Identification and Characterization of a Novel Endo-β-1,4-Xylanase from Streptomyces sp. T7 and Its Application in Xylo-Oligosaccharide Production
Abstract
:1. Introduction
2. Results and Discussion
2.1. Obtaining Xylanase Secreting Strains
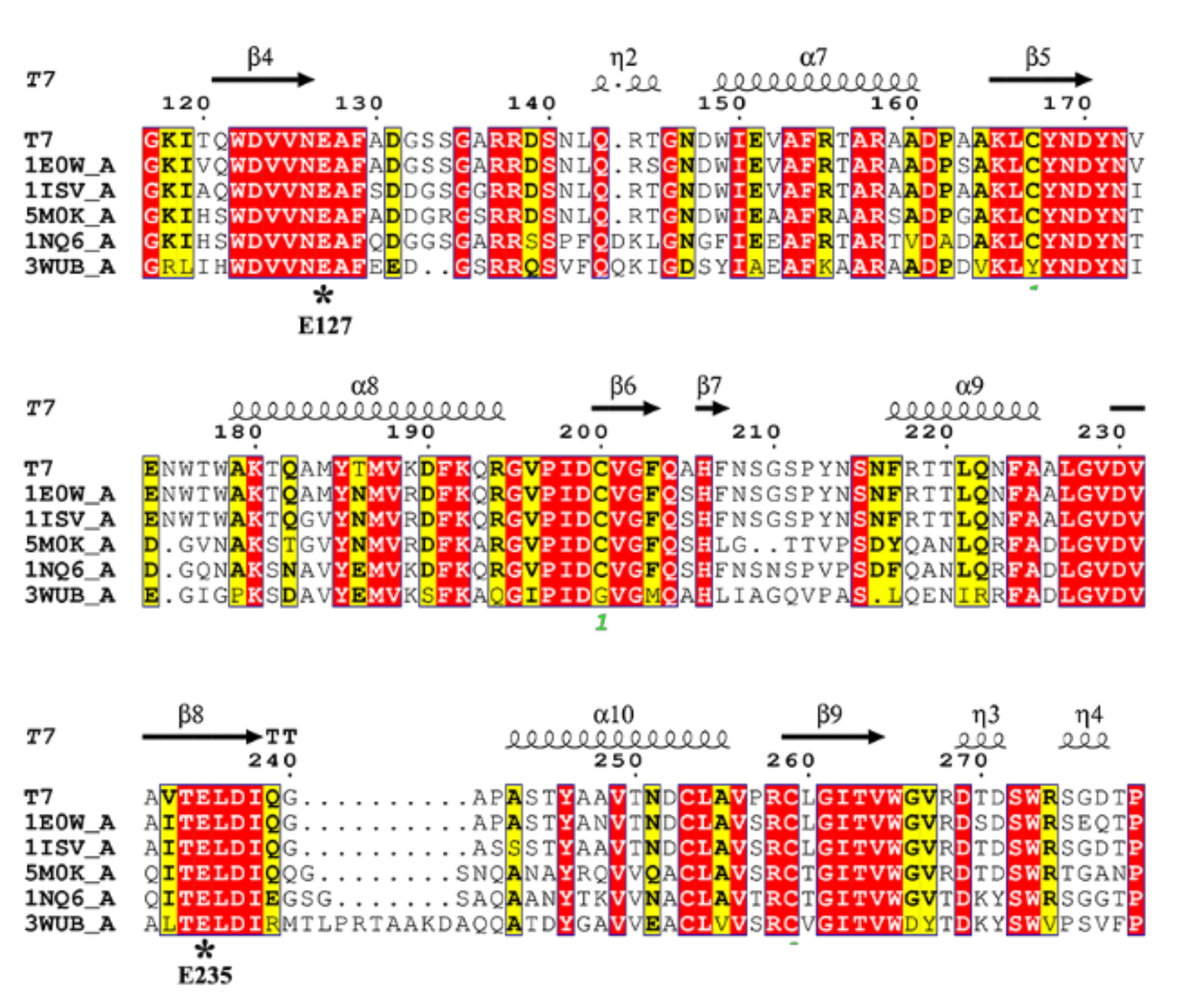
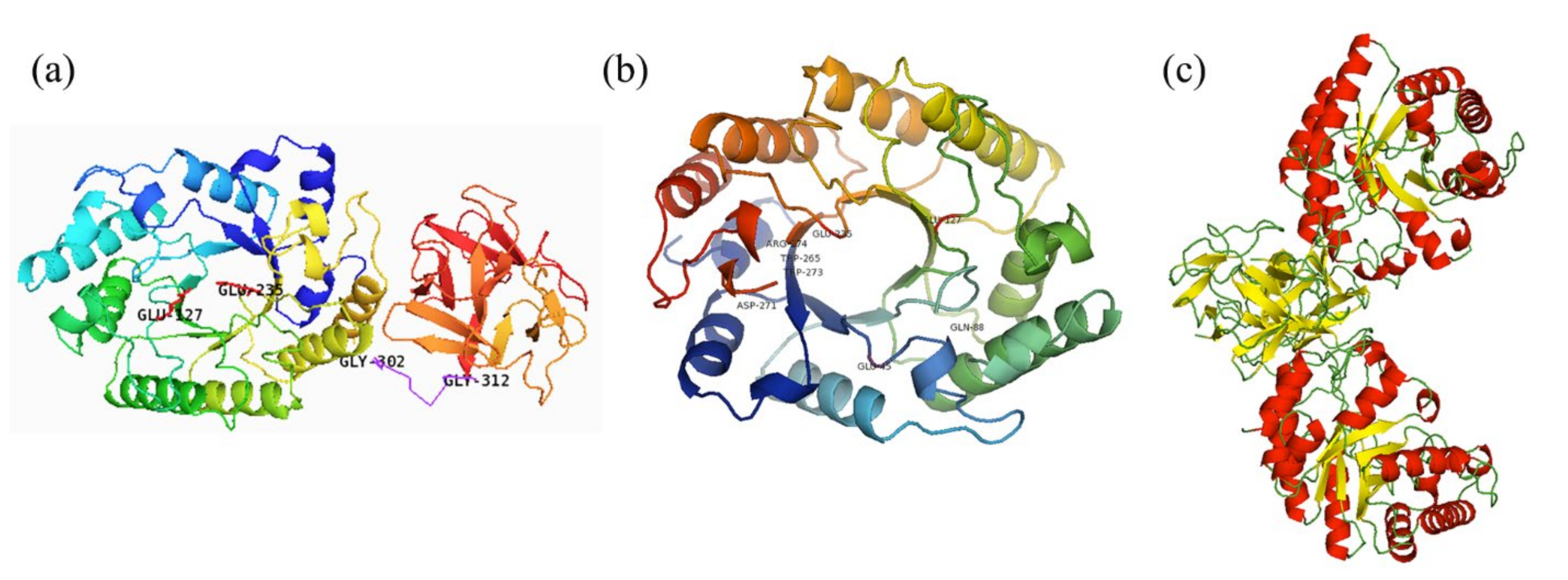
2.2. Sequence and Structure Analysis
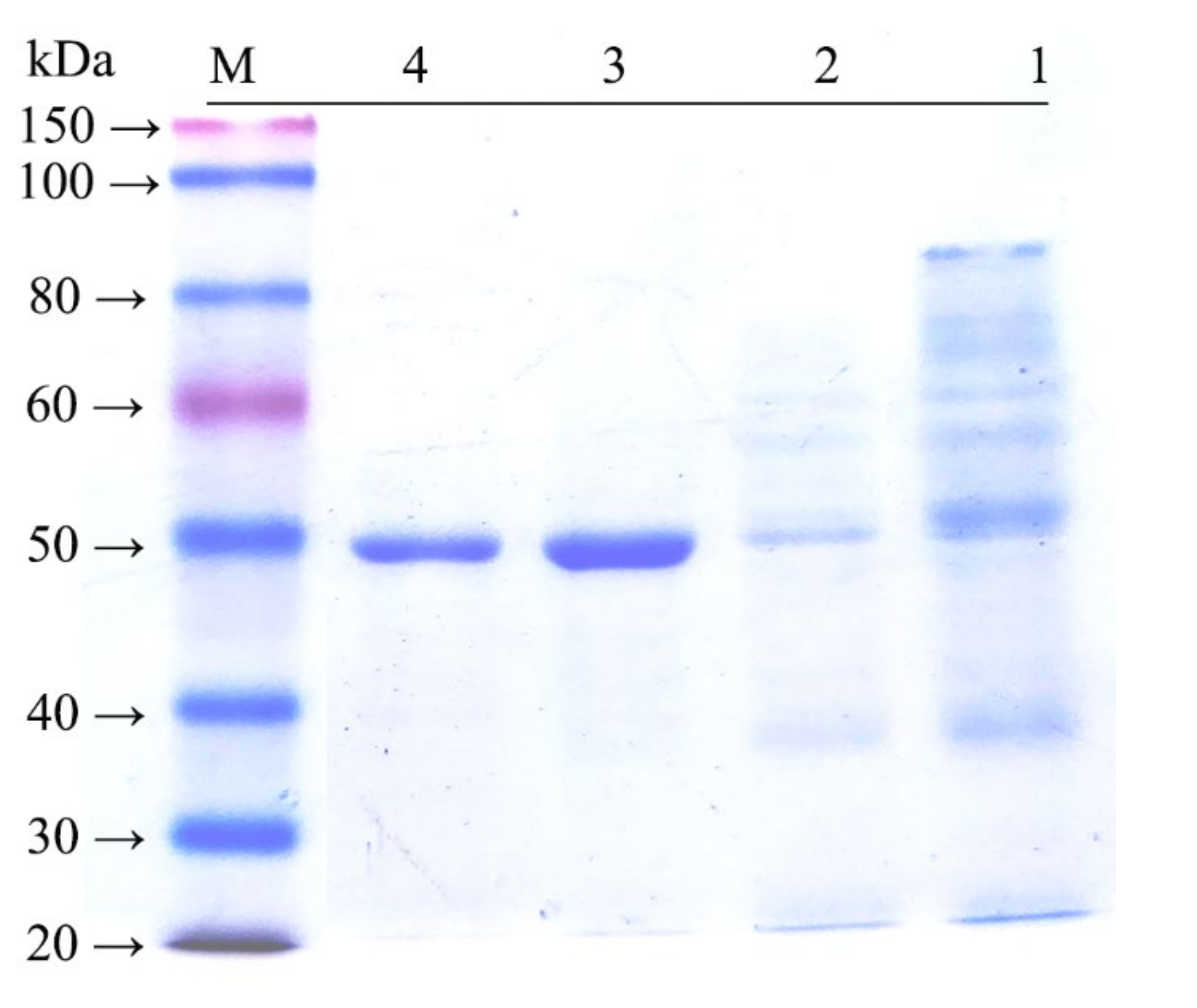
2.3. Expression and Purification
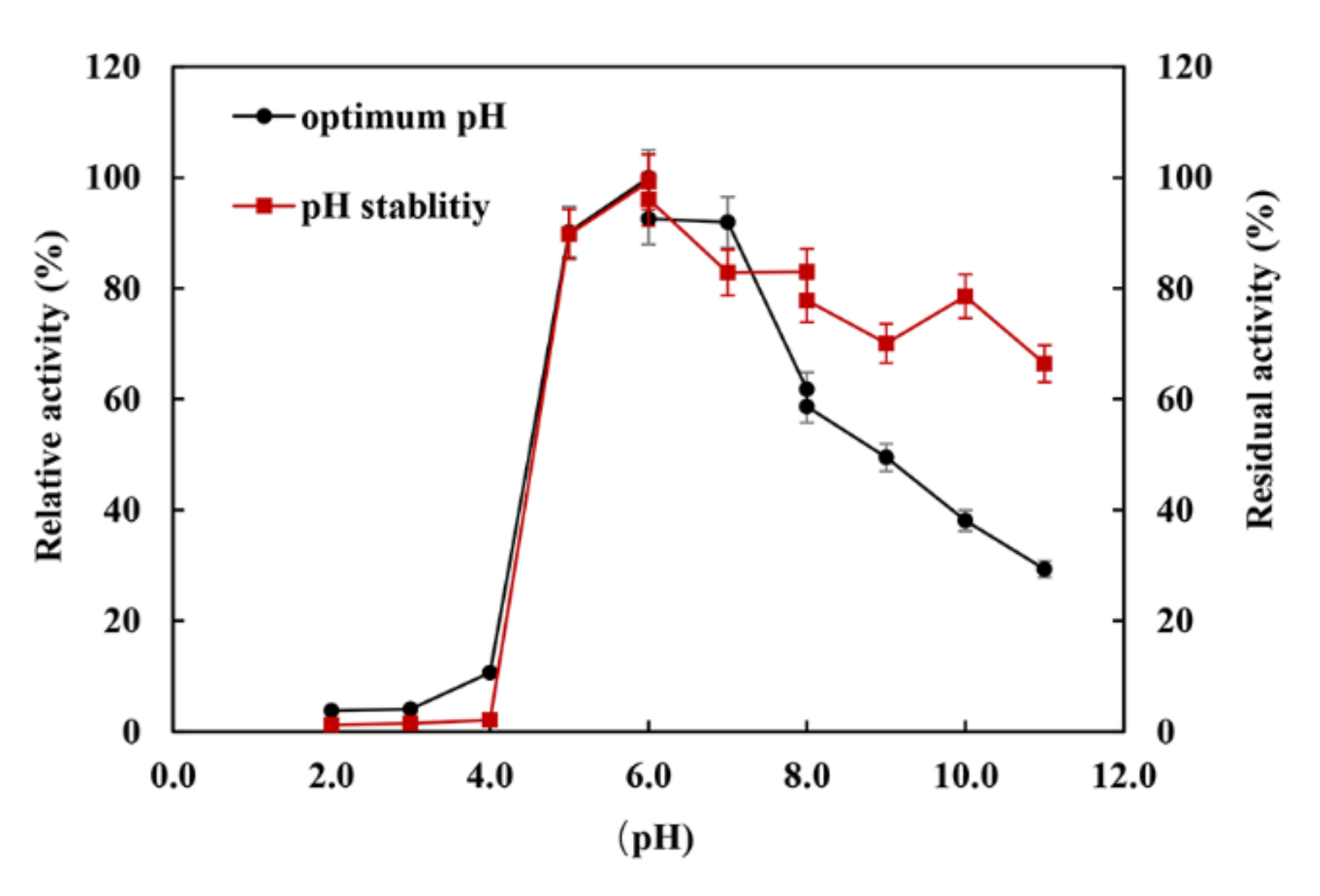
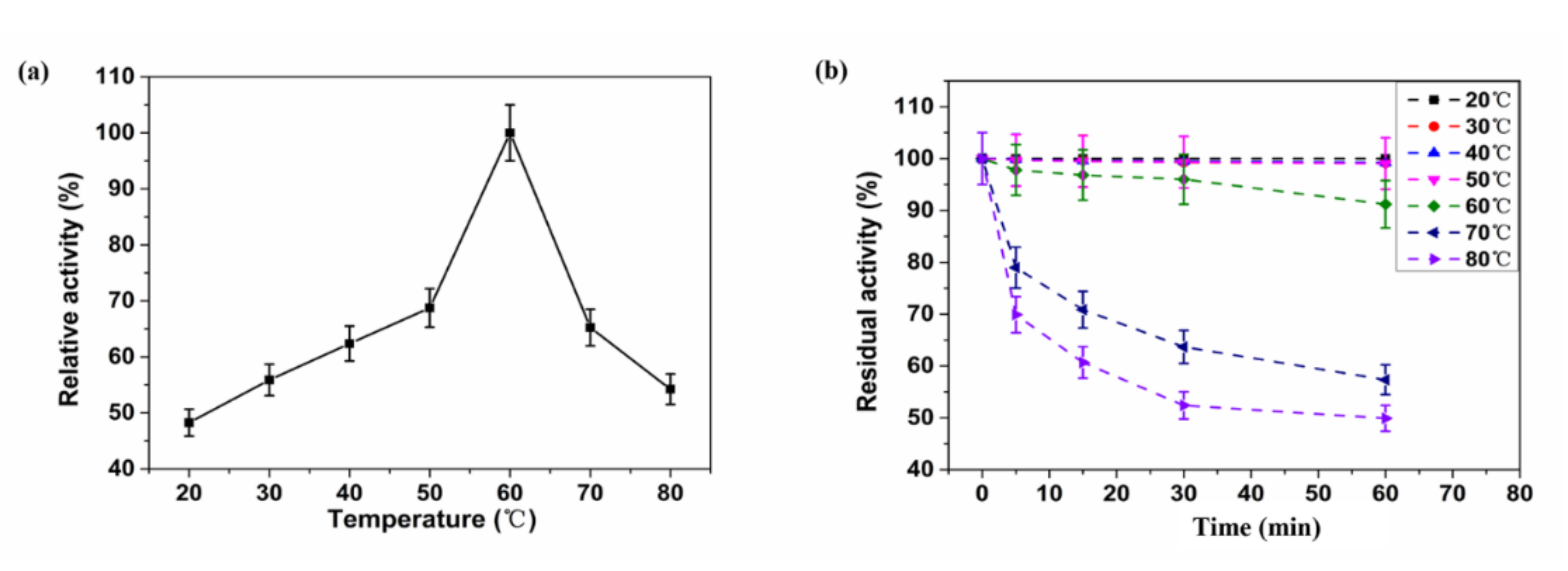
2.4. Biochemical Characteristics
2.5. Substrate Specificity and Kinetic Properties
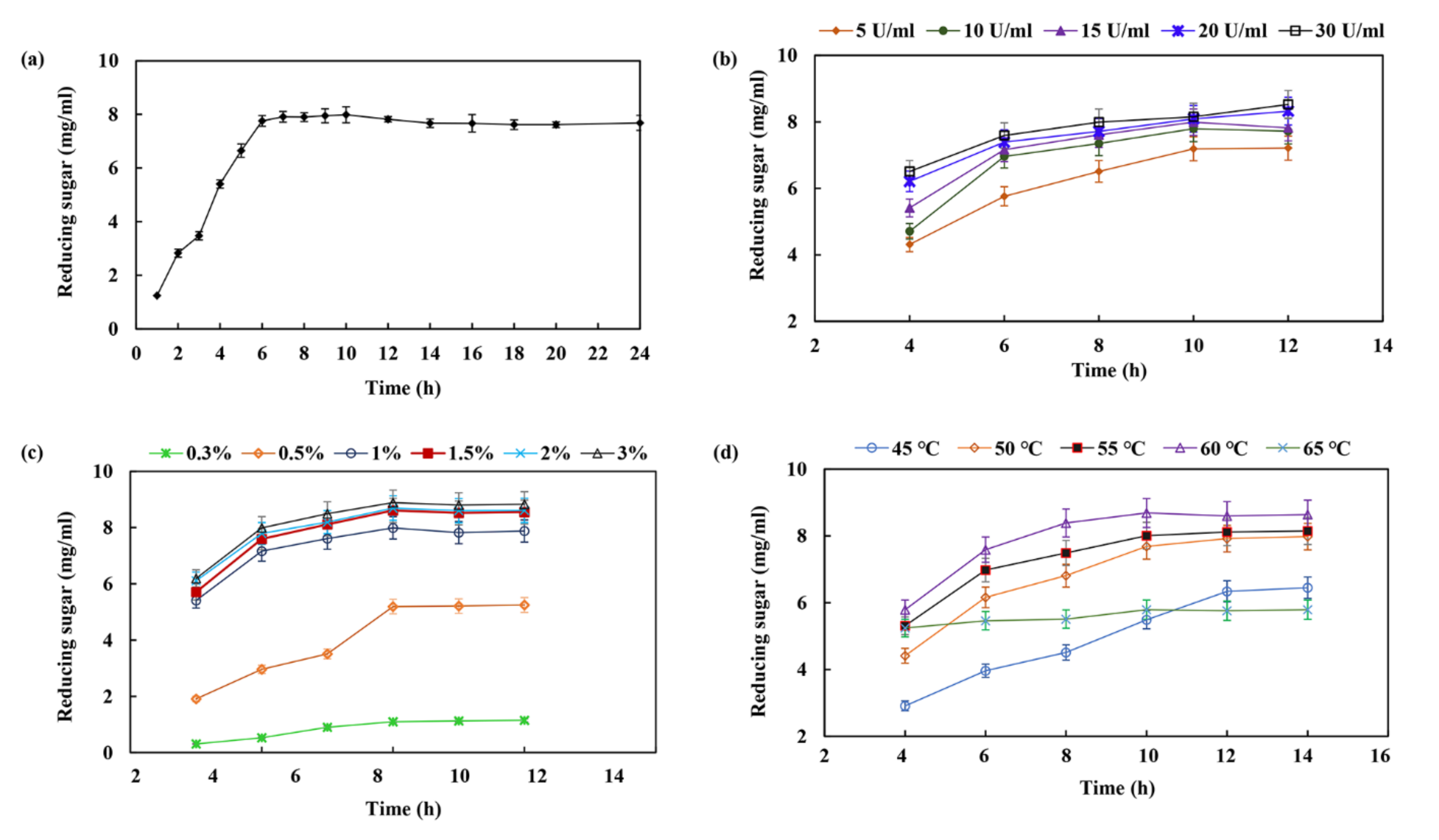
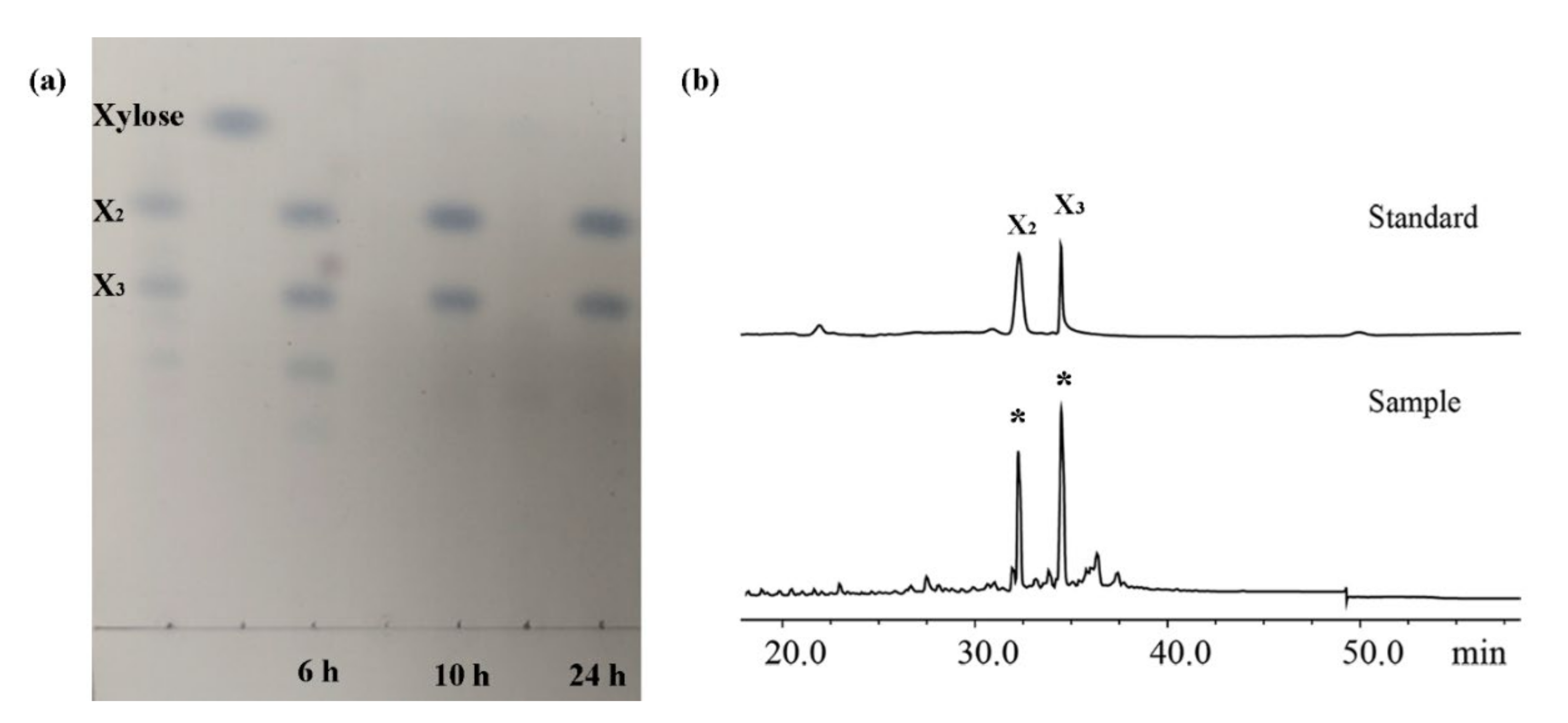
2.6. XOS Production
3. Materials and Methods
3.1. Strains, Plasmids, and Chemicals
3.2. Microorganism Isolation and Identification
3.3. Sequence and Structure Analysis
3.4. Cloning and Recombinant Expression
3.5. Purification
3.6. Enzyme Activity Assay
3.7. Biochemical and Kinetic Characteristics
3.8. Production and Analysis of XOS
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Conflicts of Interest
References
- Takahashi, D.; Zuther, E.; Hincha, D.K. Analysis of Changes in Plant Cell Wall Composition and Structure During Cold Acclimation. Methods Mol. Biol. 2020, 2156, 255–268. [Google Scholar] [PubMed]
- Lee, Y.; Kwon, E.E.; Lee, J. Polymers derived from hemicellulosic parts of lignocellulosic biomass. Rev. Environ. Sci. Biotechnol. 2019, 18, 317–334. [Google Scholar] [CrossRef]
- Biely, P.; Vrsanská, M.; Tenkanen, M.; Kluepfel, D. Endo-beta-1,4-xylanase families: Differences in catalytic properties. J. Biotechnol. 1997, 57, 151–166. [Google Scholar] [CrossRef]
- Collins, T.; Gerday, C.; Feller, G. Xylanases, xylanase families and extremophilic xylanases. FEMS Microbiol. Rev. 2005, 29, 3–23. [Google Scholar] [CrossRef] [Green Version]
- Cheng, Y.S.; Chen, C.C.; Huang, C.H.; Ko, T.P.; Luo, W.; Huang, J.W.; Liu, J.R.; Guo, R.T. Structural analysis of a glycoside hydrolase family 11 xylanase from Neocallimastix patriciarum: Insights into the molecular basis of a thermophilic enzyme. J. Biol. Chem. 2014, 289, 11020–11028. [Google Scholar] [CrossRef] [Green Version]
- Schoepe, J.; Pollet, A.; Verjans, P.; Delcour, J.A.; Strelkov, S.V.; Courtin, C.M. Crystallization and preliminary X-ray analysis of a cold-active endo-β-1,4-D-xylanase from glycoside hydrolase family 8. Acta Crystallogr. Sect. F. Struct. Biol. Cryst. Commun. 2011, 67, 150–152. [Google Scholar] [CrossRef] [Green Version]
- Verma, D.; Satyanarayana, T. Molecular approaches for ameliorating microbial xylanases. Bioresour. Technol. 2012, 7, 360–367. [Google Scholar] [CrossRef]
- Yagi, H.; Takehara, R.; Tamaki, A.; Teramoto, K.; Tsutsui, S.; Kaneko, S. Functional Characterization of the GH10 and GH11 Xylanases from Streptomyces olivaceoviridis E-86 Provide Insights into the Advantage of GH11 Xylanase in Catalyzing Biomass Degradation. J. Appl. Glycosci. 2019, 66, 29–35. [Google Scholar] [CrossRef] [Green Version]
- Alokika; Singh, B. Production, characteristics, and biotechnological applications of microbial xylanases. Appl. Microbiol. Biotechnol. 2019, 103, 8763–8784. [Google Scholar] [CrossRef]
- Linares-Pasten, J.A.; Aronsson, A.; Karlsson, E.N. Structural Considerations on the Use of Endo-Xylanases for the Production of prebiotic Xylooligosaccharides from Biomass. Curr. Protein Pept. Sci. 2018, 19, 48–67. [Google Scholar] [CrossRef] [Green Version]
- Chen, Y.; Xie, Y.; Ajuwon, K.M.; Zhong, R.; Li, T.; Chen, L.; Zhang, H.; Beckers, Y.; Everaert, N. Xylo-Oligosaccharides, Preparation and Application to Human and Animal Health: A Review. Front. Nutr. 2021, 8, 731930. [Google Scholar] [CrossRef] [PubMed]
- Walia, A.; Guleria, S.; Mehta, P.; Chauhan, A.; Parkash, J. Microbial xylanases and their industrial application in pulp and paper biobleaching: A review. 3 Biotech. 2017, 7, 11. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Marasinghe, S.D.; Jo, E.; Hettiarachchi, S.A.; Lee, Y.; Eom, T.Y.; Gang, Y.; Kang, Y.H.; Oh, C. Characterization of glycoside hydrolase family 11 xylanase from Streptomyces sp. strain J103; its synergetic effect with acetyl xylan esterase and enhancement of enzymatic hydrolysis of lignocellulosic biomass. Microb. Cell Fact. 2021, 20, 129. [Google Scholar] [CrossRef]
- Hakulinen, N.; Turunen, O.; Jänis, J.; Leisola, M.; Rouvinen, J. Three-dimensional structures of thermophilic beta-1,4-xylanases from Chaetomium thermophilum and Nonomuraea flexuosa. Comparison of twelve xylanases in relation to their thermal stability. Eur. J. Biochem. 2003, 270, 1399–1412. [Google Scholar] [CrossRef] [Green Version]
- Sriyapai, T.; Somyoonsap, P.; Matsui, K.; Kawai, F.; Chansiri, K. Cloning of a thermostable xylanase from Actinomadura sp. S14 and its expression in Escherichia coli and Pichia pastoris. J. Biosci. Bioeng. 2011, 111, 528–536. [Google Scholar] [CrossRef]
- Li, N.; Meng, K.; Wang, Y.; Shi, P.; Luo, H.; Bai, Y.; Yang, P.; Yao, B. Cloning, expression, and characterization of a new xylanase with broad temperature adaptability from Streptomyces sp. S9. Appl. Microbiol. Biotechnol. 2008, 80, 231–240. [Google Scholar] [CrossRef] [PubMed]
- Boonchuay, P.; Techapun, C.; Leksawasdi, N.; Seesuriyachan, P.; Hanmoungjai, P.; Watanabe, M.; Takenaka, S.; Chaiyaso, T. An integrated process for xylooligosaccharide and bioethanol production from corncob. Bioresour. Technol. 2018, 256, 399–407. [Google Scholar] [CrossRef]
- Fatokun, E.N.; Nwodo, U.U.; Okoh, A.I. Classical optimisation of cellulase and xylanase production by a marine Streptomyces species. Appl. Sci. 2016, 6, 286. [Google Scholar] [CrossRef] [Green Version]
- Roberge, M.; Shareck, F.; Morosoli, R.; Kluepfel, D.; Dupont, C. Characterization of two important histidine residues in the active site of xylanase A from Streptomyces lividans, a family 10 glycanase. Biochemistry 1997, 36, 776975. [Google Scholar] [CrossRef]
- Kaneko, S.; Ichinose, H.; Fujimotom, Z.; Kuno, A.; Yura, K.; Go, M.; Mizuno, H.; Kusakabe, I.; Kobayashi, H. Structure and function of a family 10 beta-xylanase chimera of Streptomyces olivaceoviridis E-86 FXYN and Cellulomonas fimi Cex. J. Biol. Chem. 2004, 279, 26619–26626. [Google Scholar] [CrossRef] [Green Version]
- Fujimoto, Z.; Kaneko, S.; Kuno, A.; Kobayashi, H.; Kusakabe, I.; Mizuno, H. Crystal structures of decorated xylooligosaccharides bound to a family 10 xylanase from Streptomyces olivaceoviridis E-86. J. Biol. Chem. 2004, 279, 9606–9614. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fujimoto, Z.; Kuno, A.; Kaneko, S.; Yoshida, S.; Kobayashi, H.; Kusakabe, I.; Mizuno, H. Crystal structure of Streptomyces olivaceoviridis E-86 beta-xylanase containing xylan-binding domain. J. Mol. Biol. 2000, 300, 575–585. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Shi, P.; Zhang, R.; Huang, H.; Meng, K.; Yang, P.; Yao, B. Symbiotic Streptomyces sp. TN119 GH 11 xylanase: A new pH-stable, protease- and SDS-resistant xylanase. J. Ind. Microbiol. Biotechnol. 2011, 38, 523–530. [Google Scholar] [CrossRef] [PubMed]
- Mamo, G.; Thunnissen, M.; Hatti-Kaul, R.; Mattiasson, B. An alkaline active xylanase: Insights into mechanisms of high pH catalytic adaptation. Biochimie 2009, 91, 1187–1196. [Google Scholar] [CrossRef] [PubMed]
- Ruiz-Arribas, A.; Fernández-Abalos, J.M.; Sánchez, P.; Garda, A.L.; Santamariá, R.I. Overproduction, purification, and biochemical characterization of a xylanase (Xys1) from Streptomyces halstedii JM8. Appl. Environ. Microbiol. 1995, 61, 2414–2419. [Google Scholar] [CrossRef] [Green Version]
- Li, N.; Yang, P.L.; Wang, Y.R.; Luo, H.Y.; Meng, K.; Wu, N.F.; Fan, Y.L.; Yao, B.J. Cloning, expression, and characterization of protease-resistant xylanase from Streptomyces fradiae var. k11. Microbiol. Biotechnol. 2008, 18, 410–416. [Google Scholar]
- Chantasingh, D.; Pootanakit, K.; Champreda, V.; Kanokratana, P.; Eurwilaichitr, L. Cloning, expression, and characterization of a xylanase 10 from Aspergillus terreus (BCC129) in Pichia pastoris. Protein Expr. Purif. 2006, 46, 143–149. [Google Scholar] [CrossRef]
- Tang, K.; Cui, Y.; Xiao, J.; Ding, M.; Chao, H.; Wu, J.; Han, Z.; Liu, J.; Li, X.; Yan, D. Molecular cloning and characterization of a novel xylanase from Microbacterium imperiale YD-01. J. Food Biochem. 2021, 45, e13988. [Google Scholar] [CrossRef]
- Kim, D.Y.; Han, M.K.; Oh, H.W.; Park, D.S.; Kim, S.J.; Lee, S.G.; Shin, D.H.; Son, K.H.; Bae, K.S.; Park, H.Y. Catalytic properties of a GH10 endo-β-1,4-xylanase from Streptomyces thermocarboxydus HY-15 isolated from the gut of Eisenia fetida. J. Mol. Catal. B Enzym. 2010, 62, 32–39. [Google Scholar] [CrossRef]
- Oh, H.W.; Heo, S.Y.; Kim, D.Y.; Park, D.S.; Bae, K.S.; Park, H.Y. Biochemical characterization and sequence analysis of a xylanase produced by an exo-symbiotic bacterium of Gryllotalpa orientalis, Cellulosimicrobium sp. HY-12. Antonie Van Leeuwenhoek 2008, 93, 437–442. [Google Scholar] [CrossRef]
- Marcolongo, L.; La Cara, F.; Morana, A.; Di Salle, A.; Del Monaco, G.; Paixão, S.M.; Alves, L.; Ionata, E. Properties of an alkali-thermo stable xylanase from Geobacillus thermodenitrificans A333 and applicability in xylooligosaccharides generation. World J. Microbiol. Biotechnol. 2015, 31, 633–648. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Georis, J.; Giannotta, F.; De Buyl, E.; Granier, B.; Frère, J. Purification and properties of three endo-beta-1,4-xylanases produced by Streptomyces sp. strain S38 which differ in their ability to enhance the bleaching of kraft pulps. Enzyme Microb. Technol. 2000, 26, 178–186. [Google Scholar] [CrossRef]
- Pollet, A.; Delcour, J.A.; Courtin, C.M. Structural determinants of the substrate specificities of xylanases from different glycoside hydrolase families. Crit. Rev. Biotechnol. 2010, 30, 176–191. [Google Scholar] [CrossRef] [PubMed]
- Chapla, D.; Pandit, P.; Shah, A. Production of xylooligosaccharides from corncob xylan by fungal xylanase and their utilization by probiotics. Bioresour. Technol. 2012, 115, 215–221. [Google Scholar] [CrossRef] [PubMed]
- Akpinar, O.; Erdogan, K.; Bostanci, S. Enzymatic production of xylooligosaccharide from selected agricultural wastes. Food Bioprod. Process. 2009, 87, 145–151. [Google Scholar] [CrossRef]
- Lafond, M.; Tauzin, A.; Desseaux, V.; Bonnin, E.; Ajandouz, E.H.; Giardina, T. GH10 xylanase D from Penicillium funiculosum: Biochemical studies and xylooligosaccharide production. Microb. Cell. Fact. 2011, 10, 20. [Google Scholar] [CrossRef] [Green Version]
- Vacilotto, M.M.; Sepulchro, A.V.; Pellegrini, V.A.; Polikarpov, I. Production of prebiotic xylooligosaccharides from arabino- and glucuronoxylan using a two-domain Jonesia denitrificans xylanase from GH10 family. Enzyme Microb. Technol. 2021, 144, 109743. [Google Scholar] [CrossRef]
- Sepulchro, A.V.; Pellegrini, V.A.; Briganti, L.; de Araujo, E.A.; de Araujo, S.S.; Polikarpov, I. Transformation of xylan into value-added biocommodities using Thermobacillus composti GH10 xylanase. Carbohydr. Polym. 2020, 247, 116714. [Google Scholar] [CrossRef]
- Vacilotto, M.M.; Pellegrini, V.O.; Sepulchro, A.G.; de Mello Capetti, C.C.; Curvelo, A.A.; Marcondes, W.F.; Arantes, V.; Polikarpov, I. Paludibacter propionicigenes GH10 xylanase as a tool for enzymatic xylooligosaccharides production from heteroxylans. Carbohydr. Polym. 2022, 275, 118684. [Google Scholar] [CrossRef]
- Liu, L.; Xu, M.; Cao, Y.; Wang, H.; Shao, J.; Xu, M.; Zhang, Y.; Wang, Y.; Zhang, W.; Meng, X.; et al. Biochemical characterization of xylanases from Streptomyces sp. B6 and their application in the xylooligosaccharide production from viscose fiber production waste. J. Agric. Food Chem. 2020, 68, 3184–3194. [Google Scholar] [CrossRef]
- Velasco, J.; Oliva, B.; Mulinari, E.J.; Quintero, L.P.; da Silva Lima, A.; Gonçalves, A.L.; Gonçalves, T.A.; Damasio, A.; Squina, F.M.; Ferreira Milagres, A.M.; et al. Heterologous expression and functional characterization of a GH10 endoxylanase from Aspergillus fumigatus var. niveus with potential biotechnological application. Biotechnol. Rep. Amst. 2019, 24, e00382. [Google Scholar] [PubMed]
- Miller, G.L. Use of dinitrosalicylic acid reagent for determining reducing sugars. Anal. Chem. 1959, 31, 426–428. [Google Scholar] [CrossRef]
- Yoon, K.Y.; Woodams, E.E.; Hang, Y.D. Enzymatic production of pentoses from the hemicellulose fraction of corn residues. LWT-Food Sci. Technol. 2006, 39, 387–391. [Google Scholar] [CrossRef]
| Metal Ions/Chemical Reagents | Relative Activity (%) | ||
|---|---|---|---|
| 5 mM | 2.5 mM | 1 mM | |
| KCl | 80.5 ± 0.59 | 87.2 ± 0.60 | 102.9 ± 0.62 |
| NaCl | 92.9 ± 0.73 | 98.9 ± 0.40 | 99.7 ± 0.40 |
| Ni2+ | 95.6 ± 0.49 | 97.7 ± 0.38 | 98.0 ± 0.17 |
| CaCl2 | 86.7 ± 0.60 | 92.0 ± 0.39 | 95.1 ± 0.20 |
| MnSO4 | 110.4 ± 1.70 | 109.9 ± 0.31 | 107.4 ± 0.29 |
| FeSO4 | 43.9 ± 0.6 | 49.9 ± 0.68 | 85.9 ± 0.38 |
| CoCl2 | 96.5 ± 0.73 | 99.5 ± 0.42 | 104.1 ± 0.38 |
| MgCl2 | 86.2 ± 2.27 | 88.0 ± 1.86 | 101.2 ± 0.24 |
| FeCl3 | 23.7 ± 2.97 | 25.8 ± 1.51 | 27.2 ± 0.28 |
| EDTA | 24.8 ± 0.92 | 64.6 ± 1.04 | 82.5 ± 0.46 |
| SDS | 23.2 ± 2.64 | 35.3 ± 1.32 | 41.7 ± 0.25 |
| Substrate | Enzyme Assay | Kinetic Parameter | |||
|---|---|---|---|---|---|
| Enzyme | Activity Units (U/mL) | Km (mg/mL) | Vmax (U/mg mL) | kcat/Km (s−1·mg−1) | |
| pNPA | α-arabinofuranosidase | <0.05 | NA | NA | NA |
| pNPX | β-xylosidase | <0.05 | NA | NA | NA |
| pNPGα | α-glucosidase | <0.05 | NA | NA | NA |
| pNPGβ | β-glucosidase | <0.05 | NA | NA | NA |
| CMC | Cellulase | <0.5 | NA | NA | NA |
| Avicel | Cellulase | <0.5 | NA | NA | NA |
| Corncob xylan | Xylanase | 235.8 ± 3.1 | 2.33 ± 0.15 | 678.41 ± 1.35 | 58.23 |
| Birchwood xylan | Xylanase | 203.3 ± 2.8 | 2.78 ± 0.21 | 596.54 ± 2.12 | 42.91 |
| Oat spelt xylan | Xylanase | 193.3 ± 2.5 | 3.12 ± 0.11 | 545.33 ±1.18 | 34.96 |
| Beechwood xylan | Xylanase | 188.6 ± 3.6 | 3.54 ± 0.13 | 495.23 ± 1.87 | 27.98 |
| Sources | pH | Thermostability | XOS Production | References | |
|---|---|---|---|---|---|
| Stability | (°C) | Main Type | Xylose | ||
| Penicillium funiculosum | 4.0–5.5 | ≤70 | X2 | – | [38] |
| Jonesia denitrificans | 6.0–9.0 | ≤55 | X4, X5, X6 | * | [39] |
| Thermobacillus composti | 5.0–9.0 | ≤65 | X2, X3 | * | [40] |
| Paludibacter propionicigenes | 4.0–9.0 | <50 | X2, X3 | * | [41] |
| Streptomyces sp. B6 | 5.0–9.0 | ≤70 | X2, X3, X4 | ** | [42] |
| Aspergillus fumigatus var. niveus | 4.5–7.0 | ≤70 | X2, X3 | – | [43] |
| Streptomyces sp. T7 | 5.0–11.0 | ≤80 | X2, X3 | – | this paper |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, Y.; Zhang, X.; Lu, C.; Lu, P.; Yin, C.; Ye, Z.; Huang, Z. Identification and Characterization of a Novel Endo-β-1,4-Xylanase from Streptomyces sp. T7 and Its Application in Xylo-Oligosaccharide Production. Molecules 2022, 27, 2516. https://doi.org/10.3390/molecules27082516
Li Y, Zhang X, Lu C, Lu P, Yin C, Ye Z, Huang Z. Identification and Characterization of a Novel Endo-β-1,4-Xylanase from Streptomyces sp. T7 and Its Application in Xylo-Oligosaccharide Production. Molecules. 2022; 27(8):2516. https://doi.org/10.3390/molecules27082516
Chicago/Turabian StyleLi, Yumei, Xinxin Zhang, Chunwen Lu, Peng Lu, Chongxu Yin, Zhengmao Ye, and Zhaosong Huang. 2022. "Identification and Characterization of a Novel Endo-β-1,4-Xylanase from Streptomyces sp. T7 and Its Application in Xylo-Oligosaccharide Production" Molecules 27, no. 8: 2516. https://doi.org/10.3390/molecules27082516
APA StyleLi, Y., Zhang, X., Lu, C., Lu, P., Yin, C., Ye, Z., & Huang, Z. (2022). Identification and Characterization of a Novel Endo-β-1,4-Xylanase from Streptomyces sp. T7 and Its Application in Xylo-Oligosaccharide Production. Molecules, 27(8), 2516. https://doi.org/10.3390/molecules27082516





