Emerging Roles of Claudins in Human Cancer
Abstract
:1. Introduction
2. Tight Junctions
3. Claudins
4. Dysregulation of Claudins in Human Cancer
4.1. Claudin Expression in Human Cancers
4.2. Regulation of Claudin Expression in Human Cancers
5. Role of Claudins in Human Cancer
5.1. Claudins in Tumor Progression
5.1.1. Claudin-1
5.1.2. Claudin-3 and Claudin-4
5.1.3. Claudin-6
5.1.4. Claudin-7
5.1.5. Claudin-11
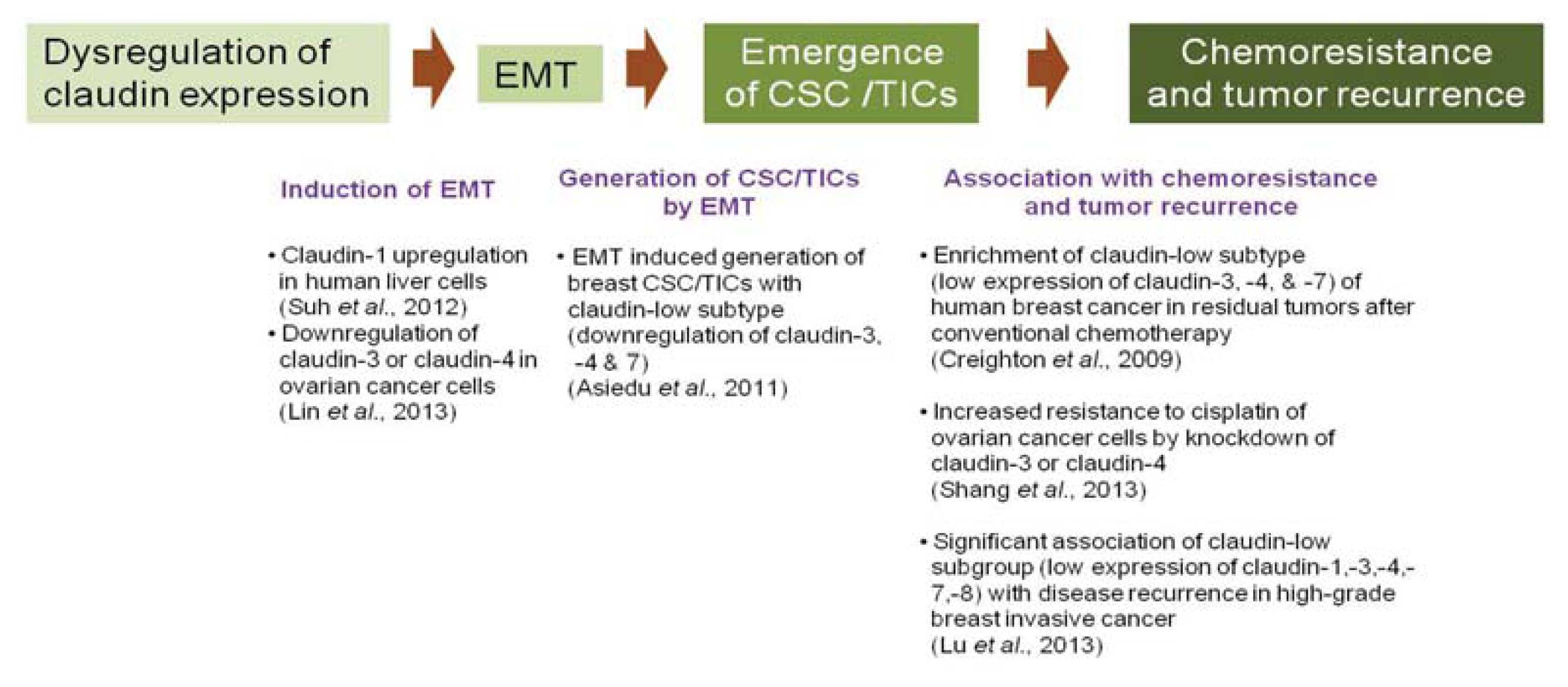
5.2. Claudins in Cancer Stem Cells or Tumor-Initiating Cells
5.3. Claudins in Chemoresistance
6. Claudins as Biomarkers and Therapeutic Targets
6.1. Prognostic Significance of Claudins in Cancer
6.2. Claudins as Drug Targets in Cancer
7. Conclusions
Acknowledgments
Conflicts of Interest
References
- Morin, P.J. Claudin proteins in human cancer: Promising new targets for diagnosis and therapy. Cancer Res 2005, 65, 9603–9606. [Google Scholar]
- Lal-Nag, M.; Morin, P.J. The claudins. Genome Biol 2009, 10, 235. [Google Scholar]
- Singh, A.B.; Sharma, A.; Dhawan, P. Claudin family of proteins and cancer: An overview. J. Oncol 2010, 2010, 541957. [Google Scholar]
- Honda, H.; Pazin, M.J.; Ji, H.; Wernyj, R.P.; Morin, P.J. Crucial roles of Sp1 and epigenetic modifications in the regulation of the CLDN4 promoter in ovarian cancer cells. J. Biol. Chem 2006, 281, 21433–21444. [Google Scholar]
- Honda, H.; Pazin, M.J.; D’Souza, T.; Ji, H.; Morin, P.J. Regulation of the CLDN3 gene in ovarian cancer cells. Cancer Biol. Ther 2007, 6, 1733–1742. [Google Scholar]
- Kwon, M.J.; Kim, S.S.; Choi, Y.L.; Jung, H.S.; Balch, C.; Kim, S.H.; Song, Y.S.; Marquez, V.E.; Nephew, K.P.; Shin, Y.K. Derepression of CLDN3 and CLDN4 during ovarian tumorigenesis is associated with loss of repressive histone modifications. Carcinogenesis 2010, 31, 974–983. [Google Scholar]
- Kwon, M.J.; Kim, S.H.; Jeong, H.M.; Jung, H.S.; Kim, S.S.; Lee, J.E.; Gye, M.C.; Erkin, O.C.; Koh, S.S.; Choi, Y.L.; et al. Claudin-4 overexpression is associated with epigenetic derepression in gastric carcinoma. Lab. Invest 2011, 91, 1652–1667. [Google Scholar]
- Baccelli, I.; Trumpp, A. The evolving concept of cancer and metastasis stem cells. J. Cell. Biol 2012, 198, 281–293. [Google Scholar]
- Singh, A.; Settleman, J. EMT, cancer stem cells and drug resistance: An emerging axis of evil in the war on cancer. Oncogene 2010, 29, 4741–4751. [Google Scholar]
- Herschkowitz, J.I.; Simin, K.; Weigman, V.J.; Mikaelian, I.; Usary, J.; Hu, Z.; Rasmussen, K.E.; Jones, L.P.; Assefnia, S.; Chandrasekharan, S. Identification of conserved gene expression features between murine mammary carcinoma models and human breast tumors. Genome Biol 2007, 8, R76. [Google Scholar]
- Hennessy, B.T.; Gonzalez-Angulo, A.M.; Stemke-Hale, K.; Gilcrease, M.Z.; Krishnamurthy, S.; Lee, J.S.; Fridlyand, J.; Sahin, A.; Agarwal, R.; Joy, C.; et al. Characterization of a naturally occurring breast cancer subset enriched in epithelial-to-mesenchymal transition and stem cell characteristics. Cancer Res 2009, 69, 4116–4124. [Google Scholar]
- Prat, A.; Parker, J.S.; Karginova, O.; Fan, C.; Livasy, C.; Herschkowitz, J.I.; He, X.; Perou, C.M. Phenotypic and molecular characterization of the claudin-low intrinsic subtype of breast cancer. Breast Cancer Res 2010, 12, R68. [Google Scholar]
- Lu, S.; Singh, K.; Mangray, S.; Tavares, R.; Noble, L.; Resnick, M.B.; Yakirevich, E. Claudin expression in high-grade invasive ductal carcinoma of the breast: Correlation with the molecular subtype. Mod. Pathol 2013, 26, 485–495. [Google Scholar]
- Turksen, K. Claudins and cancer stem cells. Stem Cell Rev 2011, 7, 797–798. [Google Scholar]
- Liu, T.; Cheng, W.; Lai, D.; Huang, Y.; Guo, L. Characterization of primary ovarian cancer cells in different culture systems. Oncol. Rep 2010, 23, 1277–1284. [Google Scholar]
- Qin, W.; Ren, Q.; Liu, T.; Huang, Y.; Wang, J. MicroRNA-155 is a novel suppressor of ovarian cancer-initiating cells that targets CLDN1. FEBS Lett 2013, 587, 1434–1439. [Google Scholar]
- Creighton, C.J.; Li, X.; Landis, M.; Dixon, J.M.; Neumeister, V.M.; Sjolund, A.; Rimm, D.L.; Wong, H.; Rodriguez, A.; Herschkowitz, J.I.; et al. Residual breast cancers after conventional therapy display mesenchymal as well as tumor-initiating features. Proc. Natl. Acad. Sci. USA 2009, 106, 13820–13825. [Google Scholar]
- Shang, X.; Lin, X.; Manorek, G.; Howell, S.B. Claudin-3 and claudin-4 regulate sensitivity to cisplatin by controlling expression of the copper and cisplatin influx transporter CTR1. Mol. Pharmacol 2013, 83, 85–94. [Google Scholar]
- Shin, K.; Fogg, V.C.; Margolis, B. Tight junctions and cell polarity. Annu. Rev. Cell Dev. Biol 2006, 22, 207–235. [Google Scholar]
- Matter, K.; Balda, M.S. Signalling to and from tight junctions. Nat. Rev. Mol. Cell Biol 2003, 4, 225–236. [Google Scholar]
- Matter, K.; Aijaz, S.; Tsapara, A.; Balda, M.S. Mammalian tight junctions in the regulation of epithelial differentiation and proliferation. Curr. Opin. Cell Biol 2005, 17, 453–458. [Google Scholar]
- Tsukita, S.; Yamazaki, Y.; Katsuno, T.; Tamura, A. Tight junction-based epithelial microenvironment and cell proliferation. Oncogene 2008, 27, 6930–6938. [Google Scholar]
- Assemat, E.; Bazellieres, E.; Pallesi-Pocachard, E.; Le Bivic, A.; Massey-Harroche, D. Polarity complex proteins. Biochim. Biophys. Acta 2008, 1778, 614–630. [Google Scholar]
- Cordenonsi, M.; Zanconato, F.; Azzolin, L.; Forcato, M.; Rosato, A.; Frasson, C.; Inui, M.; Montagner, M.; Parenti, A.R.; Poletti, A.; et al. The Hippo transducer TAZ confers cancer stem cell-related traits on breast cancer cells. Cell 2011, 147, 759–772. [Google Scholar]
- Matter, K.; Balda, M.S. Epithelial tight junctions, gene expression and nucleo-junctional interplay. J. Cell Sci 2007, 120, 1505–1511. [Google Scholar]
- Krause, G.; Winkler, L.; Mueller, S.L.; Haseloff, R.F.; Piontek, J.; Blasig, I.E. Structure and function of claudins. Biochim. Biophys. Acta 2008, 1778, 631–645. [Google Scholar]
- Fujibe, M.; Chiba, H.; Kojima, T.; Soma, T.; Wada, T.; Yamashita, T.; Sawada, N. Thr203 of claudin-1, a putative phosphorylation site for MAP kinase, is required to promote the barrier function of tight junctions. Exp. Cell Res 2004, 295, 36–47. [Google Scholar]
- Nunbhakdi-Craig, V.; Machleidt, T.; Ogris, E.; Bellotto, D.; White, C.L., III; Sontag, E. Protein phosphatase 2A associates with and regulates atypical PKC and the epithelial tight junction complex. J. Cell Biol. 2002, 158, 967–978. [Google Scholar]
- Ishizaki, T.; Chiba, H.; Kojima, T.; Fujibe, M.; Soma, T.; Miyajima, H.; Nagasawa, K.; Wada, I.; Sawada, N. Cyclic AMP induces phosphorylation of claudin-5 immunoprecipitates and expression of claudin-5 gene in blood-brain-barrier endothelial cells via protein kinase A-dependent and -independent pathways. Exp. Cell Res 2003, 290, 275–288. [Google Scholar]
- Ikari, A.; Ito, M.; Okude, C.; Sawada, H.; Harada, H.; Degawa, M.; Sakai, H.; Takahashi, T.; Sugatani, J.; Miwa, M. Claudin-16 is directly phosphorylated by protein kinase A independently of a vasodilator-stimulated phosphoprotein-mediated pathway. J. Cell Physiol 2008, 214, 221–229. [Google Scholar]
- Yamauchi, K.; Rai, T.; Kobayashi, K.; Sohara, E.; Suzuki, T.; Itoh, T.; Suda, S.; Hayama, A.; Sasaki, S.; Uchida, S. Disease-causing mutant WNK4 increases paracellular chloride permeability and phosphorylates claudins. Proc. Natl. Acad. Sci. USA 2004, 101, 4690–4694. [Google Scholar]
- Tsukita, S.; Furuse, M.; Itoh, M. Multifunctional strands in tight junctions. Nat. Rev. Mol. Cell Biol 2001, 2, 285–293. [Google Scholar]
- Turksen, K.; Troy, T.C. Barriers built on claudins. J. Cell Sci 2004, 117, 2435–2447. [Google Scholar]
- Van Itallie, C.; Rahner, C.; Anderson, J.M. Regulated expression of claudin-4 decreases paracellular conductance through a selective decrease in sodium permeability. J. Clin. Invest 2001, 107, 1319–1327. [Google Scholar]
- Karst, A.M.; Drapkin, R. Ovarian cancer pathogenesis: a model in evolution. J. Oncol 2010, 2010, 932371. [Google Scholar]
- Karst, A.M.; Levanon, K.; Drapkin, R. Modeling high-grade serous ovarian carcinogenesis from the fallopian tube. Proc. Natl. Acad. Sci. USA 2011, 108, 7547–7552. [Google Scholar]
- Levanon, K.; Ng, V.; Piao, H.Y.; Zhang, Y.; Chang, M.C.; Roh, M.H.; Kindelberger, D.W.; Hirsch, M.S.; Crum, C.P.; Marto, J.A.; et al. Primary ex vivo cultures of human fallopian tube epithelium as a model for serous ovarian carcinogenesis. Oncogene 2010, 29, 1103–1113. [Google Scholar]
- Lee, Y.; Miron, A.; Drapkin, R.; Nucci, M.R.; Medeiros, F.; Saleemuddin, A.; Garber, J.; Birch, C.; Mou, H.; Gordon, R.W.; et al. A candidate precursor to serous carcinoma that originates in the distal fallopian tube. J. Pathol 2007, 211, 26–35. [Google Scholar]
- Tokes, A.M.; Kulka, J.; Paku, S.; Szik, A.; Paska, C.; Novak, P.K.; Szilak, L.; Kiss, A.; Bogi, K.; Schaff, Z. Claudin-1, -3 and -4 proteins and mRNA expression in benign and malignant breast lesions: a research study. Breast Cancer Res 2005, 7, R296–R305. [Google Scholar]
- Kulka, J.; Szasz, A.M.; Nemeth, Z.; Madaras, L.; Schaff, Z.; Molnar, I.A.; Tokes, A.M. Expression of tight junction protein claudin-4 in basal-like breast carcinomas. Pathol. Oncol. Res 2009, 15, 59–64. [Google Scholar]
- Resnick, M.B.; Gavilanez, M.; Newton, E.; Konkin, T.; Bhattacharya, B.; Britt, D.E.; Sabo, E.; Moss, S.F. Claudin expression in gastric adenocarcinomas: A tissue microarray study with prognostic correlation. Hum. Pathol 2005, 36, 886–892. [Google Scholar]
- Soini, Y. Expression of claudins 1, 2, 3, 4, 5 and 7 in various types of tumours. Histopathology 2005, 46, 551–560. [Google Scholar]
- Kim, T.H.; Huh, J.H.; Lee, S.; Kang, H.; Kim, G.I.; An, H.J. Down-regulation of claudin-2 in breast carcinomas is associated with advanced disease. Histopathology 2008, 53, 48–55. [Google Scholar]
- Kominsky, S.L.; Vali, M.; Korz, D.; Gabig, T.G.; Weitzman, S.A.; Argani, P.; Sukumar, S. Clostridium perfringens enterotoxin elicits rapid and specific cytolysis of breast carcinoma cells mediated through tight junction proteins claudin 3 and 4. Am. J. Pathol 2004, 164, 1627–1633. [Google Scholar]
- Kominsky, S.L.; Argani, P.; Korz, D.; Evron, E.; Raman, V.; Garrett, E.; Rein, A.; Sauter, G.; Kallioniemi, O.P.; Sukumar, S. Loss of the tight junction protein claudin-7 correlates with histological grade in both ductal carcinoma in situ and invasive ductal carcinoma of the breast. Oncogene 2003, 22, 2021–2033. [Google Scholar]
- Sobel, G.; Paska, C.; Szabo, I.; Kiss, A.; Kadar, A.; Schaff, Z. Increased expression of claudins in cervical squamous intraepithelial neoplasia and invasive carcinoma. Hum. Pathol 2005, 36, 162–169. [Google Scholar]
- De Oliveira, S.S.; de Oliveira, I.M.; de Souza, W.; Morgado-Diaz, J.A. Claudins upregulation in human colorectal cancer. FEBS Lett 2005, 579, 6179–6185. [Google Scholar]
- Dhawan, P.; Singh, A.B.; Deane, N.G.; No, Y.; Shiou, S.R.; Schmidt, C.; Neff, J.; Washington, M.K.; Beauchamp, R.D. Claudin-1 regulates cellular transformation and metastatic behavior in colon cancer. J. Clin. Invest 2005, 115, 1765–1776. [Google Scholar]
- Montgomery, E.; Mamelak, A.J.; Gibson, M.; Maitra, A.; Sheikh, S.; Amr, S.S.; Yang, S.; Brock, M.; Forastiere, A.; Zhang, S.; et al. Overexpression of claudin proteins in esophageal adenocarcinoma and its precursor lesions. Appl. Immunohistochem. Mol. Morphol 2006, 14, 24–30. [Google Scholar]
- Usami, Y.; Chiba, H.; Nakayama, F.; Ueda, J.; Matsuda, Y.; Sawada, N.; Komori, T.; Ito, A.; Yokozaki, H. Reduced expression of claudin-7 correlates with invasion and metastasis in squamous cell carcinoma of the esophagus. Hum. Pathol 2006, 37, 569–577. [Google Scholar]
- Cunningham, S.C.; Kamangar, F.; Kim, M.P.; Hammoud, S.; Haque, R.; Iacobuzio-Donahue, C.A.; Maitra, A.; Ashfaq, R.; Hustinx, S.; Heitmiller, R.E.; et al. Claudin-4, mitogen-activated protein kinase kinase 4, and stratifin are markers of gastric adenocarcinoma precursor lesions. Cancer Epidemiol. Biomarkers Prev 2006, 15, 281–287. [Google Scholar]
- Johnson, A.H.; Frierson, H.F.; Zaika, A.; Powell, S.M.; Roche, J.; Crowe, S.; Moskaluk, C.A.; El-Rifai, W. Expression of tight-junction protein claudin-7 is an early event in gastric tumorigenesis. Am. J. Pathol 2005, 167, 577–584. [Google Scholar]
- Agarwal, R.; Mori, Y.; Cheng, Y.; Jin, Z.; Olaru, A.V.; Hamilton, J.P.; David, S.; Selaru, F.M.; Yang, J.; Abraham, J.M.; et al. Silencing of claudin-11 is associated with increased invasiveness of gastric cancer cells. PLoS One 2009, 4, e8002. [Google Scholar]
- Higashi, Y.; Suzuki, S.; Sakaguchi, T.; Nakamura, T.; Baba, S.; Reinecker, H.C.; Nakamura, S.; Konno, H. Loss of claudin-1 expression correlates with malignancy of hepatocellular carcinoma. J. Surg. Res 2007, 139, 68–76. [Google Scholar]
- Leotlela, P.D.; Wade, M.S.; Duray, P.H.; Rhode, M.J.; Brown, H.F.; Rosenthal, D.T.; Dissanayake, S.K.; Earley, R.; Indig, F.E.; Nickoloff, B.J.; et al. Claudin-1 overexpression in melanoma is regulated by PKC and contributes to melanoma cell motility. Oncogene 2007, 26, 3846–3856. [Google Scholar]
- Hough, C.D.; Sherman-Baust, C.A.; Pizer, E.S.; Montz, F.J.; Im, D.D.; Rosenshein, N.B.; Cho, K.R.; Riggins, G.J.; Morin, P.J. Large-scale serial analysis of gene expression reveals genes differentially expressed in ovarian cancer. Cancer Res 2000, 60, 6281–6287. [Google Scholar]
- Rangel, L.B.; Agarwal, R.; D’Souza, T.; Pizer, E.S.; Alo, P.L.; Lancaster, W.D.; Gregoire, L.; Schwartz, D.R.; Cho, K.R.; Morin, P.J. Tight junction proteins claudin-3 and claudin-4 are frequently overexpressed in ovarian cancer but not in ovarian cystadenomas. Clin. Cancer Res 2003, 9, 2567–2575. [Google Scholar]
- Heinzelmann-Schwarz, V.A.; Gardiner-Garden, M.; Henshall, S.M.; Scurry, J.; Scolyer, R.A.; Davies, M.J.; Heinzelmann, M.; Kalish, L.H.; Bali, A.; Kench, J.G.; et al. Overexpression of the cell adhesion molecules DDR1, Claudin 3, and Ep-CAM in metaplastic ovarian epithelium and ovarian cancer. Clin. Cancer Res 2004, 10, 4427–4436. [Google Scholar]
- Lu, K.H.; Patterson, A.P.; Wang, L.; Marquez, R.T.; Atkinson, E.N.; Baggerly, K.A.; Ramoth, L.R.; Rosen, D.G.; Liu, J.; Hellstrom, I.; et al. Selection of potential markers for epithelial ovarian cancer with gene expression arrays and recursive descent partition analysis. Clin. Cancer Res 2004, 10, 3291–3300. [Google Scholar]
- Santin, A.D.; Zhan, F.; Bellone, S.; Palmieri, M.; Cane, S.; Bignotti, E.; Anfossi, S.; Gokden, M.; Dunn, D.; Roman, J.J.; et al. Gene expression profiles in primary ovarian serous papillary tumors and normal ovarian epithelium: Identification of candidate molecular markers for ovarian cancer diagnosis and therapy. Int. J. Cancer 2004, 112, 14–25. [Google Scholar]
- Zhu, Y.; Brannstrom, M.; Janson, P.O.; Sundfeldt, K. Differences in expression patterns of the tight junction proteins,claudin 1, 3, 4 and 5, in human ovarian surface epithelium as compared to epithelia in inclusion cysts and epithelial ovarian tumours. Int. J. Cancer 2006, 118, 1884–1891. [Google Scholar]
- Hibbs, K.; Skubitz, K.M.; Pambuccian, S.E.; Casey, R.C.; Burleson, K.M.; Oegema, T.R., Jr; Thiele, J.J.; Grindle, S.M.; Bliss, R.L.; Skubitz, A.P. Differential gene expression in ovarian carcinoma: Identification of potential biomarkers. Am. J. Pathol. 2004, 165, 397–414. [Google Scholar]
- Tassi, R.A.; Bignotti, E.; Falchetti, M.; Ravanini, M.; Calza, S.; Ravaggi, A.; Bandiera, E.; Facchetti, F.; Pecorelli, S.; Santin, A.D. Claudin-7 expression in human epithelial ovarian cancer. Int. J. Gynecol. Cancer 2008, 18, 1262–1271. [Google Scholar]
- Shang, X.; Lin, X.; Alvarez, E.; Manorek, G.; Howell, S.B. Tight junction proteins claudin-3 and claudin-4 control tumor growth and metastases. Neoplasia 2012, 14, 974–985. [Google Scholar]
- Sheehan, G.M.; Kallakury, B.V.; Sheehan, C.E.; Fisher, H.A.; Kaufman, R.P., Jr; Ross, J.S. Loss of claudins-1 and -7 and expression of claudins-3 and -4 correlate with prognostic variables in prostatic adenocarcinomas. Hum. Pathol. 2007, 38, 564–569. [Google Scholar]
- Long, H.; Crean, C.D.; Lee, W.H.; Cummings, O.W.; Gabig, T.G. Expression of Clostridium perfringens enterotoxin receptors claudin-3 and claudin-4 in prostate cancer epithelium. Cancer Res 2001, 61, 7878–7881. [Google Scholar]
- Michl, P.; Buchholz, M.; Rolke, M.; Kunsch, S.; Lohr, M.; McClane, B.; Tsukita, S.; Leder, G.; Adler, G.; Gress, T.M. Claudin-4: A new target for pancreatic cancer treatment using Clostridium perfringens enterotoxin. Gastroenterology 2001, 121, 678–684. [Google Scholar]
- Nichols, L.S.; Ashfaq, R.; Iacobuzio-Donahue, C.A. Claudin 4 protein expression in primary and metastatic pancreatic cancer: Support for use as a therapeutic target. Am. J. Clin. Pathol 2004, 121, 226–230. [Google Scholar]
- Santin, A.D.; Zhan, F.; Cane, S.; Bellone, S.; Palmieri, M.; Thomas, M.; Burnett, A.; Roman, J.J.; Cannon, M.J.; Shaughnessy, J., Jr; et al. Gene expression fingerprint of uterine serous papillary carcinoma: identification of novel molecular markers for uterine serous cancer diagnosis and therapy. Br. J. Cancer 2005, 92, 1561–1573. [Google Scholar]
- Ikenouchi, J.; Matsuda, M.; Furuse, M.; Tsukita, S. Regulation of tight junctions during the epithelium-mesenchyme transition: direct repression of the gene expression of claudins/occludin by Snail. J. Cell Sci 2003, 116, 1959–1967. [Google Scholar]
- Martinez-Estrada, O.M.; Culleres, A.; Soriano, F.X.; Peinado, H.; Bolos, V.; Martinez, F.O.; Reina, M.; Cano, A.; Fabre, M.; Vilaro, S. The transcription factors Slug and Snail act as repressors of Claudin-1 expression in epithelial cells. Biochem. J 2006, 394, 449–457. [Google Scholar]
- Chang, T.L.; Ito, K.; Ko, T.K.; Liu, Q.; Salto-Tellez, M.; Yeoh, K.G.; Fukamachi, H.; Ito, Y. Claudin-1 has tumor suppressive activity and is a direct target of RUNX3 in gastric epithelial cells. Gastroenterology 2009, 138, 255–265. [Google Scholar]
- Bhat, A.A.; Sharma, A.; Pope, J.; Krishnan, M.; Washington, M.K.; Singh, A.B.; Dhawan, P. Caudal homeobox protein Cdx-2 cooperates with Wnt pathway to regulate claudin-1 expression in colon cancer cells. PLoS One 2012, 7, e37174. [Google Scholar]
- Di Cello, F.; Cope, L.; Li, H.; Jeschke, J.; Wang, W.; Baylin, S.B.; Zahnow, C.A. Methylation of the claudin 1 promoter is associated with loss of expression in estrogen receptor positive breast cancer. PLoS One 2013, 8, e68630. [Google Scholar]
- Litkouhi, B.; Kwong, J.; Lo, C.M.; Smedley, J.G., III; McClane, B.A.; Aponte, M.; Gao, Z.; Sarno, J.L.; Hinners, J.; Welch, W.R.; et al. Claudin-4 overexpression in epithelial ovarian cancer is associated with hypomethylation and is a potential target for modulation of tight junction barrier function using a C-terminal fragment of Clostridium perfringens enterotoxin. Neoplasia 2007, 9, 304–314. [Google Scholar]
- Sahin, U.; Koslowski, M.; Dhaene, K.; Usener, D.; Brandenburg, G.; Seitz, G.; Huber, C.; Tureci, O. Claudin-18 splice variant 2 is a pan-cancer target suitable for therapeutic antibody development. Clin. Cancer Res 2008, 14, 7624–7634. [Google Scholar]
- Ito, T.; Kojima, T.; Yamaguchi, H.; Kyuno, D.; Kimura, Y.; Imamura, M.; Takasawa, A.; Murata, M.; Tanaka, S.; Hirata, K.; et al. Transcriptional regulation of claudin-18 via specific protein kinase C signaling pathways and modification of DNA methylation in human pancreatic cancer cells. J. Cell. Biochem 2011, 112, 1761–1772. [Google Scholar]
- Krishnan, M.; Singh, A.B.; Smith, J.J.; Sharma, A.; Chen, X.; Eschrich, S.; Yeatman, T.J.; Beauchamp, R.D.; Dhawan, P. HDAC inhibitors regulate claudin-1 expression in colon cancer cells through modulation of mRNA stability. Oncogene 2009, 29, 305–312. [Google Scholar]
- Sharma, A.; Bhat, A.A.; Krishnan, M.; Singh, A.B.; Dhawan, P. Trichostatin-A modulates claudin-1 mRNA stability through the modulation of Hu antigen R and tristetraprolin in colon cancer cells. Carcinogenesis 2013. [Google Scholar] [CrossRef]
- Darido, C.; Buchert, M.; Pannequin, J.; Bastide, P.; Zalzali, H.; Mantamadiotis, T.; Bourgaux, J.F.; Garambois, V.; Jay, P.; Blache, P.; et al. Defective claudin-7 regulation by Tcf-4 and Sox-9 disrupts the polarity and increases the tumorigenicity of colorectal cancer cells. Cancer Res 2008, 68, 4258–4268. [Google Scholar]
- Michl, P.; Barth, C.; Buchholz, M.; Lerch, M.M.; Rolke, M.; Holzmann, K.H.; Menke, A.; Fensterer, H.; Giehl, K.; Lohr, M.; et al. Claudin-4 expression decreases invasiveness and metastatic potential of pancreatic cancer. Cancer Res 2003, 63, 6265–6271. [Google Scholar]
- Singh, A.B.; Harris, R.C. Epidermal growth factor receptor activation differentially regulates claudin expression and enhances transepithelial resistance in Madin-Darby canine kidney cells. J. Biol. Chem 2004, 279, 3543–3552. [Google Scholar]
- Oshima, T.; Miwa, H.; Joh, T. Aspirin induces gastric epithelial barrier dysfunction by activating p38 MAPK via claudin-7. Am. J. Physiol. Cell. Physiol 2008, 295, C800–C806. [Google Scholar]
- Lioni, M.; Brafford, P.; Andl, C.; Rustgi, A.; El-Deiry, W.; Herlyn, M.; Smalley, K.S. Dysregulation of claudin-7 leads to loss of E-cadherin expression and the increased invasion of esophageal squamous cell carcinoma cells. Am. J. Pathol 2007, 170, 709–721. [Google Scholar]
- Boireau, S.; Buchert, M.; Samuel, M.S.; Pannequin, J.; Ryan, J.L.; Choquet, A.; Chapuis, H.; Rebillard, X.; Avances, C.; Ernst, M.; et al. DNA-methylation-dependent alterations of claudin-4 expression in human bladder carcinoma. Carcinogenesis 2007, 28, 246–258. [Google Scholar]
- D’Souza, T.; Agarwal, R.; Morin, P.J. Phosphorylation of claudin-3 at threonine 192 by cAMP-dependent protein kinase regulates tight junction barrier function in ovarian cancer cells. J. Biol. Chem 2005, 280, 26233–26240. [Google Scholar]
- D’Souza, T.; Indig, F.E.; Morin, P.J. Phosphorylation of claudin-4 by PKCepsilon regulates tight junction barrier function in ovarian cancer cells. Exp. Cell Res 2007, 313, 3364–3375. [Google Scholar]
- Blanchard, A.A.; Ma, X.; Dueck, K.J.; Penner, C.; Cooper, S.C.; Mulhall, D.; Murphy, L.C.; Leygue, E.; Myal, Y. Claudin 1 expression in basal-like breast cancer is related to patient age. BMC Cancer 2013, 13, 268. [Google Scholar]
- Akasaka, H.; Sato, F.; Morohashi, S.; Wu, Y.; Liu, Y.; Kondo, J.; Odagiri, H.; Hakamada, K.; Kijima, H. Anti-apoptotic effect of claudin-1 in tamoxifen-treated human breast cancer MCF-7 cells. BMC Cancer 2010, 10, 548. [Google Scholar]
- Yoon, C.H.; Kim, M.J.; Park, M.J.; Park, I.C.; Hwang, S.G.; An, S.; Choi, Y.H.; Yoon, G.; Lee, S.J. Claudin-1 acts through c-ABl-PKCdelta signaling and has a causal role in the acquisition of invasive capacity in human liver cells. J. Biol. Chem 2009, 285, 226–233. [Google Scholar]
- Suh, Y.; Yoon, C.H.; Kim, R.K.; Lim, E.J.; Oh, Y.S.; Hwang, S.G.; An, S.; Yoon, G.; Gye, M.C.; Yi, J.M.; et al. Claudin-1 induces epithelial-mesenchymal transition through activation of the c-Abl-ERK signaling pathway in human liver cells. Oncogene 2012. [Google Scholar] [CrossRef]
- Oku, N.; Sasabe, E.; Ueta, E.; Yamamoto, T.; Osaki, T. Tight junction protein claudin-1 enhances the invasive activity of oral squamous cell carcinoma cells by promoting cleavage of laminin-5 gamma2 chain via matrix metalloproteinase (MMP)-2 and membrane-type MMP-1. Cancer Res 2006, 66, 5251–5257. [Google Scholar]
- Chao, Y.C.; Pan, S.H.; Yang, S.C.; Yu, S.L.; Che, T.F.; Lin, C.W.; Tsai, M.S.; Chang, G.C.; Wu, C.H.; Wu, Y.Y.; et al. Claudin-1 is a metastasis suppressor and correlates with clinical outcome in lung adenocarcinoma. Am. J. Respir. Crit. Care Med 2009, 179, 123–133. [Google Scholar]
- Agarwal, R.; D’Souza, T.; Morin, P.J. Claudin-3 and claudin-4 expression in ovarian epithelial cells enhances invasion and is associated with increased matrix metalloproteinase-2 activity. Cancer Res 2005, 65, 7378–7385. [Google Scholar]
- Huang, Y.H.; Bao, Y.; Peng, W.; Goldberg, M.; Love, K.; Bumcrot, D.A.; Cole, G.; Langer, R.; Anderson, D.G.; Sawicki, J.A. Claudin-3 gene silencing with siRNA suppresses ovarian tumor growth and metastasis. Proc. Natl. Acad. Sci. USA 2009, 106, 3426–3430. [Google Scholar]
- Lin, X.; Shang, X.; Manorek, G.; Howell, S.B. Regulation of the epithelial-mesenchymal transition by claudin-3 and claudin-4. PLoS One 2013, 8, e67496. [Google Scholar]
- Li, J.; Chigurupati, S.; Agarwal, R.; Mughal, M.R.; Mattson, M.P.; Becker, K.G.; Wood, W.H., III; Zhang, Y.; Morin, P.J. Possible angiogenic roles for claudin-4 in ovarian cancer. Cancer Biol. Ther. 2009, 8, 1806–1814. [Google Scholar]
- Zavala-Zendejas, V.E.; Torres-Martinez, A.C.; Salas-Morales, B.; Fortoul, T.I.; Montano, L.F.; Rendon-Huerta, E.P. Claudin-6, 7, or 9 overexpression in the human gastric adenocarcinoma cell line AGS increases its invasiveness, migration, and proliferation rate. Cancer Invest 2011, 29, 1–11. [Google Scholar]
- Osanai, M.; Murata, M.; Chiba, H.; Kojima, T.; Sawada, N. Epigenetic silencing of claudin-6 promotes anchorage-independent growth of breast carcinoma cells. Cancer Sci 2007, 98, 1557–1562. [Google Scholar]
- Wu, Q.; Liu, Y.; Ren, Y.; Xu, X.; Yu, L.; Li, Y.; Quan, C. Tight junction protein, claudin-6, downregulates the malignant phenotype of breast carcinoma. Eur. J. Cancer Prev 2010, 19, 186–194. [Google Scholar]
- Dahiya, N.; Becker, K.G.; Wood, W.H., III; Zhang, Y.; Morin, P.J. Claudin-7 is frequently overexpressed in ovarian cancer and promotes invasion. PLoS One 2011, 6, e22119. [Google Scholar]
- Lu, Z.; Ding, L.; Hong, H.; Hoggard, J.; Lu, Q.; Chen, Y.H. Claudin-7 inhibits human lung cancer cell migration and invasion through ERK/MAPK signaling pathway. Exp. Cell Res 2011, 317, 1935–1946. [Google Scholar]
- Awsare, N.S.; Martin, T.A.; Haynes, M.D.; Matthews, P.N.; Jiang, W.G. Claudin-11 decreases the invasiveness of bladder cancer cells. Oncol. Rep 2011, 25, 1503–1509. [Google Scholar]
- Webb, P.G.; Spillman, M.A.; Baumgartner, H.K. Claudins play a role in normal and tumor cell motility. BMC Cell Biol 2013, 14, 19. [Google Scholar]
- Choi, Y.L.; Kim, J.; Kwon, M.J.; Choi, J.S.; Kim, T.J.; Bae, D.S.; Koh, S.S.; In, Y.H.; Park, Y.W.; Kim, S.H.; et al. Expression profile of tight junction protein claudin 3 and claudin 4 in ovarian serous adenocarcinoma with prognostic correlation. Histol. Histopathol 2007, 22, 1185–1195. [Google Scholar]
- Szasz, A.M.; Nemeth, Z.; Gyorffy, B.; Micsinai, M.; Krenacs, T.; Baranyai, Z.; Harsanyi, L.; Kiss, A.; Schaff, Z.; Tokes, A.M.; et al. Identification of a claudin-4 and E-cadherin score to predict prognosis in breast cancer. Cancer Sci 2011, 102, 2248–2254. [Google Scholar]
- Lee, K.W.; Lee, N.K.; Kim, J.H.; Kang, M.S.; Yoo, H.Y.; Kim, H.H.; Um, S.H.; Kim, S.H. Twist1 causes the transcriptional repression of claudin-4 with prognostic significance in esophageal cancer. Biochem. Biophys. Res. Commun 2012, 423, 454–460. [Google Scholar]
- Ersoz, S.; Mungan, S.; Cobanoglu, U.; Turgutalp, H.; Ozoran, Y. Prognostic importance of Claudin-1 and Claudin-4 expression in colon carcinomas. Pathol. Res. Pract 2011, 207, 285–289. [Google Scholar]
- Matsuoka, T.; Mitomi, H.; Fukui, N.; Kanazawa, H.; Saito, T.; Hayashi, T.; Yao, T. Cluster analysis of claudin-1 and -4, E-cadherin, and beta-catenin expression in colorectal cancers. J. Surg. Oncol 2011, 103, 674–686. [Google Scholar]
- Tsutsumi, K.; Sato, N.; Tanabe, R.; Mizumoto, K.; Morimatsu, K.; Kayashima, T.; Fujita, H.; Ohuchida, K.; Ohtsuka, T.; Takahata, S.; et al. Claudin-4 expression predicts survival in pancreatic ductal adenocarcinoma. Ann. Surg. Oncol 2012, 19, S491–S499. [Google Scholar]
- Kyuno, D.; Kojima, T.; Ito, T.; Yamaguchi, H.; Tsujiwaki, M.; Takasawa, A.; Murata, M.; Tanaka, S.; Hirata, K.; Sawada, N. Protein kinase Calpha inhibitor enhances the sensitivity of human pancreatic cancer HPAC cells to Clostridium perfringens enterotoxin via claudin-4. Cell Tissue Res 2011, 346, 369–381. [Google Scholar]
- Birks, D.K.; Kleinschmidt-DeMasters, B.K.; Donson, A.M.; Barton, V.N.; McNatt, S.A.; Foreman, N.K.; Handler, M.H. Claudin 6 is a positive marker for atypical teratoid/rhabdoid tumors. Brain Pathol 2010, 20, 140–150. [Google Scholar]
- Yafang, L.; Qiong, W.; Yue, R.; Xiaoming, X.; Lina, Y.; Mingzi, Z.; Ting, Z.; Yulin, L.; Chengshi, Q. Role of estrogen receptor-alpha in the regulation of claudin-6 expression in breast cancer cells. J. Breast Cancer 2011, 14, 20–27. [Google Scholar]
- Rendon-Huerta, E.; Teresa, F.; Teresa, G.M.; Xochitl, G.S.; Georgina, A.F.; Veronica, Z.Z.; Montano, L.F. Distribution and expression pattern of claudins 6, 7, and 9 in diffuse- and intestinal-type gastric adenocarcinomas. J. Gastrointest. Cancer 2010, 41, 52–59. [Google Scholar]
- Lal-Nag, M.; Battis, M.; Santin, A.D.; Morin, P.J. Claudin-6: A novel receptor for CPE-mediated cytotoxicity in ovarian cancer. Oncogenesis 2012, 1, e33. [Google Scholar]
- Nubel, T.; Preobraschenski, J.; Tuncay, H.; Weiss, T.; Kuhn, S.; Ladwein, M.; Langbein, L.; Zoller, M. Claudin-7 regulates EpCAM-mediated functions in tumor progression. Mol. Cancer Res 2009, 7, 285–299. [Google Scholar]
- Bronstein, J.M.; Chen, K.; Tiwari-Woodruff, S.; Kornblum, H.I. Developmental expression of OSP/claudin-11. J. Neurosci. Res 2000, 60, 284–290. [Google Scholar]
- Katsushima, K.; Shinjo, K.; Natsume, A.; Ohka, F.; Fujii, M.; Osada, H.; Sekido, Y.; Kondo, Y. Contribution of microRNA-1275 to Claudin11 protein suppression via a polycomb-mediated silencing mechanism in human glioma stem-like cells. J. Biol. Chem 2012, 287, 27396–27406. [Google Scholar]
- Lehmann, B.D.; Bauer, J.A.; Chen, X.; Sanders, M.E.; Chakravarthy, A.B.; Shyr, Y.; Pietenpol, J.A. Identification of human triple-negative breast cancer subtypes and preclinical models for selection of targeted therapies. J. Clin. Invest 2011, 121, 2750–2767. [Google Scholar]
- Asiedu, M.K.; Ingle, J.N.; Behrens, M.D.; Radisky, D.C.; Knutson, K.L. TGFbeta/TNF(alpha)-mediated epithelial-mesenchymal transition generates breast cancer stem cells with a claudin-low phenotype. Cancer Res 2011, 71, 4707–4719. [Google Scholar]
- Bruna, A.; Greenwood, W.; Le Quesne, J.; Teschendorff, A.; Miranda-Saavedra, D.; Rueda, O.M.; Sandoval, J.L.; Vidakovic, A.T.; Saadi, A.; Pharoah, P.; et al. TGFbeta induces the formation of tumour-initiating cells in claudinlow breast cancer. Nat. Commun 2012, 3, 1055. [Google Scholar]
- Morel, A.P.; Hinkal, G.W.; Thomas, C.; Fauvet, F.; Courtois-Cox, S.; Wierinckx, A.; Devouassoux-Shisheboran, M.; Treilleux, I.; Tissier, A.; Gras, B.; et al. EMT inducers catalyze malignant transformation of mammary epithelial cells and drive tumorigenesis towards claudin-low tumors in transgenic mice. PLoS Genet 2012, 8, e1002723. [Google Scholar]
- Boylan, K.L.; Misemer, B.; Derycke, M.S.; Andersen, J.D.; Harrington, K.M.; Kalloger, S.E.; Gilks, C.B.; Pambuccian, S.E.; Skubitz, A.P. Claudin 4 is differentially expressed between ovarian cancer subtypes and plays a role in spheroid formation. Int. J. Mol. Sci 2011, 12, 1334–1358. [Google Scholar]
- Zhang, G.J.; Xiao, H.X.; Tian, H.P.; Liu, Z.L.; Xia, S.S.; Zhou, T. Upregulation of microRNA-155 promotes the migration and invasion of colorectal cancer cells through the regulation of claudin-1 expression. Int. J. Mol. Med 2013, 31, 1375–1380. [Google Scholar]
- Wang, L.; Xue, Y.; Shen, Y.; Li, W.; Cheng, Y.; Yan, X.; Shi, W.; Wang, J.; Gong, Z.; Yang, G.; et al. Claudin 6: A novel surface marker for characterizing mouse pluripotent stem cells. Cell. Res 2012, 22, 1082–1085. [Google Scholar]
- Turksen, K.; Troy, T.C. Claudin-6: A novel tight junction molecule is developmentally regulated in mouse embryonic epithelium. Dev. Dyn 2001, 222, 292–300. [Google Scholar]
- Ben-David, U.; Nudel, N.; Benvenisty, N. Immunologic and chemical targeting of the tight-junction protein Claudin-6 eliminates tumorigenic human pluripotent stem cells. Nat. Commun 2013, 4, 1992. [Google Scholar]
- Longley, D.B.; Johnston, P.G. Molecular mechanisms of drug resistance. J. Pathol 2005, 205, 275–292. [Google Scholar]
- Stewart, J.J.; White, J.T.; Yan, X.; Collins, S.; Drescher, C.W.; Urban, N.D.; Hood, L.; Lin, B. Proteins associated with Cisplatin resistance in ovarian cancer cells identified by quantitative proteomic technology and integrated with mRNA expression levels. Mol. Cell. Prot 2006, 5, 433–443. [Google Scholar]
- Yoshida, H.; Sumi, T.; Zhi, X.; Yasui, T.; Honda, K.; Ishiko, O. Claudin-4: A potential therapeutic target in chemotherapy-resistant ovarian cancer. Anticancer Res 2011, 31, 1271–1277. [Google Scholar]
- Casagrande, F.; Cocco, E.; Bellone, S.; Richter, C.E.; Bellone, M.; Todeschini, P.; Siegel, E.; Varughese, J.; Arin-Silasi, D.; Azodi, M.; et al. Eradication of chemotherapy-resistant CD44+ human ovarian cancer stem cells in mice by intraperitoneal administration of Clostridium perfringens enterotoxin. Cancer 2011, 117, 5519–5528. [Google Scholar]
- Santin, A.D.; Cane, S.; Bellone, S.; Palmieri, M.; Siegel, E.R.; Thomas, M.; Roman, J.J.; Burnett, A.; Cannon, M.J.; Pecorelli, S. Treatment of chemotherapy-resistant human ovarian cancer xenografts in C.B-17/SCID mice by intraperitoneal administration of Clostridium perfringens enterotoxin. Cancer Res 2005, 65, 4334–4342. [Google Scholar]
- Kim, C.J.; Lee, J.W.; Choi, J.J.; Choi, H.Y.; Park, Y.A.; Jeon, H.K.; Sung, C.O.; Song, S.Y.; Lee, Y.Y.; Choi, C.H.; et al. High claudin-7 expression is associated with a poor response to platinum-based chemotherapy in epithelial ovarian carcinoma. Eur. J. Cancer 2011, 47, 918–925. [Google Scholar]
- Thuma, F.; Zoller, M. EpCAM-associated claudin-7 supports lymphatic spread and drug resistance in rat pancreatic cancer. Int. J. Cancer 2013, 133, 855–866. [Google Scholar]
- Hoggard, J.; Fan, J.; Lu, Z.; Lu, Q.; Sutton, L.; Chen, Y.H. Claudin-7 increases chemosensitivity to cisplatin through the upregulation of caspase pathway in human NCI-H522 lung cancer cells. Cancer Sci 2013, 104, 611–618. [Google Scholar]
- Li, M.; Balch, C.; Montgomery, J.S.; Jeong, M.; Chung, J.H.; Yan, P.; Huang, T.H.; Kim, S.; Nephew, K.P. Integrated analysis of DNA methylation and gene expression reveals specific signaling pathways associated with platinum resistance in ovarian cancer. BMC Med. Genomics 2009, 2, 34. [Google Scholar]
- Fortier, A.M.; Asselin, E.; Cadrin, M. Keratin 8 and 18 loss in epithelial cancer cells increases collective cell migration and cisplatin sensitivity through claudin1 up-regulation. J. Biol. Chem 2013, 288, 11555–11571. [Google Scholar]
- Li, J.; Sherman-Baust, C.A.; Tsai-Turton, M.; Bristow, R.E.; Roden, R.B.; Morin, P.J. Claudin-containing exosomes in the peripheral circulation of women with ovarian cancer. BMC Cancer 2009, 9, 244. [Google Scholar]
- Resnick, M.B.; Konkin, T.; Routhier, J.; Sabo, E.; Pricolo, V.E. Claudin-1 is a strong prognostic indicator in stage II colonic cancer: A tissue microarray study. Mod. Pathol 2005, 18, 511–518. [Google Scholar]
- Morohashi, S.; Kusumi, T.; Sato, F.; Odagiri, H.; Chiba, H.; Yoshihara, S.; Hakamada, K.; Sasaki, M.; Kijima, H. Decreased expression of claudin-1 correlates with recurrence status in breast cancer. Int. J. Mol. Med 2007, 20, 139–143. [Google Scholar]
- Kolokytha, P.; Yiannou, P.; Keramopoulos, D.; Kolokythas, A.; Nonni, A.; Patsouris, E.; Pavlakis, K. Claudin-3 and Claudin-4: Distinct prognostic significance in triple-negative and luminal breast cancer. Appl. Immunohistochem. Mol. Morphol. 2013. [Google Scholar] [CrossRef]
- Lanigan, F.; McKiernan, E.; Brennan, D.J.; Hegarty, S.; Millikan, R.C.; McBryan, J.; Jirstrom, K.; Landberg, G.; Martin, F.; Duffy, M.J.; et al. Increased claudin-4 expression is associated with poor prognosis and high tumour grade in breast cancer. Int. J. Cancer 2009, 124, 2088–2097. [Google Scholar]
- Bernardi, M.A.; Logullo, A.F.; Pasini, F.S.; Nonogaki, S.; Blumke, C.; Soares, F.A.; Brentani, M.M. Prognostic significance of CD24 and claudin-7 immunoexpression in ductal invasive breast cancer. Oncol. Rep 2012, 27, 28–38. [Google Scholar]
- Yamamoto, T.; Oshima, T.; Yoshihara, K.; Yamanaka, S.; Nishii, T.; Arai, H.; Inui, K.; Kaneko, T.; Nozawa, A.; Woo, T.; et al. Reduced expression of claudin-7 is associated with poor outcome in non-small cell lung cancer. Oncol. Lett 2010, 1, 501–505. [Google Scholar]
- Yoshizawa, K.; Nozaki, S.; Kato, A.; Hirai, M.; Yanase, M.; Yoshimoto, T.; Kimura, I.; Sugiura, S.; Okamune, A.; Kitahara, H.; et al. Loss of claudin-7 is a negative prognostic factor for invasion and metastasis in oral squamous cell carcinoma. Oncol. Rep 2013, 29, 445–450. [Google Scholar]
- Cheung, S.T.; Leung, K.L.; Ip, Y.C.; Chen, X.; Fong, D.Y.; Ng, I.O.; Fan, S.T.; So, S. Claudin-10 expression level is associated with recurrence of primary hepatocellular carcinoma. Clin. Cancer Res 2005, 11, 551–556. [Google Scholar]
- Ohtani, S.; Terashima, M.; Satoh, J.; Soeta, N.; Saze, Z.; Kashimura, S.; Ohsuka, F.; Hoshino, Y.; Kogure, M.; Gotoh, M. Expression of tight-junction-associated proteins in human gastric cancer: Downregulation of claudin-4 correlates with tumor aggressiveness and survival. Gastric Cancer 2009, 12, 43–51. [Google Scholar]
- Romani, C.; Comper, F.; Bandiera, E.; Ravaggi, A.; Bignotti, E.; Tassi, R.A.; Pecorelli, S.; Santin, A.D. Development and characterization of a human single-chain antibody fragment against claudin-3: A novel therapeutic target in ovarian and uterine carcinomas. Am. J. Obstet. Gynecol 2009, 201, e71–e79. [Google Scholar]
- Suzuki, M.; Kato-Nakano, M.; Kawamoto, S.; Furuya, A.; Abe, Y.; Misaka, H.; Kimoto, N.; Nakamura, K.; Ohta, S.; Ando, H. Therapeutic antitumor efficacy of monoclonal antibody against Claudin-4 for pancreatic and ovarian cancers. Cancer Sci 2009, 100, 1623–1630. [Google Scholar]
- Kato-Nakano, M.; Suzuki, M.; Kawamoto, S.; Furuya, A.; Ohta, S.; Nakamura, K.; Ando, H. Characterization and evaluation of the antitumour activity of a dual-targeting monoclonal antibody against claudin-3 and claudin-4. Anticancer Res 2010, 30, 4555–4562. [Google Scholar]
- Yuan, X.; Lin, X.; Manorek, G.; Kanatani, I.; Cheung, L.H.; Rosenblum, M.G.; Howell, S.B. Recombinant CPE fused to tumor necrosis factor targets human ovarian cancer cells expressing the claudin-3 and claudin-4 receptors. Mol. Cancer Ther 2009, 8, 1906–1915. [Google Scholar]
| Gene | Chromosomal location | Transcript | Protein size (amino acids) | Molecular weight (Da) |
|---|---|---|---|---|
| CLDN1 | 3q28–q29 | NM_021101.3 | 211 | 22,744 |
| CLDN2 | Xq22.3–q23 | NM_020384.2 | 230 | 24,549 |
| CLDN3 | 7q11.23 | NM_001306.3 | 220 | 23,319 |
| CLDN4 | 7q11.23 | NM_001305.3 | 209 | 22,077 |
| CLDN5 | 22q11.21 | NM_001130861.1 | 218 | 23,147 |
| NM_003277.3 | ||||
| CLDN6 | 16p13.3 | NM_021195.4 | 220 | 23,292 |
| CLDN7 | 17p13 | NM_001307.4 | 211 | 22,390 |
| 158 | 16,837 | |||
| CLDN8 | 21q22.11 | NM_199328.1 | 225 | 24,845 |
| CLDN9 | 16p13.3 | NM_020982.2 | 217 | 22,848 |
| CLDN10 | 13q31–q34 | NM_182848.2 | 226 | 24,251 |
| NM_006984.3 | 228 | 24,488 | ||
| CLDN11 | 3q26.2–q26.3 | NM_005602.5 | 207 | 21,993 |
| CLDN12 | 7q21 | NM_012129.2 | 244 | 27,110 |
| CLDN14 | 21q22.3 | NM_144492.1 | 239 | 25,699 |
| NM_012130.2 | ||||
| CLDN15 | 7q11.22 | NM_014343.1 | 228 | 24,356 |
| CLDN16 | 3q28 | NM_006580.2 | 305 | 33,836 |
| CLDN17 | 21q22.11 | NM_012131.1 | 224 | 24,603 |
| CLDN18 | 3q22.3 | NM_001002026.2 | 261 | 27,856 |
| NM_016369.3 | 261 | 27,720 | ||
| CLDN19 | 1p34.2 | NM_001123395.1 | 224 | 23,229 |
| NM_148960.2 | 211 | 22,077 | ||
| CLDN20 | 6q25 | NM_001001346.2 | 219 | 23,515 |
| CLDN21 | 4q35.1 | NA | NA | NA |
| CLDN22 | 4q35.1 | NM_001111319.1 | 220 | 24,509 |
| CLDN23 | 8p23.1 | NM_194284.2 | 292 | 31,915 |
| CLDN24 | 4q35.1 | XM_001714660.1 | 205 | 22,802 |
| Cancer | Claudin-1 | Claudin-2 | Claudin-3 | Claudin-4 | Claudin-7 | Claudin-11 |
|---|---|---|---|---|---|---|
| Breast | Down [39] (invasive breast carcinoma) Up [13] (basal type of high-grade invasive ductal breast carcinoma) | Down [42,43] | Up [44] | Up [44] Up [13,40] (basal-like type of high-grade invasive ductal breast carcinoma) Down [39] (invasive ductal breast carcinoma of grade 1) | Down [45] (invasive ductal breast carcinoma) Up [13] (luminal type of high-grade invasive ductal breast carcinoma) | |
| Cervical | Up [46] | Up [46] | Up [46] | Up [46] | ||
| Colorectal | Up [47,48] | Up [47] | Up [47] | |||
| Esophageal | Up [49] | Up [49] | Down [50] | |||
| Gastric | Up [41] | Up [41] | Up [41,51] | Up [52] | Down [53] | |
| Liver | Down [54] | |||||
| Melanoma | Up [55] | |||||
| Ovarian | Up [56–61] (normal: ovarian surface epithelium) Down [64] (normal: fallopian tube) | Up [56,57,60–62] Down [64] (normal: fallopian tube) | Up [63] | |||
| Prostate | Down [65] | Down [42] | Up [66] | Up [66] | Down [65] | |
| Pancreatic | Up [67,68] | |||||
| Uterine | Up [69] | Up [69] |
| Claudins | Cancer | Function | In vitro or in vivo | Role | References |
|---|---|---|---|---|---|
| Claudin-1 | Breast | Increase of cell migration | In vitro | Cancer promoting | [88] |
| Breast | Anti-apoptotic effect | In vitro | Cancer promoting | [89] | |
| Colon | Increase of invasion and metastatic behavior | In vitro & in vivo | Cancer promoting | [48] | |
| Liver | Increase of invasion | In vitro | Cancer promoting | [90] | |
| Liver | Induction of EMT | In vitro | Cancer promoting | [91] | |
| Melanoma | Increase of cell motility and invasion | In vitro | Cancer promoting | [55] | |
| Oral | Increase of invasion | In vitro | Cancer promoting | [92] | |
| Gastric | Inhibition of tumorigenicity | In vivo | Tumor suppressive | [72] | |
| Lung | Inhibition of cell migration and invasion, in vivo metastasis | In vitro & in vivo | Tumor suppressive | [93] | |
| Claudin-3 | Ovarian | Increase of invasion | In vitro | Cancer promoting | [94] |
| Ovarian | Promoting in vivo tumor growth and metastasis | In vivo | Cancer promoting | [95] | |
| Ovarian | Inhibition of in vivo tumor growth and metastasis | In vitro & in vivo | Tumor suppressive | [64] | |
| Ovarian | Suppression of EMT | In vitro & in vivo | Tumor suppressive | [96] | |
| Claudin-4 | Ovarian | Increase of invasion | In vitro | Cancer promoting | [94] |
| Ovarian | Stimulation of angiogenesis | In vitro & in vivo | Cancer promoting | [97] | |
| Gastric | Inhibition of migration and invasion | In vitro | Tumor suppressive | [7] | |
| Ovarian | Suppression of EMT | In vitro & in vivo | Tumor suppressive | [96] | |
| Pancreatic | Suppression of cell invasion and metastasis | In vitro & in vivo | Tumor suppressive | [81] | |
| Claudin-6 | Gastric | Increase of proliferation, migration and invasion | In vitro | Cancer promoting | [98] |
| Breast | Inhibition of anchorageindependent growth | In vitro | Tumor suppressive | [99] | |
| Breast | Inhibition of anchorageindependent growth, migration and invasion | In vitro | Tumor suppressive | [100] | |
| Claudin-7 | Colorectal | Increase of cell proliferation and tumorigenicity | In vitro & in vivo | Cancer promoting | [80] |
| Ovarian | Increase of invasion | In vitro | Cancer promoting | [101] | |
| Esophageal | Decrease of cell growth and invasion | In vitro | Tumor suppressive | [84] | |
| Lung | Inhibition of migration and invasion, in vivo tumor growth | In vitro & in vivo | Tumor suppressive | [102] | |
| Claudin-11 | Bladder | Inhibition of cell invasion | In vitro | Tumor suppressive | [103] |
| Gastric | Inhibition of cell invasion | In vitro | Tumor suppressive | [53] | |
| Claudins | Cancer types | Association of claudin protein expression based on IHC and patient survival | Prognosis | Independent prognostic factor | Reference |
|---|---|---|---|---|---|
| Claudin-1 | Breast | Low expression→shorter disease-free interval | Low→poor | [140] | |
| Colon | Low expression→shorter overall survival and disease-free survival | Low→poor | Independent predictor of recurrence and positive prognostic factor | [139] | |
| Colorectal | Low expression→shorter cancer-specific and disease-free survival | Low→poor | [109] | ||
| Prostate | Low expression→shorter recurrence-free survival | Low→poor | Independent predictor of recurrence | [65] | |
| Claudin-3 | Breast | Positive expression→shorter disease-free survival (triple-negative) Positive expression→longer disease-free survival (luminal) | High→poor (triple-negative) High→better (luminal) | [141] | |
| Ovarian | High expression→shorter survival | High→poor | Independent negative prognostic factor | [105] | |
| Claudin-4 | Breast | High expression→shorter cancer-specific survival, overall survival and recurrence-free survival | High→poor | Independent negative prognostic factor | [142] |
| Breast | Positive expression→longer disease-free survival (triple-negative) Positive expression→shorter disease-free survival (luminal type) | High→better (triple-negative) High→poor (luminal ) | [141] | ||
| Colorectal | Low expression shorter cancer-specific and disease-free survival | Low→poor | [109] | ||
| Esophageal | Twist1 high/claudin-4 low expression→shorter overall survival | Twist1 high/claudin-4 low→poor | Independent positive prognostic factor | [107] | |
| Gastric | Moderate to strong staining→shorter overall survival | High→poor | Independent negative prognostic factor | [41] | |
| Gastric | High membranous expression→longer overall survival | High→better | Independent positive prognostic factor | [7] | |
| Pancreatic | Low gene expression *→shorter overall survival | Low→poor | Independent positive prognostic factor | [110] | |
| Claudin-7 | Breast | Positive expression→shorter recurrence-free survival | High→poor | [143] | |
| Lung | Low expression→shorter overall survival | Low→poor | [144] | ||
| Oral | Positive expression→better prognosis | High→better | [145] | ||
| Claudin-10 | Liver | High gene expression *→shorter disease-free survival | High→poor | Independent prognostic factor for disease recurrence | [146] |
© 2013 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Kwon, M.J. Emerging Roles of Claudins in Human Cancer. Int. J. Mol. Sci. 2013, 14, 18148-18180. https://doi.org/10.3390/ijms140918148
Kwon MJ. Emerging Roles of Claudins in Human Cancer. International Journal of Molecular Sciences. 2013; 14(9):18148-18180. https://doi.org/10.3390/ijms140918148
Chicago/Turabian StyleKwon, Mi Jeong. 2013. "Emerging Roles of Claudins in Human Cancer" International Journal of Molecular Sciences 14, no. 9: 18148-18180. https://doi.org/10.3390/ijms140918148
APA StyleKwon, M. J. (2013). Emerging Roles of Claudins in Human Cancer. International Journal of Molecular Sciences, 14(9), 18148-18180. https://doi.org/10.3390/ijms140918148




