.83 | Nuclear |
| GrDEAD24 | Gorai.007G097600.1 | Chr07: 7210939-7217193 | 494 | 56.51 | 8.4 | Nuclear |
| GrDEAD25 | Gorai.007G107800.1 | Chr07: 8134802-8139693 | 792 | 88.91 | 9.46 | Nuclear |
| GrDEAD26 | Gorai.007G226800.1 | Chr07: 27332174-27337564 | 615 | 68.61 | 9.2 | Nuclear |
| GrDEAD27 | Gorai.007G304300.1 | Chr07: 51812182-51818395 | 444 | 49.54 | 8.97 | Cytoplasmic |
| GrDEAD28 | Gorai.007G309800.1 | Chr07: 52334743-52340317 | 820 | 90.34 | 8.33 | Chloroplast |
| GrDEAD29 | Gorai.007G362700.1 | Chr07: 59429333-59434032 | 617 | 67.37 | 8.16 | Nuclear |
| GrDEAD30 | Gorai.008G282800.1 | Chr08: 55902284-55906923 | 705 | 77.12 | 9.54 | Nuclear |
| GrDEAD31 | Gorai.008G245100.1 | Chr08: 52991359-52996142 | 835 | 94.62 | 9.51 | Nuclear |
| GrDEAD32 | Gorai.009G043400.1 | Chr09: 3162403-3166053 | 412 | 46.76 | 5.37 | Cytoplasmic |
| GrDEAD33 | Gorai.009G126600.1 | Chr09: 9548083-9551194 | 587 | 65.9 | 6.7 | Cytoplasmic |
| GrDEAD34 | Gorai.009G140700.1 | Chr09: 10637675-10642934 | 505 | 55.88 | 8.97 | Cytoplasmic |
| GrDEAD35 | Gorai.009G142800.1 | Chr09: 10823077-10826736 | 413 | 47.01 | 5.29 | Cytoplasmic |
| GrDEAD36 | Gorai.009G193900.1 | Chr09: 14896221-14903727 | 833 | 94.11 | 7.12 | Nuclear |
| GrDEAD37 | Gorai.009G410800.1 | Chr09: 61968396-61972555 | 501 | 55.79 | 5.65 | Cytoplasmic |
| GrDEAD38 | Gorai.009G437700.1 | Chr09: 68810677-68815318 | 752 | 84.97 | 8.96 | Nuclear |
| GrDEAD39 | Gorai.010G098500.1 | Chr10: 17146619-17149615 | 413 | 46.77 | 5.37 | Cytoplasmic |
| GrDEAD40 | Gorai.010G134100.1 | Chr10: 29846928-29852855 | 620 | 68.47 | 9.87 | Mitochondrial |
| GrDEAD41 | Gorai.011G063400.1 | Chr11: 5308916-5312265 | 413 | 46.92 | 5.29 | Cytoplasmic |
| GrDEAD42 | Gorai.011G066000.1 | Chr11: 5584414-5590538 | 505 | 55.72 | 9.04 | Mitochondrial |
| GrDEAD43 | Gorai.011G148000.1 | Chr11: 24134100-24136116 | 661 | 76.36 | 8.62 | Nuclear |
| GrDEAD44 | Gorai.011G261800.1 | Chr11: 59292567-59298642 | 1104 | 125.79 | 5.78 | Nuclear |
| GrDEAD45 | Gorai.011G288300.1 | Chr11: 61987801-61991449 | 493 | 55.02 | 6.3 | Cytoplasmic |
| GrDEAD46 | Gorai.012G015300.1 | Chr12: 1700644-1705443 | 598 | 68.17 | 9.48 | Nuclear |
| GrDEAD47 | Gorai.012G039900.1 | Chr12: 4943976-4950340 | 643 | 69.16 | 9.51 | Nuclear |
| GrDEAD48 | Gorai.012G063000.1 | Chr12: 8891463-8896643 | 688 | 75.16 | 9.8 | Nuclear |
| GrDEAD49 | Gorai.012G079600.1 | Chr12: 12903543-12907932 | 593 | 66.23 | 9.11 | Mitochondrial |
| GrDEAD50 | Gorai.013G142600.1 | Chr13: 38926934-38932273 | 444 | 49.59 | 8.97 | Cytoplasmic |
| GrDEAD51 | Gorai.013G247600.1 | Chr13: 56620576-56625972 | 616 | 66.79 | 8.66 | Nuclear |
| GrDEAH1 | Gorai.001G141100.1 | Chr01: 19085241-19093500 | 1328 | 148.92 | 1.97 | Nuclear |
| GrDEAH2 | Gorai.002G190800.1 | Chr02: 51591530-51599433 | 979 | 109.02 | 9.01 | Nuclear |
| GrDEAH3 | Gorai.003G041600.1 | Chr03: 5046943-5054254 | 1750 | 196.93 | 6.75 | Nuclear |
| GrDEAH4 | Gorai.004G081200.1 | Chr04: 9735695-9741428 | 758 | 85.98 | 6.61 | Nuclear |
| GrDEAH5 | Gorai.004G164900.1 | Chr04: 45891531-45901380 | 1615 | 182.54 | 7.34 | Nuclear |
| GrDEAH6 | Gorai.004G265100.1 | Chr04: 60082397-60092032 | 711 | 80.98 | 6.59 | Nuclear |
| GrDEAH7 | Gorai.005G058400.1 | Chr05: 6046005-6052692 | 1063 | 123.42 | 5.87 | Nuclear |
| GrDEAH8 | Gorai.005G157800.1 | Chr05: 45129943-45137098 | 734 | 83.31 | 5.09 | Nuclear |
| GrDEAH9 | Gorai.005G161600.1 | Chr05: 46720561-46734800 | 1028 | 114.74 | 5.99 | Nuclear |
| GrDEAH10 | Gorai.005G170100.1 | Chr05: 49756283-49764261 | 700 | 78.71 | 7.56 | Nuclear |
| GrDEAH11 | Gorai.005G213400.1 | Chr05: 59588117-59596596 | 1067 | 119.04 | 6.37 | Nuclear |
| GrDEAH12 | Gorai.006G000100.1 | Chr06: 50440-58229 | 825 | 93.92 | 8.85 | Cytoplasmic |
| GrDEAH13 | Gorai.006G125500.1 | Chr06: 37710924-37727582 | 1747 | 199.38 | 5.8 | Nuclear |
| GrDEAH14 | Gorai.006G135500.1 | Chr06: 39140704-39151672 | 1536 | 174.79 | 8.3 | Nuclear |
| GrDEAH15 | Gorai.006G248800.1 | Chr06: 49317977-49323365 | 1112 | 126.26 | 8.08 | Nuclear |
| GrDEAH16 | Gorai.006G264900.1 | Chr06: 50379562-50386054 | 1065 | 123.47 | 5.93 | Nuclear |
| GrDEAH17 | Gorai.007G040200.1 | Chr07: 2796320-2806189 | 1142 | 128.14 | 7.07 | PlasmaMembrane |
| GrDEAH18 | Gorai.007G136500.1 | Chr07: 11144557-11150427 | 758 | 86.08 | 7.26 | Nuclear |
| GrDEAH19 | Gorai.007G190100.1 | Chr07: 18534829-18540845 | 732 | 81.93 | 9.15 | Nuclear |
| GrDEAH20 | Gorai.007G273900.1 | Chr07: 46703371-46719556 | 1484 | 168.81 | 5.48 | Nuclear |
| GrDEAH21 | Gorai.007G299200.1 | Chr07: 51082328-51088099 | 882 | 99.76 | 8.49 | Nuclear |
| GrDEAH22 | Gorai.007G305400.1 | Chr07: 51890852-51895676 | 611 | 68.07 | 8.18 | Nuclear |
| GrDEAH23 | Gorai.008G137800.1 | Chr08: 38782533-38786182 | 725 | 82.66 | 7.22 | Nuclear |
| GrDEAH24 | Gorai.008G154400.1 | Chr08: 41323487-41330255 | 885 | 100.52 | 7.91 | Nuclear |
| GrDEAH25 | Gorai.008G168800.1 | Chr08: 44099126-44107869 | 731 | 82.91 | 5.38 | Nuclear |
| GrDEAH26 | Gorai.008G176500.1 | Chr08: 45286639-45294517 | 718 | 80.5 | 8.88 | Nuclear |
| GrDEAH27 | Gorai.008G197200.1 | Chr08: 48161550-48167878 | 1067 | 121.28 | 5.64 | Nuclear |
| GrDEAH28 | Gorai.009G117300.1 | Chr09: 8628265-8633063 | 1189 | 135.53 | 6.56 | Nuclear |
| GrDEAH29 | Gorai.009G171000.1 | Chr09: 13221402-13230043 | 1391 | 153.37 | 5.43 | Nuclear |
| GrDEAH30 | Gorai.009G186700.1 | Chr09: 14354401-14365207 | 1234 | 137.83 | 8.29 | Nuclear |
| GrDEAH31 | Gorai.009G267400.1 | Chr09: 22217004-22220722 | 1006 | 113.77 | 5.31 | Nuclear |
| GrDEAH32 | Gorai.010G093100.1 | Chr10: 15093632-15104933 | 1675 | 187.55 | 6.05 | Nuclear |
| GrDEAH33 | Gorai.010G151100.1 | Chr10: 40671899-40680492 | 1325 | 146.14 | 5.3 | Nuclear |
| GrDEAH34 | Gorai.010G228200.1 | Chr10: 59997688-60013893 | 1021 | 115 | 8.53 | Nuclear |
| GrDEAH35 | Gorai.011G007600.1 | Chr11: 569884-574434 | 1184 | 135.34 | 6.68 | Nuclear |
| GrDEAH36 | Gorai.011G032400.1 | Chr11: 2409712-2417022 | 697 | 77.95 | 8.37 | PlasmaMembrane |
| GrDEAH37 | Gorai.011G075000.1 | Chr11: 7156546-7161392 | 913 | 103.07 | 7.79 | Nuclear |
| GrDEAH38 | Gorai.011G111000.1 | Chr11: 13326515-13334229 | 1760 | 197.96 | 7.62 | Nuclear |
| GrDEAH39 | Gorai.011G187000.1 | Chr11: 44528408-44535161 | 1060 | 123.2 | 5.95 | Nuclear |
| GrDEAH40 | Gorai.011G206500.1 | Chr11: 49795790-49814474 | 1767 | 199.65 | 7.03 | Nuclear |
| GrDEAH41 | Gorai.012G017400.1 | Chr12: 1996523-2003747 | 1345 | 151.14 | 8.44 | Nuclear |
| GrDEAH42 | Gorai.012G136000.1 | Chr12: 30913260-30920633 | 1038 | 114.76 | 6.6 | Nuclear |
| GrDEAH43 | Gorai.012G176400.1 | Chr12: 34564794-34567329 | 658 | 74.12 | 6.1 | Nuclear |
| GrDEAH44 | Gorai.013G001200.1 | Chr13: 68544-73180 | 863 | 96.21 | 8.9 | Chloroplast |
| GrDEAH45 | Gorai.013G038600.1 | Chr13: 3074495-3083714 | 1203 | 135.39 | 7.84 | Nuclear |
| GrDEAH46 | Gorai.013G043100.1 | Chr13: 3742541-3750413 | 971 | 107.92 | 6.2 | Nuclear |
| GrDEAH47 | Gorai.013G133100.1 | Chr13: 35035921-35039609 | 990 | 112.45 | 5.64 | Nuclear |
| GrDEAH48 | Gorai.013G139300.1 | Chr13: 37542094-37555083 | 1371 | 152.68 | 5.82 | Nuclear |
| GrDEAH49 | Gorai.013G141900.1 | Chr13: 38687077-38695377 | 1232 | 139.18 | 6.13 | Nuclear |
| GrDEAH50 | Gorai.013G218600.1 | Chr13: 53839220-53854330 | 2161 | 238.25 | 6.56 | Nuclear |
| GrDEAH51 | Gorai.013G252300.1 | Chr13: 56979166-56992254 | 1773 | 202.64 | 5.77 | Nuclear |
| GrDEAH52 | Gorai.013G258600.1 | Chr13: 57397754-57406240 | 1037 | 115.47 | 7.45 | Nuclear |
| GrDExD/H1 | Gorai.001G036900.1 | Chr01: 3403877-3420237 | 1455 | 165.33 | 5.25 | Nuclear |
| GrDExD/H2 | Gorai.001G150400.1 | Chr01: 20836533-20846178 | 1950 | 218.72 | 5.96 | Nuclear |
| GrDExD/H3 | Gorai.001G168700.1 | Chr01: 24030814-24041949 | 2238 | 251.24 | 9 | Nuclear |
| GrDExD/H4 | Gorai.002G016100.1 | Chr02: 1050235-1053354 | 405 | 45.9 | 5.99 | Cytoplasmic |
| GrDExD/H5 | Gorai.002G143000.1 | Chr02: 25828154-25839065 | 1393 | 158.07 | 6.64 | PlasmaMembrane |
| GrDExD/H6 | Gorai.002G216000.1 | Chr02: 56556123-56573745 | 1470 | 163.86 | 6.4 | Nuclear |
| GrDExD/H7 | Gorai.003G100200.1 | Chr03: 30861094-30875047 | 1287 | 144.52 | 6.5 | Nuclear |
| GrDExD/H8 | Gorai.003G125100.1 | Chr03: 37290357-37302757 | 1340 | 149.47 | 5.67 | Cytoplasmic |
| GrDExD/H9 | Gorai.003G156900.1 | Chr03: 42640922-42647039 | 796 | 90.08 | 8.15 | Nuclear |
| GrDExD/H10 | Gorai.004G209000.1 | Chr04: 54157901-54160653 | 548 | 59.42 | 9.63 | Mitochondrial |
| GrDExD/H11 | Gorai.004G221400.1 | Chr04: 55605032-55610980 | 1225 | 137.54 | 6.4 | Nuclear |
| GrDExD/H12 | Gorai.004G264200.1 | Chr04: 59938105-59956863 | 2042 | 231.37 | 5.47 | Nuclear |
| GrDExD/H13 | Gorai.004G292500.1 | Chr04: 62043821-62048421 | 428 | 48.46 | 5.4 | Cytoplasmic |
| GrDExD/H14 | Gorai.005G137100.1 | Chr05: 35497500-35505536 | 926 | 104.62 | 8.77 | Nuclear |
| GrDExD/H15 | Gorai.005G170000.1 | Chr05: 49691064-49693215 | 401 | 45.32 | 9.41 | Mitochondrial |
| GrDExD/H16 | Gorai.005G194600.1 | Chr05: 56476340-56494984 | 1217 | 137.04 | 7.12 | Nuclear |
| GrDExD/H17 | Gorai.006G005000.1 | Chr06: 967752-981201 | 1063 | 120.14 | 5.98 | Nuclear |
| GrDExD/H18 | Gorai.006G017900.1 | Chr06: 4179363-4184250 | 851 | 97.23 | 6.43 | Nuclear |
| GrDExD/H19 | Gorai.006G241400.1 | Chr06: 48762774-48767537 | 563 | 63.2 | 8.72 | Mitochondrial |
| GrDExD/H20 | Gorai.006G257000.1 | Chr06: 49880967-49896698 | 2586 | 283.31 | 6.46 | Nuclear |
| GrDExD/H21 | Gorai.007G006300.1 | Chr07: 523949-533327 | 1196 | 14.44 | 8.96 | Nuclear |
| GrDExD/H22 | Gorai.007G023800.1 | Chr07: 1694751-1699635 | 770 | 86.86 | 8.08 | Nuclear |
| GrDExD/H23 | Gorai.007G100800.1 | Chr07: 7468905-7480135 | 2258 | 253.19 | 8.82 | Nuclear |
| GrDExD/H24 | Gorai.007G374300.1 | Chr07: 60633068-60637091 | 532 | 59.72 | 9.39 | Nuclear |
| GrDExD/H25 | Gorai.008G031000.1 | Chr08: 3714524-3722015 | 930 | 105.31 | 6.29 | Nuclear |
| GrDExD/H26 | Gorai.008G049300.1 | Chr08: 6883759-6898577 | 2377 | 263.01 | 7.2 | Nuclear |
| GrDExD/H27 | Gorai.008G068400.1 | Chr08: 11250164-11254784 | 748 | 85.76 | 5.76 | Nuclear |
| GrDExD/H28 | Gorai.008G121100.1 | Chr08: 35933836-35944775 | 2250 | 252.23 | 8.95 | Nuclear |
| GrDExD/H29 | Gorai.008G145400.1 | Chr08: 39788075-39795065 | 1017 | 115.15 | 6.02 | PlasmaMembrane |
| GrDExD/H30 | Gorai.008G149500.1 | Chr08: 40485456-40488718 | 408 | 46.16 | 5.98 | Nuclear |
| GrDExD/H31 | Gorai.008G198400.1 | Chr08: 48292290-48321142 | 2090 | 236.59 | 6.08 | Nuclear |
| GrDExD/H32 | Gorai.009G053300.1 | Chr09: 3872981-3886515 | 1654 | 186.18 | 6.7 | Nuclear |
| GrDExD/H33 | Gorai.009G057600.1 | Chr09: 4145298-4149685 | 428 | 48.49 | 5.41 | Cytoplasmic |
| GrDExD/H34 | Gorai.009G153000.1 | Chr09: 11690907-11694880 | 514 | 57.39 | 9.54 | Nuclear |
| GrDExD/H35 | Gorai.009G345500.1 | Chr09: 41630420-41637165 | 1179 | 133.41 | 5.3 | Nuclear |
| GrDExD/H36 | Gorai.009G374900.1 | Chr09: 50970390-50979491 | 1395 | 158.66 | 6.56 | PlasmaMembrane |
| GrDExD/H37 | Gorai.009G393800.1 | Chr09: 54260282-54269764 | 1035 | 115.37 | 8.81 | Nuclear |
| GrDExD/H38 | Gorai.010G114300.1 | Chr10: 22016499-22022004 | 428 | 48.44 | 5.41 | Cytoplasmic |
| GrDExD/H39 | Gorai.010G134200.1 | Chr10: 29886669-29889520 | 409 | 45.73 | 9.2 | Nuclear |
| GrDExD/H40 | Gorai.010G135100.1 | Chr10: 30499026-30514000 | 688 | 77 | 8.43 | Cytoplasmic |
| GrDExD/H41 | Gorai.010G175700.1 | Chr10: 51517626-51529268 | 991 | 112.56 | 6.05 | Nuclear |
| GrDExD/H42 | Gorai.010G194700.1 | Chr10: 55411136-55418053 | 1268 | 146.35 | 8.06 | Nuclear |
| GrDExD/H43 | Gorai.010G244100.1 | Chr10: 61318853-61332614 | 1462 | 166.45 | 5.32 | Nuclear |
| GrDExD/H44 | Gorai.011G082500.1 | Chr11: 8316574-8321873 | 427 | 48.38 | 5.4 | Cytoplasmic |
| GrDExD/H45 | Gorai.011G088700.1 | Chr11: 9302151-9305910 | 535 | 58.4 | 8.55 | Nuclear |
| GrDExD/H46 | Gorai.011G165900.1 | Chr11: 32740783-32745005 | 481 | 54.19 | 9 | Nuclear |
| GrDExD/H47 | Gorai.011G166100.1 | Chr11: 32787373-32798664 | 1114 | 128.03 | 6.23 | Nuclear |
| GrDExD/H48 | Gorai.011G185100.1 | Chr11: 43975253-43992851 | 3361 | 365.5 | 5.33 | Nuclear |
| GrDExD/H49 | Gorai.011G217300.1 | Chr11: 52330942-52338804 | 2177 | 247.23 | 5.53 | Cytoplasmic |
| GrDExD/H50 | Gorai.012G023100.1 | Chr12: 2830126-2846019 | 1138 | 128.76 | 8.88 | Nuclear |
| GrDExD/H51 | Gorai.012G039300.1 | Chr12: 4882407-4892116 | 1221 | 134.91 | 6.11 | Nuclear |
| GrDExD/H52 | Gorai.012G063400.1 | Chr12: 8945198-8952950 | 1210 | 135.5 | 8.5 | Nuclear |
| GrDExD/H53 | Gorai.013G034700.1 | Chr13: 2677455-2684701 | 982 | 111.62 | 7.1 | Nuclear |
| GrDExD/H54 | Gorai.013G070700.1 | Chr13: 8189231-8192867 | 406 | 45.98 | 5.98 | Nuclear |
| GrDExD/H55 | Gorai.013G146500.1 | Chr13: 40255533-40278859 | 1256 | 139.06 | 7.25 | Nuclear |
| GrDExD/H56 | Gorai.013G197700.1 | Chr13: 50405230-50425237 | 2054 | 227.78 | 5.83 | Nuclear |
| GrDExD/H57 | Gorai.013G211300.1 | Chr13: 52317728-52328526 | 990 | 111.41 | 6.26 | Cytoplasmic |
| GrDExD/H58 | Gorai.013G215200.1 | Chr13: 53550658-53555657 | 428 | 48.44 | 5.4 | Cytoplasmic |
© 2014 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Chen, J.; Zhang, Y.; Liu, J.; Xia, M.; Wang, W.; Shen, F. Genome-Wide Analysis of the RNA Helicase Gene Family in Gossypium raimondii . Int. J. Mol. Sci. 2014, 15, 4635-4656. https://doi.org/10.3390/ijms15034635
Chen J, Zhang Y, Liu J, Xia M, Wang W, Shen F. Genome-Wide Analysis of the RNA Helicase Gene Family in Gossypium raimondii . International Journal of Molecular Sciences. 2014; 15(3):4635-4656. https://doi.org/10.3390/ijms15034635
Chicago/Turabian StyleChen, Jie, Yujuan Zhang, Jubo Liu, Minxuan Xia, Wei Wang, and Fafu Shen. 2014. "Genome-Wide Analysis of the RNA Helicase Gene Family in Gossypium raimondii " International Journal of Molecular Sciences 15, no. 3: 4635-4656. https://doi.org/10.3390/ijms15034635
APA StyleChen, J., Zhang, Y., Liu, J., Xia, M., Wang, W., & Shen, F. (2014). Genome-Wide Analysis of the RNA Helicase Gene Family in Gossypium raimondii . International Journal of Molecular Sciences, 15(3), 4635-4656. https://doi.org/10.3390/ijms15034635




