Differential Analysis of Sesquiterpenoids of Atractylodes macrocephalus from Different Origins Based on Transcriptomics
Abstract
1. Introduction
2. Results and Analysis
2.1. Transcriptome Data Assembly
2.2. Unigene Annotation and Classification
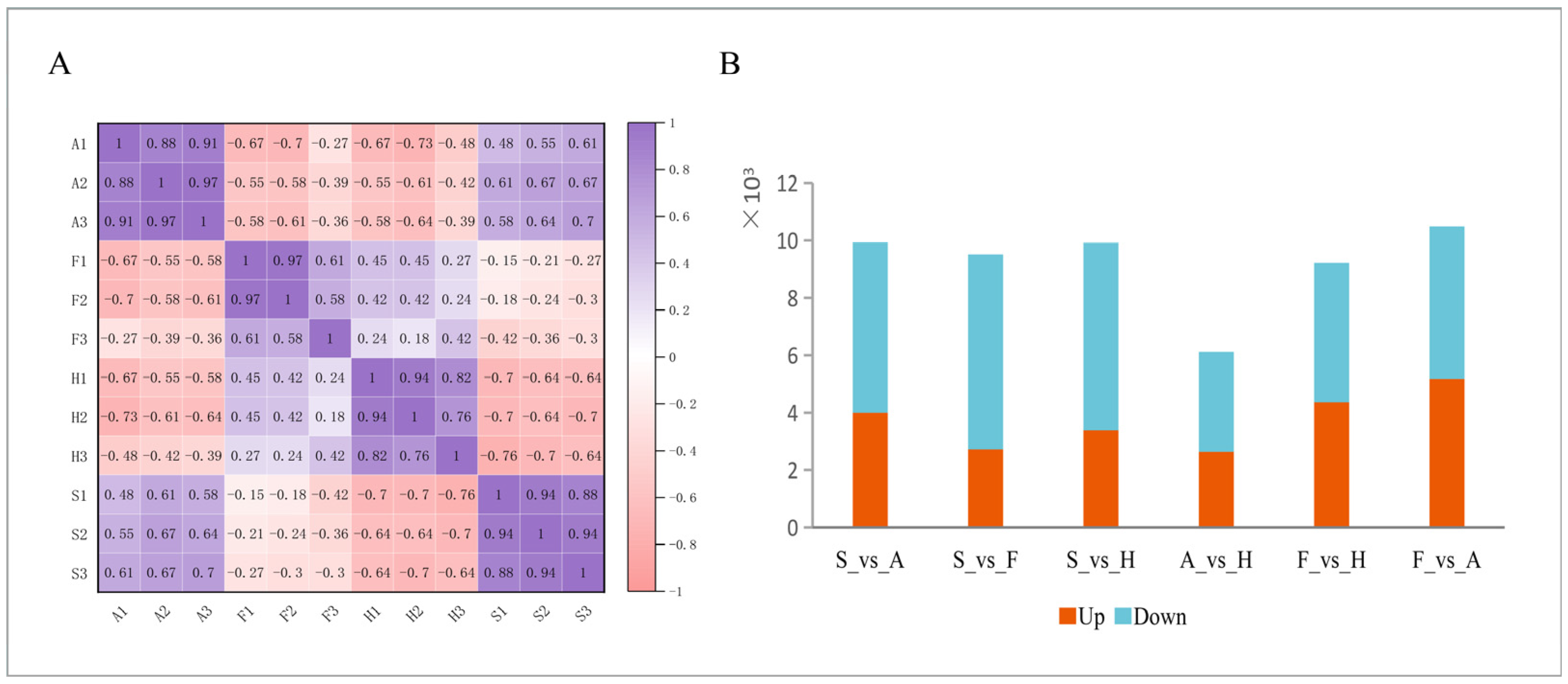
2.3. Different Origins Correlation Analysis Between Samples
2.4. Differential Expression Analysis
2.5. Gene Ontology Analysis of Differentially Expressed Genes
2.6. Kyoto Encyclopedia of Genes and Genomes Metabolic Pathway Analysis of Differentially Expressed Genes
2.7. Integrated Analysis of Terpenoid Biosynthesis Transcriptome
2.8. Transcriptome Validation
2.9. Correlation Between Terpenoid Compound Content
3. Discussion and Outlook
4. Materials and Methods
4.1. Sample Collection
4.2. RNA Extraction and Library Construction
4.3. Evaluation of Sequencing Results and Transcript Splicing
4.4. Functional Gene Annotation
4.5. Analysis of Differentially Expressed Genes
4.6. Quantitative Real-Time Polymerase Chain Reaction Analysis
4.7. High-Performance Liquid Chromatography Analysis of Terpenoid Compounds
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| A.M. | Atractylodes macrocephala |
| bp | base pair |
| qRT-PCR | quantitative real-time polymerase chain reaction |
| HPLC | high-performance liquid chromatography |
| BLAST | Basic Local Alignment Search Tool |
| GO | gene ontology |
| NR | NCBI non-redundant protein database |
| CDD | Conserved Domain Database |
| PFAM | Protein Families Database |
| KOG | euKaryotic Orthologous Groups |
| Swissprot | Swiss-Prot Protein Sequence Database |
| NT | Nucleotide Sequence Database |
| KEGG | Kyoto Encyclopedia of Genes and Genomes |
| DEGs | differentially expressed genes |
| MVA | mevalonate |
| MEP | methylerythritol phosphate |
| IPP | isopentenyl pyrophosphate |
| FPP | farnesyl pyrophosphate |
Appendix A
| Sample | Raw Reads | Raw Bases (bp) | Total Clean Reads | Clean Bases (bp) | GC Base Ratio% | Q20 (%) | Q30 (%) |
|---|---|---|---|---|---|---|---|
| A1 | 58,513,266 | 8,776,989,900 | 56,554,110 | 8,083,340,053 | 46.90% | 98.68 | 95.09 |
| A2 | 48,741,824 | 7,311,273,600 | 47,615,740 | 6,914,157,200 | 46.41% | 98.79 | 95.39 |
| A3 | 68,116,068 | 10,217,410,200 | 66,441,512 | 9,645,298,208 | 46.44% | 98.78 | 95.38 |
| F1 | 66,118,908 | 9,917,836,200 | 63,846,718 | 8,967,518,770 | 46.07% | 98.71 | 95.17 |
| F2 | 61,448,460 | 9,217,269,000 | 59,914,550 | 8,632,686,915 | 46.24% | 98.78 | 95.37 |
| F3 | 66,995,486 | 10,049,322,900 | 64,339,162 | 9,144,040,803 | 46.44% | 98.59 | 94.81 |
| H1 | 62,200,254 | 9,330,038,100 | 60,835,706 | 8,836,916,202 | 45.20% | 98.89 | 95.68 |
| H2 | 67,874,028 | 10,181,104,200 | 66,255,006 | 9,575,996,019 | 45.12% | 98.73 | 95.20 |
| H3 | 66,325,174 | 9,948,776,100 | 64,629,984 | 9,366,665,180 | 44.98% | 98.80 | 95.40 |
| S1 | 61,571,838 | 9,235,775,700 | 60,117,158 | 8,719,798,400 | 46.40% | 98.75 | 95.29 |
| S2 | 76,311,598 | 11,446,739,700 | 74,287,334 | 10,739,613,623 | 46.40% | 98.72 | 95.18 |
| S3 | 74,683,862 | 11,202,579,300 | 72,738,186 | 10,514,272,054 | 46.49% | 98.73 | 95.22 |
References
- Sun, Z.; Cheng, D.; Liu, C.; Zhong, X.; Chen, Z.; Zhang, M. National Product of Geographical Indication of Geo-authentic Crude Drug—Xianfeng Largehead Atractylodes Rhizome. J. Li-Shizhen Tradit. Chin. Med. 2010, 21, 2650–2652. [Google Scholar] [CrossRef]
- Shin, I.S.; Jeon, W.Y.; Shin, H.K.; Cha, S.W.; Lee, M.Y. Banhabaekchulchunma-tang, a Traditional Herbal Formula, Attenuates Absolute Ethanol-Induced Gastric Injury by Enhancing the Antioxidant Status. BMC Complement. Altern. Med. 2013, 13, 170. [Google Scholar] [CrossRef] [PubMed]
- Matsumoto, H.; Matsumoto, A.; Miyata, S.; Tomogane, Y.; Minami, H.; Masuda, A.; Yamaura, I.; Yoshida, Y.; Hirata, Y. The Effect of Japanese Herbal Medicines (Kampo) Goreisan and Saireito on the Prevention of Recurrent Chronic Subdural Hematoma: A Prospective Randomized Study. Neurosurgery 2024, 94, 80–89. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.H.; Jung, H.N.; Lee, K.Y.; Kim, J.; Lee, J.C.; Jang, Y.S. Suppression of Th2-Type Immune Response-Mediated Allergic Diarrhea Following Oral Administration of Traditional Korean Medicine: Atractylodes macrocephala Koidz. Immunopharmacol. Immunotoxicol. 2005, 27, 331–343. [Google Scholar] [CrossRef]
- Commission, N.P. Pharmacopoeia of the People’s Republic of China; China Medical Science Press: Beijing, China, 2020. [Google Scholar]
- Anonymous. Shennong’s Classic of Materia Medica; Inner Mongolia People’s Publishing House: Hohhot, China, 2006. [Google Scholar]
- Liu, C.; Wang, S.; Xiang, Z.; Xu, T.; He, M.; Xue, Q.; Song, H.; Gao, P.; Cong, Z. The Chemistry and Efficacy Benefits of Polysaccharides from Atractylodes macrocephala Koidz. Front. Pharmacol. 2022, 13, 952061. [Google Scholar] [CrossRef]
- Wang, C.; Su, J.; Tian, Y.; Li, T.; Zhong, Y.; Chen, S.; Lv, G. The Rhizomes of Atractylodes macrocephala Koidz Improve Gastrointestinal Health and Pregnancy Outcomes in Pregnant Mice via Modulating Intestinal Barrier and Water-Fluid Metabolism. J. Ethnopharmacol. 2024, 326, 117971. [Google Scholar] [CrossRef]
- Choi, N.R.; Choi, W.G.; Lee, J.H.; Park, J.; Kim, Y.T.; Das, R.; Woo, J.H.; Kim, B.J. Atractylodes macrocephala Koidz Alleviates Symptoms in Zymosan-Induced Irritable Bowel Syndrome Mouse Model through TRPV1, NaV1.5, and NaV1.7 Channel Modulation. Nutrients 2024, 16, 1683. [Google Scholar] [CrossRef]
- Mazza, G.; Cottrell, T. Volatile Components of Roots, Stems, Leaves, and Flowers of Echinacea Species. J. Agric. Food Chem. 1999, 47, 3081–3085. [Google Scholar] [CrossRef]
- Luo, J.P.; Liu, Y.P.; Feng, Y.F.; Guo, X.L.; Cao, H. Two Chemotypes of Pogostemon cablin and Influence of Region of Cultivation and Harvesting Time on Volatile Oil Composition. Acta Pharm. Sin. 2003, 38, 307–310. [Google Scholar]
- Gu, S.; Li, L.; Huang, H.; Wang, B.; Zhang, T. Antitumor, Antiviral, and Anti-Inflammatory Efficacy of Essential Oils from Atractylodes macrocephala Koidz. Produced with Different Processing Methods. Molecules 2019, 24, 2956. [Google Scholar] [CrossRef]
- Wang, P.; Xie, Z.S.; Song, J.Y.; Zeng, H.H.; Dai, L.P.; E., H.C.; Ye, Z.P.; Gao, S.; Xu, J.Y.; Zhang, Z.Q. Four New Sesquiterpene Lactones from Atractylodes macrocephala and Their CREB Agonistic Activities. Fitoterapia 2020, 147, 104730. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Yu, H.; Hou, A.; Man, W.; Wang, S.; Zhang, J.; Wang, X.; Zheng, S.; Jiang, H.; Kuang, H. A Review of the Ethnopharmacology, Phytochemistry, Pharmacology, Application, Quality Control, Processing, Toxicology, and Pharmacokinetics of the Dried Rhizome of Atractylodes macrocephala. Front. Pharmacol. 2021, 12, 727154. [Google Scholar] [CrossRef] [PubMed]
- Jing, L.; Mu, H.; Ling, L.; Tong, Z. Research Progress on Chemical Constituents and Pharmacological Effects of Volatile Oil from Atractylodes macrocephala. Chin. Tradit. Herb. Drugs 2024, 46, 881–889. [Google Scholar]
- Wang, K.T.; Chen, L.G.; Chou, D.S.; Liang, W.L.; Wang, C.C. Anti-Oxidative Abilities of Essential Oils from Atractylodes ovata Rhizome. Evid. Based Complement. Altern. Med. 2011, 2011, 204892. [Google Scholar] [CrossRef]
- Tian, D.; Mengqi, S.; Yun, W. Volatile Oil from Forsythia suspense: Chemical Constituents and Pharmacological Effects. Nat. Prod. Res. Dev. 2018, 30, 1834–1842. [Google Scholar] [CrossRef]
- Kim, H.Y.; Kim, J.H. Sesquiterpenoids Isolated from the Rhizomes of Genus Atractylodes. Chem. Biodivers. 2022, 19, e202200703. [Google Scholar] [CrossRef]
- Xie, Z.; Lin, M.; He, X.; Dong, Y.; Chen, Y.; Li, B.; Chen, S.; Lv, G. Chemical Constitution, Pharmacological Effects and the Underlying Mechanism of Atractylenolides: A Review. Molecules 2023, 28, 3987. [Google Scholar] [CrossRef]
- Arizmendi, N.; Alam, S.B.; Azyat, K.; Makeiff, D.; Befus, A.D.; Kulka, M. The Complexity of Sesquiterpene Chemistry Dictates Its Pleiotropic Biologic Effects on Inflammation. Molecules 2022, 27, 2450. [Google Scholar] [CrossRef]
- Zhou, T.; Luo, X.; Zhang, C.; Xu, X.; Yu, C.; Jiang, Z.; Zhang, L.; Yuan, H.; Zheng, B.; Pi, E.; et al. Comparative Metabolomic Analysis Reveals the Variations in Taxoids and Flavonoids Among Three Taxus Species. BMC Plant Biol. 2019, 19, 529. [Google Scholar] [CrossRef]
- Zhao, X.; Yan, Y.; Zhou, W.H.; Feng, R.Z.; Shuai, Y.K.; Yang, L.; Liu, M.J.; He, X.Y.; Wei, Q. Transcriptome and Metabolome Reveal the Accumulation of Secondary Metabolites in Different Varieties of Cinnamomum longepaniculatum. BMC Plant Biol. 2022, 22, 243. [Google Scholar] [CrossRef]
- Shun, L.; Yuxing, W.; Jing, F.; Xuemei, M.; Yanli, F. Metabolomics Analysis of Terpenoid Metabolites in Leaves of Lycium barbarum from Different Origins in Ningxia. Food Ind. 2024, 45, 75–79. [Google Scholar] [CrossRef]
- Qi, Y.X.; Liu, Y.B.; Rong, W.H. RNA-Seq and Its Applications: A New Technology for Transcriptomics. Hereditas 2011, 33, 1191–1202. [Google Scholar] [CrossRef] [PubMed]
- Hai, C.T.; Luyen, N.T.; Giang, D.H.; Minh, B.Q.; Trung, N.Q.; Chinh, P.T.; Hau, D.V.; Dat, N.T. Atractylodes macrocephala Rhizomes Contain Anti-inflammatory Sesquiterpenes. Chem. Pharm. Bull. 2023, 71, 451–453. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Y.H.; Zhang, P.; Tao, Y.; Liu, Y.; Cao, G.; Zhou, L.; Yang, C.H. Banxia Baizhu Tianma Decoction Attenuates Obesity-Related Hypertension. J. Ethnopharmacol. 2021, 266, 113453. [Google Scholar] [CrossRef]
- Li, Y.; Zhang, L.; Zhang, Y.; Miao, Z.; Liu, Z.; Zhou, G.; He, J.; Ding, N.; Zhou, H.; Zhou, T.; et al. Potential Molecular Mechanism of Guiqi Baizhu Decoction in Radiation-Induced Intestinal Edema by Regulating HIF-1α, AQP4 and Na+/K+-ATPase. Phytomedicine 2022, 107, 154445. [Google Scholar] [CrossRef]
- Lemons, J.M.S.; Narrowe, A.B.; Liu, L.; Firrman, J.; Mahalak, K.K.; Van den Abbeele, P.; Baudot, A.; Deyaert, S.; Li, Y.; Yu, L.L. Impact of Baizhu, Daqingye, and Hehuanhua Extracts on the Human Gut Microbiome. Front. Cell Infect. Microbiol. 2023, 13, 1298392. [Google Scholar] [CrossRef]
- Luo, Y.; Zhu, F.; Wu, J.; Wu, J.; Wu, P.; Liu, Y. Effect of Shenling Baizhu San on Intestinal Flora in a Rat Model of Ulcerative Colitis with Spleen Deficiency and Dampness. Evid. Based Complement. Altern. Med. 2022, 2022, 9985147. [Google Scholar] [CrossRef]
- Xie, G.; Zhou, R.; Huang, L.; Zhang, S.; Tan, Z. In Vitro Biotransformation of Total Glycosides in Qiwei Baizhu Powder by the Gut Microbiota of Normal and Diarrheal Mice: Novel Insight Into the Biotransformation of Multi-Glycosides by the Gut Microbiota. Front. Chem. 2022, 10, 907886. [Google Scholar] [CrossRef]
- Alberts, B.; Johnson, A.; Lewis, J. Molecular Biology of the Cell; Garland Science: New York, NY, USA, 1983. [Google Scholar]
- Tao, H.; Lauterbach, L.; Bian, G.; Chen, R.; Hou, A.; Mori, T.; Cheng, S.; Hu, B.; Lu, L.; Mu, X.; et al. Discovery of non-squalene triterpenes. Nature 2022, 606, 414–419. [Google Scholar] [CrossRef]
- Dong, Y.; Zhang, W.; Ling, Z.; Li, J.; Bai, H.; Li, H.; Shi, L. Advances in Transcription Factors Regulating Plant Terpenoids Biosynthesis. Chin. Bull. Bot. 2020, 55, 340–350. [Google Scholar] [CrossRef]
- Chen, F.; Tholl, D.; Bohlmann, J.; Pichersky, E. The Family of Terpene Synthases in Plants: A Mid-Size Family of Genes for Specialized Metabolism That Is Highly Diversified throughout the Kingdom. Plant J. 2011, 66, 212–229. [Google Scholar] [CrossRef]
- Nagegowda, D.A.; Gupta, P. Advances in Biosynthesis, Regulation, and Metabolic Engineering of Plant Specialized Terpenoids. Plant Sci. 2020, 294, 110457. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Yang, B.; Li, M.; Li, Z.; Tu, Y.; Tang, L.; He, G. Research on Germplasm Diversity of Amomum villosum Lour. in Genuine Producing Areas. PLoS ONE 2022, 17, e0268246. [Google Scholar] [CrossRef] [PubMed]
- Zhou, T.; Wang, Y.; Qin, J.; Zhao, S.; Cao, D.; Zhu, M.; Jiang, Y. Potential Risk, Spatial Distribution, and Soil Identification of Potentially Toxic Elements in Lycium barbarum L. (Wolfberry) Fruits and Soil System in Ningxia, China. Int. J. Environ. Res. Public Health 2022, 19, 16186. [Google Scholar] [CrossRef] [PubMed]
- Vranová, E.; Coman, D.; Gruissem, W. Network Analysis of the MVA and MEP Pathways for Isoprenoid Synthesis. Annu. Rev. Plant Biol. 2013, 64, 665–700. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Zhang, W.; Liao, Y.; Ye, J.; Xu, F. Contemporary Understanding of Transcription Factor Regulation of Terpenoid Biosynthesis in Plants. Planta 2023, 259, 2. [Google Scholar] [CrossRef] [PubMed]
- Xue, Y.; Cao, X.; Chen, X.; Deng, X.; Deng, X.W.; Ding, Y.; Dong, A.; Duan, C.G.; Fang, X.; Gong, L.; et al. Epigenetics in the Modern Era of Crop Improvements. Sci. China Life Sci. 2025, 68, 1570–1609. [Google Scholar] [CrossRef]
- Tholl, D. Terpene Synthases and the Regulation, Diversity and Biological Roles of Terpene Metabolism. Curr. Opin. Plant Biol. 2006, 9, 297–304. [Google Scholar] [CrossRef]
- Liao, P.; Hemmerlin, A.; Bach, T.J.; Chye, M.L. The Potential of the Mevalonate Pathway for Enhanced Isoprenoid Production. Biotechnol. Adv. 2016, 34, 697–713. [Google Scholar] [CrossRef]
- Abbas, F.; Ke, Y.; Yu, R.; Yue, Y.; Amanullah, S.; Jahangir, M.M.; Fan, Y. Volatile Terpenoids: Multiple Functions, Biosynthesis, Modulation and Manipulation by Genetic Engineering. Planta 2017, 246, 803–816. [Google Scholar] [CrossRef]
- Zamani, Z.; Tamartash, R.; Heydari, Q.; Jelodar, Z.J. Impact of Soil Factors and Climate on Phytochemical Characteristics of Stachys lavandulifolia Vahl. Chem. Biodivers. 2025, 22, e202402352. [Google Scholar] [CrossRef] [PubMed]
- Xiong, R.; He, X.; Gao, N.; Li, Q.; Qiu, Z.; Hou, Y.; Shen, W. Soil pH Amendment Alters the Abundance, Diversity, and Composition of Microbial Communities in Two Contrasting Agricultural Soils. Microbiol. Spectr. 2024, 12, e0416523. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Yang, M.; Zhou, M.; Zhan, J.; Liu, M.; Chen, H.; Zheng, T. Effect of Rhizospheric Fe2+ on Terpenoid Biosynthesis and Accumulation within Conyza blinii H. Lév. Trop. Plants 2024, 3, E003. [Google Scholar] [CrossRef]
- Jan, R.; Asaf, S.; Numan, M.; Lubna; Kim, K.-M. Plant Secondary Metabolite Biosynthesis and Transcriptional Regulation in Response to Biotic and Abiotic Stress Conditions. Agronomy 2021, 11, 968. [Google Scholar] [CrossRef]
- Alvin, A.; Miller, K.I.; Neilan, B.A. Exploring the Potential of Endophytes from Medicinal Plants as Sources of Antimycobacterial Compounds. Microbiol. Res. 2014, 169, 483–495. [Google Scholar] [CrossRef]
- Kunyang, L.; Jiancai, X.; Hongyang, W.; Jinzhen, Q.; Xiufu, W.; Binbin, Y.; Chuanzhi, K.; Kai, S.; Yan, Z.; Lanping, G. Mechanism and Application Prospects of Microbial Fertilizers in Regulating Quality Formation of Medicinal Plants. China J. Chin. Mater. Medica 2024, 49, 912–923. [Google Scholar] [CrossRef]
- Han, J.Y.; Kim, M.J.; Ban, Y.W.; Hwang, H.S.; Choi, Y.E. The Involvement of β-Amyrin 28-Oxidase (CYP716A52v2) in Oleanane-Type Ginsenoside Biosynthesis in Panax ginseng. Plant Cell Physiol. 2013, 54, 2034–2046. [Google Scholar] [CrossRef]
- Yasumoto, S.; Seki, H.; Shimizu, Y.; Fukushima, E.O.; Muranaka, T. Functional Characterization of CYP716 Family P450 Enzymes in Triterpenoid Biosynthesis in Tomato. Front. Plant Sci. 2017, 8, 21. [Google Scholar] [CrossRef]
- Shulse, C.N.; Cole, B.J.; Ciobanu, D.; Lin, J.; Yoshinaga, Y.; Gouran, M.; Turco, G.M.; Zhu, Y.; O’Malley, R.C.; Brady, S.M.; et al. High-Throughput Single-Cell Transcriptome Profiling of Plant Cell Types. Cell Rep. 2019, 27, 2241–2247.e2244. [Google Scholar] [CrossRef]
- Chaonan, M.; Jin, Z.; Junning, Z. Quality Evaluation of the Anti-Shock and Anti-Sluggish Arrhythmia Effects of the Concocted Products of Radix et Rhizoma Pinelliae from Different Places of Origin. Pharmacol. Clin. Chin. Mater. 2014, 30, 67–70. [Google Scholar] [CrossRef]
- Zang, Y.; Li, Y.; Qiao, J.; Chen, H.; Liu, C. Mechanism of Genuineness of Glycyrrhiza uralensis Based on SNP of β-Amyrin Synthase Gene. Acta Pharm. Sin. 2015, 50, 906–909. [Google Scholar] [CrossRef]
- Kliebenstein, D.J. Plant Defense Compounds: Systems Approaches to Metabolic Analysis. Annu. Rev. Phytopathol. 2012, 50, 155–173. [Google Scholar] [CrossRef]
- Wong, M.L.; Medrano, J.F. Real-time PCR for mRNA Quantitation. Biotechniques 2005, 39, 75–85. [Google Scholar] [CrossRef]




| Categories | Transcripts | Unigenes |
|---|---|---|
| Total sequence number | 915,152 | 374,475 |
| Length ≥ 500 bp | 390,703 | 128,529 |
| Length ≥ 1000 bp | 174,160 | 49,105 |
| Min length | 201 | 201 |
| Max length | 15,628 | 15,628 |
| Total length | 630,146,258 | 215,050,079 |
| Average length | 688.57 | 574.27 |
| N50 length (bp) | 1003 | 760 |
| N90 length (bp) | 760 | 256 |
| Group | Top Five Class KEGG Pathway Names | Number |
|---|---|---|
| S vs. A | Ribosomes | 77 |
| Carbon metabolism | 52 | |
| Starch and sucrose metabolism | 43 | |
| S vs. F | Ribosomes | 256 |
| Carbon metabolism | 162 | |
| Biosynthesis of amino acids | 121 | |
| S vs. H | Ribosomes | 200 |
| Oxidative phosphorylation | 110 | |
| Carbon metabolism | 117 | |
| A vs. H | Ribosomes | 60 |
| Protein processing in the endoplasmic reticulum | 38 | |
| Carbon metabolism/RNA transport | 36 | |
| F vs. H | Oxidative phosphorylation | 67 |
| Carbon metabolism/Neurotrophin signaling pathway | 62 | |
| Ribosomes | 54 | |
| F vs. A | Ribosomes | 144 |
| Starch and sucrose metabolism | 78 | |
| Carbon metabolism | 76 |
| Key Enzymes | Gene Annotation Information | A vs. H | F vs. A | F vs. H | S vs. A | S vs. F | S vs. H |
|---|---|---|---|---|---|---|---|
| DXS | TRINITY_DN98338_c5_g1 | 15.53 | −6.54 | —— | −15.53 | —— | —— |
| TRINITY_DN118127_c2_g1 | —— | 15.75 | 15.17 | —— | —— | ||
| ISPF | TRINITY_DN102344_c2_g1 | 3.67 | —— | 3.87 | —— | —— | 3.39 |
| TRINITY_DN113213_c0_g5 | —— | —— | —— | —— | −16.44 | —— | |
| ISPS | TRINITY_DN113106_c1_g2 | −6.0 | —— | —— | 6.23 | —— | —— |
| TRINITY_DN121289_c2_g5 | 4.23 | −5.32 | —— | −4.7 | —— | —— | |
| TRINITY_DN121289_c2_g1 | 4.94 | −6.04 | —— | −5.42 | —— | —— | |
| TRINITY_DN121289_c3_g2 | 4.77 | —— | —— | −4.59 | —— | —— | |
| TRINITY_DN103012_c0_g1 | —— | 8.63 | 4.34 | 6.94 | —— | —— | |
| TRINITY_DN124089_c3_g2 | —— | −5.44 | —— | —— | —— | —— | |
| ISPE | TRINITY_DN114236_c2_g1 | —— | 3.44 | —— | —— | —— | —— |
| GCPE | TRINITY_DN119478_c1_g1 | —— | —— | 4.55 | —— | —— | 5.33 |
| FDPS | TRINITY_DN58900_c0_g1 | −13.6 | —— | −7.22 | —— | —— | —— |
| TRINITY_DN108354_c1_g1 | —— | 4.6 | 5.19 | 4.55 | —— | 5.15 | |
| TRINITY_DN90804_c0_g1 | —— | —— | —— | −3.89 | —— | —— | |
| TRINITY_DN88578_c0_g1 | —— | —— | —— | —— | —— | −12.23 | |
| ACAT | TRINITY_DN120516_c1_g3 | —— | —— | —— | 6.69 | 3.82 | —— |
| TRINITY_DN92952_c0_g1 | —— | —— | —— | —— | −6.18 | —— | |
| TRINITY_DN90745_c0_g2 | —— | —— | —— | —— | −12.82 | —— | |
| HMGCR | TRINITY_DN100174_c0_g1 | —— | —— | —— | −3.36 | —— | —— |
| TRINITY_DN112315_c0_g2 | —— | —— | —— | 14.5 | 14.5 | 14.5 | |
| MVD | TRINITY_DN102480_c1_g1 | —— | —— | —— | 3.37 | —— | —— |
| PCME | TRINITY_DN105029_c1_g2 | —— | —— | —— | −4.28 | —— | —— |
| TRINITY_DN90804_c0_g1 | —— | —— | —— | −3.98 | —— | —— | |
| STE24 | TRINITY_DN87551_c0_g1 | —— | —— | —— | —— | −4.11 | —— |
| FNTB | TRINITY_DN110526_c2_g2 | —— | —— | —— | —— | —— | 14.1 |
| Gene | Forward Primers | Reverse Primers |
|---|---|---|
| Ubiquitin | GATTTGCATACCACCACGAAG | AAGTAGAAAGCTCCGACACCAT |
| IPSE | GGACTACTTCTTTGATGAAAACTGA | TTTGGGTCTACCTGACACTGG |
| GCPE | GTGGGTGTTGTTTGAACTGCTC | ACAGACCCGAAATAAAGAACCTC |
| HMGCR | AACACCCTCACACCAGCAGAT | GGACCCTGCCCAAAACATC |
| FDPS | CTGCAACAATGAGCACCGAT | ACCAACCAAGGGCACACG |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Sun, A.; Yu, X.; Lu, S.; Wu, T.; Meng, K.-Y.; Hao, J.-W.; Zhao, N.; Chai, J.-H.; He, T.-T. Differential Analysis of Sesquiterpenoids of Atractylodes macrocephalus from Different Origins Based on Transcriptomics. Molecules 2026, 31, 1075. https://doi.org/10.3390/molecules31071075
Sun A, Yu X, Lu S, Wu T, Meng K-Y, Hao J-W, Zhao N, Chai J-H, He T-T. Differential Analysis of Sesquiterpenoids of Atractylodes macrocephalus from Different Origins Based on Transcriptomics. Molecules. 2026; 31(7):1075. https://doi.org/10.3390/molecules31071075
Chicago/Turabian StyleSun, Ao, Xin Yu, Shan Lu, Tong Wu, Ke-Yi Meng, Jing-Wei Hao, Nan Zhao, Jun-Hong Chai, and Ting-Ting He. 2026. "Differential Analysis of Sesquiterpenoids of Atractylodes macrocephalus from Different Origins Based on Transcriptomics" Molecules 31, no. 7: 1075. https://doi.org/10.3390/molecules31071075
APA StyleSun, A., Yu, X., Lu, S., Wu, T., Meng, K.-Y., Hao, J.-W., Zhao, N., Chai, J.-H., & He, T.-T. (2026). Differential Analysis of Sesquiterpenoids of Atractylodes macrocephalus from Different Origins Based on Transcriptomics. Molecules, 31(7), 1075. https://doi.org/10.3390/molecules31071075








