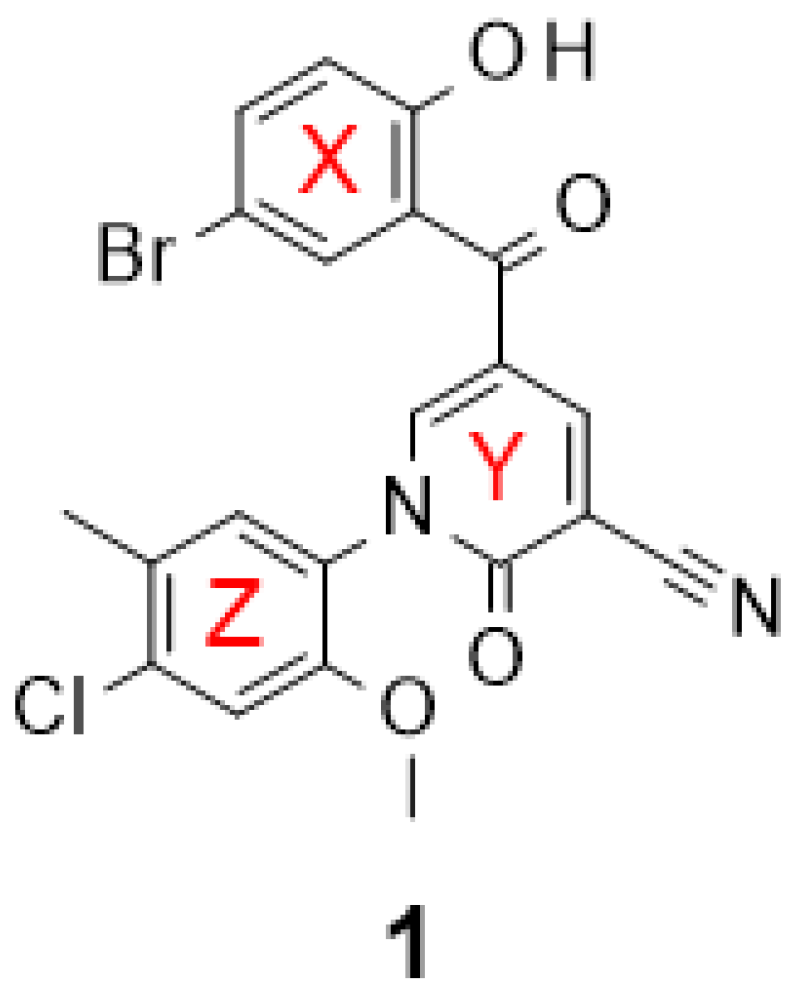
Discovery of Substituted 5-(2-Hydroxybenzoyl)-2-Pyridone Analogues as Inhibitors of the Human Caf1/CNOT7 Ribonuclease
Abstract
1. Introduction
2. Results and Discussion
3. Materials and Methods
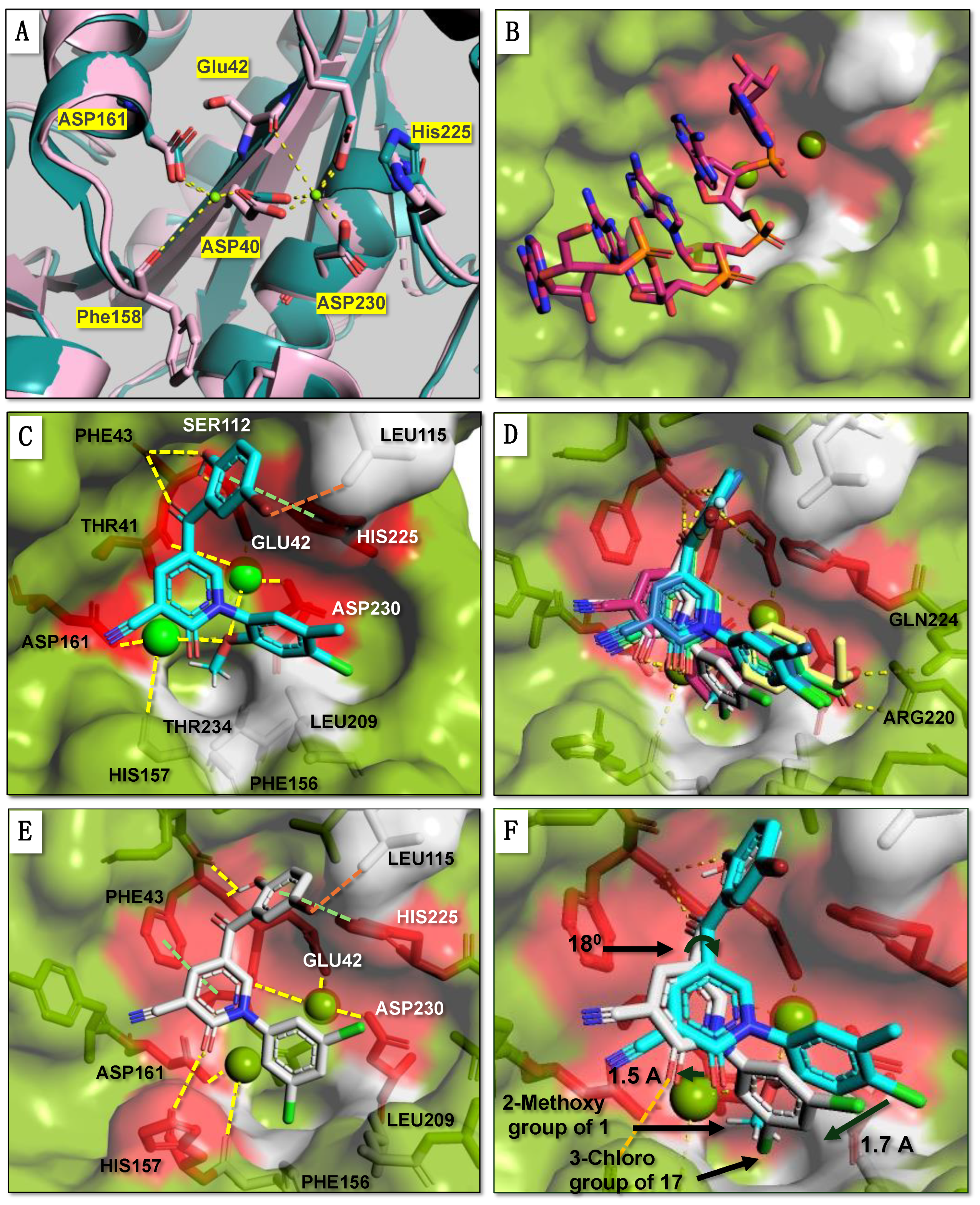
Molecular Docking
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Bianchin, C.; Mauxion, F.; Sentis, S.; Seraphin, B.; Corbo, L. Conservation of the deadenylase activity of proteins of the Caf1 family in human. RNA 2005, 11, 487–494. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Goldstrohm, A.C.; Wickens, M. Multifunctional deadenylase complexes diversify mRNA control. Nat. Rev. Mol. Cell Biol. 2008, 9, 337–344. [Google Scholar] [CrossRef] [PubMed]
- Wahle, E.; Winkler, G.S. RNA decay machines: Deadenylation by the Ccr4-Not and Pan2-Pan3 complexes. Biochim. Biophys. Acta 2013, 1829, 561–570. [Google Scholar] [CrossRef] [PubMed]
- Raisch, T.; Valkov, E. Regulation of the multisubunit CCR4-NOT deadenylase in the initiation of mRNA degradation. Curr. Opin. Struct. Biol. 2022, 77, 102460. [Google Scholar] [CrossRef]
- Krempl, C.; Lazzaretti, D.; Sprangers, R. A structural biology view on the enzymes involved in eukaryotic mRNA turnover. Biol. Chem. 2023, 404, 1101–1121. [Google Scholar] [CrossRef]
- Yang, W. Nucleases: Diversity of structure, function and mechanism. Q. Rev. Biophys. 2011, 44, 1–93. [Google Scholar] [CrossRef]
- Zuo, Y.; Deutscher, M.P. Exoribonuclease superfamilies: Structural analysis and phylogenetic distribution. Nucleic Acids Res. 2001, 29, 1017–1026. [Google Scholar] [CrossRef]
- Horiuchi, M.; Takeuchi, K.; Noda, N.; Muroya, N.; Suzuki, T.; Nakamura, T.; Kawamura-Tsuzuku, J.; Takahasi, K.; Yamamoto, T.; Inagaki, F. Structural basis for the antiproliferative activity of the Tob-hCaf1 complex. J. Biol. Chem. 2009, 284, 13244–13255. [Google Scholar] [CrossRef]
- Jonstrup, A.T.; Andersen, K.R.; Van, L.B.; Brodersen, D.E. The 1.4-A crystal structure of the S. pombe Pop2p deadenylase subunit unveils the configuration of an active enzyme. Nucleic Acids Res. 2007, 35, 3153–3164. [Google Scholar] [CrossRef]
- Basquin, J.; Roudko, V.V.; Rode, M.; Basquin, C.; Seraphin, B.; Conti, E. Architecture of the nuclease module of the yeast Ccr4-not complex: The Not1-Caf1-Ccr4 interaction. Mol. Cell 2012, 48, 207–218. [Google Scholar] [CrossRef]
- Petit, A.P.; Wohlbold, L.; Bawankar, P.; Huntzinger, E.; Schmidt, S.; Izaurralde, E.; Weichenrieder, O. The structural basis for the interaction between the CAF1 nuclease and the NOT1 scaffold of the human CCR4-NOT deadenylase complex. Nucleic Acids Res. 2012, 40, 11058–11072. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.; Pavanello, L.; Potapov, A.; Bartlam, M.; Winkler, G.S. Structure of the human Ccr4-Not nuclease module using X-ray crystallography and electron paramagnetic resonance spectroscopy distance measurements. Protein Sci. 2022, 31, 758–764. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Khazina, E.; Izaurralde, E.; Weichenrieder, O. Crystal structure and functional properties of the human CCR4-CAF1 deadenylase complex. Nucleic Acids Res. 2021, 49, 6489–6510. [Google Scholar] [CrossRef] [PubMed]
- Tucker, M.; Valencia-Sanchez, M.A.; Staples, R.R.; Chen, J.; Denis, C.L.; Parker, R. The transcription factor associated Ccr4 and Caf1 proteins are components of the major cytoplasmic mRNA deadenylase in Saccharomyces cerevisiae. Cell 2001, 104, 377–386. [Google Scholar] [CrossRef] [PubMed]
- Tucker, M.; Staples, R.R.; Valencia-Sanchez, M.A.; Muhlrad, D.; Parker, R. Ccr4p is the catalytic subunit of a Ccr4p/Pop2p/Notp mRNA deadenylase complex in Saccharomyces cerevisiae. Embo J. 2002, 21, 1427–1436. [Google Scholar] [CrossRef]
- Chen, J.; Chiang, Y.C.; Denis, C.L. CCR4, a 3′-5′ poly(A) RNA and ssDNA exonuclease, is the catalytic component of the cytoplasmic deadenylase. Embo J. 2002, 21, 1414–1426. [Google Scholar] [CrossRef]
- Temme, C.; Zaessinger, S.; Meyer, S.; Simonelig, M.; Wahle, E. A complex containing the CCR4 and CAF1 proteins is involved in mRNA deadenylation in Drosophila. Embo J. 2004, 23, 2862–2871. [Google Scholar] [CrossRef]
- Yamashita, A.; Chang, T.C.; Yamashita, Y.; Zhu, W.; Zhong, Z.; Chen, C.Y.; Shyu, A.B. Concerted action of poly(A) nucleases and decapping enzyme in mammalian mRNA turnover. Nat. Struct. Mol. Biol. 2005, 12, 1054–1063. [Google Scholar] [CrossRef]
- Dupressoir, A.; Morel, A.P.; Barbot, W.; Loireau, M.P.; Corbo, L.; Heidmann, T. Identification of four families of yCCR4- and Mg2+-dependent endonuclease-related proteins in higher eukaryotes, and characterization of orthologs of yCCR4 with a conserved leucine-rich repeat essential for hCAF1/hPOP2 binding. BMC Genom. 2001, 2, 9. [Google Scholar] [CrossRef]
- Airhihen, B.; Pavanello, L.; Jadhav, G.P.; Fischer, P.M.; Winkler, G.S. 1-Hydroxy-xanthine derivatives inhibit the human Caf1 nuclease and Caf1-containing nuclease complexes via Mg2+-dependent binding. FEBS Open Bio 2019, 9, 717–727. [Google Scholar] [CrossRef]
- Maryati, M.; Airhihen, B.; Winkler, G.S. The enzyme activities of Caf1 and Ccr4 are both required for deadenylation by the human Ccr4-Not nuclease module. Biochem. J. 2015, 469, 169–176. [Google Scholar] [CrossRef] [PubMed]
- Pekovic, F.; Rammelt, C.; Kubikova, J.; Metz, J.; Jeske, M.; Wahle, E. RNA binding proteins Smaug and Cup induce CCR4-NOT-dependent deadenylation of the nanos mRNA in a reconstituted system. Nucleic Acids Res. 2023, 51, 3950–3970. [Google Scholar] [CrossRef] [PubMed]
- Niinuma, S.; Fukaya, T.; Tomari, Y. CCR4 and CAF1 deadenylases have an intrinsic activity to remove the post-poly(A) sequence. RNA 2016, 22, 1550–1559. [Google Scholar] [CrossRef] [PubMed]
- Raisch, T.; Chang, C.T.; Levdansky, Y.; Muthukumar, S.; Raunser, S.; Valkov, E. Reconstitution of recombinant human CCR4-NOT reveals molecular insights into regulated deadenylation. Nat. Commun. 2019, 10, 3173. [Google Scholar] [CrossRef]
- Webster, M.W.; Chen, Y.H.; Stowell, J.A.W.; Alhusaini, N.; Sweet, T.; Graveley, B.R.; Coller, J.; Passmore, L.A. mRNA Deadenylation Is Coupled to Translation Rates by the Differential Activities of Ccr4-Not Nucleases. Mol. Cell 2018, 70, 1089–1100 e1088. [Google Scholar] [CrossRef]
- Yi, H.; Park, J.; Ha, M.; Lim, J.; Chang, H.; Kim, V.N. PABP Cooperates with the CCR4-NOT Complex to Promote mRNA Deadenylation and Block Precocious Decay. Mol. Cell 2018, 70, 1081–1088 e1085. [Google Scholar] [CrossRef]
- Aslam, A.; Mittal, S.; Koch, F.; Andrau, J.C.; Winkler, G.S. The Ccr4-Not Deadenylase Subunits CNOT7 and CNOT8 Have Overlapping Roles and Modulate Cell Proliferation. Mol. Biol. Cell 2009, 20, 3840–3850. [Google Scholar] [CrossRef]
- Mostafa, D.; Takahashi, A.; Yanagiya, A.; Yamaguchi, T.; Abe, T.; Kureha, T.; Kuba, K.; Kanegae, Y.; Furuta, Y.; Yamamoto, T.; et al. Essential functions of the CNOT7/8 catalytic subunits of the CCR4-NOT complex in mRNA regulation and cell viability. RNA Biol. 2020, 17, 403–416. [Google Scholar] [CrossRef]
- Stoney, P.N.; Yanagiya, A.; Nishijima, S.; Yamamoto, T. CNOT7 Outcompetes Its Paralog CNOT8 for Integration into The CCR4-NOT Complex. J. Mol. Biol. 2022, 434, 167523. [Google Scholar] [CrossRef]
- Shirai, Y.T.; Suzuki, T.; Morita, M.; Takahashi, A.; Yamamoto, T. Multifunctional roles of the mammalian CCR4-NOT complex in physiological phenomena. Front. Genet. 2014, 5, 286. [Google Scholar] [CrossRef]
- Song, X.H.; Liao, X.Y.; Zheng, X.Y.; Liu, J.Q.; Zhang, Z.W.; Zhang, L.N.; Yan, Y.B. Human Ccr4 and Caf1 Deadenylases Regulate Proliferation and Tumorigenicity of Human Gastric Cancer Cells via Modulating Cell Cycle Progression. Cancers 2021, 13, 834. [Google Scholar] [CrossRef] [PubMed]
- Washio-Oikawa, K.; Nakamura, T.; Usui, M.; Yoneda, M.; Ezura, Y.; Ishikawa, I.; Nakashima, K.; Noda, T.; Yamamoto, T.; Noda, M. Cnot7-null mice exhibit high bone mass phenotype and modulation of BMP actions. J. Bone Miner. Res. 2007, 22, 1217–1223. [Google Scholar] [CrossRef]
- Washio-Oikawa, K.; Nakamura, T.; Usui, M.; Yoneda, M.; Ezura, Y.; Ishikawa, I.; Nakashima, K.; Yamamoto, T.; Noda, M. Expression analysis of LacZ gene placed in the locus of Cnot7 exhibits its activity in osteoblasts in vivo and in mineralized nodules in vitro. J. Cell Biochem. 2006, 99, 538–544. [Google Scholar] [CrossRef] [PubMed]
- Levy, R.; Mott, R.F.; Iraqi, F.A.; Gabet, Y. Collaborative cross mice in a genetic association study reveal new candidate genes for bone microarchitecture. BMC Genom. 2015, 16, 1013. [Google Scholar] [CrossRef] [PubMed]
- Faraji, F.; Hu, Y.; Wu, G.; Goldberger, N.E.; Walker, R.C.; Zhang, J.; Hunter, K.W. An integrated systems genetics screen reveals the transcriptional structure of inherited predisposition to metastatic disease. Genome Res. 2014, 24, 227–240. [Google Scholar] [CrossRef] [PubMed]
- Faraji, F.; Hu, Y.; Yang, H.H.; Lee, M.P.; Winkler, G.S.; Hafner, M.; Hunter, K.W. Post-transcriptional Control of Tumor Cell Autonomous Metastatic Potential by CCR4-NOT Deadenylase CNOT7. PLoS Genet. 2016, 12, e1005820. [Google Scholar] [CrossRef]
- Maryati, M.; Kaur, I.; Gopal, J.; Olotu-Umoren, L.; Oveh, B.; Hashmi, L.; Fischer, P.M.; Winkler, G.S. A fluorescence-based assay suitable for quantitative analysis of deadenylase enzyme activity. Nucleic Acids Res. 2014, 42, e30. [Google Scholar] [CrossRef]
- Jadhav, G.P.; Kaur, I.; Maryati, M.; Airhihen, B.; Fischer, P.M.; Winkler, G.S. Discovery, synthesis and biochemical profiling of purine-2,6-dione derivatives as inhibitors of the human poly(A)-selective ribonuclease Caf1. Bioorganic Med. Chem. Lett. 2015, 25, 4219–4224. [Google Scholar] [CrossRef]
- Exell, J.C.; Thompson, M.J.; Finger, L.D.; Shaw, S.J.; Debreczeni, J.; Ward, T.A.; McWhirter, C.; Sioberg, C.L.; Martinez Molina, D.; Abbott, W.M.; et al. Cellularly active N-hydroxyurea FEN1 inhibitors block substrate entry to the active site. Nat. Chem. Biol. 2016, 12, 815–821. [Google Scholar] [CrossRef]
- Klumpp, K.; Hang, J.Q.; Rajendran, S.; Yang, Y.; Derosier, A.; Wong Kai In, P.; Overton, H.; Parkes, K.E.; Cammack, N.; Martin, J.A. Two-metal ion mechanism of RNA cleavage by HIV RNase H and mechanism-based design of selective HIV RNase H inhibitors. Nucleic Acids Res. 2003, 31, 6852–6859. [Google Scholar] [CrossRef]
- Parkes, K.E.; Ermert, P.; Fassler, J.; Ives, J.; Martin, J.A.; Merrett, J.H.; Obrecht, D.; Williams, G.; Klumpp, K. Use of a pharmacophore model to discover a new class of influenza endonuclease inhibitors. J. Med. Chem. 2003, 46, 1153–1164. [Google Scholar] [CrossRef] [PubMed]
- Tumey, L.N.; Bom, D.; Huck, B.; Gleason, E.; Wang, J.; Silver, D.; Brunden, K.; Boozer, S.; Rundlett, S.; Sherf, B.; et al. The identification and optimization of a N-hydroxy urea series of flap endonuclease 1 inhibitors. Bioorganic Med. Chem. Lett. 2005, 15, 277–281. [Google Scholar] [CrossRef] [PubMed]
- Trott, O.; Olson, A.J. AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 2010, 31, 455–461. [Google Scholar] [CrossRef] [PubMed]
- Eberhardt, J.; Santos-Martins, D.; Tillack, A.F.; Forli, S. AutoDock Vina 1.2.0: New Docking Methods, Expanded Force Field, and Python Bindings. J. Chem. Inf. Model. 2021, 61, 3891–3898. [Google Scholar] [CrossRef] [PubMed]
- Tang, T.T.L.; Stowell, J.A.W.; Hill, C.H.; Passmore, L.A. The intrinsic structure of poly(A) RNA determines the specificity of Pan2 and Caf1 deadenylases. Nat. Struct. Mol. Biol. 2019, 26, 433–442. [Google Scholar] [CrossRef]
- Airhihen, B.; Pavanello, L.; Maryati, M.; Winkler, G.S. Quantitative Biochemical Analysis of Deadenylase Enzymes Using Fluorescence and Chemiluminescence-Based Assays. Methods Mol. Biol. 2024, 2723, 55–68. [Google Scholar]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Couch, G.S.; Greenblatt, D.M.; Meng, E.C.; Ferrin, T.E. UCSF Chimera--a visualization system for exploratory research and analysis. J. Comput. Chem. 2004, 25, 1605–1612. [Google Scholar] [CrossRef]
- Diedrich, K.; Krause, B.; Berg, O.; Rarey, M. PoseEdit: Enhanced ligand binding mode communication by interactive 2D diagrams. J. Comput. Aided Mol. Des. 2023, 37, 491–503. [Google Scholar] [CrossRef]
- Schoning-Stierand, K.; Diedrich, K.; Fahrrolfes, R.; Flachsenberg, F.; Meyder, A.; Nittinger, E.; Steinegger, R.; Rarey, M. ProteinsPlus: Interactive analysis of protein-ligand binding interfaces. Nucleic Acids Res. 2020, 48, W48–W53. [Google Scholar] [CrossRef]
- Schrödinger, L.L.C. The PyMol Molecular Graphics System; version 2.5.0 (open source); De-Lano Scientific: San Carlos, CA, USA, 2021. [Google Scholar]
- SwissADME: A free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci. Rep. 2017, 7, 42717. [CrossRef]
- iLOGP: A simple, robust, and efficient description of n-octanol/water partition coefficient for drug design using the GB/SA approach. J. Chem. Inf. Model. 2014, 54, 3284–3301. [CrossRef]
- A BOILED-Egg to predict gastrointestinal absorption and brain penetration of small molecules. ChemMedChem 2016, 11, 1117–1121. [CrossRef]
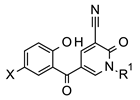 | Docking Score b | 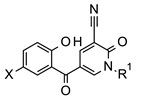 | Docking Score b | ||||||
|---|---|---|---|---|---|---|---|---|---|
| Cmpd | X | R1 | IC50 (μM) a | Cmpd | X | R1 | IC50 (μM) a | ||
| 1 | Br | 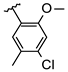 | 24.1 ± 3.3 (n = 3) c | −8.17 | |||||
| 2 | Br | 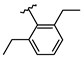 | 39.4 ± 3.1 (n = 4) | - | 10 | Br | 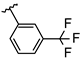 | 42.1 ± 1.4 (n = 4) | - |
| 3 | Br | 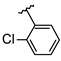 | 65.6 ± 3.3 (n = 4) | - | 11 | F |  | 33.6 ± 1.1 (n = 4) | −8.40 |
| 4 | Br |  | 39.3 ± 1.6 (n = 4) | - | 12 | Br | 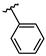 | 108.3 ± 4.0 (n = 4) | −8.11 |
| 5 | Br |  | 81.1 ± 5.0 (n = 4) | - | 13 | Br | 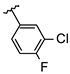 | 47.0 ± 2.7 (n = 4) | - |
| 6 | Br |  | 80.9 ± 9.2 (n = 4) | - | 14 | Br | 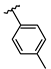 | 57.5 ± 4.7 (n = 4) | - |
| 7 | Br | 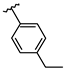 | 39.0 ± 2.6 (n = 4) | - | 15 | Br | 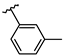 | 31.8 ± 1.8 (n = 3) | −9.06 |
| 8 | Br | 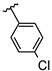 | 32.2 ± 1.6 (n = 4) | −8.53 | 16 | Br | 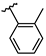 | 44.5 ± 3.6 (n = 3) | - |
| 9 | Br |  | 33.2 ± 0.5 (n = 4) | −8.97 | 17 | Br | 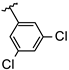 | 19.2 ± 1.1 (n = 3) | −8.72 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kaur, I.; Jadhav, G.P.; Fischer, P.M.; Winkler, G.S. Discovery of Substituted 5-(2-Hydroxybenzoyl)-2-Pyridone Analogues as Inhibitors of the Human Caf1/CNOT7 Ribonuclease. Molecules 2024, 29, 4351. https://doi.org/10.3390/molecules29184351
Kaur I, Jadhav GP, Fischer PM, Winkler GS. Discovery of Substituted 5-(2-Hydroxybenzoyl)-2-Pyridone Analogues as Inhibitors of the Human Caf1/CNOT7 Ribonuclease. Molecules. 2024; 29(18):4351. https://doi.org/10.3390/molecules29184351
Chicago/Turabian StyleKaur, Ishwinder, Gopal P. Jadhav, Peter M. Fischer, and Gerlof Sebastiaan Winkler. 2024. "Discovery of Substituted 5-(2-Hydroxybenzoyl)-2-Pyridone Analogues as Inhibitors of the Human Caf1/CNOT7 Ribonuclease" Molecules 29, no. 18: 4351. https://doi.org/10.3390/molecules29184351
APA StyleKaur, I., Jadhav, G. P., Fischer, P. M., & Winkler, G. S. (2024). Discovery of Substituted 5-(2-Hydroxybenzoyl)-2-Pyridone Analogues as Inhibitors of the Human Caf1/CNOT7 Ribonuclease. Molecules, 29(18), 4351. https://doi.org/10.3390/molecules29184351







