Whole-Cell Biocatalysis for the Production of Structurally Diverse Methoxydihydrochalcones: Broad Activity of the Yarrowia Clade
Abstract
1. Introduction
2. Results and Discussion
3. Materials and Methods
3.1. Substrates
3.2. Microorganisms
3.3. Screening
3.4. Preparative Scale
3.5. TLC and NMR Analysis
3.6. UPLC
3.7. Bioinformatics Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- de Carvalho, C.C.C.R. Whole Cell Biocatalysts: Essential Workers from Nature to the Industry. Microb. Biotechnol. 2017, 10, 250–263. [Google Scholar] [CrossRef]
- Lin, B.; Tao, Y. Whole-Cell Biocatalysts by Design. Microb. Cell Factories 2017, 16, 106. [Google Scholar] [CrossRef]
- Wachtmeister, J.; Rother, D. Recent Advances in Whole Cell Biocatalysis Techniques Bridging from Investigative to Industrial Scale. Curr. Opin. Biotechnol. 2016, 42, 169–177. [Google Scholar] [CrossRef]
- Park, Y.-K.; Ledesma-Amaro, R. What Makes Yarrowia lipolytica Well Suited for Industry? Trends Biotechnol. 2023, 41, 242–254. [Google Scholar] [CrossRef]
- Qiao, K.; Wasylenko, T.M.; Zhou, K.; Xu, P.; Stephanopoulos, G. Lipid Production in Yarrowia lipolytica Is Maximized by Engineering Cytosolic Redox Metabolism. Nat. Biotechnol. 2017, 35, 173–177. [Google Scholar] [CrossRef]
- Fukuda, R. Utilization of N-Alkane and Roles of Lipid Transfer Proteins in Yarrowia lipolytica. World J. Microbiol. Biotechnol. 2023, 39, 97. [Google Scholar] [CrossRef] [PubMed]
- Fickers, P.; Benetti, P.; Wache, Y.; Marty, A.; Mauersberger, S.; Smit, M.; Nicaud, J. Hydrophobic Substrate Utilisation by the Yeast and Its Potential Applications. FEMS Yeast Res. 2005, 5, 527–543. [Google Scholar] [CrossRef] [PubMed]
- Fickers, P.; Cheng, H.; Lin, C.S.K. Sugar Alcohols and Organic Acids Synthesis in Yarrowia lipolytica: Where Are We? Microorganisms 2020, 8, 574. [Google Scholar] [CrossRef] [PubMed]
- Carvalho, A.S.S.; Sales, J.C.S.; Nascimento, F.V.d.; Ribeiro, B.D.; Souza, C.E.C.d.; Lemes, A.C.; Coelho, M.A.Z. Lipase Production by Yarrowia lipolytica in Solid-State Fermentation Using Amazon Fruit By-Products and Soybean Meal as Substrate. Catalysts 2023, 13, 289. [Google Scholar] [CrossRef]
- Blazeck, J.; Hill, A.; Liu, L.; Knight, R.; Miller, J.; Pan, A.; Otoupal, P.; Alper, H.S. Harnessing Yarrowia lipolytica Lipogenesis to Create a Platform for Lipid and Biofuel Production. Nat. Commun. 2014, 5, 3131. [Google Scholar] [CrossRef]
- Arnesen, J.A.; Belmonte Del Ama, A.; Jayachandran, S.; Dahlin, J.; Rago, D.; Andersen, A.J.C.; Borodina, I. Engineering of Yarrowia lipolytica for the Production of Plant Triterpenoids: Asiatic, Madecassic, and Arjunolic Acids. Metab. Eng. Commun. 2022, 14, e00197. [Google Scholar] [CrossRef]
- EFSA Panel on Nutrition, Novel Foods and Food Allergens (NDA); Turck, D.; Bohn, T.; Castenmiller, J.; De Henauw, S.; Hirsch-Ernst, K.I.; Maciuk, A.; Mangelsdorf, I.; McArdle, H.J.; Naska, A.; et al. Safety of an Extension of Use of Yarrowia lipolytica Yeast Biomass as a Novel Food Pursuant to Regulation (EU) 2015/2283. EFSA J. 2023, 21, e8416. [Google Scholar] [CrossRef]
- EFSA Panel on Nutrition, Novel Foods and Food Allergens (NDA); Turck, D.; Castenmiller, J.; de Henauw, S.; Hirsch-Ernst, K.; Kearney, J.; Maciuk, A.; Mangelsdorf, I.; McArdle, H.J.; Naska, A.; et al. Safety of Yarrowia lipolytica Yeast Biomass as a Novel Food Pursuant to Regulation (EU) 2015/2283. EFSA J. 2019, 17, e05594. [Google Scholar] [CrossRef] [PubMed]
- Dias, M.C.; Pinto, D.C.G.A.; Silva, A.M.S. Plant Flavonoids: Chemical Characteristics and Biological Activity. Molecules 2021, 26, 5377. [Google Scholar] [CrossRef] [PubMed]
- Lee, D.-S.; Li, B.; Im, N.-K.; Kim, Y.-C.; Jeong, G.-S. 4,2′,5′-Trihydroxy-4′-Methoxychalcone from Dalbergia odorifera Exhibits Anti-Inflammatory Properties by Inducing Heme Oxygenase-1 in Murine Macrophages. Int. Immunopharmacol. 2013, 16, 114–121. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.F.; Wu, S.N.; Gao, J.M.; Liao, Z.Y.; Tseng, Y.T.; Fülöp, F.; Chang, F.R.; Lo, Y.C. The Antioxidant, Anti-Inflammatory, and Neuroprotective Properties of the Synthetic Chalcone Derivative AN07. Molecules 2020, 25, 2907. [Google Scholar] [CrossRef]
- Tseng, Y.T.; Tsai, Y.H.; Fülöp, F.; Chang, F.R.; Lo, Y.C. 2-Iodo-40-Methoxychalcone Attenuates Methylglyoxal-Induced Neurotoxicity by Activation of GLP-1 Receptor and Enhancement of Neurotrophic Signal, Antioxidant Defense and Glyoxalase Pathway. Molecules 2019, 24, 2249. [Google Scholar] [CrossRef]
- Marques, B.C.; Santos, M.B.; Anselmo, D.B.; Monteiro, D.A.; Gomes, E.; Saiki, M.F.C.; Rahal, P.; Rosalen, P.L.; Sardi, J.C.O.; Regasini, L.O. Methoxychalcones: Effect of Methoxyl Group on the Antifungal, Antibacterial and Antiproliferative Activities. Med. Chem. 2020, 16, 881–891. [Google Scholar] [CrossRef]
- Nowakowska, Z. A Review of Anti-Infective and Anti-Inflammatory Chalcones. Eur. J. Med. Chem. 2007, 42, 125–137. [Google Scholar] [CrossRef]
- Johnson, R.; Dludla, P.V.; Muller, C.J.F.; Huisamen, B.; Essop, M.F.; Louw, J. The Transcription Profile Unveils the Cardioprotective Effect of Aspalathin against Lipid Toxicity in an In Vitro H9c2 Model. Molecules 2017, 22, 219. [Google Scholar] [CrossRef]
- Zhang, G.; Yang, G.; Liu, J. Phloretin Attenuates Behavior Deficits and Neuroinflammatory Response in MPTP Induced Parkinson’s Disease in Mice. Life Sci. 2019, 232, 116600. [Google Scholar] [CrossRef]
- Jesus, A.R.; Vila-Viçosa, D.; Machuqueiro, M.; Marques, A.P.; Dore, T.M.; Rauter, A.P. Targeting Type 2 Diabetes with C-Glucosyl Dihydrochalcones as Selective Sodium Glucose Co-Transporter 2 (SGLT2) Inhibitors: Synthesis and Biological Evaluation. J. Med. Chem. 2017, 60, 568–579. [Google Scholar] [CrossRef]
- Olender, D.; Kujawski, J.; Skóra, B.; Baranowska-Wójcik, E.; Sowa-Kasprzak, K.; Pawełczyk, A.; Zaprutko, L.; Szwajgier, D.; Szychowski, K.A. Bis-Chalcones Obtained via One-Pot Synthesis as the Anti-Neurodegenerative Agents and Their Effect on the HT-22 Cell Line. Heliyon 2024, 10, e37147. [Google Scholar] [CrossRef] [PubMed]
- Jackson, P.A.; Widen, J.C.; Harki, D.A.; Brummond, K.M. Covalent Modifiers: A Chemical Perspective on the Reactivity of α,β-Unsaturated Carbonyls with Thiols via Hetero-Michael Addition Reactions. J. Med. Chem. 2017, 60, 839–885. [Google Scholar] [CrossRef] [PubMed]
- Łużny, M.; Kaczanowska, D.; Gawdzik, B.; Wzorek, A.; Pawlak, A.; Obmińska-Mrukowicz, B.; Dymarska, M.; Kozłowska, E.; Kostrzewa-Susłow, E.; Janeczko, T. Regiospecific Hydrogenation of Bromochalcone by Unconventional Yeast Strains. Molecules 2022, 27, 3681. [Google Scholar] [CrossRef]
- Łużny, M.; Krzywda, M.; Kozłowska, E.; Kostrzewa-Susłow, E.; Janeczko, T. Effective Hydrogenation of 3-(2”-Furyl)- and 3-(2”-Thienyl)-1-(2′-Hydroxyphenyl)-Prop-2-En-1-One in Selected Yeast Cultures. Molecules 2019, 24, 3185. [Google Scholar] [CrossRef]
- Chlipała, P.; Janeczko, T.; Mazur, M. Effect of Sunlight-Induced Isomerisation on the Biotransformation of 4′-Hydroxychalcones by Yarrowia lipolytica KCh 71. Int. J. Mol. Sci. 2025, 26, 9027. [Google Scholar] [CrossRef]
- Łużny, M.; Kozłowska, E.; Kostrzewa-Susłow, E.; Janeczko, T. Highly Effective, Regiospecific Hydrogenation of Methoxychalcone by Yarrowia lipolytica Enables Production of Food Sweeteners. Catalysts 2020, 10, 1135. [Google Scholar] [CrossRef]
- Chlipała, P.; Bienia, J.; Mazur, M.; Dymarska, M.; Janeczko, T. Efficient Production of 4′-Hydroxydihydrochalcones Using Non-Conventional Yeast Strains. Int. J. Mol. Sci. 2024, 25, 10735. [Google Scholar] [CrossRef] [PubMed]
- Chlipała, P.; Janeczko, T.; Mazur, M. Bioreduction of 4′-Hydroxychalcone in Deep Eutectic Solvents: Optimization and Efficacy with Various Yeast Strains. Int. J. Mol. Sci. 2024, 25, 7152. [Google Scholar] [CrossRef]
- Toogood, H.S.; Gardiner, J.M.; Scrutton, N.S. Biocatalytic Reductions and Chemical Versatility of the Old Yellow Enzyme Family of Flavoprotein Oxidoreductases. ChemCatChem 2010, 2, 892–914. [Google Scholar] [CrossRef]
- Wu, S.; Wang, Q.; Ma, X.; Qiu, L.; Yan, H. Modulation of the Catalytic Performance of OYE3 by Engineering Key Residues at the Entrance of the Catalytic Pocket. Biotechnol. Appl. Biochem. 2023, 70, 1720–1730. [Google Scholar] [CrossRef] [PubMed]
- Fryszkowska, A.; Toogood, H.; Sakuma, M.; Gardiner, J.M.; Stephens, G.M.; Scrutton, N.S. Asymmetric Reduction of Activated Alkenes by Pentaerythritol Tetranitrate Reductase: Specificity and Control of Stereochemical Outcome by Reaction Optimisation. Adv. Synth. Catal. 2009, 351, 2976–2990. [Google Scholar] [CrossRef]
- Toogood, H.S.; Scrutton, N.S. Discovery, Characterization, Engineering and Applications of Ene-Reductases for Industrial Biocatalysis. ACS Catal. 2018, 8, 4333. [Google Scholar] [CrossRef]
- Ding, Y.; Perez-Ortiz, G.; Peate, J.; Barry, S.M. Redesigning Enzymes for Biocatalysis: Exploiting Structural Understanding for Improved Selectivity. Front. Mol. Biosci. 2022, 9, 908285. [Google Scholar] [CrossRef]
- Gatti, F.G.; Parmeggiani, F.; Sacchetti, A. Synthetic Strategies Based on C=C Bioreductions for the Preparation of Biologically Active Molecules. In Synthetic Methods for Biologically Active Molecules; Brenna, E., Ed.; Wiley: Hoboken, NJ, USA, 2013; pp. 49–84. [Google Scholar]
- Thevenieau, F.; Beopoulos, A.; Desfougeres, T.; Sabirova, J.; Albertin, K.; Zinjarde, S.; Nicaud, J.-M. Uptake and Assimilation of Hydrophobic Substrates by the Oleaginous Yeast Yarrowia lipolytica. In Handbook of Hydrocarbon and Lipid Microbiology; Timmis, K.N., Ed.; Springer: Berlin/Heidelberg, Germany, 2010; pp. 1513–1527. [Google Scholar]
- Stueckler, C.; Mueller, N.J.; Winkler, C.K.; Glueck, S.M.; Gruber, K.; Steinkellner, G.; Faber, K. Bioreduction of α-Methylcinnamaldehyde Derivatives: Chemo-Enzymatic Asymmetric Synthesis of LilialTM and HelionalTM. Dalton Trans. 2010, 39, 8472. [Google Scholar] [CrossRef]
- Sánchez-Moreno, I.; Iturrate, L.; Doyagüez, E.G.; Martínez, J.A.; Fernández-Mayoralas, A.; García-Junceda, E. Activated α,β-Unsaturated Aldehydes as Substrate of Dihydroxyacetone Phosphate (DHAP)-Dependent Aldolases in the Context of a Multienzyme System. Adv. Synth. Catal. 2009, 351, 2967–2975. [Google Scholar] [CrossRef]
- Fan, X.-Y.; Yu, Y.; Yao, Y.; Li, W.-D.; Tao, F.-Y.; Wang, N. Applications of Ene-Reductases in the Synthesis of Flavors and Fragrances. J. Agric. Food Chem. 2024, 72, 18305–18320. [Google Scholar] [CrossRef] [PubMed]
- Brenna, E.; Crotti, M.; Gatti, F.G.; Manfredi, A.; Monti, D.; Parmeggiani, F.; Santangelo, S.; Zampieri, D. Enantioselective Synthesis of (R)-2-Arylpropanenitriles Catalysed by Ene-Reductases in Aqueous Media and in Biphasic Ionic Liquid–Water Systems. ChemCatChem 2014, 6, 2425–2431. [Google Scholar] [CrossRef]
- Du, F.; Zhang, F.; Hang, Y.; Jing, H.; Zheng, Y.; Ma, W.; Sun, X.; Huang, H. Advances in Production of Customized Functional Unsaturated Fatty Acids in Yarrowia lipolytica. Agric. Prod. Process. Storage 2025, 1, 14. [Google Scholar] [CrossRef]
- Laribi, A.; Zieniuk, B.; Bouchedja, D.N.; Hafid, K.; Elmechta, L.; Becila, S. Valorization of Olive Mill Wastewater via Yarrowia lipolytica: Sustainable Production of High-Value Metabolites and Biocompounds—A Review. Fermentation 2025, 11, 326. [Google Scholar] [CrossRef]
- Liu, X.; Wang, C.; Tang, S.; Wang, G.; Huang, Y.; Yang, F.; Tan, X.; Bai, J.; Huang, L. Comparative Study on the Alleviating Effect of Neohesperidin Dihydrochalcones and Its Synthetic Precursor Neohesperidin on Ovalbumin-Induced Food Allergy. Food Res. Int. 2025, 212, 116436. [Google Scholar] [CrossRef] [PubMed]
- Liu, M.; Liu, C.; Zhaxi, P.; Kou, X.; Liu, Y.; Xue, Z. Research Progress on Hypoglycemic Effects and Molecular Mechanisms of Flavonoids: A Review. Antioxidants 2025, 14, 378. [Google Scholar] [CrossRef]
- Hall, M.; Stueckler, C.; Hauer, B.; Stuermer, R.; Friedrich, T.; Breuer, M.; Kroutil, W.; Faber, K. Asymmetric Bioreduction of Activated C=C Bonds Using Zymomonas Mobilis NCR Enoate Reductase and Old Yellow Enzymes OYE 1–3 from Yeasts. Eur. J. Org. Chem. 2008, 2008, 1511–1516. [Google Scholar] [CrossRef]
- Toogood, H.S.; Fryszkowska, A.; Hare, V.; Fisher, K.; Roujeinikova, A.; Leys, D.; Gardiner, J.M.; Stephens, G.M.; Scrutton, N.S. Structure-Based Insight into the Asymmetric Bioreduction of the C=C Double Bond of α,β-Unsaturated Nitroalkenes by Pentaerythritol Tetranitrate Reductase. Adv. Synth. Catal. 2008, 350, 2789–2803. [Google Scholar] [CrossRef] [PubMed]

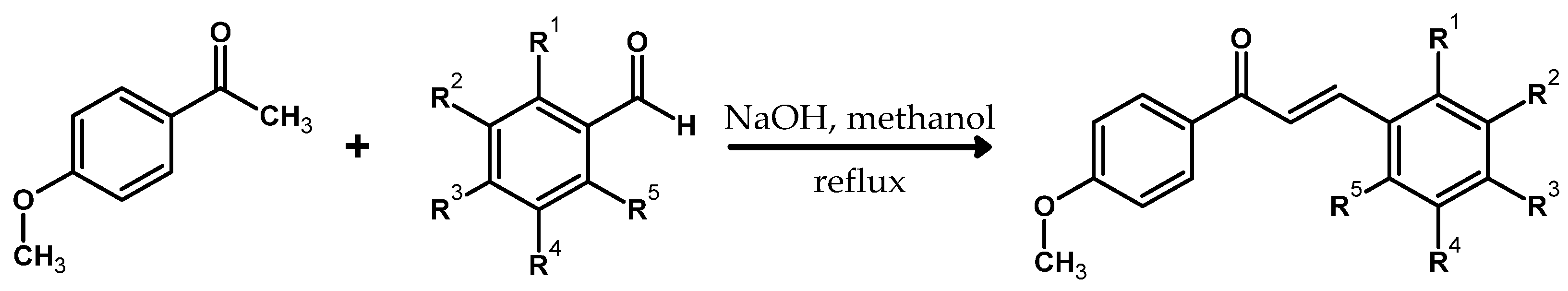
| Number | Strain | Abbreviation |
|---|---|---|
| 1 | Yarrowia lipolytica W-29 (control) | YALI |
| 2 | Candida osloensis CBS10146 | YAOS |
| 3 | Yarrowia phangngaensis CBS10407 | YAPH |
| 4 | Yarrowia parophonii CBS12427 | YAPA |
| 5 | Candida hispaniensis CBS9996 | OLHI |
| 6 | Yarrowia bubula CBS12934 | YABU |
| 7 | Candida alimentaria CBS10151 | YAAL |
| 8 | Yarrowia keelungensis CBS11062 | YAKE |
| 9 | Yarrowia brassicae CBS15225 | YABR |
| 10 | Yarrowia deformans CBS2071 | YADE |
| 11 | Yarrowia divulgata CBS11013 | YADI |
| 12 | Yarrowia yakushimensis CBS10253 | YAYA |
| 13 | Yarrowia lipolytica 24lIV | 24lIV |
| 14 | Yarrowia lipolytica 24lI | 24lI |
| 15 | Yarrowia lipolytica KCh 71 | KCh71 |
| 16 | Yarrowia lipolytica H222 | H222 |
| 17 | Yarrowia lipolytica ClY29-26-3 | ClY29-26-3 |
| 18 | Yarrowia lipolytica A101 | A101 |
| 19 | Candida galli CBS9722 | YAGA |
| 20 | Yarrowia porcina CBS12932 | YAPO |
| 21 | Candida hollandica CBS4855 | YAHO |
| Strain | 1 h | 12 h | 24 h | ||||||
|---|---|---|---|---|---|---|---|---|---|
| 1a | 1b | 1c | 1a | 1b | 1c | 1a | 1b | 1c | |
| YALI | 10.9 ± 7.7 | 12.6 ± 9.6 | 76.6 ± 8.5 | 3.0 ± 0.8 | 16.2 ± 3.4 | 80.7 ± 4.2 | 2.4 ± 0.5 | 13.9 ± 2.8 | 83.7 ± 3.3 |
| YAOS | 15.2 ± 1.6 | 11.6 ± 3.2 | 73.3 ± 4.7 | 5.0 ± 0.4 | 21.9 ± 2.4 | 73.1 ± 2.0 | 3.4 ± 0.4 | 19.5 ± 2.4 | 77.1 ± 2.8 |
| YAPH | 10.0 ± 6.4 | 15.8 ± 4.6 | 74.2 ± 2.8 | 3.6 ± 0.4 | 21.5 ± 2.9 | 74.9 ± 3.3 | 2.5 ± 0.0 | 14.1 ± 0.4 | 83.4 ± 0.4 |
| YAPA | 16.0 ± 4.9 | 12.6 ± 2.3 | 71.4 ± 3.7 | 4.1 ± 1.1 | 17.1 ± 3.3 | 78.8 ± 4.5 | 2.7 ± 0.5 | 15.6 ± 2.8 | 81.7 ± 3.3 |
| OLHI | 62.1 ± 3.2 | 5.3 ± 1.5 | 32.6 ± 1.6 | 11.2 ± 1.1 | 37.0 ± 2.0 | 51.8 ± 1.0 | 4.9 ± 1.0 | 25.8 ± 5.1 | 69.2 ± 6.0 |
| YABU | 21.8 ± 2.9 | 14.4 ± 2.8 | 63.7 ± 1.3 | 6.6 ± 1.8 | 29.6 ± 0.4 | 63.8 ± 1.4 | 4.8 ± 1.3 | 26.5 ± 6.7 | 68.7 ± 8.0 |
| YAAL | 20.3 ± 1.3 | 17.9 ± 7.3 | 61.9 ± 6.4 | 3.9 ± 0.8 | 22.8 ± 3.7 | 73.3 ± 4.5 | 1.6 ± 0.6 | 9.1 ± 2.9 | 89.3 ± 3.6 |
| YAKE | 7.0 ± 6.1 | 11.4 ± 9.1 | 81.5 ± 7.8 | 3.2 ± 0.8 | 15.0 ± 4.0 | 81.8 ± 4.7 | 1.4 ± 0.5 | 8.7 ± 3.5 | 89.9 ± 4.0 |
| YABR | 10.8 ± 2.7 | 40.7 ± 7.6 | 48.5 ± 7.5 | 7.4 ± 0.2 | 30.0 ± 4.2 | 62.6 ± 4.5 | 3.8 ± 0.3 | 22.0 ± 1.9 | 74.2 ± 2.2 |
| YADE | 6.9 ± 5.5 | 18.9 ± 4.2 | 74.2 ± 9.7 | 3.8 ± 1.3 | 22.1 ± 6.5 | 74.1 ± 7.8 | 2.5 ± 1.1 | 13.7 ± 5.8 | 83.9 ± 6.9 |
| YADI | 16.7 ± 4.5 | 21.7 ± 7.9 | 61.6 ± 9.3 | 3.6 ± 1.2 | 19.8 ± 5.1 | 76.7 ± 6.3 | 2.1 ± 1.3 | 12.1 ± 7.8 | 85.8 ± 9.2 |
| YAYA | 37.7 ± 6.2 | 32.1 ± 2.1 | 30.3 ± 8.2 | 14.5 ± 1.2 | 62.1 ± 0.8 | 23.3 ± 1.2 | 10.3 ± 1.9 | 50.4 ± 6.7 | 39.3 ± 6.9 |
| 24lIV | 27.9 ± 18.3 | 22.3 ± 6.8 | 49.8 ± 11.7 | 9.8 ± 1.8 | 33.1 ± 3.4 | 57.1 ± 4.6 | 3.1 ± 1.1 | 17.9 ± 6.2 | 79.0 ± 7.2 |
| 24lI | 17.6 ± 7.4 | 29.1 ± 3.0 | 53.3 ± 9.0 | 5.2 ± 0.5 | 30.4 ± 2.1 | 64.4 ± 2.6 | 5.5 ± 0.6 | 31.8 ± 2.8 | 62.7 ± 3.4 |
| KCh 71 | 9.0 ± 5.1 | 21.4 ± 10.7 | 69.7 ± 5.6 | 3.3 ± 0.3 | 20.4 ± 2.2 | 76.3 ± 2.5 | 2.9 ± 0.0 | 16.1 ± 0.5 | 81.0 ± 0.5 |
| H222 | 24.5 ± 2.1 | 28.7 ± 5.2 | 46.9 ± 4.2 | 8.7 ± 0.2 | 50.7 ± 1.1 | 40.7 ± 1.3 | 6.5 ± 0.8 | 37.7 ± 4.8 | 55.9 ± 5.6 |
| ClY29-26-3 | 4.0 ± 2.1 | 7.5 ± 3.9 | 88.5 ± 6.0 | 1.6 ± 0.1 | 9.7 ± 0.5 | 88.7 ± 0.6 | 0.8 ± 0.1 | 4.6 ± 1.0 | 94.6 ± 1.1 |
| A101 | 6.7 ± 1.8 | 23.5 ± 5.8 | 69.8 ± 4.7 | 2.6 ± 0.7 | 16.1 ± 3.1 | 81.3 ± 3.8 | 1.5 ± 0.5 | 8.3 ± 3.0 | 90.3 ± 3.5 |
| YAGA | 8.3 ± 0.3 | 16.4 ± 9.3 | 75.3 ± 9.5 | 3.5 ± 1.5 | 20.2 ± 8.0 | 76.2 ± 9.6 | 2.3 ± 1.1 | 13.4 ± 6.6 | 84.3 ± 7.8 |
| YAPO | 9.3 ± 3.7 | 31.2 ± 9.5 | 59.5 ± 5.8 | 5.7 ± 0.3 | 31.3 ± 1.3 | 63.0 ± 1.6 | 5.3 ± 1.1 | 30.3 ± 5.3 | 64.4 ± 6.4 |
| YAHO | 7.4 ± 5.6 | 32.5 ± 8.7 | 60.1 ± 14.0 | 4.0 ± 2.0 | 24.7 ± 10.8 | 71.3 ± 12.8 | 1.5 ± 0.9 | 8.5 ± 4.9 | 89.9 ± 5.8 |
| Compound | |||||||
|---|---|---|---|---|---|---|---|
| Atom No. | 1a | 2a | 3a | 4a | 5a | 6a | 7a |
| C=O | 188.16 | 188.44 | 188.22 | 188.13 | 188.44 | 188.25 | 189.49 |
| C-α | 122.88 | 122.89 | 123.08 | 120.35 | 120.21 | 123.27 | 121.81 |
| C-β | 143.95 | 139.00 | 143.99 | 143.91 | 139.21 | 144.14 | 135.37 |
| C-1 | 136.25 | 124.73 | 137.59 | 128.80 | 117.66 | 138.12 | 106.90 |
| C-1′ | 132.01 | 132.19 | 131.95 | 132.23 | 132.48 | 131.94 | 132.92 |
| C-2 | 129.40 | 159.06 | 114.05 | 131.17 | 161.20 | 107.21 | 162.63 |
| C-2′/C-6′ | 131.63 | 131.76 | 131.65 | 131.48 | 131.36 | 131.66 | 131.16 |
| C-3 | 129.80 | 112.35 | 161.13 | 115.23 | 130.93 | 162.21 | 91.65 |
| C-3′/C-5′ | 114.75 | 114.70 | 114.73 | 114.66 | 114.62 | 114.54 | 114.57 |
| C-4 | 129.80 | 132.60 | 122.06 | 162.58 | 106.91 | 103.32 | 164.32 |
| C-4′ | 164.53 | 164.40 | 164.52 | 164.33 | 164.17 | 164.53 | 163.96 |
| C-5 | 129.80 | 121.56 | 130.77 | 115.23 | 164.18 | 162.21 | 91.65 |
| C-6 | 129.40 | 129.37 | 117.10 | 131.17 | 99.02 | 107.21 | 162.63 |
| C-2 OCH3 | — | 56.06 | — | — | 56.12 | — | 56.36 |
| C-3 OCH3 | — | — | 55.96 | — | — | 55.83 | — |
| C-4 OCH3 | — | — | — | 55.78 | — | — | 55.86 |
| C-4′ OCH3 | 55.97 | 55.93 | 55.70 | 55.93 | 56.09 | 55.97 | 55.88 |
| C-5 OCH3 | — | — | — | — | 55.89 | 55.83 | — |
| C-6 OCH3 | — | — | — | — | — | — | 56.36 |
| Compound | ||||||
|---|---|---|---|---|---|---|
| Atom No. | 1c | 2c | 3c | 4c | 5c | 6c |
| C=O | 197.75 | 198.21 | 197.76 | 197.87 | 198.36 | 197.80 |
| C-α | 40.41 | 39.00 | 40.31 | 40.67 | 39.30 | 40.26 |
| C-β | 30.89 | 26.32 | 30.93 | 30.03 | 25.77 | 31.22 |
| C-1 | 142.71 | 130.44 | 144.24 | 134.47 | 131.09 | 144.99 |
| C-1′ | 131.06 | 131.06 | 131.02 | 131.05 | 122.5 | 131.06 |
| C-2 | 129.33 | 158.52 | 115.02 | 130.21 | 159.30 | 107.32 |
| C-2′/C-6′ | 131.06 | 131.06 | 131.04 | 131.02 | 131.04 | 131.06 |
| C-3 | 129.18 | 111.23 | 160.78 | 114.56 | – | 161.92 |
| C-3′/C-5′ | 114.57 | 114.57 | 114.55 | 114.54 | 114.54 | 114.57 |
| C-4 | 126.70 | 128.25 | 121.51 | 158.97 | 105.09 | 98.59 |
| C-4′ | 164.41 | 164.36 | 164.37 | 164.34 | 164.31 | 164.40 |
| C-5 | 129.18 | 121.21 | 130.12 | 114.56 | 160.59 | 161.92 |
| C-6 | 129.33 | 130.73 | 112.09 | 130.21 | 130.99 | 107.32 |
| C-2 OCH3 | - | 55.63 | – | – | 55.67 | – |
| C-3 OCH3 | - | – | 55.33 | – | – | 55.46 |
| C-4 OCH3 | - | – | – | 55.41 | – | – |
| C-4′ OCH3 | 55.91 | 55.90 | 55.89 | 55.88 | 55.89 | 55.91 |
| C-5 OCH3 | - | – | – | – | 55.54 | 55.46 |
| C-6 OCH3 | - | – | – | – | – | – |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Chlipała, P.; Mazur, M.; Kancelista, A.; Lazar, Z.; Janeczko, T. Whole-Cell Biocatalysis for the Production of Structurally Diverse Methoxydihydrochalcones: Broad Activity of the Yarrowia Clade. Molecules 2026, 31, 1049. https://doi.org/10.3390/molecules31061049
Chlipała P, Mazur M, Kancelista A, Lazar Z, Janeczko T. Whole-Cell Biocatalysis for the Production of Structurally Diverse Methoxydihydrochalcones: Broad Activity of the Yarrowia Clade. Molecules. 2026; 31(6):1049. https://doi.org/10.3390/molecules31061049
Chicago/Turabian StyleChlipała, Paweł, Marcelina Mazur, Anna Kancelista, Zbigniew Lazar, and Tomasz Janeczko. 2026. "Whole-Cell Biocatalysis for the Production of Structurally Diverse Methoxydihydrochalcones: Broad Activity of the Yarrowia Clade" Molecules 31, no. 6: 1049. https://doi.org/10.3390/molecules31061049
APA StyleChlipała, P., Mazur, M., Kancelista, A., Lazar, Z., & Janeczko, T. (2026). Whole-Cell Biocatalysis for the Production of Structurally Diverse Methoxydihydrochalcones: Broad Activity of the Yarrowia Clade. Molecules, 31(6), 1049. https://doi.org/10.3390/molecules31061049







