In Vitro and In Silico Anti-Acetylcholinesterase Activity from Macaranga tanarius and Syzygium jambos
Abstract
1. Introduction
2. Results and Discussion
2.1. Isolation and Identification of Active Constituents
2.2. In Vitro Anti-AChE Activity
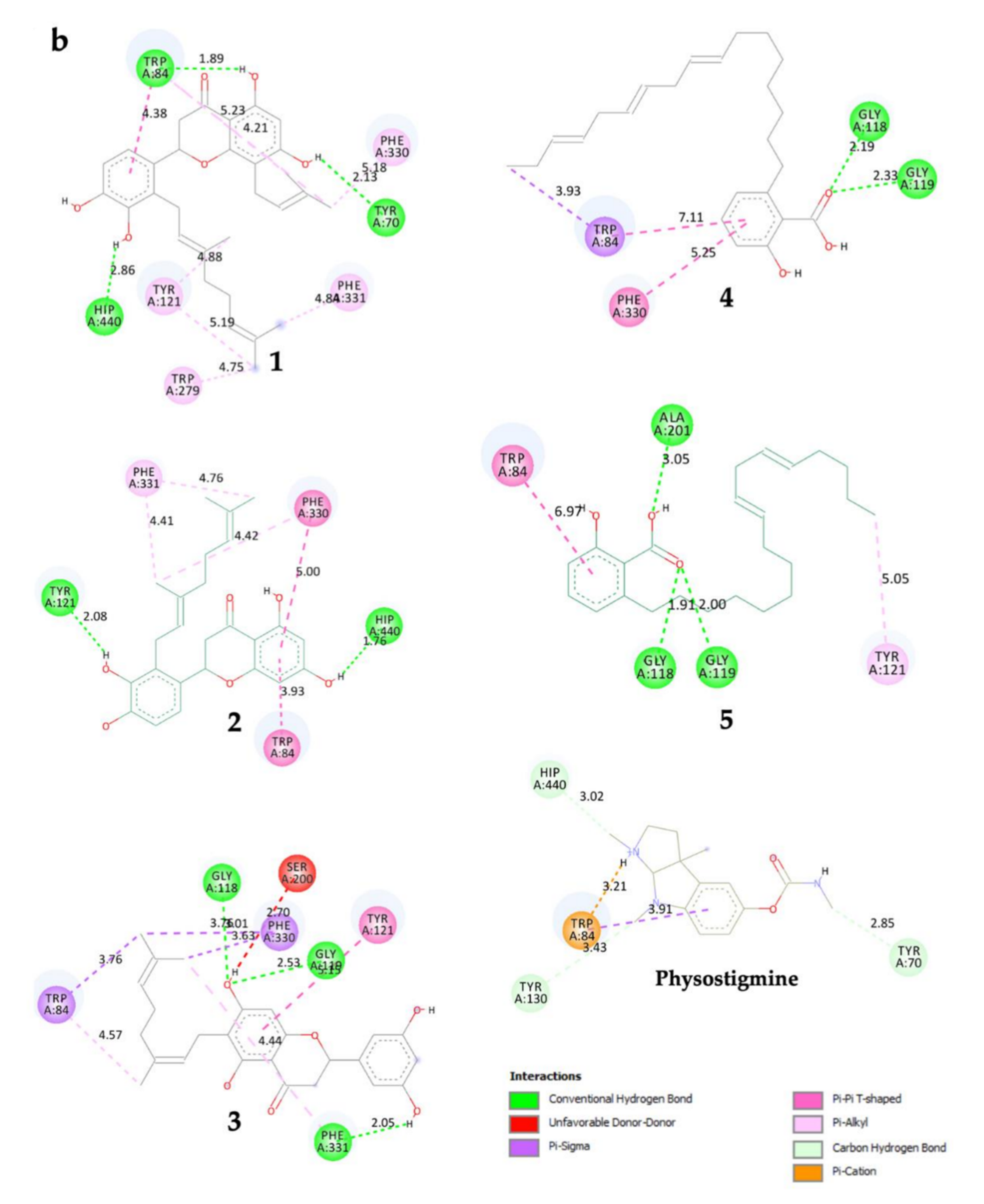
2.3. In Silico Molecular Docking
3. Materials and Methods
3.1. Chemicals and Instruments
3.2. Plant Materials
3.3. Plant Extraction and Partition
3.4. Isolation of Bioactive Constituents
3.5. In Vitro Anti-Acetylcholinesterase Assay
3.6. In Silico Molecular Docking
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Sample Availability
References
- Zakaria, R.; Rosely, N.F.N.; Mansor, M.; Zakaria, M.Y. The distribution of Macaranga, genus (family euphorbiaceae) in Penang Island, Peninsular Malaysia. J. Biosci. 2008, 19, 91–99. [Google Scholar]
- Lim, T.Y.; Lim, Y.Y.; Yule, C.M. Evaluation of antioxidant, antibacterial and anti-tyrosinase activities of four Macaranga species. Food Chem. 2009, 114, 594–599. [Google Scholar] [CrossRef]
- Phommart, S.; Sutthivaiyakit, P.; Chimnoi, N.; Ruchirawat, S.; Sutthivaiyakit, S. Constituents of the leaves of Macaranga tanarius. J. Nat. Prod. 2005, 68, 927–930. [Google Scholar] [CrossRef]
- Shahinozzaman, M.; Obanda, D.N.; Tawata, S. Chemical composition and pharmacological properties of Macaranga-type Pacific propolis: A Review. Phyther. Res. 2021, 35, 207–222. [Google Scholar] [CrossRef]
- Hendra, P.; Fenty, J.J. Evaluation of antidiabetic and antihuperlidemic activities of Macaranga tanarius in rats feed with high glucose-fructose diet. Int. J. Pharm. Pharm. Sci. 2016, 8, 462–463. [Google Scholar]
- Arung, E.T.; Amirta, R.; Zhu, Q.; Amen, Y.; Shimizu, K. effect of wood, bark, and leaf extracts of Macaranga trees on cytotoxic activity in some cancer and normal cell lines. J. Indian Acad. Wood Sci. 2018, 15, 115–119. [Google Scholar] [CrossRef]
- Marliana, E.; Ruga, R.; Hairani, R.; Tjahjandarie, T.S.; Tanjung, M. Antioxidant activity of flavonoid constituents from the leaves of Macaranga tanarius. Proc. J. Phys. Conf. Ser. 2019, 1277, 12014. [Google Scholar] [CrossRef]
- Ochieng, M.A.; Bakrim, B.; Bitchagno, G.T.M.; Mahmoud, M.F.; Sobeh, M. Syzygium jambos L. Alston: An insight into its phytochemistry, traditional uses, and pharmacological properties. Front. Pharmacol. 2022, 13, 786712. [Google Scholar] [CrossRef]
- Lim, T.K. Syzygium jambos. In Edible Medicinal and Non Medicinal Plants; Springer: Dordrecht, The Netherlands, 2012; pp. 760–766. [Google Scholar]
- Mahmoud, M.F.; Abdelaal, S.; Mohammed, H.O.; El-Shazly, A.M.; Daoud, R.; El Raey, M.A.; Sobeh, M. Syzygium jambos extract mitigates pancreatic oxidative stress, inflammation and apoptosis and modulates hepatic IRS-2/AKT/GLUT4 signaling pathway in streptozotocin-induced diabetic rats. Biomed. Pharmacother. 2021, 142, 112085. [Google Scholar] [CrossRef]
- Sobeh, M.; Esmat, A.; Petruk, G.; Abdelfattah, M.A.O.; Dmirieh, M.; Monti, D.M.; Abdel-Naim, A.B.; Wink, M. Phenolic compounds from Syzygium jambos (myrtaceae) exhibit distinct antioxidant and hepatoprotective activities in vivo. J. Funct. Foods 2018, 41, 223–231. [Google Scholar] [CrossRef]
- Apaza Ticona, L.; Souto Pérez, B.; Martín Alejano, V.; Slowing, K. Anti-inflammatory and anti-arthritic activities of glycosylated flavonoids from Syzygium jambos in edematogenic agent-induced paw edema in mice. Rev. Bras. Farmacogn. 2021, 31, 429–441. [Google Scholar] [CrossRef]
- Rawa, M.S.A.; Hassan, Z.; Murugaiyah, V.; Nogawa, T.; Wahab, H.A. Anti-cholinesterase potential of diverse botanical families from Malaysia: Evaluation of crude extracts and fractions from liquid-liquid extraction and acid-base fractionation. J. Ethnopharmacol. 2019, 245, 112160. [Google Scholar] [CrossRef]
- Taylor, P. The cholinesterases. J. Biol. Chem. 1991, 266, 4025–4028. [Google Scholar] [CrossRef]
- Sharma, K. Cholinesterase inhibitors as Alzheimer’s therapeutics (review). Mol. Med. Rep. 2019, 20, 1479–1487. [Google Scholar] [CrossRef]
- Inui, S.; Shimamura, Y.; Masuda, S.; Shirafuji, K.; Moli, R.T.; Kumazawa, S. A new prenylflavonoid isolated from propolis collected in the Solomon islands. Biosci. Biotechnol. Biochem. 2012, 76, 1038–1040. [Google Scholar] [CrossRef]
- Murphy, B.T.; Cao, S.; Norris, A.; Miller, J.S.; Ratovoson, F.; Andriantsiferana, R.; Rasamison, V.E.; Kingston, D.G.I. Cytotoxic compounds of Schizolaena hystrix from the Madagascar rainforest. Planta Med. 2006, 72, 1235–1238. [Google Scholar] [CrossRef]
- Coates, N.J.; Gilpin, M.L.; Gwynn, M.N.; Lewis, D.E.; Milner, P.H.; Spear, S.R.; Tyler, J.W. SB-202742, a Novel β-Lactamase Inhibitor Isolated from Spondias Mombin. J. Nat. Prod. 1994, 57, 654–657. [Google Scholar] [CrossRef]
- Irie, J.; Murata, M.; Homma, S. Glycerol-3-phosphate dehydrogenase inhibitors, anacardic acids, from Ginkgo biloba. Biosci. Biotechnol. Biochem. 1996, 60, 240–243. [Google Scholar] [CrossRef]
- Morais, S.M.; Silva, K.A.; Araujo, H.; Vieira, I.G.P.; Alves, D.R.; Fontenelle, R.O.S.; Silva, A. Anacardic acid constituents from cashew nut shell liquid: NMR characterization and the effect of unsaturation on its biological activities. Pharmaceuticals 2017, 10, 31. [Google Scholar] [CrossRef]
- Greenblatt, H.M.; Guillou, C.; Guénard, D.; Argaman, A.; Botti, S.; Badet, B.; Thal, C.; Silman, I.; Sussman, J.L. The complex of a bivalent derivative of galanthamine with Torpedo acetylcholinesterase displays drastic deformation of the active-site gorge: Implications for structure-based drug design. J. Am. Chem. Soc. 2004, 126, 15405–15411. [Google Scholar] [CrossRef]
- Dvir, H.; Silman, I.; Harel, M.; Rosenberry, T.L.; Sussman, J.L. Acetylcholinesterase: From 3D structure to function. Chem. Biol. Interact. 2010, 187, 10–22. [Google Scholar] [CrossRef]
- Wiesner, J.; Kříž, Z.; Kuča, K.; Jun, D.; Koča, J. Acetylcholinesterases–the structural similarities and differences. J. Enzyme Inhib. Med. Chem. 2007, 22, 417–424. [Google Scholar] [CrossRef]
- Kua, J.; Zhang, Y.; Eslami, A.C.; Butler, J.R.; McCammon, J.A. Studying the roles of W86, E202, and Y337 in binding of acetylcholine to acetylcholinesterase using a combined molecular dynamics and multiple docking approach. Protein Sci. 2003, 12, 2675–2684. [Google Scholar] [CrossRef]


| Compound | AChE Inhibition (μM) |
|---|---|
| 1 | 1.0 |
| 2 | 33 |
| 3 | 6.8 |
| 4 | 2.4 |
| 5 | 0.54 |
| Physostigmine | 0.10 |
| Ligand | Free Energy Binding (FEB) in kcal/mol | Type of TcAChE Active Site | TcAChE Key Residue | Type of Binding Interaction (Distance in Å) |
|---|---|---|---|---|
| 1 | −12.58 | PAS | TYR 70 | H-bond (2.13) |
| CAS | TRP 84 | H-bond (1.89) | ||
| π–π stacked (4.38) | ||||
| π–alkyl (4.21) | ||||
| PAS | TYR 121 | π–alkyl (4.88, 5.19) | ||
| PAS | TRP 279 | π–alkyl (4.75) | ||
| PAS | PHE 330 | π–alkyl (5.18) | ||
| PAS | PHE 331 | π–alkyl (4.84) | ||
| ES | HIS 440 | H–bond (2.86) | ||
| 2 | −10.60 | CAS | TRP 84 | π–π stacked (3.93) |
| PAS | TYR 121 | H-bond (2.08) | ||
| PAS | PHE 330 | π–π T-shaped (5.00) | ||
| π–alkyl (4.42) | ||||
| PAS | PHE 331 | π–alkyl (4.41, 4.76) | ||
| ES | HIS 440 | H-bond (1.76) | ||
| 3 | −10.52 | CAS | TRP 84 | π–σ (3.76) |
| π–alkyl (4.57) | ||||
| H-bond (3.75) | ||||
| OH | GLY 118 | H-bond (2.53) | ||
| OH | GLY 119 | π–π T-shaped (5.15) | ||
| PAS | TYR 121 | π–σ (3.01, 3.63) | ||
| PAS | PHE 330 | H-bond (2.05) | ||
| PAS | PHE 331 | π–alkyl (4.44) | ||
| 4 | −7.94 | CAS | TRP 84 | π–σ (3.93) |
| π–π T (7.11) | ||||
| OH | GLY 118 | H-bond (2.19) | ||
| OH | GLY 119 | H-bond (2.33) | ||
| PAS | PHE 330 | π–π T-shaped (5.25) | ||
| 5 | −7.25 | CAS | TRP 84 | π–π T (6.97) |
| OH | GLY 118 | H-bond (1.91) | ||
| OH | GLY 119 | H-bond (2.00) | ||
| PAS | TYR 121 | π–alkyl (5.05) | ||
| OH | ALA 201 | H-bond (3.05) | ||
| Physostigmine (+control) | −8.81 | PAS | TYR 70 | C–H-bond (2.85) |
| CAS | TRP 84 | π–cation (3.21) | ||
| π–σ (3.91) | ||||
| CAS | TYR 130 | C–H-bond (3.43) | ||
| ES | HIS 440 | C–H-bond (3.02) | ||
| Galanthamine (redocked ligand) | −8.68 (reference RMSD: 0.71) | CAS | TRP 84 | π–alkyl (5.05, 5.48) |
| ES | SER 200 | H-bond (2.19) | ||
| C–H-bond (3.15) | ||||
| OH | GLY 118 | H-bond (2.58) | ||
| PAS | ASP 72 | C–H-bond (3.77) | ||
| PAS | PHE 330 | π–alkyl (4.67) | ||
| PAS | PHE 331 | π–π T-shaped (5.30) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Amir Rawa, M.S.; Nurul Azman, N.A.; Mohamad, S.; Nogawa, T.; Wahab, H.A. In Vitro and In Silico Anti-Acetylcholinesterase Activity from Macaranga tanarius and Syzygium jambos. Molecules 2022, 27, 2648. https://doi.org/10.3390/molecules27092648
Amir Rawa MS, Nurul Azman NA, Mohamad S, Nogawa T, Wahab HA. In Vitro and In Silico Anti-Acetylcholinesterase Activity from Macaranga tanarius and Syzygium jambos. Molecules. 2022; 27(9):2648. https://doi.org/10.3390/molecules27092648
Chicago/Turabian StyleAmir Rawa, Mira Syahfriena, Nurul Amira Nurul Azman, Suriani Mohamad, Toshihiko Nogawa, and Habibah A. Wahab. 2022. "In Vitro and In Silico Anti-Acetylcholinesterase Activity from Macaranga tanarius and Syzygium jambos" Molecules 27, no. 9: 2648. https://doi.org/10.3390/molecules27092648
APA StyleAmir Rawa, M. S., Nurul Azman, N. A., Mohamad, S., Nogawa, T., & Wahab, H. A. (2022). In Vitro and In Silico Anti-Acetylcholinesterase Activity from Macaranga tanarius and Syzygium jambos. Molecules, 27(9), 2648. https://doi.org/10.3390/molecules27092648








