8-Oxoguanine Disrupts G-Quadruplex DNA Stability and Modulates FANCJ AKKQ Peptide Binding
Abstract
1. Introduction
2. Results
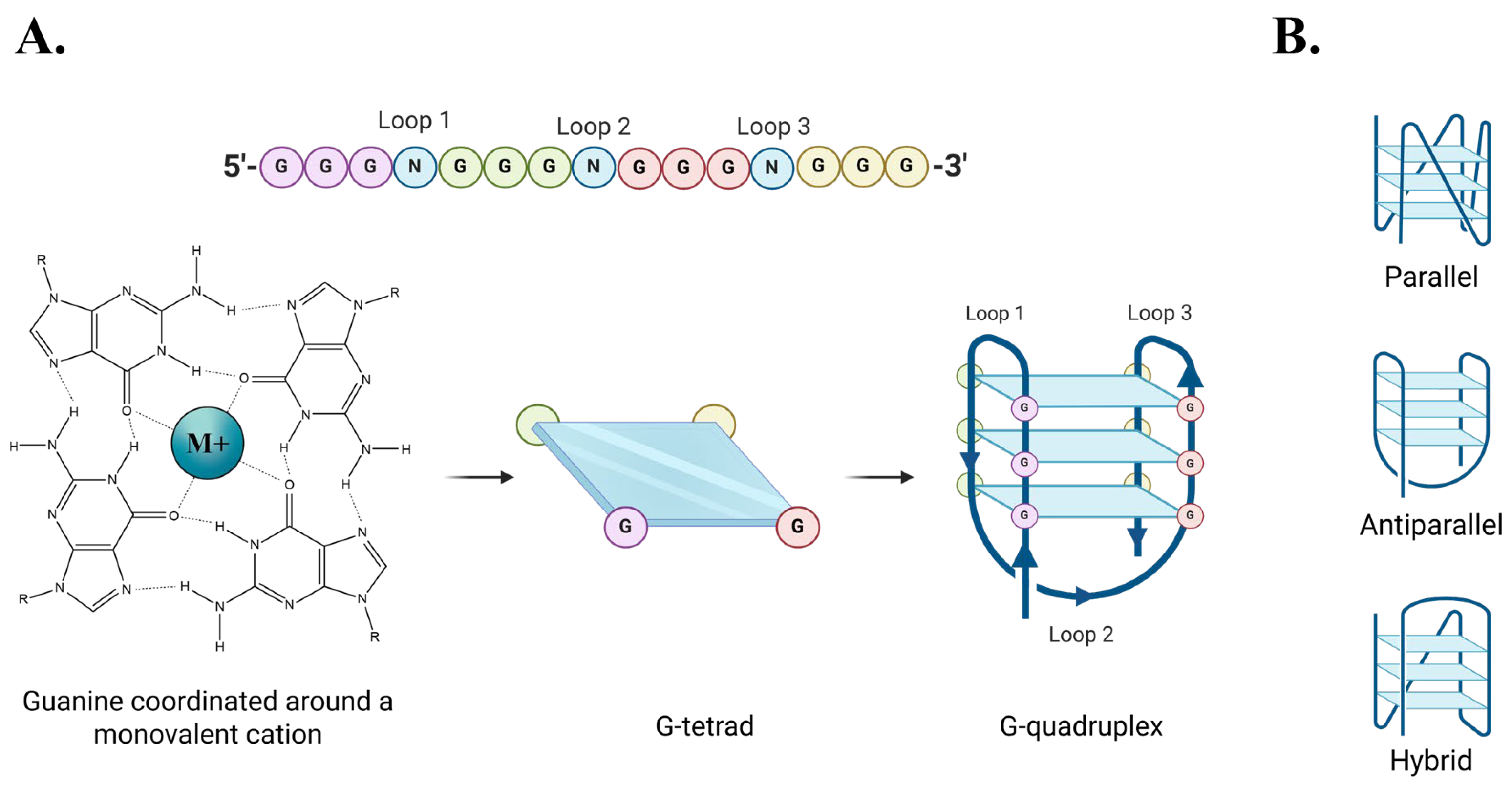
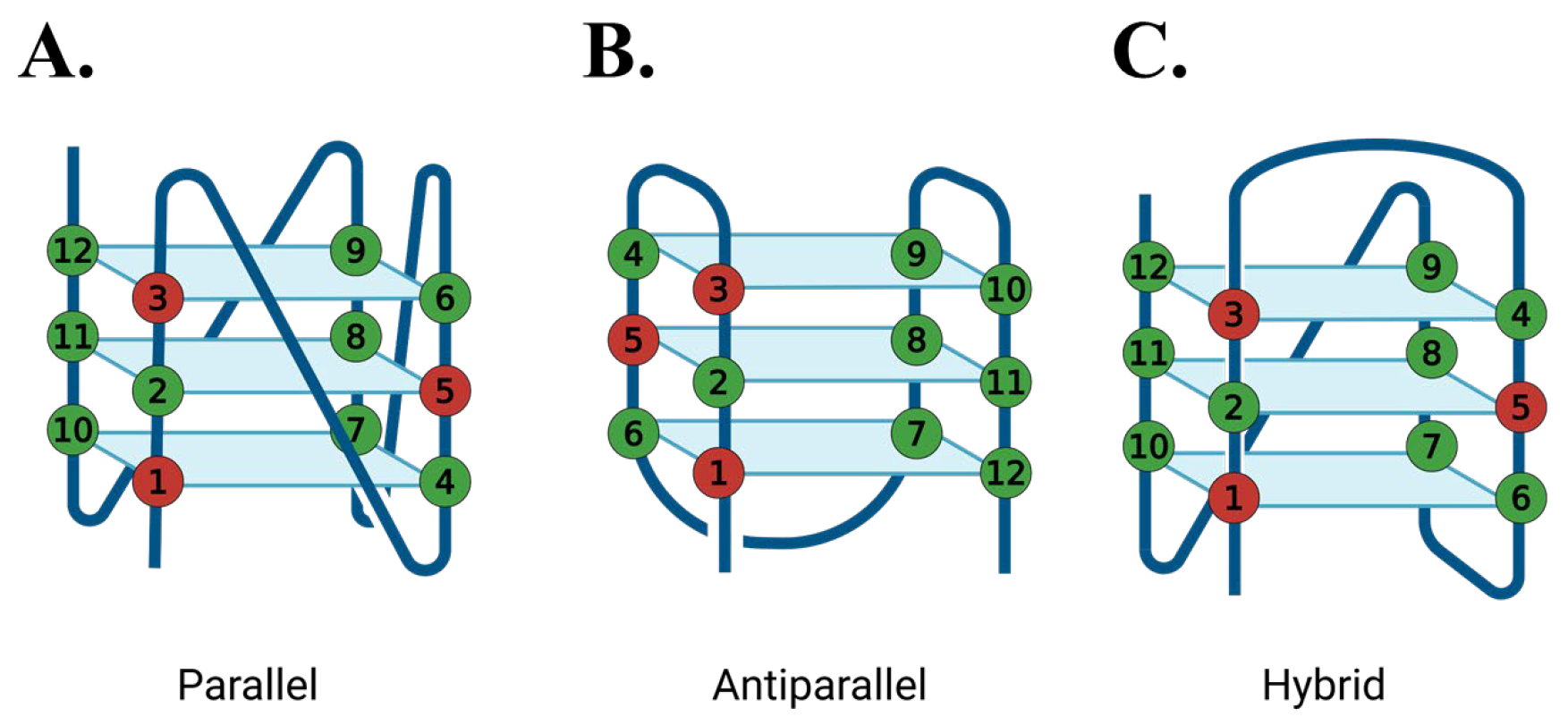
2.1. G-Quadruplex Conformation and Thermal Stability
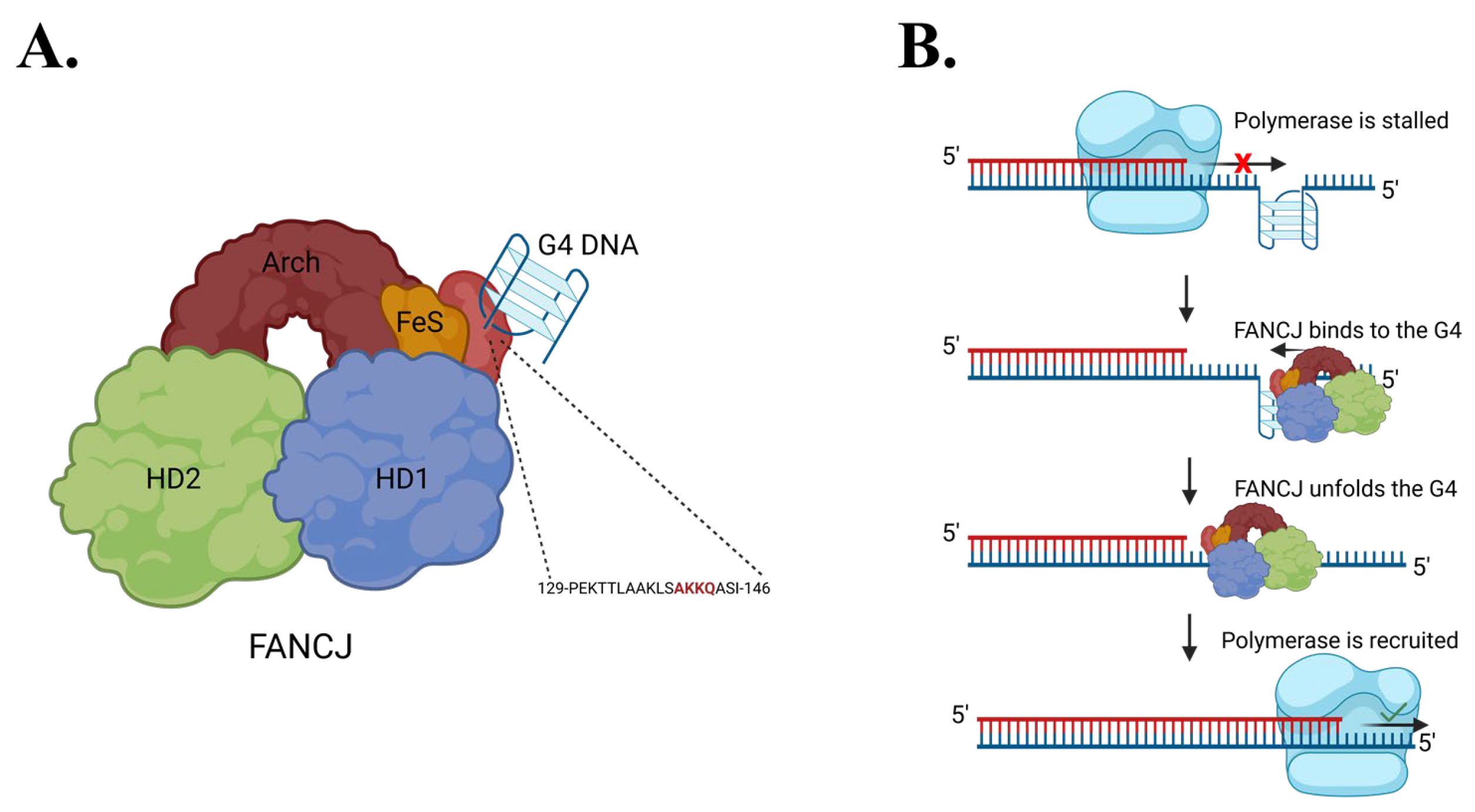
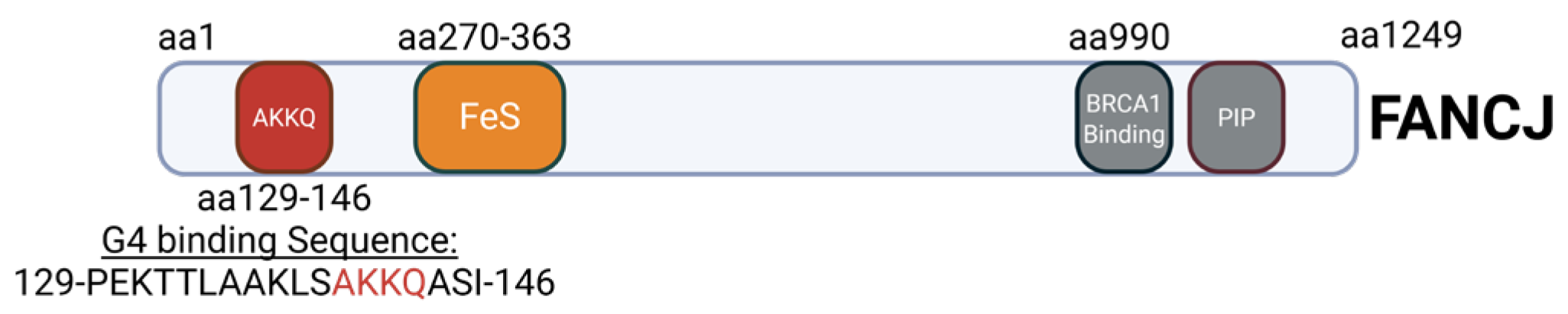
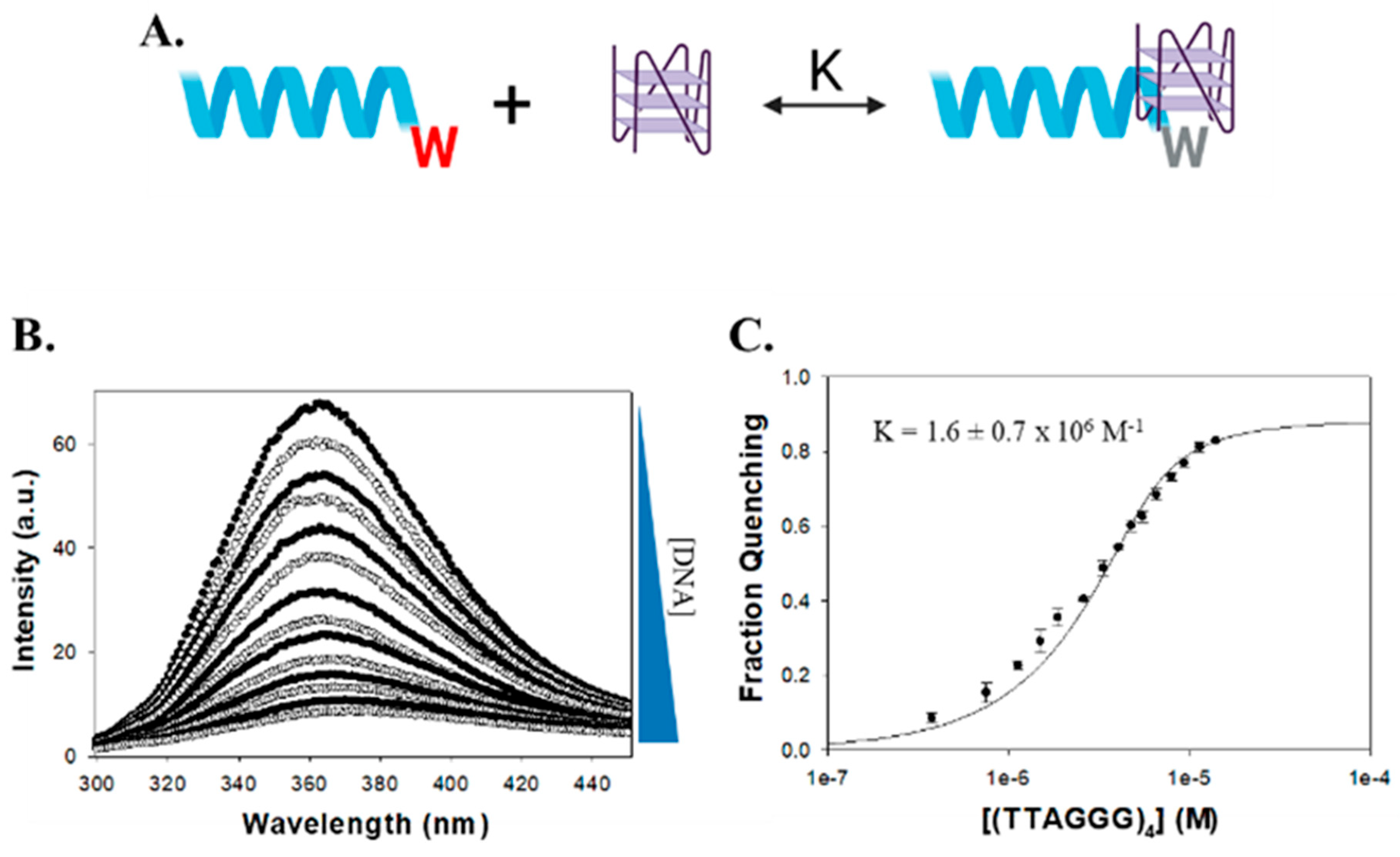
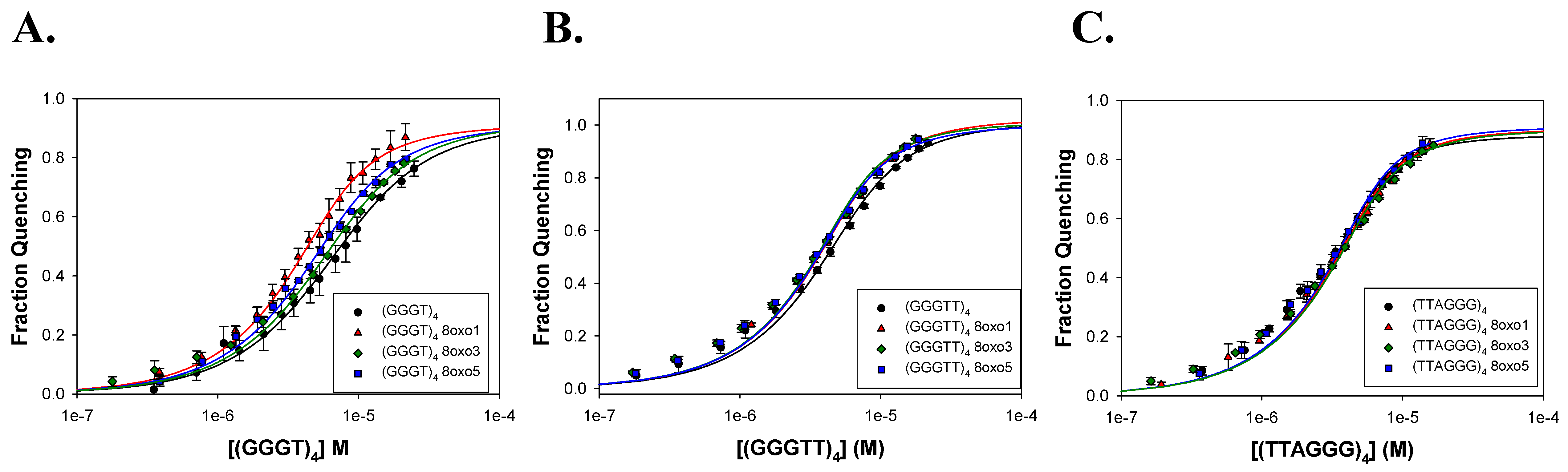
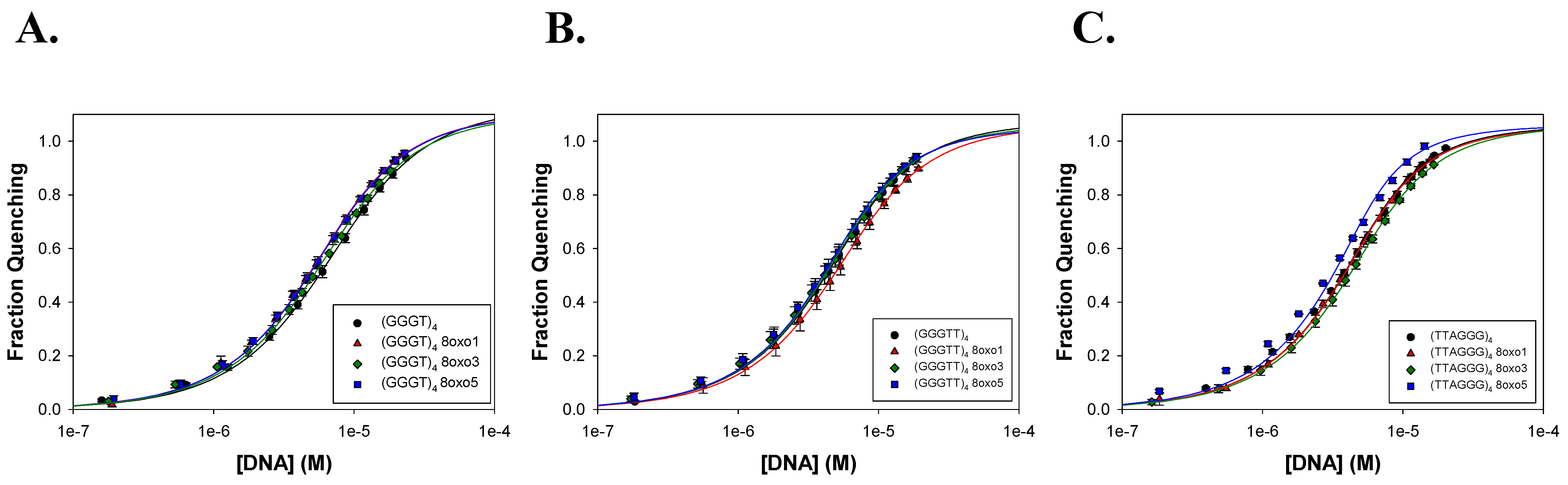
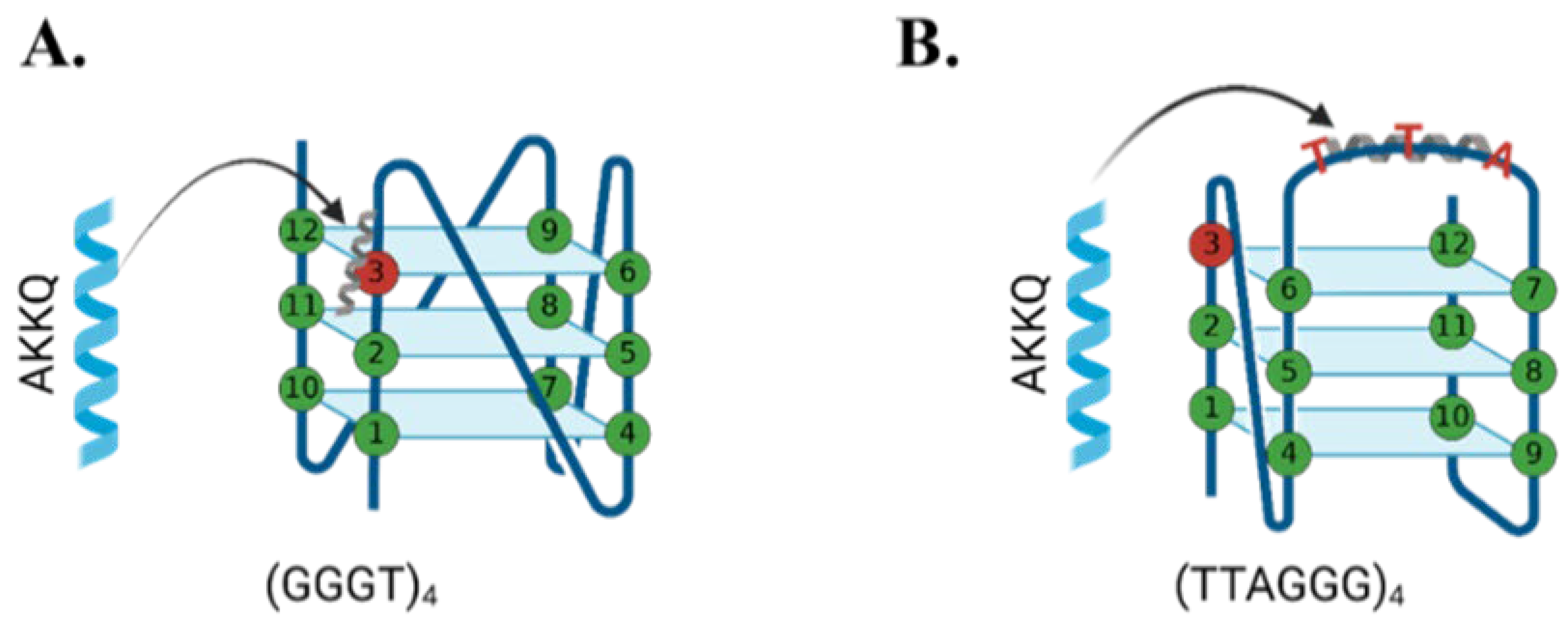
2.2. FANCJ AKKQ Binding to 8oxoG4s
3. Discussion
3.1. G4 Conformation and Stability
3.2. FANCJ AKKQ Binding to 8oxoG4s
3.3. Biological Implications and Future Directions
4. Materials and Methods
4.1. Buffers and Reagents
4.2. Peptide and DNA Oligos
4.3. CD Spectroscopy
4.4. Fluorescence Spectroscopy
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| G4 | G-quadruplex |
| 8oxoG | 8-oxoguanine |
| CD | Circular dichroism |
References
- Bochman, M.L.; Paeschke, K.; Zakian, V.A. DNA secondary structures: Stability and function of G-quadruplex structures. Nat. Rev. Genet. 2012, 13, 770–780. [Google Scholar] [CrossRef]
- Nový, J.; Böhm, S.; Králová, J.; Král, V.; Urbanová, M. Formation and temperature stability of G-quadruplex structures studied by electronic and vibrational circular dichroism spectroscopy combined with ab initio calculations. Biopolymers 2007, 89, 144–152. [Google Scholar] [CrossRef] [PubMed]
- Chaires, J.B. Human telomeric G-quadruplex: Thermodynamic and kinetic studies of telomeric quadruplex stability. FEBS J. 2010, 277, 1098–1106. [Google Scholar] [CrossRef]
- Guédin, A.; Gros, J.; Alberti, P.; Mergny, J.L. How long is too long? Effects of loop size on G-quadruplex stability. Nucleic Acids Res. 2010, 38, 7858–7868. [Google Scholar] [CrossRef]
- Huppert, J.L.; Balasubramanian, S. Prevalence of quadruplexes in the human genome. Nucleic Acids Res. 2005, 33, 2908–2916. [Google Scholar] [CrossRef]
- Lombardi, E.P.; Holmes, A.; Verga, D.; Teulade-Fichou, M.P.; Nicolas, A.; Londoño-Vallejo, A. Thermodynamically stable and genetically unstable G-quadruplexes are depleted in genomes across species. Nucleic Acids Res. 2019, 47, 6098–6113. [Google Scholar] [CrossRef]
- Hazel, P.; Huppert, J.; Balasubramanian, S.; Neidle, S. Loop-Length-Dependent Folding of G-Quadruplexes. J. Am. Chem. Soc. 2004, 126, 16405–16415. [Google Scholar] [CrossRef]
- Hao, F.; Ma, Y.; Guan, Y. Effects of Central Loop Length and Metal Ions on the Thermal Stability of G-Quadruplexes. Molecules 2019, 24, 1863. [Google Scholar] [CrossRef]
- Burge, S.; Parkinson, G.N.; Hazel, P.; Todd, A.K.; Neidle, S. Quadruplex DNA: Sequence, topology and structure. Nucleic Acids Res. 2006, 34, 5402–5415. [Google Scholar] [CrossRef] [PubMed]
- Villar-Guerra, R.D.; Trent, J.O.; Chaires, J.B. G-Quadruplex Secondary Structure Obtained from Circular Dichroism Spectros-copy. Angewandte Chemie. Angew. Chem. Int. Ed. 2018, 57, 7171–7175. [Google Scholar] [CrossRef] [PubMed]
- Heddi, B.; Cheong, V.V.; Martadinata, H.; Phan, A.T. Insights into G-quadruplex specific recognition by the DEAH-box helicase RHAU: Solution structure of a peptide–quadruplex complex. Proceedings of the National Academy of Sciences of the United States of America. Proc. Natl. Acad. Sci. USA 2015, 112, 9608–9613. [Google Scholar] [CrossRef]
- Budhathoki, J.B.; Maleki, P.; Roy, W.A.; Janscak, P.; Yodh, J.G.; Balci, H. A Comparative Study of G-Quadruplex Unfolding and DNA Reeling Activities of Human RECQ5 Helicase. Biophys. J. 2016, 110, 2585–2596. [Google Scholar] [CrossRef] [PubMed]
- Tippana, R.; Hwang, H.; Opresko, P.L.; Bohr, V.A.; Myong, S. Single-molecule imaging reveals a common mechanism shared by G-quadruplex–resolving helicases. Proc. Natl. Acad. Sci. USA 2016, 113, 8448–8453. [Google Scholar] [CrossRef]
- Zheng, K.W.; Zhang, J.Y.; He, Y.D.; Gong, J.Y.; Wen, C.J.; Chen, J.N.; Hao, Y.H.; Zhao, Y.; Tan, Z. Detection of genomic G-quadruplexes in living cells using a small artificial protein. Nucleic Acids Res. 2020, 48, 11706–11720. [Google Scholar] [CrossRef]
- Wang, L.; Wang, Q.-M.; Wang, Y.-R.; Xi, X.-G.; Hou, X.-M. DNA-unwinding activity of Saccharomyces cerevisiae Pif1 is modulated by thermal stability, folding conformation, and loop lengths of G-quadruplex DNA. J. Biol. Chem. 2018, 293, 18504–18513. [Google Scholar] [CrossRef] [PubMed]
- Rhodes, D.; Lipps, H.J. G-quadruplexes and their regulatory roles in biology. Nucleic Acids Res. 2015, 43, 8627–8637. [Google Scholar] [CrossRef]
- Huppert, J.L.; Balasubramanian, S. G-quadruplexes in promoters throughout the human genome. Nucleic Acids Res. 2006, 35, 406–413. [Google Scholar] [CrossRef]
- Beaudoin, J.-D.; Perreault, J.-P. 5’-UTR G-quadruplex structures acting as translational repressors. Nucleic Acids Res. 2010, 38, 7022–7036. [Google Scholar] [CrossRef]
- Smith, J.S.; Chen, Q.A.; Yatsunyk, L.; Nicoludis, J.M.; Garcia, M.S.; Kranaster, R.; Balasubramanian, S.; Monchaud, D.; Teulade-Fichou, M.-P.; Abramowitz, L.; et al. Rudimentary G-quadruplex–based telomere capping in Saccharomyces cerevisiae. Nat. Struct. Mol. Biol. 2011, 18, 478–485. [Google Scholar] [CrossRef]
- Georgakopoulos-Soares, I.; Parada, G.E.; Wong, H.Y.; Medhi, R.; Furlan, G.; Munita, R.; Miska, E.A.; Kwok, C.K.; Hemberg, M. Alternative splicing modulation by G-quadruplexes. Nat. Commun. 2022, 13, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Xu, H.; Di-Antonio, M.; McKinney, S.; Mathew, V.; Ho, B.; O’Neil, N.J.; Dos Santos, N.; Silvester, J.; Wei, V.; Garcia, J.; et al. CX-5461 is a DNA G-quadruplex stabilizer with se-lective lethality in BRCA1/2 deficient tumours. Nat Commun. 2017, 8, 14432. [Google Scholar] [CrossRef] [PubMed]
- Steenken, S.; Jovanovic, S.V. How Easily Oxidizable Is DNA? One-Electron Reduction Potentials of Adenosine and Guanosine Radicals in Aqueous Solution. J. Am. Chem. Soc. 1997, 119, 617–618. [Google Scholar] [CrossRef]
- Oikawa, S. Sequence-specific DNA damage by reactive oxygen species: Implications for carcinogenesis and aging. Environ Health Prev. Med. 2005, 10, 65–71. [Google Scholar] [CrossRef]
- Neeley, W.L.; Essigmann, J.M. Mechanisms of Formation, Genotoxicity, and Mutation of Guanine Oxidation Products. Chem. Res. Toxicol. 2006, 19, 491–505. [Google Scholar] [CrossRef]
- Ohno, M.; Sakumi, K.; Fukumura, R.; Furuichi, M.; Iwasaki, Y.; Hokama, M.; Ikemura, T.; Tsuzuki, T.; Gondo, Y.; Nakabeppu, Y. 8-oxoguanine causes spontaneous de novo germline mutations in mice. Sci. Rep. 2014, 4, 4689. [Google Scholar] [CrossRef]
- Bielskutė, S.; Plavec, J.; Podbevšek, P. Impact of Oxidative Lesions on the Human Telomeric G-Quadruplex. J. Am. Chem. Soc. 2019, 141, 2594–2603. [Google Scholar] [CrossRef]
- Vorlícková, M.; Tomasko, M.; Sagi, A.J.; Bednarova, K.; Sagi, J. 8-oxoguanine in a quadruplex of the human telomere DNA se-quence. FEBS J. 2012, 279, 29–39. [Google Scholar] [CrossRef]
- Broth, R.M., Jr.; Cantor, S.B. Molecular and cellular functions of the FANCJ DNA helicase defective in cancer and in Fanconi anemia. Front. Genet. 2014, 5, 372. [Google Scholar] [CrossRef] [PubMed]
- Palovcak, A.; Liu, W.; Yuan, F.; Zhang, Y. Maintenance of genome stability by Fanconi anemia proteins. Cell Biosci. 2017, 7, 1–18. [Google Scholar] [CrossRef]
- Datta, A.; Robert, M.B., Jr. Holding All the Cards—How Fanconi Anemia Proteins Deal with Replication Stress and Preserve Genomic Stability. Genes 2019, 10, 170. [Google Scholar] [CrossRef]
- Wu, Y.; Shin-Ya, K.; Brosh, R.M., Jr. FANCJ Helicase Defective in Fanconia Anemia and Breast Cancer Unwinds G-Quadruplex DNA To Defend Genomic Stability. Mol. Cell. Biol. 2008, 28, 4116–4128. [Google Scholar] [CrossRef]
- Cantor, S.B.; Xie, J. Assessing the link between BACH1/FANCJ and MLH1 in DNA crosslink repair. Environ. Mol. Mutagen. 2010, 51, 500–507. [Google Scholar] [CrossRef]
- Sarkies, P.; Murat, P.; Phillips, L.G.; Patel, K.; Balasubramanian, S.; Sale, J.E. FANCJ coordinates two pathways that maintain epigenetic stability at G-quadruplex DNA. Nucleic Acids Res. 2011, 40, 1485–1498. [Google Scholar] [CrossRef]
- Bosch, P.C.; Segura-Bayona, S.; Koole, W.; van Heteren, J.T.; Dewar, J.M.; Tijsterman, M.; Knipscheer, P. FANCJ promotes DNA synthesis through G-quadruplex structures. EMBO J. 2014, 33, 2521–2533. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.G.; Spies, M. G-quadruplex recognition and remodeling by the FANCJ helicase. Nucleic Acids Res. 2016, 44, 8742–8753. [Google Scholar] [CrossRef] [PubMed]
- Lowran, K.; Campbell, L.; Popp, P.; Wu, C.G. Assembly of a G-Quadruplex Repair Complex by the FANCJ DNA Helicase and the REV1 Polymerase. Genes 2019, 11, 5. [Google Scholar] [CrossRef]
- Lattmann, S.; Giri, B.; Vaughn, J.P.; Akman, S.A.; Nagamine, Y. Role of the amino terminal RHAU-specific motif in the recognition and resolution of guanine quadruplex-RNA by the DEAH-box RNA helicase RHAU. Nucleic Acids Res. 2010, 38, 6219–6233. [Google Scholar] [CrossRef]
- Rachwal, P.A.; Fox, K.R. Quadruplex melting. Methods 2007, 43, 291–301. [Google Scholar] [CrossRef] [PubMed]
- Do, N.Q.; Lim, K.W.; Teo, M.H.; Heddi, B.; Phan, A.T. Stacking of G-quadruplexes: NMR structure of a G-rich oligonucleotide with potential anti-HIV and anticancer activity†. Nucleic Acids Res. 2011, 39, 9448–9457. [Google Scholar] [CrossRef]
- Guédin, A.; De Cian, A.; Gros, J.; Lacroix, L.; Mergny, J.-L. Sequence effects in single-base loops for quadruplexes. Biochimie 2008, 90, 686–696. [Google Scholar] [CrossRef]
- Lin, C.; Yang, D. Human Telomeric G-Quadruplex Structures and G-Quadruplex-Interactive Compounds. Methods Mol. Biol. 2018, 2035, 171–196. [Google Scholar] [CrossRef]
- Hardin, C.C.; Watson, T.; Corregan, M.; Bailey, C. Cation-dependent transition between the quadruplex and Watson-Crick hairpin forms of d(CGCG3GCG). Biochemistry 1992, 31, 833–841. [Google Scholar] [CrossRef] [PubMed]
- Gaur, P.; Bain, F.E.; Honda, M.; Granger, S.L.; Spies, M. Single-Molecule Analysis of the Improved Variants of the G-Quadruplex Recognition Protein G4P. Int. J. Mol. Sci. 2023, 24, 10274. [Google Scholar] [CrossRef] [PubMed]
- Eddy, S.; Ketkar, A.; Zafar, M.K.; Maddukuri, L.; Choi, J.-Y.; Eoff, R.L. Human Rev1 polymerase disrupts G-quadruplex DNA. Nucleic Acids Res. 2013, 42, 3272–3285. [Google Scholar] [CrossRef]
- Greenfield, N.J. Using circular dichroism collected as a function of temperature to determine the thermodynamics of protein unfolding and binding interactions. Nat. Protoc. 2006, 1, 2527–2535. [Google Scholar] [CrossRef]

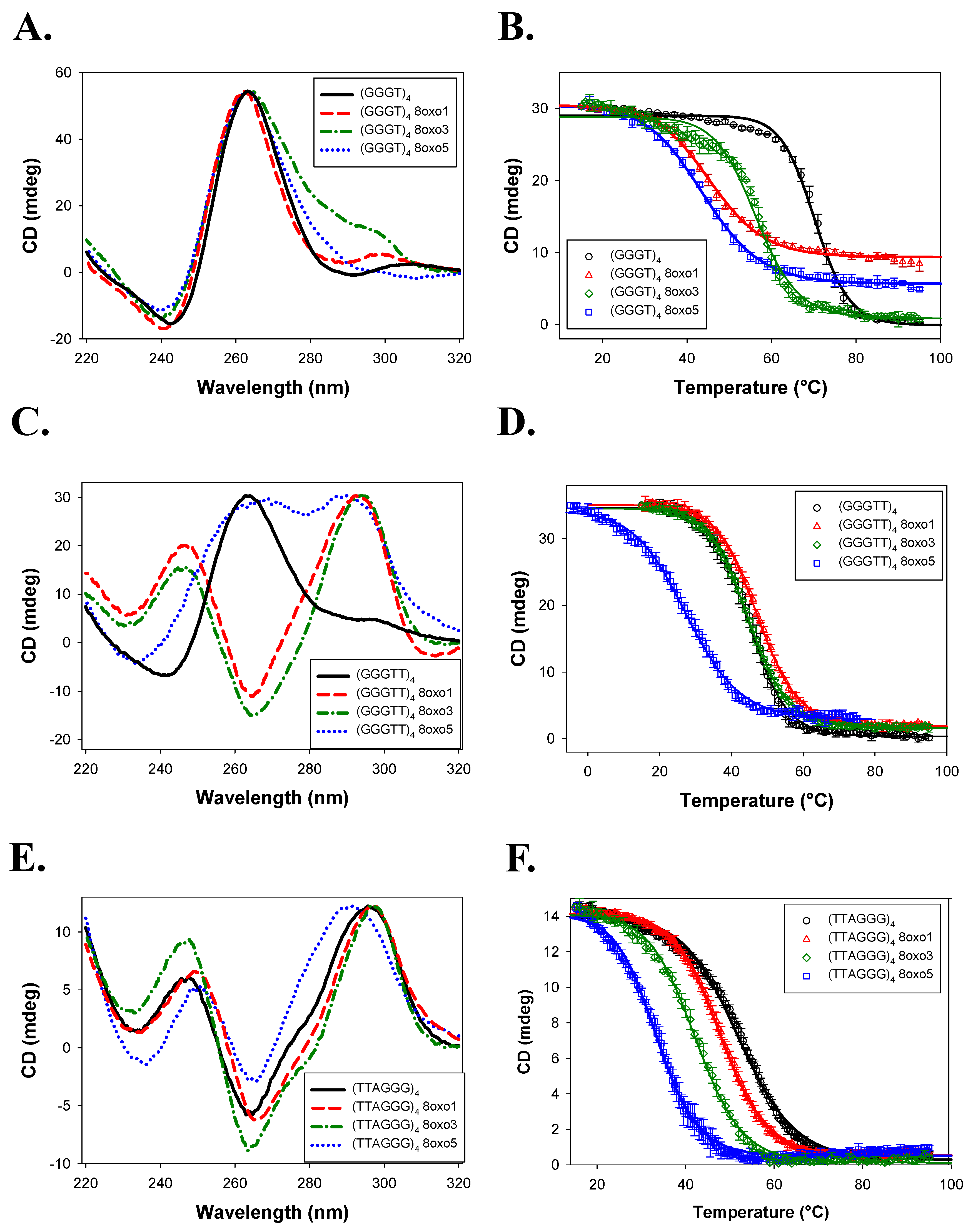
| Name | Conformation | Tm (°C) | ΔTm (°C) |
|---|---|---|---|
| (GGGT)4 | Parallel | >88.4 ± 8.4 | - |
| (GGGT)4 8oxo1 | Parallel | 69.4 ± 0.6 | −19.0 ± 8.4 |
| (GGGT)4 8oxo3 | Parallel | 80.3 ± 0.4 | −8.1 ± 8.4 |
| (GGGT)4 8oxo5 | Parallel | 71.6 ± 0.4 | −16.8 ± 8.4 |
| (GGGTT)4 | Parallel | 83.5 ± 0.7 | - |
| (GGGTT)4 8oxo1 | Parallel | 52.0 ± 1.4 | −31.5 ± 1.6 |
| (GGGTT)4 8oxo3 | Antiparallel | 57.2 ± 0.3 | −26.3 ± 0.7 |
| (GGGTT)4 8oxo5 | Parallel | 55.9 ± 0.3 | −27.6 ± 0.8 |
| (TTAGGG)4 | Hybrid | 62.5 ± 0.5 | - |
| (TTAGGG)4 8oxo1 | Hybrid-esque | 54.5 ± 0.5 | −8.0 ± 0.7 |
| (TTAGGG)4 8oxo3 | Antiparallel | 49.5 ± 0.3 | −13.0 ± 0.6 |
| (TTAGGG)4 8oxo5 | Hybrid-esque | 38.9 ± 0.6 | −23.6 ± 0.8 |
| Name | Conformation | Tm (°C) | ΔTm (°C) |
|---|---|---|---|
| (GGGT)4 | Parallel | 70.3 ± 0.7 | - |
| (GGGT)4 8oxo1 | Parallel | 42.5 ± 0.5 | −27.8 ± 0.9 |
| (GGGT)4 8oxo3 | Parallel | 56.4 ± 0.4 | −13.9 ± 0.8 |
| (GGGT)4 8oxo5 | Parallel | 44.0 ± 0.3 | −26.3 ± 0.8 |
| (GGGTT)4 | Parallel | 44.8 ± 1.2 | - |
| (GGGTT)4 8oxo1 | Antiparallel | 47.2 ± 0.3 | 2.4 ± 1.2 |
| (GGGTT)4 8oxo3 | Antiparallel | 44.6 ± 0.2 | −0.2 ± 1.2 |
| (GGGTT)4 8oxo5 | Hybrid-esque | 27.5 ± 0.9 | −17.3 ± 1.4 |
| (TTAGGG)4 | Antiparallel | 52.1 ± 0.3 | - |
| (TTAGGG)4 8oxo1 | Antiparallel | 47.8 ± 0.6 | −4.3 ± 0.7 |
| (TTAGGG)4 8oxo3 | Antiparallel | 42.2 ± 0.7 | −9.9 ± 0.8 |
| (TTAGGG)4 8oxo5 | Antiparallel | 33.3 ± 0.4 | −18.8 ± 0.5 |
| Name | K (M−1) | R2 |
|---|---|---|
| GGGT)4 | 2.7 ± 0.4 × 105 | 0.997 |
| (GGGT)4 8oxo1 | 7.7 ± 0.7 × 105 | 0.998 |
| (GGGT)4 8oxo3 | 3.2 ± 0.1 × 105 | 0.997 |
| (GGGT)4 8oxo5 | 4.4 ± 0.6 × 105 | 0.997 |
| (GGGTT)4 | 6.0 ± 0.9 × 105 | 0.997 |
| (GGGTT)4 8oxo1 | 8.2 ± 1.0 × 105 | 0.997 |
| (GGGTT)4 8oxo3 | 9.6 ± 1.0 × 105 | 0.996 |
| (GGGTT)4 8oxo5 | 9.3 ± 0.8 × 105 | 0.997 |
| (TTAGGG)4 | 1.6 ± 0.7 × 106 | 0.996 |
| (TTAGGG)4 8oxo1 | 1.3 ± 0.4 × 106 | 0.997 |
| (TTAGGG)4 8oxo3 | 1.2 ± 0.1 × 106 | 0.996 |
| (TTAGGG)4 8oxo5 | 1.4 ± 0.5 × 106 | 0.997 |
| Name | K (M−1) | R2 |
|---|---|---|
| (GGGT)4 | 2.3 ± 0.3 × 105 | 0.9997 |
| (GGGT)4 8oxo1 | 3.4 ± 0.5 × 105 | 0.9996 |
| (GGGT)4 8oxo3 | 2.9 ± 0.5 × 105 | 0.9994 |
| (GGGT)4 8oxo5 | 3.4 ± 0.5 × 105 | 0.9997 |
| (GGGTT)4 | 4.9 ± 2.0 × 105 | 0.9995 |
| (GGGTT)4 8oxo1 | 3.9 ± 1.3 × 105 | 0.9993 |
| (GGGTT)4 8oxo3 | 5.2 ± 1.7 × 105 | 0.9994 |
| (GGGTT)4 8oxo5 | 5.6 ± 1.1 × 105 | 0.9992 |
| (TTAGGG)4 | 6.5 ± 0.6 × 105 | 0.9991 |
| (TTAGGG)4 8oxo1 | 6.5 ± 1.4 × 105 | 0.9996 |
| (TTAGGG)4 8oxo3 | 5.1 ± 1.1 × 105 | 0.9998 |
| (TTAGGG)4 8oxo5 | 1.1 ± 0.1 × 106 | 0.9974 |
| Name | Sequence (5′→3′) | Molar Extinction Coefficient (M−1·cm−1) | Molecular Weight (g/mol) |
|---|---|---|---|
| (GGGT)4 | GGGTGGGTGGGTGGGT | 157,200 | 5105.3 |
| (GGGT)4 8oxo1 | T/i8oxodG/GGTGGGTGGGTGGGT | 159,100 | 5425.5 |
| (GGGT)4 8oxo3 | GG/i8oxodG/TGGGTGGGTGGGT | 151,600 | 5121.3 |
| (GGGT)4 8oxo5 | GGGTG/i8oxodG/GTGGGTGGGT | 151,600 | 5121.3 |
| (GGGTT)4 | GGGTTGGGTTGGGTTGGGTT | 189,600 | 6322.1 |
| (GGGTT)4 8oxo1 | /i8oxodG/GGTTGGGTTGGGTTGGGTT | 184,000 | 6338.1 |
| (GGGTT)4 8oxo3 | GG/i8oxodG/TTGGGTGGGTTGGGTT | 184,000 | 6338.1 |
| (GGGTT)4 8oxo5 | GGGTTG/i8oxodG/GTTGGGTTGGGTT | 184,000 | 6338.1 |
| (TTAGGG)4 | TTAGGGTTAGGGTTAGGGTTAGGG | 244,600 | 7575.0 |
| (TTAGGG)4 8oxo1 | TTA/i8oxodG/GGTTAGGGTTAGGGTTAGGG | 239,000 | 7590.9 |
| (TTAGGG)4 8oxo3 | TTAGG/i8oxodG/TTAGGGTTAGGGTTAGGG | 239,000 | 7590.9 |
| (TTAGGG)4 8oxo5 | TTAGGGTTAG/i8oxodG/GTTAGGGTTAGGG | 239,000 | 7590.9 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Campbell, L.; Lowran, K.; Cismas, E.; Wu, C.G. 8-Oxoguanine Disrupts G-Quadruplex DNA Stability and Modulates FANCJ AKKQ Peptide Binding. Molecules 2025, 30, 3424. https://doi.org/10.3390/molecules30163424
Campbell L, Lowran K, Cismas E, Wu CG. 8-Oxoguanine Disrupts G-Quadruplex DNA Stability and Modulates FANCJ AKKQ Peptide Binding. Molecules. 2025; 30(16):3424. https://doi.org/10.3390/molecules30163424
Chicago/Turabian StyleCampbell, Laura, Kaitlin Lowran, Emma Cismas, and Colin G. Wu. 2025. "8-Oxoguanine Disrupts G-Quadruplex DNA Stability and Modulates FANCJ AKKQ Peptide Binding" Molecules 30, no. 16: 3424. https://doi.org/10.3390/molecules30163424
APA StyleCampbell, L., Lowran, K., Cismas, E., & Wu, C. G. (2025). 8-Oxoguanine Disrupts G-Quadruplex DNA Stability and Modulates FANCJ AKKQ Peptide Binding. Molecules, 30(16), 3424. https://doi.org/10.3390/molecules30163424







