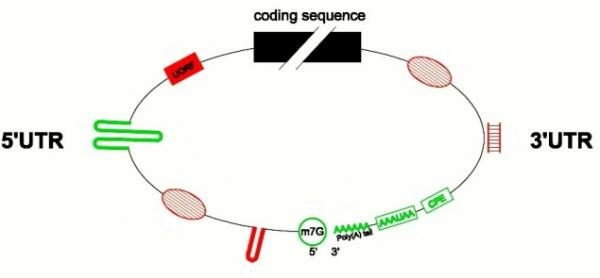
Post-transcriptional Regulation of Gene Expression in Plants during Abiotic Stress
Abstract
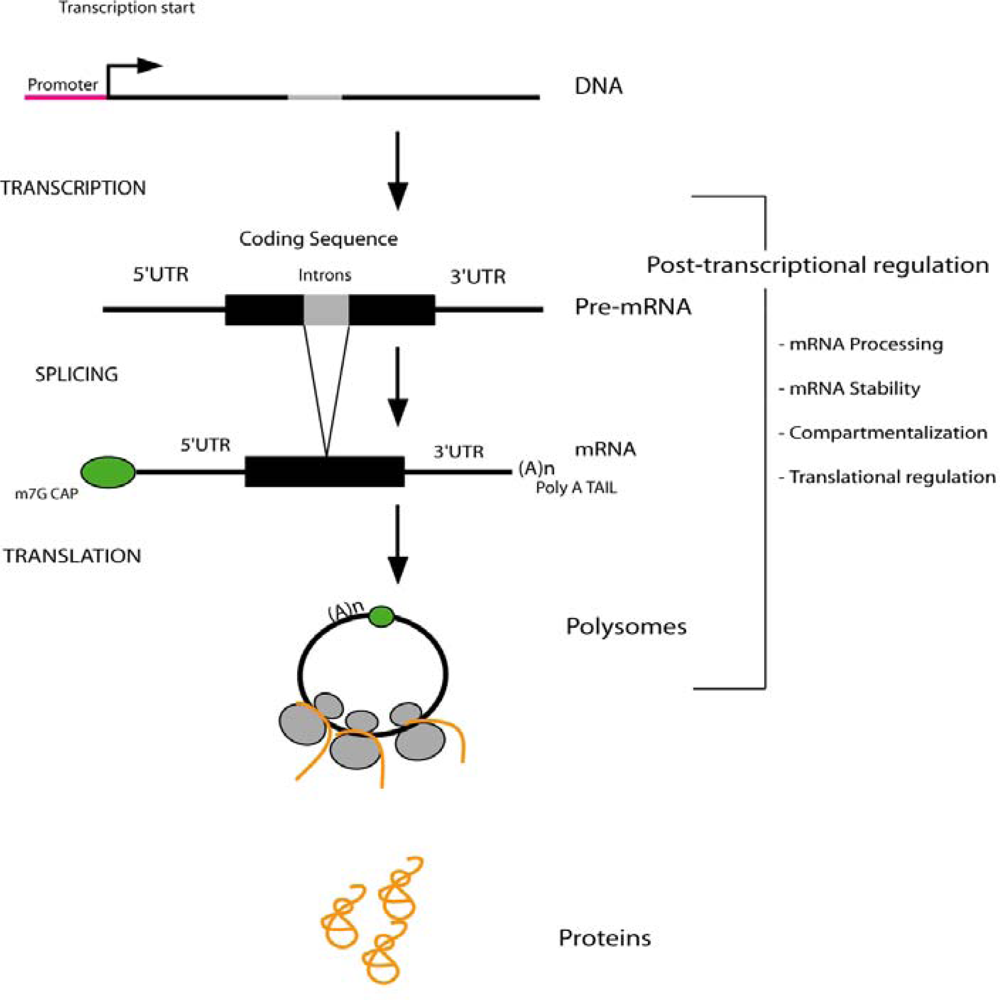
:1. Introduction
2. Regulation of mRNA Processing
3. Regulation of mRNA Stability by RNA Silencing
3.1. Implication of sRNAs in Abiotic Stress
3.2. sRNAs Up-Regulated by Stress
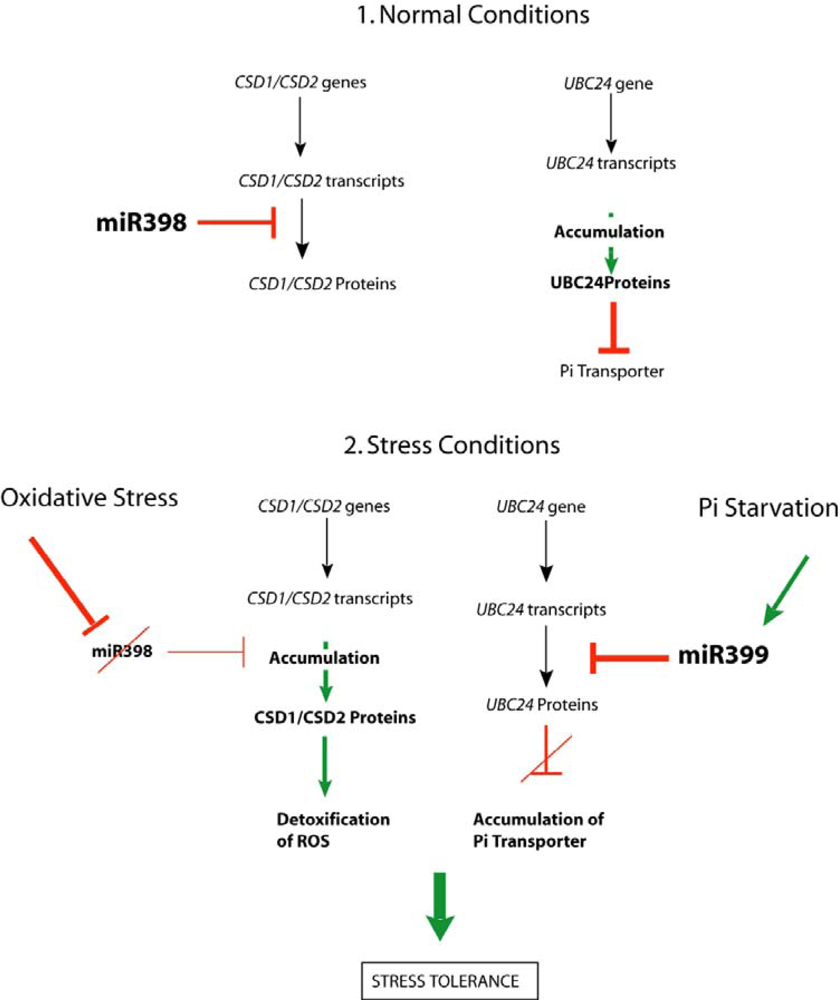
3.2.1. miR399 and Phosphate Starvation
3.2.2. Natural Antisense sRNAs and Salt Stress
3.3. sRNAs Down Regulated by Stress (miR398)
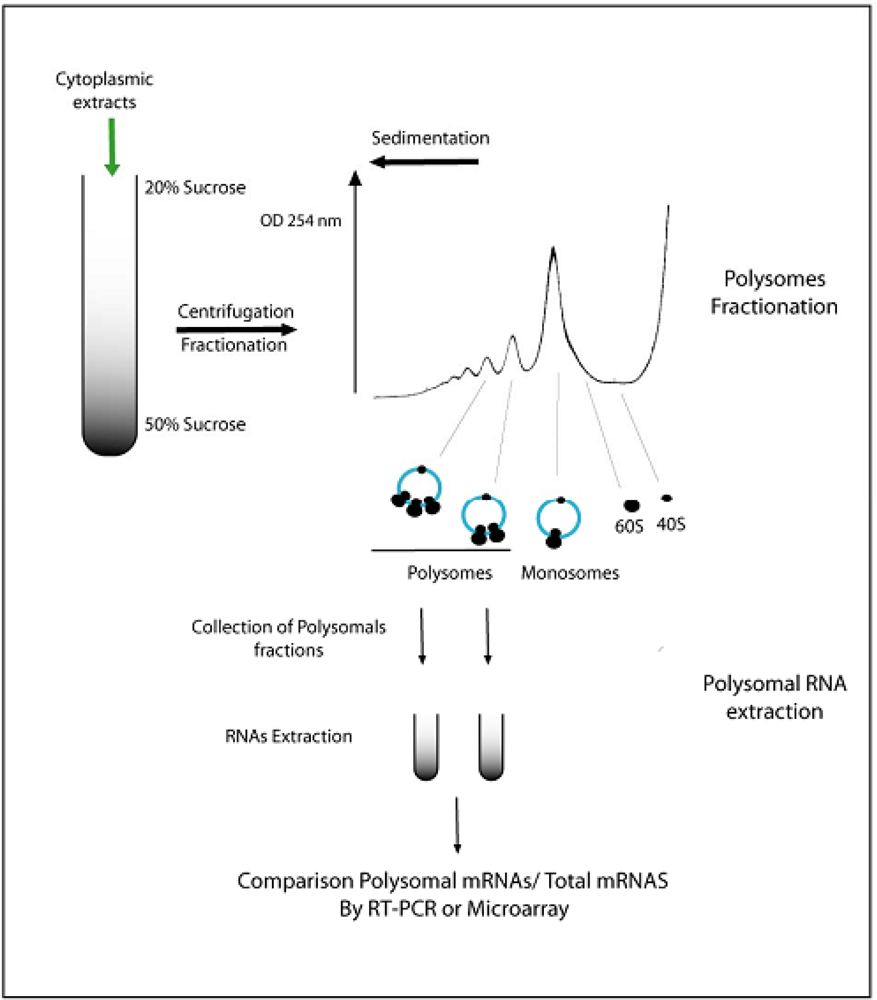
4. General Evidences of Translational Regulations in Plants
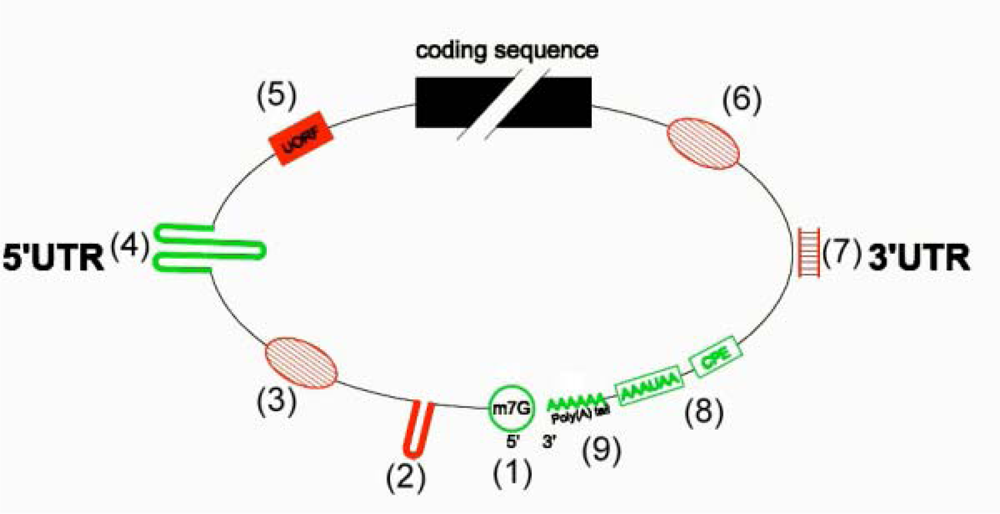
5. Involvement of Untranslated Regions (UTRs) in Translational Regulations
5.1. Upstream Open Reading Frames (uORFs) in Plants
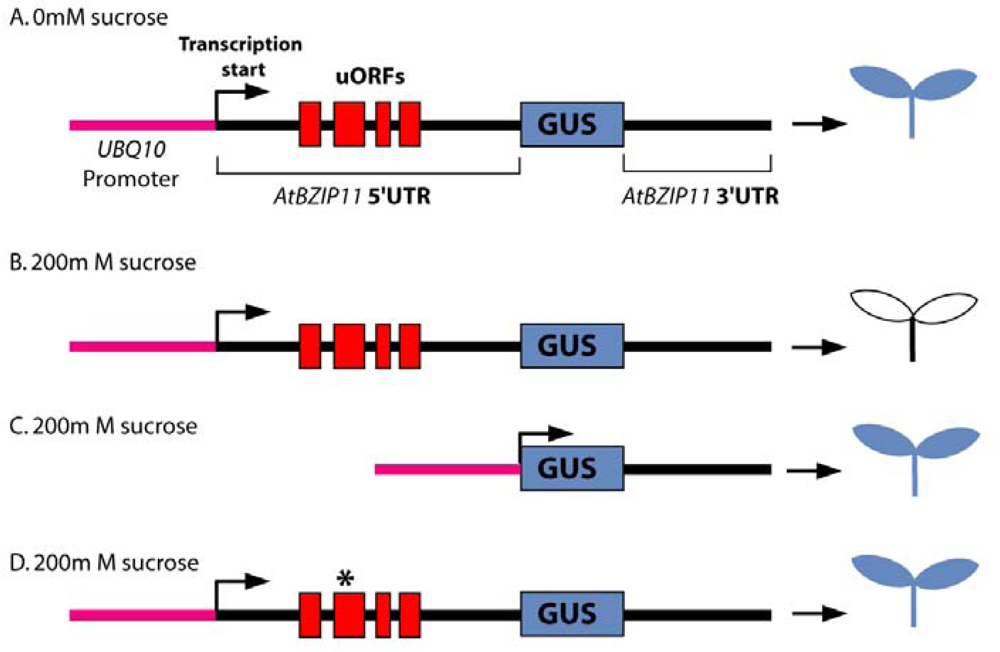
5.2. Examples of Translational Regulation Mediated by uORF
6. Compartmentalization of mRNAs in the Cytoplasm
7. Conclusions
Acknowledgments
References and Notes
- Matlin, AJ; Clark, F; Smith, CW. Understanding alternative splicing: towards a cellular code. Nat. Rev. Mol. Cell Biol 2005, 6, 386–398. [Google Scholar]
- Reddy, AS. Alternative splicing of pre-messenger RNAs in plants in the genomic era. Annu. Rev. Plant Biol 2007, 58, 267–294. [Google Scholar]
- Palusa, SG; Ali, GS; Reddy, AS. Alternative splicing of pre-mRNAs of Arabidopsis serine/arginine-rich proteins: regulation by hormones and stresses. Plant J 2007, 49, 1091–1107. [Google Scholar]
- Lee, BH; Kapoor, A; Zhu, J; Zhu, JK. STABILIZED1, a stress-upregulated nuclear protein, is required for pre-mRNA splicing, mRNA turnover, and stress tolerance in Arabidopsis. Plant Cell 2006, 18, 1736–1749. [Google Scholar]
- Lorkovic, ZJ; Barta, A. Genome analysis: RNA recognition motif (RRM) and K homology (KH) domain RNA-binding proteins from the flowering plant Arabidopsis thaliana. Nucleic Acids Res 2002, 30, 623–635. [Google Scholar]
- Kim, JY; Park, SJ; Jang, B; Jung, CH; Ahn, SJ; Goh, CH; Cho, K; Han, O; Kang, H. Functional characterization of a glycine-rich RNA-binding protein 2 in Arabidopsis thaliana under abiotic stress conditions. Plant J 2007, 50, 439–451. [Google Scholar]
- Zhu, J; Dong, CH; Zhu, JK. Interplay between cold-responsive gene regulation, metabolism and RNA processing during plant cold acclimation. Curr. Opin. Plant Biol 2007, 10, 290–295. [Google Scholar]
- Lambermon, MH; Simpson, GG; Wieczorek Kirk, DA; Hemmings-Mieszczak, M; Klahre, U; Filipowicz, W. UBP1, a novel hnRNP-like protein that functions at multiple steps of higher plant nuclear pre-mRNA maturation. EMBO J 2000, 19, 1638–1649. [Google Scholar]
- Lambermon, MH; Fu, Y; Wieczorek Kirk, DA; Dupasquier, M; Filipowicz, W; Lorkovic, ZJ. UBA1 and UBA2, two proteins that interact with UBP1, a multifunctional effector of pre-mRNA maturation in plants. Mol. Cell Biol 2002, 22, 4346–4357. [Google Scholar]
- Li, J; Kinoshita, T; Pandey, S; Ng, CK; Gygi, SP; Shimazaki, K; Assmann, SM. Modulation of an RNA-binding protein by abscisic-acid-activated protein kinase. Nature 2002, 418, 793–797. [Google Scholar]
- Li, J; Wang, XQ; Watson, MB; Assmann, SM. Regulation of abscisic acid-induced stomatal closure and anion channels by guard cell AAPK kinase. Science 2000, 287, 300–303. [Google Scholar]
- Zhang, J; Addepalli, B; Yun, KY; Hunt, AG; Xu, R; Rao, S; Li, QQ; Falcone, DL. A polyadenylation factor subunit implicated in regulating oxidative signaling in Arabidopsis thaliana. PLoS ONE 2008, 3, e2410. [Google Scholar]
- Addepalli, B; Hunt, AG. A novel endonuclease activity associated with the Arabidopsis ortholog of the 30-kDa subunit of cleavage and polyadenylation specificity factor. Nucleic Acids Res 2007, 35, 4453–4463. [Google Scholar]
- Kidner, CA; Martienssen, RA. The role of ARGONAUTE1 (AGO1) in meristem formation and identity. Dev. Biol 2005, 280, 504–517. [Google Scholar]
- Brodersen, P; Voinnet, O. The diversity of RNA silencing pathways in plants. Trends Genet 2006, 22, 268–280. [Google Scholar]
- Brodersen, P; Sakvarelidze-Achard, L; Bruun-Rasmussen, M; Dunoyer, P; Yamamoto, YY; Sieburth, L; Voinnet, O. Widespread translational inhibition by plant miRNAs and siRNAs. Science 2008, 320, 1185–1190. [Google Scholar]
- Lanet, E; Delannoy, E; Sormani, R; Floris, M; Brodersen, P; Crete, P; Voinnet, O; Robaglia, C. Biochemical Evidence for Translational Repression by Arabidopsis MicroRNAs. Plant Cell 2009. 10.1105/tpc.108.063412. [Google Scholar]
- Voinnet, O. Origin, biogenesis, and activity of plant microRNAs. Cell 2009, 136, 669–687. [Google Scholar]
- Jones-Rhoades, MW; Bartel, DP. Computational identification of plant microRNAs and their targets, including a stress-induced miRNA. Mol. Cell 2004, 14, 787–799. [Google Scholar]
- Sunkar, R; Zhu, JK. Novel and stress-regulated microRNAs and other small RNAs from Arabidopsis. Plant Cell 2004, 16, 2001–2019. [Google Scholar]
- Liu, HH; Tian, X; Li, YJ; Wu, CA; Zheng, CC. Microarray-based analysis of stress-regulated microRNAs in Arabidopsis thaliana. RNA 2008, 14, 836–843. [Google Scholar]
- Zhou, X; Wang, G; Sutoh, K; Zhu, JK; Zhang, W. Identification of cold-inducible microRNAs in plants by transcriptome analysis. Biochim. Biophys. Acta 2008, 1779, 780–788. [Google Scholar]
- Abdel-Ghany, SE; Pilon, M. MicroRNA-mediated systemic down-regulation of copper protein expression in response to low copper availability in Arabidopsis. J. Biol. Chem 2008, 283, 15932–15945. [Google Scholar]
- Kraft, E; Stone, SL; Ma, L; Su, N; Gao, Y; Lau, OS; Deng, XW; Callis, J. Genome analysis and functional characterization of the E2 and RING-type E3 ligase ubiquitination enzymes of Arabidopsis. Plant Physiol 2005, 139, 1597–1611. [Google Scholar]
- Chiou, TJ; Aung, K; Lin, SI; Wu, CC; Chiang, SF; Su, CL. Regulation of phosphate homeostasis by MicroRNA in Arabidopsis. Plant Cell 2006, 18, 412–421. [Google Scholar]
- Fujii, H; Chiou, TJ; Lin, SI; Aung, K; Zhu, JK. A miRNA involved in phosphate-starvation response in Arabidopsis. Curr. Biol 2005, 15, 2038–2043. [Google Scholar]
- Bari, R; Datt Pant, B; Stitt, M; Scheible, WR. PHO2, microRNA399, and PHR1 define a phosphate-signaling pathway in plants. Plant Physiol 2006, 141, 988–999. [Google Scholar]
- Borsani, O; Zhu, J; Verslues, PE; Sunkar, R; Zhu, JK. Endogenous siRNAs derived from a pair of natural cis-antisense transcripts regulate salt tolerance in Arabidopsis. Cell 2005, 123, 1279–1291. [Google Scholar]
- Wang, L; Wessler, SR. Role of mRNA secondary structure in translational repression of the maize transcriptional activator Lc(1,2). Plant Physiol 2001, 125, 1380–1387. [Google Scholar]
- Wang, XJ; Gaasterland, T; Chua, NH. Genome-wide prediction and identification of cis-natural antisense transcripts in Arabidopsis thaliana. Genome Biol 2005, 6, R30. [Google Scholar]
- Sunkar, R; Kapoor, A; Zhu, JK. Posttranscriptional induction of two Cu/Zn superoxide dismutase genes in Arabidopsis is mediated by downregulation of miR398 and important for oxidative stress tolerance. Plant Cell 2006, 18, 2051–2065. [Google Scholar]
- Mittler, R. Oxidative stress, antioxidants and stress tolerance. Trends Plant Sci 2002, 7, 405–410. [Google Scholar]
- Kawaguchi, R; Bailey-Serres, J. Regulation of translational initiation in plants. Curr. Opin. Plant Biol 2002, 5, 460–465. [Google Scholar]
- Noll, H. The discovery of polyribosomes. Bioessays 2008, 30, 1220–1234. [Google Scholar]
- Horiguchi, G; Fuse, T; Kawakami, N; Kodama, H; Iba, K. Temperature-dependent translational regulation of the ER omega-3 fatty acid desaturase gene in wheat root tips. Plant J 2000, 24, 805–813. [Google Scholar]
- Hua, XJ; Van de Cotte, B; Van Montagu, M; Verbruggen, N. The 5’ untranslated region of the At-P5R gene is involved in both transcriptional and post-transcriptional regulation. Plant J 2001, 26, 157–169. [Google Scholar]
- Wood, AJ; Joel Duff, R; Oliver, MJ. The translational apparatus of Tortula ruralis: polysomal retention of transcripts encoding the ribosomal proteins RPS14, RPS16 and RPL23 in desiccated and rehydrated gametophytes. J. Exp. Bot 2000, 51, 1655–1662. [Google Scholar]
- Bailey-Serres, J. Selective translation of cytoplasmic mRNAs in plants. Trends Plant Sci 1999, 4, 142–148. [Google Scholar]
- Nicolai, M; Roncato, MA; Canoy, AS; Rouquie, D; Sarda, X; Freyssinet, G; Robaglia, C. Large-scale analysis of mRNA translation states during sucrose starvation in arabidopsis cells identifies cell proliferation and chromatin structure as targets of translational control. Plant Physiol 2006, 141, 663–673. [Google Scholar]
- Kawaguchi, R; Girke, T; Bray, EA; Bailey-Serres, J. Differential mRNA translation contributes to gene regulation under non-stress and dehydration stress conditions in Arabidopsis thaliana. Plant J 2004, 38, 823–839. [Google Scholar]
- Branco-Price, C; Kaiser, KA; Jang, CJ; Larive, CK; Bailey-Serres, J. Selective mRNA translation coordinates energetic and metabolic adjustments to cellular oxygen deprivation and reoxygenation in Arabidopsis thaliana. Plant J 2008, 56, 743–755. [Google Scholar]
- Kawaguchi, R; Bailey-Serres, J. mRNA sequence features that contribute to translational regulation in Arabidopsis. Nucleic Acids Res 2005, 33, 955–965. [Google Scholar]
- Bailey-Serres, J; Dawe, RK. Both 5’ and 3’ sequences of maize adh1 mRNA are required for enhanced translation under low-oxygen conditions. Plant Physiol 1996, 112, 685–695. [Google Scholar]
- Mardanova, ES; Zamchuk, LA; Skulachev, MV; Ravin, NV. The 5’ untranslated region of the maize alcohol dehydrogenase gene contains an internal ribosome entry site. Gene 2008, 420, 11–16. [Google Scholar]
- Pitto, L; Gallie, DR; Walbot, V. Role of the Leader Sequence during Thermal Repression of Translation in Maize, Tobacco, and Carrot Protoplasts. Plant Physiol 1992, 100, 1827–1833. [Google Scholar]
- Dickey, LF; Nguyen, TT; Allen, GC; Thompson, WF. Light modulation of ferredoxin mRNA abundance requires an open reading frame. Plant Cell 1994, 6, 1171–1176. [Google Scholar]
- Dickey, LF; Petracek, ME; Nguyen, TT; Hansen, ER; Thompson, WF. Light regulation of Fed-1 mRNA requires an element in the 5’ untranslated region and correlates with differential polyribosome association. Plant Cell 1998, 10, 475–484. [Google Scholar]
- Wiese, A; Elzinga, N; Wobbes, B; Smeekens, S. A conserved upstream open reading frame mediates sucrose-induced repression of translation. Plant Cell 2004, 16, 1717–1729. [Google Scholar]
- Lovett, PS; Rogers, EJ. Ribosome regulation by the nascent peptide. Microbiol. Rev 1996, 60, 366–385. [Google Scholar]
- Abastado, JP; Miller, PF; Jackson, BM; Hinnebusch, AG. Suppression of ribosomal reinitiation at upstream open reading frames in amino acid-starved cells forms the basis for GCN4 translational control. Mol. Cell Biol 1991, 11, 486–496. [Google Scholar]
- Meijer, HA; Thomas, AA. Control of eukaryotic protein synthesis by upstream open reading frames in the 5’-untranslated region of an mRNA. Biochem. J 2002, 367, 1–11. [Google Scholar]
- Kozak, M. Pushing the limits of the scanning mechanism for initiation of translation. Gene 2002, 299, 1–34. [Google Scholar]
- Wang, XQ; Rothnagel, JA. 5’-untranslated regions with multiple upstream AUG codons can support low-level translation via leaky scanning and reinitiation. Nucleic Acids Res 2004, 32, 1382–1391. [Google Scholar]
- Franceschetti, M; Hanfrey, C; Scaramagli, S; Torrigiani, P; Bagni, N; Burtin, D; Michael, AJ. Characterization of monocot and dicot plant S-adenosyl-l-methionine decarboxylase gene families including identification in the mRNA of a highly conserved pair of upstream overlapping open reading frames. Biochem. J 2001, 353, 403–409. [Google Scholar]
- Locatelli, F; Magnani, E; Vighi, C; Lanzanova, C; Coraggio, I. Inhibitory effect of myb7 uORF on downstream gene expression in homologous (rice) and heterologous (tobacco) systems. Plant Mol. Biol 2002, 48, 309–318. [Google Scholar]
- Lohmer, S; Maddaloni, M; Motto, M; Salamini, F; Thompson, RD. Translation of the mRNA of the maize transcriptional activator Opaque-2 is inhibited by upstream open reading frames present in the leader sequence. Plant Cell 1993, 5, 65–73. [Google Scholar]
- Nishimura, T; Wada, T; Yamamoto, KT; Okada, K. The Arabidopsis STV1 protein, responsible for translation reinitiation, is required for auxin-mediated gynoecium patterning. Plant Cell 2005, 17, 2940–2953. [Google Scholar]
- Hayden, CA; Jorgensen, RA. Identification of novel conserved peptide uORF homology groups in Arabidopsis and rice reveals ancient eukaryotic origin of select groups and preferential association with transcription factor-encoding genes. BMC Biol 2007, 5, 32. [Google Scholar]
- Tran, MK; Schultz, CJ; Baumann, U. Conserved upstream open reading frames in higher plants. BMC Genomics 2008, 9, 361. [Google Scholar]
- Pegg, AE; Stanley, B; Pajunen, A; Crozat, A; Janne, OA. Properties of human and rodent S-adenosylmethionine decarboxylase. Adv. Exp. Med. Biol 1988, 250, 101–109. [Google Scholar]
- Hanfrey, C; Franceschetti, M; Mayer, MJ; Illingworth, C; Michael, AJ. Abrogation of upstream open reading frame-mediated translational control of a plant S-adenosylmethionine decarboxylase results in polyamine disruption and growth perturbations. J. Biol. Chem 2002, 277, 44131–44139. [Google Scholar]
- Hu, WW; Gong, H; Pua, EC. The pivotal roles of the plant S-adenosylmethionine decarboxylase 5’ untranslated leader sequence in regulation of gene expression at the transcriptional and posttranscriptional levels. Plant Physiol 2005, 138, 276–286. [Google Scholar]
- Rook, F; Gerrits, N; Kortstee, A; van Kampen, M; Borrias, M; Weisbeek, P; Smeekens, S. Sucrose-specific signalling represses translation of the Arabidopsis ATB2 bZIP transcription factor gene. Plant J 1998, 15, 253–263. [Google Scholar]
- Duret, L; Mouchiroud, D; Gautier, C. Statistical analysis of vertebrate sequences reveals that long genes are scarce in GC-rich isochores. J. Mol. Evol 1995, 40, 308–317. [Google Scholar]
- Rahmani, F; Hummel, M; Schuurmans, J; Wiese-Klinkenberg, A; Smeekens, S; Hanson, J. Sucrose control of translation mediated by a uORF encoded peptide. Plant Physiol 2009. 10.1104/pp.109.136036.. [Google Scholar]
- Nover, L; Scharf, KD; Neumann, D. Cytoplasmic heat shock granules are formed from precursor particles and are associated with a specific set of mRNAs. Mol. Cell Biol 1989, 9, 1298–1308. [Google Scholar]
- Weber, C; Nover, L; Fauth, M. Plant stress granules and mRNA processing bodies are distinct from heat stress granules. Plant J 2008, 56, 517–530. [Google Scholar]
- Anderson, P; Kedersha, N. RNA granules: post-transcriptional and epigenetic modulators of gene expression. Nat. Rev. Mol. Cell Biol 2009, 10, 430–436. [Google Scholar]
- Marx, J. Molecular biology. P-bodies mark the spot for controlling protein production. Science 2005, 310, 764–765. [Google Scholar]
- Combier, JP; de Billy, F; Gamas, P; Niebel, A; Rivas, S. Trans-regulation of the expression of the transcription factor MtHAP2-1 by a uORF controls root nodule development. Genes Dev 2008, 22, 1549–1559. [Google Scholar]
- Combier, JP; Frugier, F; de Billy, F; Boualem, A; El-Yahyaoui, F; Moreau, S; Vernie, T; Ott, T; Gamas, P; Crespi, M; Niebel, A. MtHAP2-1 is a key transcriptional regulator of symbiotic nodule development regulated by microRNA169 in Medicago truncatula. Genes Dev 2006, 20, 3084–3088. [Google Scholar]

© 2009 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Floris, M.; Mahgoub, H.; Lanet, E.; Robaglia, C.; Menand, B. Post-transcriptional Regulation of Gene Expression in Plants during Abiotic Stress. Int. J. Mol. Sci. 2009, 10, 3168-3185. https://doi.org/10.3390/ijms10073168
Floris M, Mahgoub H, Lanet E, Robaglia C, Menand B. Post-transcriptional Regulation of Gene Expression in Plants during Abiotic Stress. International Journal of Molecular Sciences. 2009; 10(7):3168-3185. https://doi.org/10.3390/ijms10073168
Chicago/Turabian StyleFloris, Maïna, Hany Mahgoub, Elodie Lanet, Christophe Robaglia, and Benoît Menand. 2009. "Post-transcriptional Regulation of Gene Expression in Plants during Abiotic Stress" International Journal of Molecular Sciences 10, no. 7: 3168-3185. https://doi.org/10.3390/ijms10073168
APA StyleFloris, M., Mahgoub, H., Lanet, E., Robaglia, C., & Menand, B. (2009). Post-transcriptional Regulation of Gene Expression in Plants during Abiotic Stress. International Journal of Molecular Sciences, 10(7), 3168-3185. https://doi.org/10.3390/ijms10073168