Marine-Derived Compounds as Potential Inhibitors of Hsp90 for Anticancer and Antimicrobial Drug Development: A Comprehensive In Silico Study
Abstract
:1. Introduction
2. Results
2.1. Generation and Validation of Target-Based Pharmacophore Model
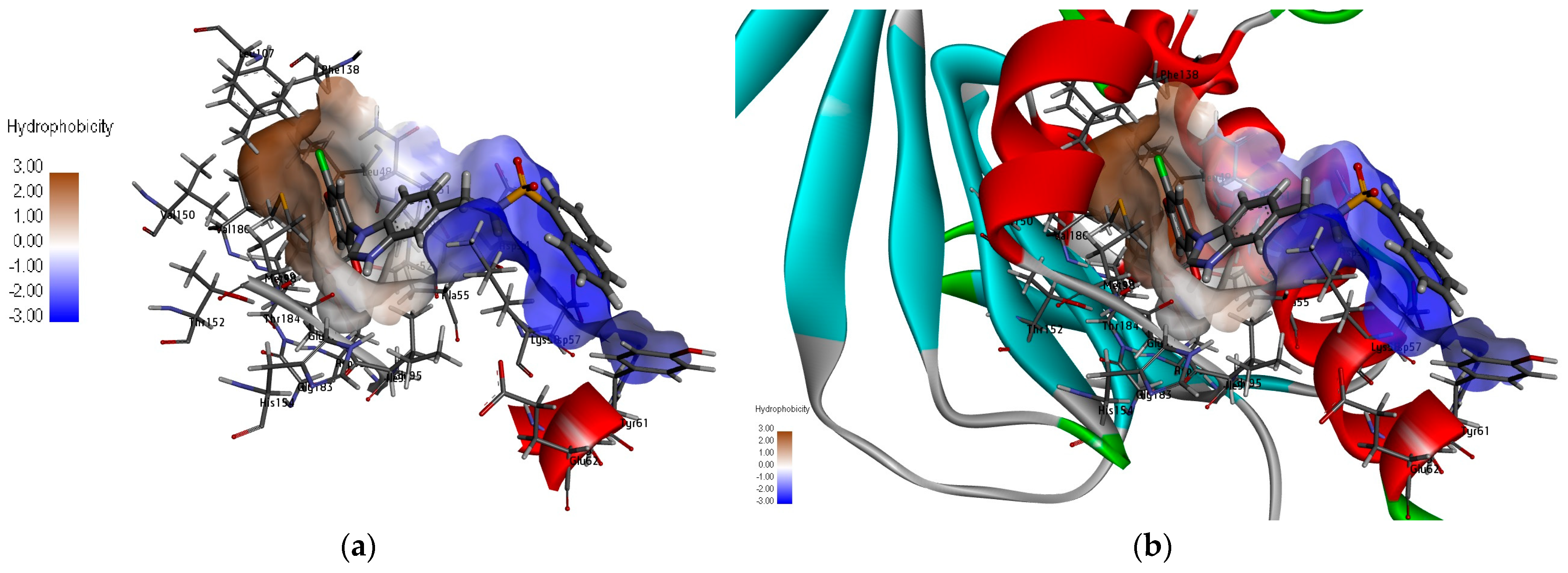
2.2. Target-Based Virtual Screening (TBVS)
2.3. Pharmacokinetic Properties and Drug Likeness
2.3.1. Molecular Properties
2.3.2. Medicinal Chemistry
2.3.3. Absorption
2.3.4. Distribution
2.3.5. Metabolism and Excretion
2.3.6. Prediction of Toxicity
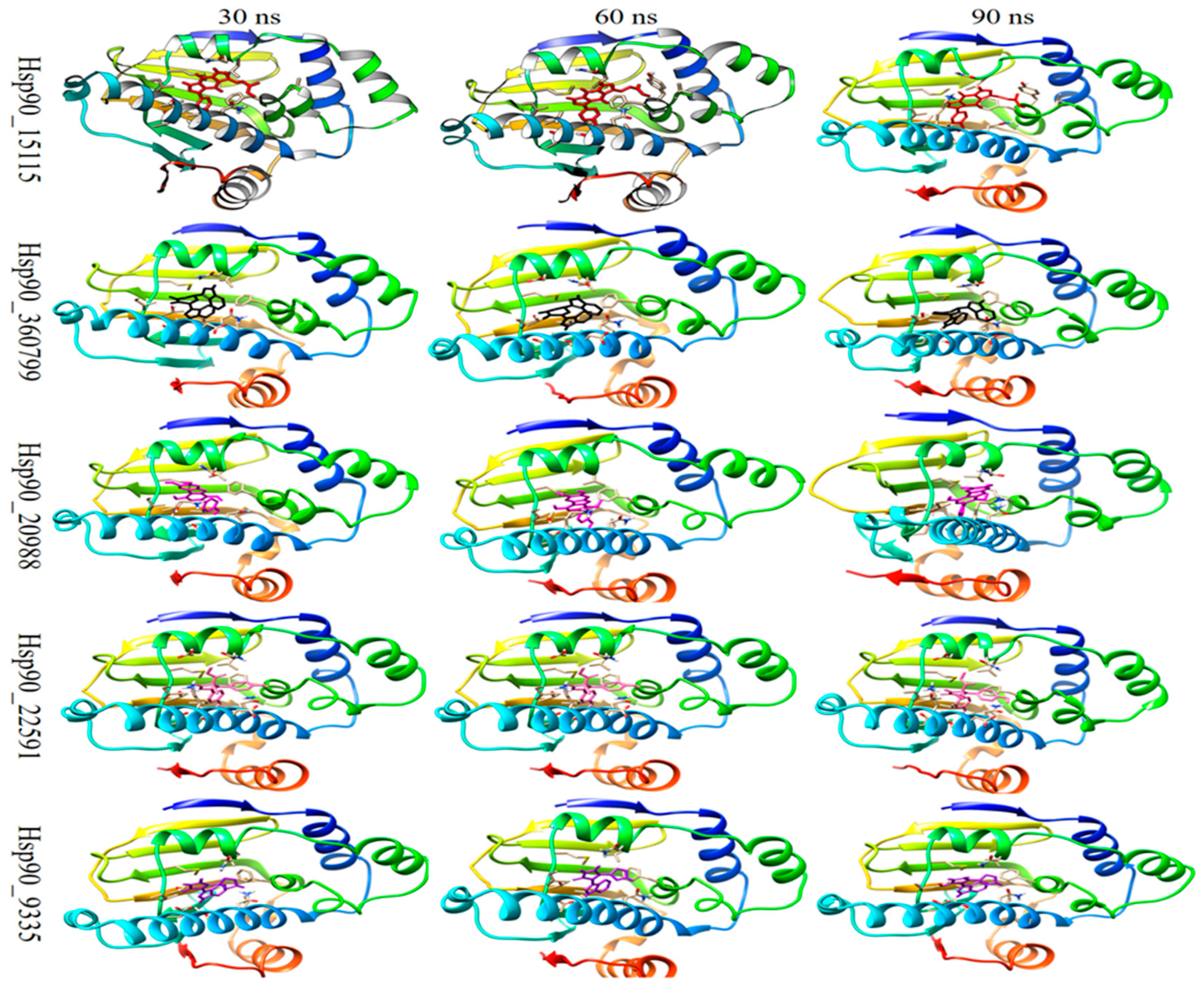
2.4. Molecular Dynamics Analysis
2.4.1. RMSD of HSP90 and Its Complexes
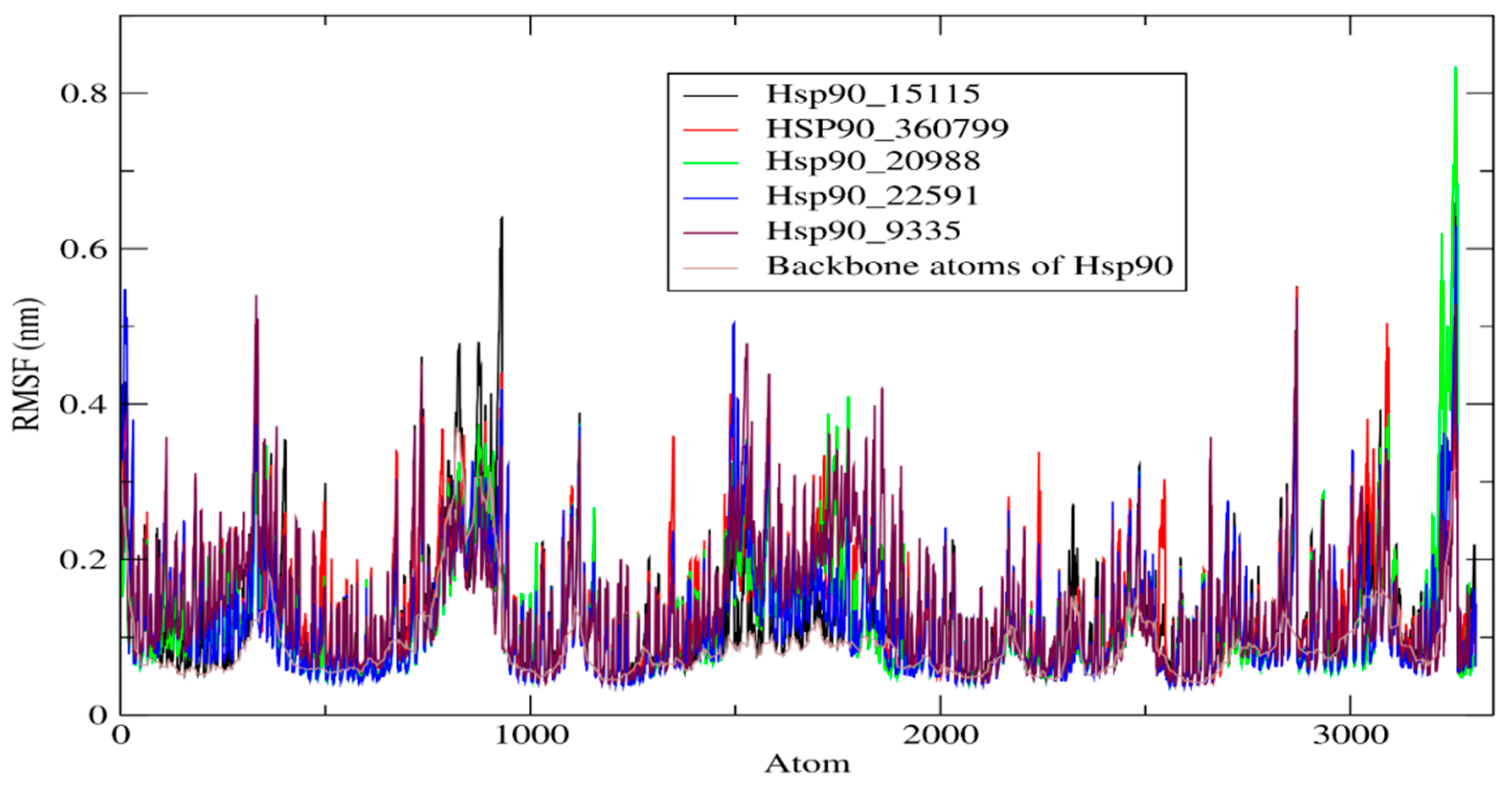
2.4.2. RMSF of HSP90 and Its Complexes
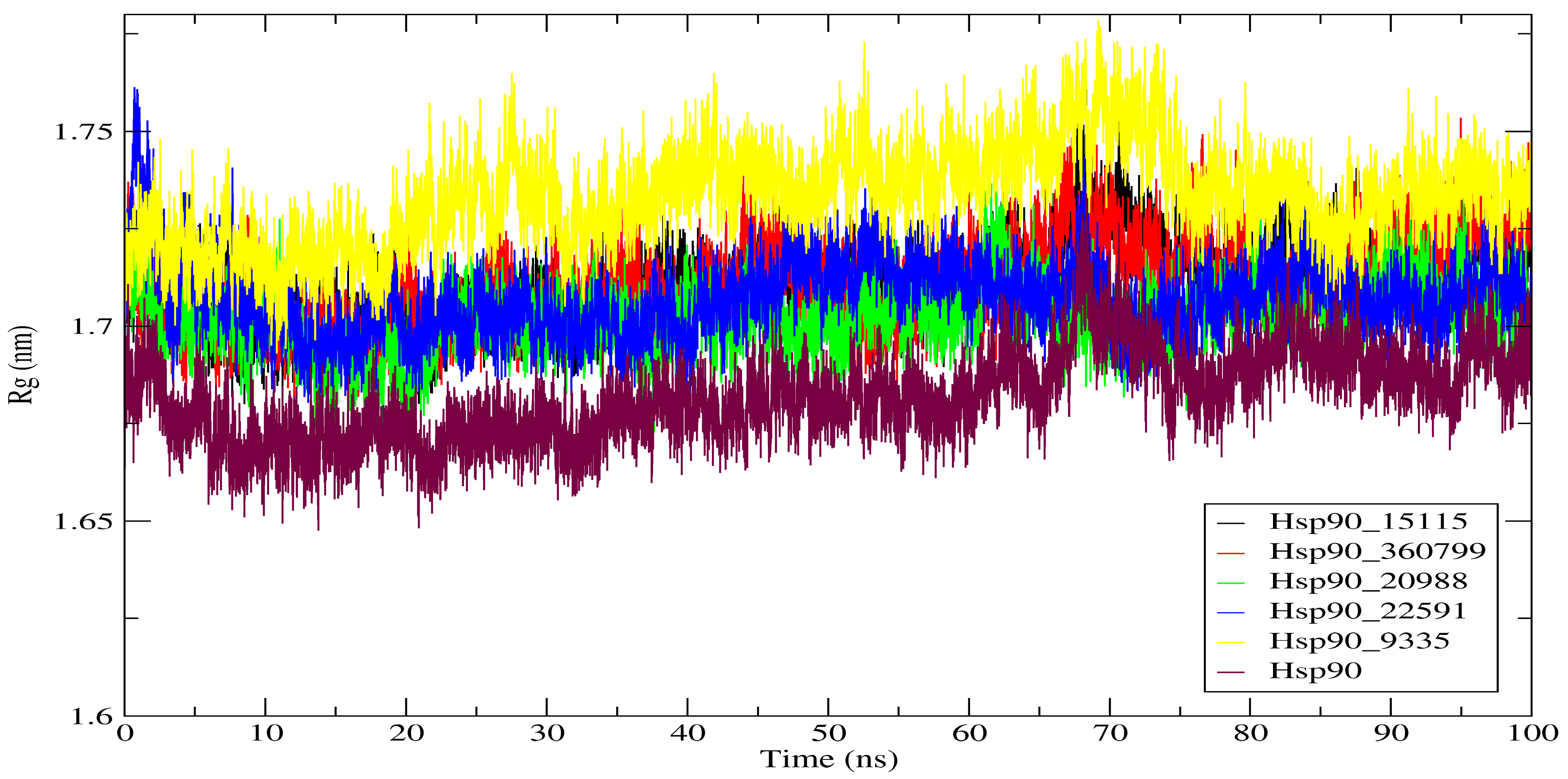
2.4.3. Radius of Gyration (Rg)
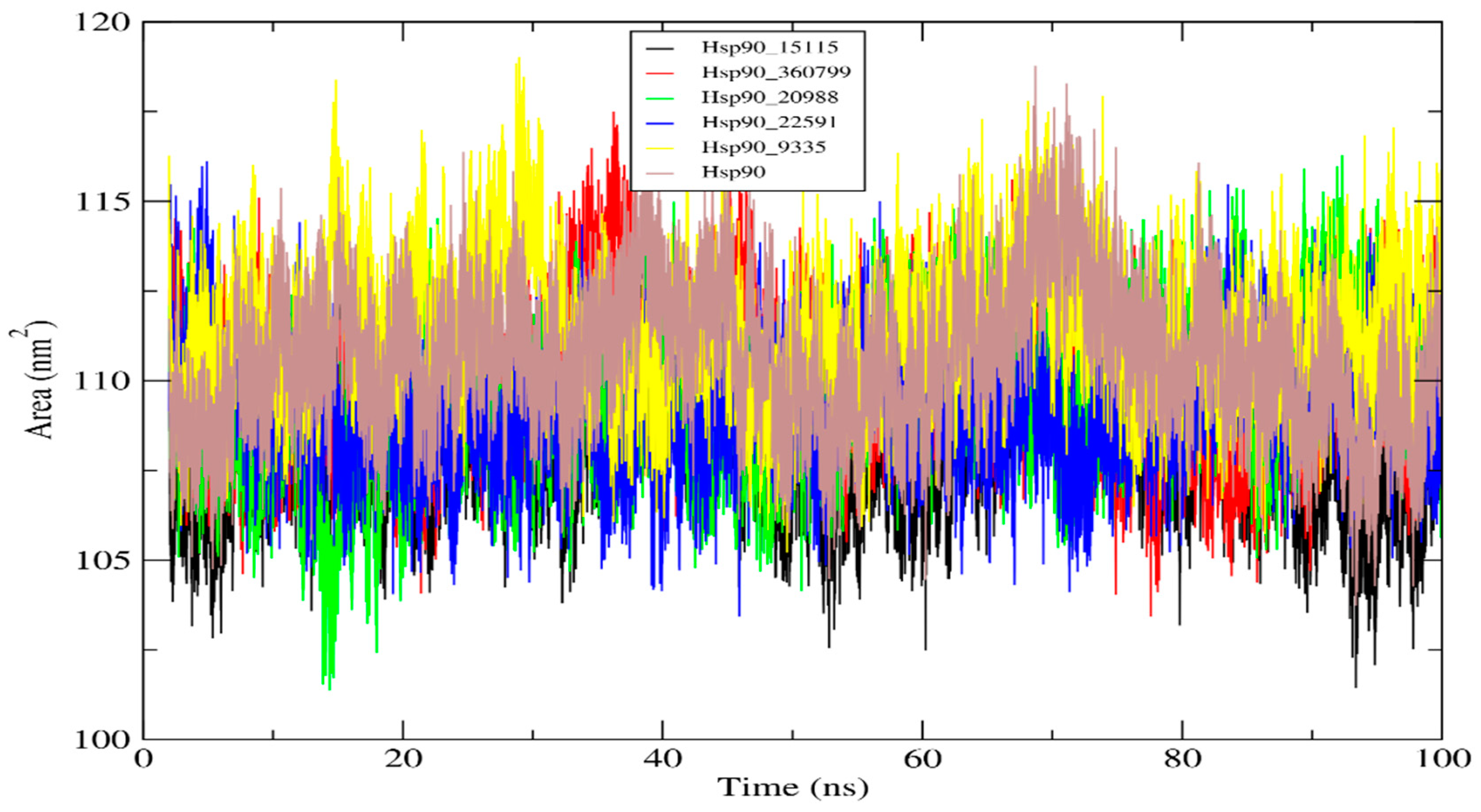
2.4.4. Solvent Accessible Surface Area (SASA) Analysis
2.4.5. MM-PBSA Calculations
3. Discussion
4. Materials and Methods
4.1. Preparation of Library of Natural Compounds
4.2. Optimizing Protein Crystal Structures
4.3. Molecular Docking Studies
4.4. Pharmacophore Modeling and Enrichment Study
4.5. Virtual Screening
4.6. In Silico and ADME-Tox and Drug-Likeness Prediction
4.7. Molecular Dynamics Investigation
4.8. MM-PBSA Binding Free Energy Calculations
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Kwon, J.H.; Powderly, W.G. The post-antibiotic era is here. Am. Assoc. Adv. Sci. 2021, 373, 471. [Google Scholar] [CrossRef]
- Harini, R.; Pavithra, P.; Nirmala, E.; Pavazhaviji, P. Challenges faced in the discovery of new antibiotics: A focus on antimicrobial resistance. World J. Pharm. Pharm. 2023, 12, 293–305. [Google Scholar]
- Wang, L.; Song, Y.; Wang, H.; Zhang, X.; Wang, M.; He, J.; Li, S.; Zhang, L.; Li, K.; Cao, L. Advances of Artificial Intelligence in Anti-Cancer Drug Design: A Review of the Past Decade. Pharmaceuticals 2023, 16, 253. [Google Scholar] [CrossRef]
- Grandori, C.; Kemp, C.J. Personalized cancer models for target discovery and precision medicine. Trends Cancer 2018, 4, 634–642. [Google Scholar] [CrossRef] [PubMed]
- Karthikeyan, A.; Joseph, A.; Nair, B.G. Promising bioactive compounds from the marine environment and their potential effects on various diseases. J. Genet. Eng. Biotechnol. 2022, 20, 14. [Google Scholar] [CrossRef]
- Matsumoto, R.; Fujii, Y.; Kawsar, S.M.A.; Kanaly, R.A.; Yasumitsu, H.; Koide, Y.; Hasan, I.; Iwahara, C.; Ogawa, Y.; Im, C.H.; et al. Cytotoxicity and glycan-binding properties of an 18 kDa lectin isolated from the marine sponge Halichondria okadai. Toxins 2012, 4, 323–338. [Google Scholar] [CrossRef]
- Fujii, Y.; Kawsar, S.M.A.; Matsumoto, R.; Yasumitsu, H.; Ishizaki, N.; Dogasaki, C.; Hosono, M.; Nitta, K.; Hamako, J.; Taei, M.; et al. A D-galactose-binding lectin purified from coronate moon turban, Turbo (Lunella) coreensis, with a unique amino acid sequence and the ability to recognize lacto-series glycosphingolipids. Comp. Biochem. Physiol. Part B Biochem. Mol. Biol. 2011, 158, 30–37. [Google Scholar] [CrossRef] [PubMed]
- Kang, H.K.; Choi, M.-C.; Seo, C.H.; Park, Y. Therapeutic properties and biological benefits of marine-derived anticancer peptides. Int. J. Mol. Sci. 2018, 19, 919. [Google Scholar] [CrossRef]
- Bajpai, V.K. Antimicrobial bioactive compounds from marine algae: A mini review. Indian J. Geo-Mar. Sci. 2016, 45, 1076–1085. [Google Scholar]
- Birbo, B.; Madu, E.E.; Madu, C.O.; Jain, A.; Lu, Y. Role of HSP90 in Cancer. Int. J. Mol. Sci. 2021, 22, 10317. [Google Scholar] [CrossRef]
- Neckers, L.; Ivy, S.P. Heat shock protein 90. Curr. Opin. Oncol. 2003, 15, 419–424. [Google Scholar] [CrossRef] [PubMed]
- Vo, C.D.; Shebert, H.L.; Zikovich, S.; Dryer, R.A.; Huang, T.P.; Moran, L.J.; Cho, J.; Wassarman, D.R.; Falahee, B.E.; Young, P.D. Repurposing Hsp90 inhibitors as antibiotics targeting histidine kinases. Bioorganic Med. Chem. Lett. 2017, 27, 5235–5244. [Google Scholar] [CrossRef] [PubMed]
- Eisele, F.; Eisele-Bürger, A.M.; Hao, X.; Berglund, L.L.; Höög, J.L.; Liu, B.; Nyström, T. An Hsp90 co-chaperone links protein folding and degradation and is part of a conserved protein quality control. Cell Rep. 2021, 35, 109328. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Li, H.; Liu, Y.; Zhao, K.; Wei, S.; Sugarman, E.T.; Liu, L.; Zhang, G. Targeting HSP90 as a novel therapy for cancer: Mechanistic insights and translational relevance. Cells 2022, 11, 2778. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.; Zhang, R.; Yang, H.; Xue, N.; Chen, X.; Yu, X. Rational design, synthesis, and biological evaluation of novel C6-modified geldanamycin derivatives as potent Hsp90 inhibitors and anti-tumor agents. Chin. Chem. Lett. 2023, 34, 107529. [Google Scholar] [CrossRef]
- Li, Y.; Zhang, T.; Schwartz, S.J.; Sun, D. New developments in Hsp90 inhibitors as anti-cancer therapeutics: Mechanisms, clinical perspective and more potential. Drug Resist. Updates 2009, 12, 17–27. [Google Scholar] [CrossRef]
- Schulte, T.W.; Akinaga, S.; Soga, S.; Sullivan, W.; Stensgard, B.; Toft, D.; Neckers, L.M. Antibiotic radicicol binds to the N-terminal domain of Hsp90 and shares important biologic activities with geldanamycin. Cell Stress Chaperones 1998, 3, 100. [Google Scholar] [CrossRef]
- Costa, T.E.; Raghavendra, N.M.; Penido, C. Natural heat shock protein 90 inhibitors in cancer and inflammation. Eur. J. Med. Chem. 2020, 189, 112063. [Google Scholar] [CrossRef]
- Neckers, L.; Workman, P. Hsp90 molecular chaperone inhibitors: Reaching the destination? Clin. Cancer Res. 2012, 18, 64–76. [Google Scholar] [CrossRef]
- Workman, P.; Burrows, F.; Neckers, L.; Rosen, N. Drugging the cancer chaperone HSP90: Exploiting oncogene addiction and tumor stress therapeutically. Ann. NY Acad Sci. 2007, 1113, 202–216. [Google Scholar] [CrossRef]
- Whitesell, L.; Lindquist, S.L. HSP90 and the chaperoning of cancer. Nat. Rev. Cancer 2005, 5, 761–772. [Google Scholar] [CrossRef]
- Kamal, A.; Thao, L.; Sensintaffar, J.; Zhang, L.; Boehm, M.F.; Fritz, L.C.; Burrows, F.J. A high-affinity conformation of Hsp90 confers tumor selectivity on Hsp90 inhibitors. Nature 2003, 425, 407–410. [Google Scholar] [CrossRef]
- Eustace, B.K.; Sakurai, T.; Stewart, J.K.; Yimlamai, D.; Unger, C.; Zehetmeier, C.; Lain, B.; Torella, C.; Henning, S.W.; Beste, G.; et al. Functional proteomic screens reveal an essential extracellular role for Hsp90 alpha in cancer cell invasiveness. Nat. Cell Biol. 2004, 6, 507–514. [Google Scholar] [CrossRef]
- Kitchen, D.B.; Decornez, H.; Furr, J.R.; Bajorath, J. Docking and scoring in virtual screening for drug discovery: Methods and applications. Nat. Rev. Drug Discov. 2004, 3, 935–949. [Google Scholar] [CrossRef]
- Blunt, J.W.; Copp, B.R.; Hu, W.-P.; Munro, M.H.; Northcote, P.T.; Prinsep, M.R. Marine natural products. Nat. Prod. Rep. 2007, 24, 31–86. [Google Scholar] [CrossRef]
- Faulkner, D.J. Marine natural products. Nat. Prod. Rep. 2001, 18, 1R–49R. [Google Scholar] [CrossRef] [PubMed]
- Walters, W.P. Going Further than Lipinski’s Rule in Drug Design. Expert Opin. Drug Discov. 2012, 7, 99–107. [Google Scholar] [CrossRef]
- Ouassaf, M.; Qais, F.A.; Belaidi, S.; Bakhouch, M.; Mohamed, A.S.; Chtita, S. Combined Pharmacophore Modeling, 3D-QSAR, Molecular Docking and Molecular Dynamics Study on Indolyl-aryl-sulfone Derivatives as New HIV1 Inhibitors. Acta Chim. Slov. 2022, 69, 489–506. [Google Scholar] [CrossRef] [PubMed]
- Ouassaf, M.; Daoui, O.; Alam, S.; Elkhattabi, S.; Belaidi, S.; Chtita, S. Pharmacophore-Based Virtual Screening, Molecular Docking, and Molecular Dynamics Studies for the Discovery of Novel FLT3 Inhibitors. J. Biomol. Struct. Dyn. 2023, 41, 7712–7724. [Google Scholar] [CrossRef]
- Lotfi, B.; Mebarka, O.; Khan, S.U.; Htar, T.T. Pharmacophore-based virtual screening, molecular docking and molecular dynamics studies for the discovery of novel neuraminidase inhibitors. J. Biomol. Struct. Dyn. 2023, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Abbasi, M.; Amanlou, M.; Aghaei, M.; Hassanzadeh, F.; Sadeghi-Aliabadi, H. Identification of new Hsp90 inhibitors: Structure based virtual screening, molecular dynamic simulation, synthesis and biological evaluation. Anti-Cancer Agents Med. Chem. (Former. Curr. Med. Chem.-Anti-Cancer Agents) 2021, 21, 2583–2591. [Google Scholar] [CrossRef] [PubMed]
- Chen, D.; Oezguen, N.; Urvil, P.; Ferguson, C.; Dann, S.M.; Savidge, T.C. Regulation of protein-ligand binding affinity by hydrogen bond pairing. Sci. Adv. 2016, 2, e1501240. [Google Scholar] [CrossRef] [PubMed]
- Ertl, P.; Schuffenhauer, A. Estimation of Synthetic Accessibility Score of Drug-like Molecules Based on Molecular Complexity and Fragment Contributions. J. Cheminformatics 2009, 1, 8. [Google Scholar] [CrossRef] [PubMed]
- Gleeson, M.P. Generation of a set of simple, interpretable ADMET rules of thumb. J. Med. Chem. 2008, 51, 817–834. [Google Scholar] [CrossRef] [PubMed]
- Lipinski, C.A. Lead-and drug-like compounds: The rule-of-five revolution. Drug Discov. Today Technol. 2004, 1, 337–341. [Google Scholar] [CrossRef] [PubMed]
- Juvale, I.I.A.; Hamid, A.A.A.; Abd Halim, K.B.; Has, A.T.C. P-glycoprotein: New insights into structure, physiological function, regulation and alterations in disease. Heliyon 2022, 8, e0977. [Google Scholar]
- Cho, M.J.; Thompson, D.P.; Cramer, C.T.; Vidmar, T.J.; Scieszka, J.F. The Madin Darby canine kidney (MDCK) epithelial cell monolayer as a model cellular transport barrier. Pharm. Res. 1989, 6, 71–77. [Google Scholar] [CrossRef]
- Baell, J.B.; Nissink, J.W.M. Seven Year Itch: Pan-Assay Interference Compounds (PAINS) in 2017 Utility and Limitations. ACS Chem. Biol. 2018, 13, 36–44. [Google Scholar] [CrossRef]
- Yang, Y.; Hu, B.; Lill, M.A. Analysis of factors influencing hydration site prediction based on molecular dynamics simulations. J. Chem. Inf. Model. 2014, 54, 2987–2995. [Google Scholar] [CrossRef]
- Miyata, Y.; Nakamoto, H.; Neckers, L. The therapeutic target Hsp90 and cancer hallmarks. Curr. Pharm. Des. 2013, 19, 347–365. [Google Scholar] [CrossRef]
- Santos, L.H.; Ferreira, R.S.; Caffarena, E.R. Integrating molecular docking and molecular dynamics simulations. Docking Screens Drug Discov. 2019, 2053, 13–34. [Google Scholar]
- Rana, K.M.; Maowa, J.; Alam, A.; Dey, S.; Hosen, A.; Hasan, I.; Fujii, Y.; Ozeki, Y.; Kawsar, S.M. In silico DFT study, molecular docking, and ADMET predictions of cytidine analogs with antimicrobial and anticancer properties. In Silico Pharmacol. 2021, 9, 1–24. [Google Scholar] [CrossRef]
- Caroli, A.; Ballante, F.; Wickersham, R.B., III; Corelli, F.; Ragno, R. Hsp90 inhibitors, part 2: Combining ligand-based and structure-based approaches for virtual screening application. J. Chem. Inf. Model. 2014, 54, 970–977. [Google Scholar] [CrossRef] [PubMed]
- Antony, P.; Baby, B.; Aleissaee, H.M.; Vijayan, R. A Molecular Modeling Investigation of the Therapeutic Potential of Marine Compounds as DPP-4 Inhibitors. Mar. Drugs 2022, 20, 777. [Google Scholar] [CrossRef] [PubMed]
- Cowen, L.E. Hsp90 orchestrates stress response signaling governing fungal drug resistance. PLoS Pathog. 2009, 5, e1000471. [Google Scholar] [CrossRef]
- Harder, E.; Damm, W.; Maple, J.; Wu, C.; Reboul, M.; Xiang, J.Y.; Wang, L.; Lupyan, D.; Dahlgren, M.K.; Knight, J.L. OPLS3: A force field providing broad coverage of drug-like small molecules and proteins. J. Chem. Theory Comput. 2016, 12, 281–296. [Google Scholar] [CrossRef] [PubMed]
- Bruncko, M.; Tahir, S.K.; Song, X.; Chen, J.; Ding, H.; Huth, J.R.; Jin, S.; Judge, R.A.; Madar, D.J.; Park, C.H.; et al. N-Aryl-Benzimidazolones as Novel Small Molecule HSP90 Inhibitors. Bioorganic Med. Chem. Lett. 2010, 20, 7503–7506. [Google Scholar] [CrossRef]
- McDonald, E.; Jones, K.; Brough, P.A.; Drysdale, M.J.; Workman, P. Discovery and Development of Pyrazole-Scaffold Hsp90 Inhibitors. Curr. Top. Med. Chem. 2006, 6, 1193–1203. [Google Scholar] [CrossRef]
- Karplus, M.; McCammon, J.A. Molecular dynamics simulations of biomolecules. Nat. Struct. Biol. 2002, 9, 646–652. [Google Scholar] [CrossRef]
- Wu, G.; Robertson, D.H.; Brooks, C.L.; Vieth, M. Detailed analysis of grid-based molecular docking: A case study of CDOCKER—A CHARMm-based MD docking algorithm. J. Comput. Chem. 2003, 24, 1549–1562. [Google Scholar] [CrossRef]
- El-Zahabi, H.S.; Khalifa, M.M.; Gado, Y.M.; Farrag, A.M.; Elaasser, M.M.; Safwat, N.A.; AbdelRaouf, R.R.; Arafa, R.K. New thiobarbituric acid scaffold-based small molecules: Synthesis, cytotoxicity, 2D-QSAR, pharmacophore modelling and in-silico ADME screening. Eur. J. Pharm. Sci. 2019, 130, 124–136. [Google Scholar] [CrossRef] [PubMed]
- Shams ul Hassan, S.; Abbas, S.Q.; Hassan, M.; Jin, H.-Z. Computational exploration of anti-cancer potential of guaiane dimers from Xylopia vielana by targeting B-RAF kinase using chemo-informatics, molecular docking, and MD simulation studies. Anti-Cancer Agents Med. Chem. (Former. Curr. Med. Chem.-Anti-Cancer Agents) 2022, 22, 731–746. [Google Scholar] [CrossRef] [PubMed]
- Alvarez, J.C. High-throughput docking as a source of novel drug leads. Curr. Opin. Chem. Biol. 2004, 8, 365–370. [Google Scholar] [CrossRef] [PubMed]
- Zoete, V.; Daina, A.; Bovigny, C.; Michielin, O. SwissSimilarity: A Web Tool for Low to Ultra High throughput Ligand-Based Virtual Screening; ACS Publications: Washington, DC, USA, 2016. [Google Scholar]
- Roy, K.; Kar, S.; Das, R.N. A primer on QSAR/QSPR Modeling: Fundamental Concepts; Springer: Berlin/Heidelberg, Germany, 2015. [Google Scholar]
- Wang, E.; Sun, H.; Wang, J.; Wang, Z.; Liu, H.; Zhang, J.Z.; Hou, T. End-point binding free energy calculation with MM/PBSA and MM/GBSA: Strategies and applications in drug design. Chem. Rev. 2019, 119, 9478–9508. [Google Scholar] [CrossRef]



| Compound CMNPD | SP (kcal/mol) | XP (kcal/mol) |
|---|---|---|
| 22591 | −8.97 | −9.03 |
| 9335 | −9.04 | −8.98 |
| 15115 | −8.71 | −8.73 |
| 20988 | −8.86 | −8.43 |
| 10015 | −7.91 | −8.35 |
| 360799 | −7.62 | −8.74 |
| MEY | −8.84 | −8.32 |
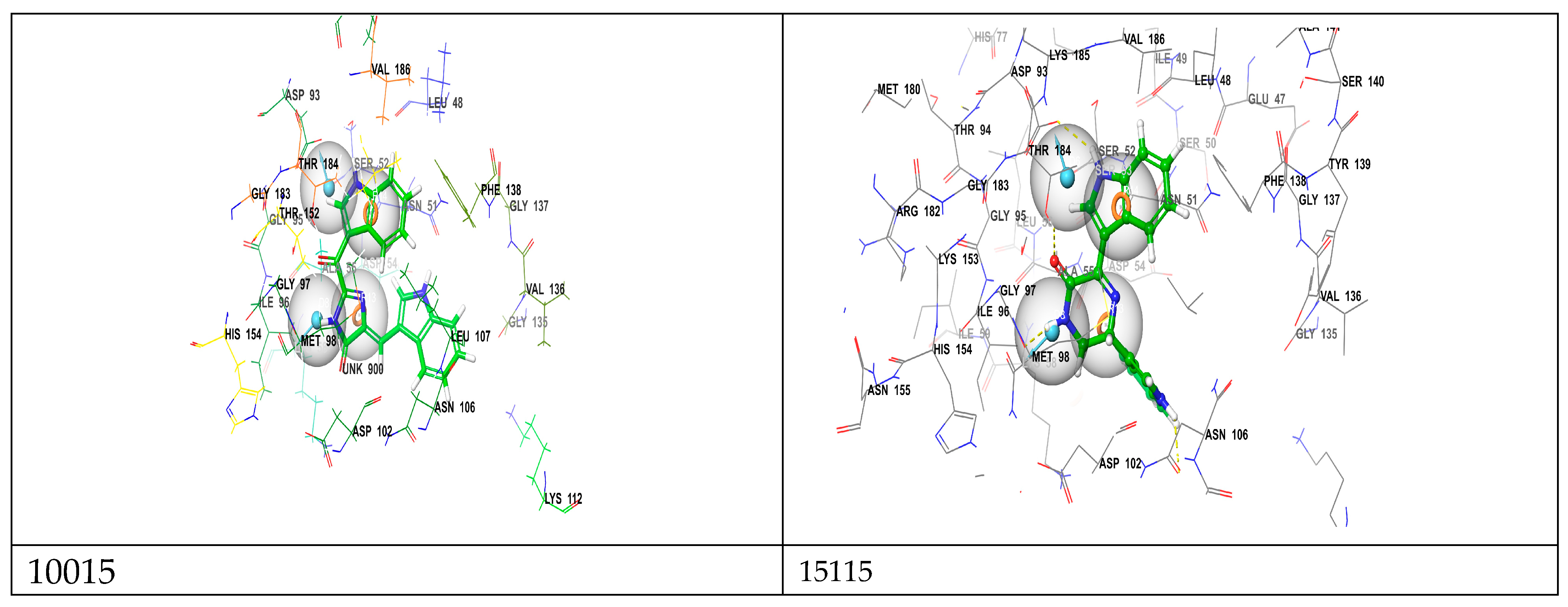
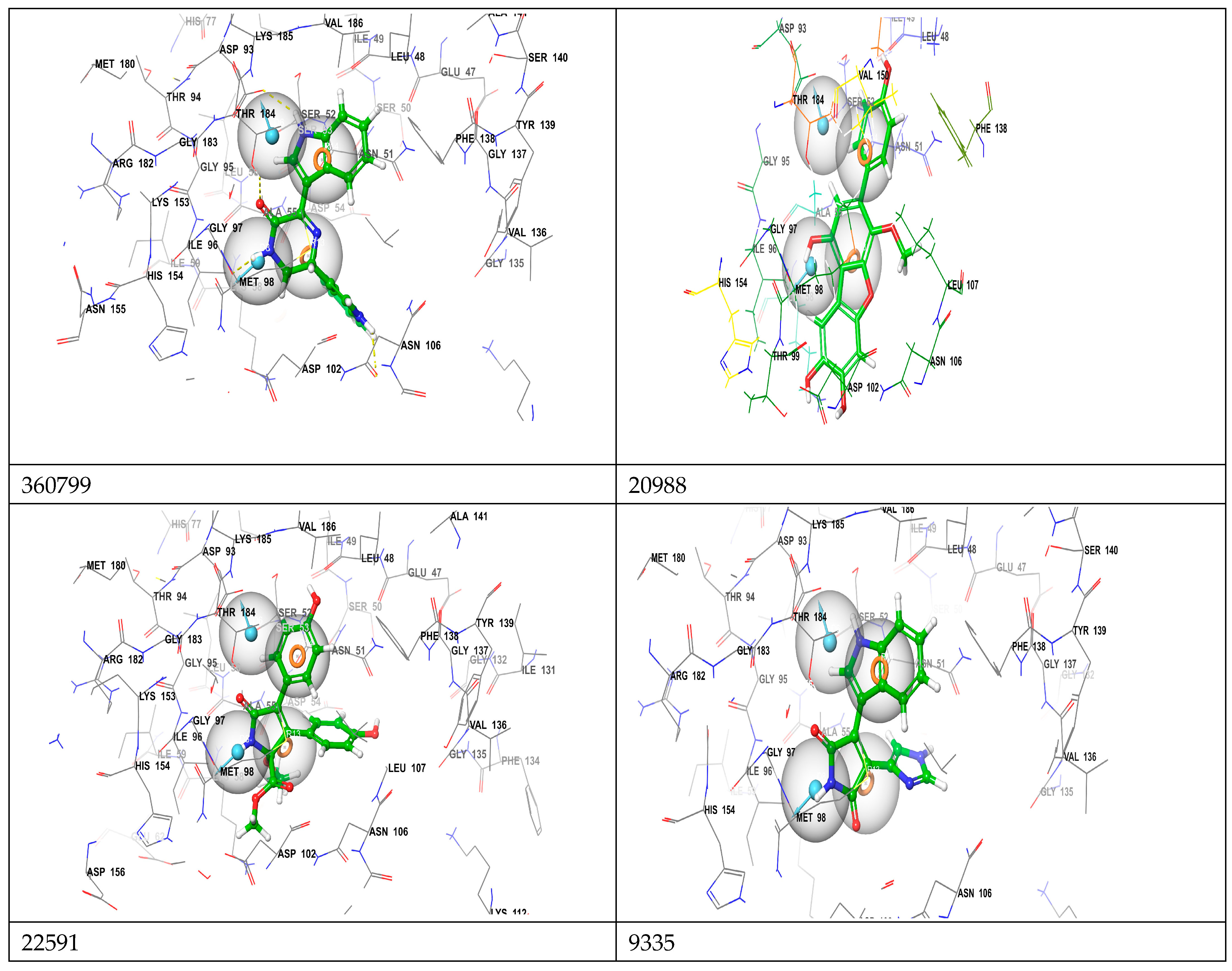
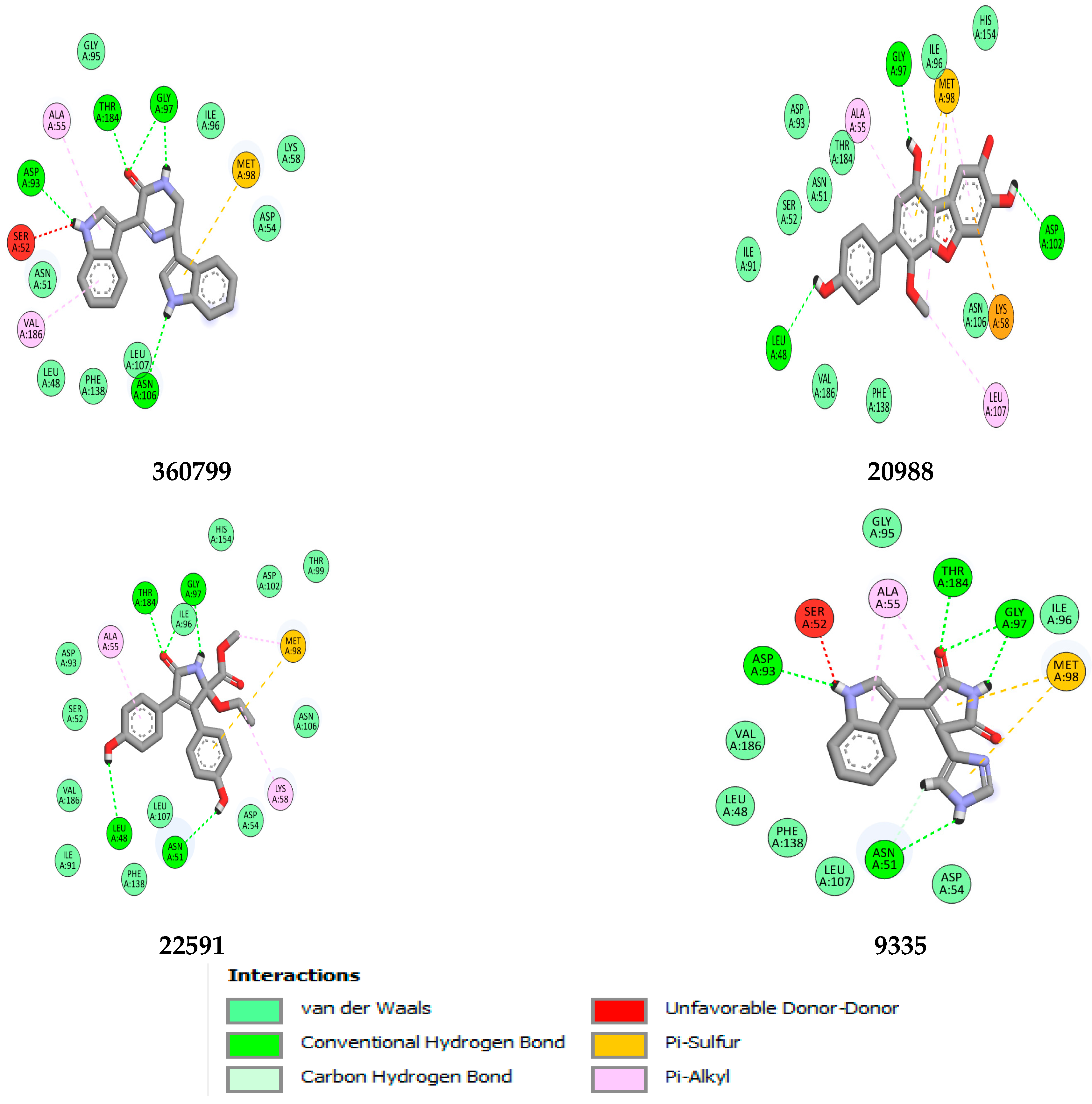
| Compound Name | H-Bonds | Hydrophobic | Others | |||
|---|---|---|---|---|---|---|
| Amino Acid Residues | Number of H-Bonds | H-Bond Distance(Å) | Amino Acid Residues | Number of Hydrophobic Bonds | ||
| 10015 | Gly97, Thr184, Asn51, Asp93 | 6 | 1.83–2.77 | Ala55, met98 |
3 3 | / |
| 15115 | Gly137, Phe138, Gly97, Leu48, Ser52, Asn106, Val136 | 10 | 2.10–2.92 | Ala55 | 2 | Met98 |
| 360799 | Gly97, Thr184, Asp93 | 6 | 1.83–2.75 | Ala55, Val86 | 2 | Met98 |
| 20988 | Gly97, Asn102, Leu48 | 3 | 1.83–2.51 | Ala55, Val186 | 2 | Met98, Lys58 |
| 22591 | Gly97, Thr184, Leu48, Asn51 | 6 | 2.05–2.76 | Ala55, Met98 | 2 | Met98 |
| 9335 | Glu97, Thr98, Asp39, Asn51 | 7 | 1.90–2.91 | Ala55 | 2 | Met98 |
| MEY | Thr184, Gly97, Asp54 | 4 | 1.70–3.05 | Phe138, Lys58, Met98, Leu108, Ala55 | 7 | / |
| 10015 | 360799 | 15115 | 20988 | 9335 | 22591 | MEY | |
|---|---|---|---|---|---|---|---|
| Molecular Weight (MW) | 354.1 | 328.1 | 363.12 | 338.0 | 278.08 | 369.12 | 490.07 |
| Volume | 358.9 | 338.1 | 362.03 | 331.96 | 271.60 | 368.87 | 459.9 |
| nRot | 3 | 2 | 5 | 2 | 2 | 6 | 5 |
| nRing | 5 | 5 | 4 | 4 | 4 | 3 | 5 |
| nHet | 6 | 5 | 7 | 6 | 6 | 7 | 10 |
| Flexibility | 0.017 | 0.074 | 0. 107 | 0.095 | 0.091 | 0.316 | 0.167 |
| TPSA | 90.11 | 73.04 | 106.26 | 103.2 | 90.64 | 105.09 | 124.42 |
| logS | −4.645 | −4.511 | −5.348 | −4.29 | −3.630 | −3.337 | −4.776 |
| logP | 2.611 | 3.404 | 2.503 | 3.869 | 1.182 | 2.938 | 4.562 |
| logD | 2.616 | 3.172 | 2.100 | 3.060 | 1.413 | 2.931 | 3.492 |
| 10015 | 360799 | 15115 | 20988 | 9335 | 22591 | MEY | |
|---|---|---|---|---|---|---|---|
| QED | 0.389 | 0.510 | 0.472 | 0.411 | 0.455 | 0.620 | 0.293 |
| SAscore | 2.821 | 3.253 | 3.206 | 2.592 | 2.715 | 2.635 | 2.615 |
| Pfizer Rule | Accepted | Rejected | Accepted | Accepted | Accepted | Accepted | Accepted |
| GSK Rule | Accepted | Accepted | Accepted | Accepted | Accepted | Accepted | Rejected |
| Golden Triangle | Accepted | Accepted | Accepted | Accepted | Accepted | Accepted | Accepted |
| PAINS | 0 alert | 0 alert | 1 alert | 0 alert | 0 alert | 0 alert | 0 alert |
| BMS Rule | 0 alert | 0 alert | 0 alert | 1alert | 0 alert | 0 alert | 0 alert |
| Chelator Rule | 0 alert | 0 alert | 0 alert | 0 alert | 0 alert | 0 alert | 0 alert |
| Absorption | 10015 | 360799 | 15115 | 20988 | 9335 | 22591 | MEY |
|---|---|---|---|---|---|---|---|
| Caco-2 Permeability | −5.172 | −5.112 | −5.323 | −4.694 | −5.200 | −4.621 | −5.856 |
| MDCK Permeability | 5.2 × 10-6 | 4.5×10-6 | 5.4×10-6 | 2.8×10-5 | 6.9 ×10-6 | 1.5 ×10-5 | 1.3 ×10-5 |
| Pgp-inhibitor | Poor | medium | excellent | poor | excellent | excellent | medium |
| HIA | excellent | excellent | excellent | excellent | excellent | excellent | medium |
| F20% | excellent | excellent | excellent | excellent | excellent | excellent | excellent |
| Distribution | 10015 | 360799 | 15115 | 20988 | 9335 | 22591 | MEY |
|---|---|---|---|---|---|---|---|
| PPB | 98.924% | 95.931% | 86.725% | 88.424% | 88.245% | 86.444% | 98.622 |
| VD L/kg | 0.345 | 0.966 | 1.747 | 0.459 | 1.131 | 0.460 | 0.429 |
| BBB Penetration | excellent | excellent | excellent | poor | excellent | excellent | excellent |
| Fu | 0.998% | 3.499% | 14.958% | 2.918% | 18.876% | 8.108% | 0.663 |
| Metabolism | 10015 | 360799 | 15115 | 20988 | 9335 | 22591 | MEY |
|---|---|---|---|---|---|---|---|
| CYP1A2 inhibitor | yes | yes | yes | yes | yes | yes | yes |
| CYP2C1 inhibitor | yes | yes | yes | yes | yes | yes | yes |
| CYP2C9 inhibitor | yes | yes | yes | yes | no | yes | yes |
| CYP2D6 inhibitor | no | no | yes | no | yes | no | yes |
| CYP3A4 inhibitor | yes | yes | yes | yes | yes | yes | yes |
| Excretion | |||||||
| CL ml/min/kg | 1.592 | 2.302 | 10.457 | 13.908 | 3.311 | 10.986 | 4.720 |
| T1/2 | 0.693 | 0.408 | 0.316 | 0.842 | 0.895 | 0.892 | 0.322 |
| 10015 | 360799 | 15115 | 20988 | 9335 | 22591 | MEY | |
|---|---|---|---|---|---|---|---|
| hERG Blockers | no | no | no | no | no | no | no |
| Rat Oral Acute Toxicity | yes | yes | no | no | no | no | no |
| Skin Sensitization | no | no | no | yes | no | no | no |
| Respiratory Toxicity | yes | no | no | no | no | no | no |
| Complexes | RMSD (nm) | RMSF (nm) | Rg (nm) | SASA (nm2) |
|---|---|---|---|---|
| Hsp90_15115 | 0.295 | 0.136 | 1.711 | 109.26 |
| Hsp90_360799 | 0.223 | 0.142 | 1.711 | 110.08 |
| Hsp90_20988 | 0.195 | 0.130 | 1.702 | 109.44 |
| Hsp90_22591 | 0.094 | 0.127 | 1.707 | 109.26 |
| Hsp90_9335 | 0.147 | 0.141 | 1.733 | 111.44 |
| Hsp90 | 0.179 | 0.093 | 1.733 | 110.61 |
|
Ligand_Hsp90 Complexes |
ΔEVDW (kJ/mol) |
ΔEEEL (kJ/mol) |
ΔEGB (kJ/mol) |
ΔESURF (kJ/mol) |
ΔGGAS (kJ/mol) |
ΔGSOLV (kJ/mol) |
ΔTOTAL (kJ/mol) |
|---|---|---|---|---|---|---|---|
| Hsp90_15115 | −34.59 | −33.26 | 48.63 | −5.39 | −67.85 | 43.23 | −24.61 |
| Hsp90_360799 | −34.89 | −21.88 | 36.59 | −4.70 | −56.77 | 31.89 | −24.88 |
| Hsp90_20988 | −35.58 | −26.15 | 43.70 | −5.08 | −61.73 | 38.62 | −23.11 |
| Hsp90_22591 | −41.86 | −29.51 | 44.06 | −5.47 | −71.37 | 38.59 | −32.78 |
| Hsp90_9335 | −29.83 | −45.69 | 52.75 | −4.17 | −75.51 | 48.57 | −26.94 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ouassaf, M.; Bourougaa, L.; Al-Mijalli, S.H.; Abdallah, E.M.; Bhat, A.R.; A. Kawsar, S.M. Marine-Derived Compounds as Potential Inhibitors of Hsp90 for Anticancer and Antimicrobial Drug Development: A Comprehensive In Silico Study. Molecules 2023, 28, 8074. https://doi.org/10.3390/molecules28248074
Ouassaf M, Bourougaa L, Al-Mijalli SH, Abdallah EM, Bhat AR, A. Kawsar SM. Marine-Derived Compounds as Potential Inhibitors of Hsp90 for Anticancer and Antimicrobial Drug Development: A Comprehensive In Silico Study. Molecules. 2023; 28(24):8074. https://doi.org/10.3390/molecules28248074
Chicago/Turabian StyleOuassaf, Mebarka, Lotfi Bourougaa, Samiah Hamad Al-Mijalli, Emad M. Abdallah, Ajmal R. Bhat, and Sarkar M. A. Kawsar. 2023. "Marine-Derived Compounds as Potential Inhibitors of Hsp90 for Anticancer and Antimicrobial Drug Development: A Comprehensive In Silico Study" Molecules 28, no. 24: 8074. https://doi.org/10.3390/molecules28248074
APA StyleOuassaf, M., Bourougaa, L., Al-Mijalli, S. H., Abdallah, E. M., Bhat, A. R., & A. Kawsar, S. M. (2023). Marine-Derived Compounds as Potential Inhibitors of Hsp90 for Anticancer and Antimicrobial Drug Development: A Comprehensive In Silico Study. Molecules, 28(24), 8074. https://doi.org/10.3390/molecules28248074










