Hit Compounds and Associated Targets in Intracellular Mycobacterium tuberculosis
Abstract
1. Introduction
2. Results and Discussion
2.1. Identification of Hit Compounds
2.2. Generation of Mutants, Mutant Characterization, and WGS
2.3. Identification of Candidate Intracellular Drug Targets in Resistant Mutants
3. Materials and Methods
3.1. Preparation of Chemical Compounds
3.2. Libraries
3.3. HTS
3.4. Bacterial Culture and Mutant Conditions
3.5. WGS of Mutants
3.6. Bioinformatics Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Sample Availability
References
- Guthrie, J.L.; Gardy, J.L. A Brief Primer on Genomic Epidemiology: Lessons Learned from Mycobacterium tuberculosis. Ann. N. Y. Acad. Sci. 2017, 1388, 59–77. [Google Scholar] [CrossRef] [PubMed]
- Hmama, Z.; Peña-Díaz, S.; Joseph, S.; Av-Gay, Y. Immunoevasion and Immunosuppression of the Macrophage by Mycobacterium tuberculosis. Immunol. Rev. 2015, 264, 220–232. [Google Scholar] [CrossRef] [PubMed]
- Deoghare, S. Bedaquiline: A New Drug Approved for Treatment of Multidrug-Resistant Tuberculosis. Indian J. Pharmacol. 2013, 45, 536–537. [Google Scholar] [CrossRef] [PubMed]
- Pethe, K.; Sequeira, P.C.; Agarwalla, S.; Rhee, K.; Kuhen, K.; Phong, W.Y.; Patel, V.; Beer, D.; Walker, J.R.; Duraiswamy, J.; et al. A Chemical Genetic Screen in Mycobacterium tuberculosis Identifies Carbon-Source-Dependent Growth Inhibitors Devoid of in Vivo Efficacy. Nat. Commun. 2010, 1, 57. [Google Scholar] [CrossRef]
- Zheng, X.; Av-Gay, Y. System for Efficacy and Cytotoxicity Screening of Inhibitors Targeting Intracellular Mycobacterium tuberculosis. J. Vis. Exp. 2017, 122, e55273. [Google Scholar] [CrossRef]
- Sorrentino, F.; Gonzalez del Rio, R.; Zheng, X.; Presa Matilla, J.; Torres Gomez, P.; Martinez Hoyos, M.; Perez Herran, M.E.; Mendoza Losana, A.; Av-Gay, Y. Development of an Intracellular Screen for New Compounds Able To Inhibit Mycobacterium tuberculosis Growth in Human Macrophages. Antimicrob. Agents Chemother. 2016, 60, 640–645. [Google Scholar] [CrossRef]
- Galagan, J.E. Genomic Insights into Tuberculosis. Nat. Rev. Genet. 2014, 15, 307–320. [Google Scholar] [CrossRef]
- Zeng, X.; Kwok, J.S.-L.; Yang, K.Y.; Leung, K.S.-S.; Shi, M.; Yang, Z.; Yam, W.-C.; Tsui, S.K.-W. Whole Genome Sequencing Data of 1110 Mycobacterium tuberculosis Isolates Identifies Insertions and Deletions Associated with Drug Resistance. BMC Genom. 2018, 19, 365. [Google Scholar] [CrossRef]
- Gardy, J.L.; Johnston, J.C.; Ho Sui, S.J.; Cook, V.J.; Shah, L.; Brodkin, E.; Rempel, S.; Moore, R.; Zhao, Y.; Holt, R.; et al. Whole-Genome Sequencing and Social-Network Analysis of a Tuberculosis Outbreak. N. Engl. J. Med. 2011, 364, 730–739. [Google Scholar] [CrossRef]
- Farhat, M.R.; Shapiro, B.J.; Kieser, K.J.; Sultana, R.; Jacobson, K.R.; Victor, T.C.; Warren, R.M.; Streicher, E.M.; Calver, A.; Sloutsky, A.; et al. Genomic Analysis Identifies Targets of Convergent Positive Selection in Drug-Resistant Mycobacterium tuberculosis. Nat. Genet. 2013, 45, 1183–1189. [Google Scholar] [CrossRef]
- Coll, F.; Phelan, J.; Hill-Cawthorne, G.A.; Nair, M.B.; Mallard, K.; Ali, S.; Abdallah, A.M.; Alghamdi, S.; Alsomali, M.; Ahmed, A.O.; et al. Genome-Wide Analysis of Multi- and Extensively Drug-Resistant Mycobacterium tuberculosis. Nat. Genet. 2018, 50, 307–316. [Google Scholar] [CrossRef] [PubMed]
- Mortimer, T.D.; Weber, A.M.; Pepperell, C.S. Signatures of Selection at Drug Resistance Loci in Mycobacterium tuberculosis. Msystems 2018, 3, e00108-17. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Zhang, S.; Cui, P.; Shi, W.; Zhang, W.; Zhang, Y. Identification of Novel Mutations Associated with Cycloserine Resistance in Mycobacterium tuberculosis. J. Antimicrob. Chemother. 2017, 72, 3272–3276. [Google Scholar] [CrossRef] [PubMed]
- Allué-Guardia, A.; Garcia, J.I.; Torrelles, J.B. Evolution of Drug-Resistant Mycobacterium tuberculosis Strains and Their Adaptation to the Human Lung Environment. Front. Microbiol. 2021, 12, 612675. [Google Scholar] [CrossRef]
- Gurcha, S.S.; Usha, V.; Cox, J.A.G.; Fütterer, K.; Abrahams, K.A.; Bhatt, A.; Alderwick, L.J.; Reynolds, R.C.; Loman, N.J.; Nataraj, V.; et al. Biochemical and Structural Characterization of Mycobacterial Aspartyl-tRNA Synthetase AspS, a Promising TB Drug Target. PLoS ONE 2014, 9, e113568. [Google Scholar] [CrossRef]
- Abrahams, K.A.; Chung, C.-W.; Ghidelli-Disse, S.; Rullas, J.; Rebollo-López, M.J.; Gurcha, S.S.; Cox, J.A.G.; Mendoza, A.; Jiménez-Navarro, E.; Martínez-Martínez, M.S.; et al. Identification of KasA as the Cellular Target of an Anti-Tubercular Scaffold. Nat. Commun. 2016, 7, 12581. [Google Scholar] [CrossRef]
- Abrahams, K.A.; Cox, J.A.G.; Fütterer, K.; Rullas, J.; Ortega-Muro, F.; Loman, N.J.; Moynihan, P.J.; Pérez-Herrán, E.; Jiménez, E.; Esquivias, J.; et al. Inhibiting Mycobacterial Tryptophan Synthase by Targeting the Inter-Subunit Interface. Sci. Rep. 2017, 7, 9430. [Google Scholar] [CrossRef]
- Abrahams, K.A.; Cox, J.A.G.; Spivey, V.L.; Loman, N.J.; Pallen, M.J.; Constantinidou, C.; Fernández, R.; Alemparte, C.; Remuiñán, M.J.; Barros, D.; et al. Identification of Novel imidazo[1,2-A]pyridine Inhibitors Targeting M. tuberculosis QcrB. PLoS ONE 2012, 7, e52951. [Google Scholar] [CrossRef]
- Baulard, A.R.; Betts, J.C.; Engohang-Ndong, J.; Quan, S.; McAdam, R.A.; Brennan, P.J.; Locht, C.; Besra, G.S. Activation of the pro-Drug Ethionamide Is Regulated in Mycobacteria. J. Biol. Chem. 2000, 275, 28326–28331. [Google Scholar] [CrossRef]
- DeBarber, A.E.; Mdluli, K.; Bosman, M.; Bekker, L.G.; Barry, C.E., 3rd. Ethionamide Activation and Sensitivity in Multidrug-Resistant Mycobacterium tuberculosis. Proc. Natl. Acad. Sci. USA 2000, 97, 9677–9682. [Google Scholar] [CrossRef]
- Dover, L.G.; Alahari, A.; Gratraud, P.; Gomes, J.M.; Bhowruth, V.; Reynolds, R.C.; Besra, G.S.; Kremer, L. EthA, a Common Activator of Thiocarbamide-Containing Drugs Acting on Different Mycobacterial Targets. Antimicrob. Agents Chemother. 2007, 51, 1055–1063. [Google Scholar] [CrossRef] [PubMed]
- Tatum, N.J.; Liebeschuetz, J.W.; Cole, J.C.; Frita, R.; Herledan, A.; Baulard, A.R.; Willand, N.; Pohl, E. New Active Leads for Tuberculosis Booster Drugs by Structure-Based Drug Discovery. Org. Biomol. Chem. 2017, 15, 10245–10255. [Google Scholar] [CrossRef] [PubMed]
- Parish, T.; Smith, D.A.; Kendall, S.; Casali, N.; Bancroft, G.J.; Stoker, N.G. Deletion of Two-Component Regulatory Systems Increases the Virulence of Mycobacterium tuberculosis. Infect. Immun. 2003, 71, 1134–1140. [Google Scholar] [CrossRef] [PubMed]
- Slinger, B.L.; Deiorio-Haggar, K.; Anthony, J.S.; Gilligan, M.M.; Meyer, M.M. Discovery and Validation of Novel and Distinct RNA Regulators for Ribosomal Protein S15 in Diverse Bacterial Phyla. BMC Genom. 2014, 15, 657. [Google Scholar] [CrossRef]
- Grant, S.S.; Wellington, S.; Kawate, T.; Desjardins, C.A.; Silvis, M.R.; Wivagg, C.; Thompson, M.; Gordon, K.; Kazyanskaya, E.; Nietupski, R.; et al. Baeyer-Villiger Monooxygenases EthA and MymA Are Required for Activation of Replicating and Non-Replicating Mycobacterium Tuberculosis Inhibitors. Cell Chem. Biol. 2016, 23, 666–677. [Google Scholar] [CrossRef]
- Moure, A.L.; Narula, G.; Sorrentino, F.; Bojang, A.; Tsui, C.K.M.; Sao Emani, C.; Porras-De Francisco, E.; Díaz, B.; Rebollo-López, M.J.; Torres-Gómez, P.A.; et al. MymA Bioactivated Thioalkylbenzoxazole Prodrug Family Active against Mycobacterium tuberculosis. J. Med. Chem. 2020, 63, 4732–4748. [Google Scholar] [CrossRef]
- Grzegorzewicz, A.E.; Pham, H.; Gundi, V.A.K.B.; Scherman, M.S.; North, E.J.; Hess, T.; Jones, V.; Gruppo, V.; Born, S.E.M.; Korduláková, J.; et al. Inhibition of Mycolic Acid Transport across the Mycobacterium tuberculosis Plasma Membrane. Nat. Chem. Biol. 2012, 8, 334–341. [Google Scholar] [CrossRef]
- Meneghetti, F.; Villa, S.; Gelain, A.; Barlocco, D.; Chiarelli, L.R.; Pasca, M.R.; Costantino, L. Iron Acquisition Pathways as Targets for Antitubercular Drugs. Curr. Med. Chem. 2016, 23, 4009–4026. [Google Scholar] [CrossRef]
- Tahlan, K.; Wilson, R.; Kastrinsky, D.B.; Arora, K.; Nair, V.; Fischer, E.; Barnes, S.W.; Walker, J.R.; Alland, D.; Barry, C.E., 3rd; et al. SQ109 Targets MmpL3, a Membrane Transporter of Trehalose Monomycolate Involved in Mycolic Acid Donation to the Cell Wall Core of Mycobacterium tuberculosis. Antimicrob. Agents Chemother. 2012, 56, 1797–1809. [Google Scholar] [CrossRef]
- Xu, Z.; Meshcheryakov, V.A.; Poce, G.; Chng, S.-S. MmpL3 Is the Flippase for Mycolic Acids in Mycobacteria. Proc. Natl. Acad. Sci. USA 2017, 114, 7993–7998. [Google Scholar] [CrossRef]
- Nowak, E.; Panjikar, S.; Morth, J.P.; Jordanova, R.; Svergun, D.I.; Tucker, P.A. Structural and Functional Aspects of the Sensor Histidine Kinase PrrB from Mycobacterium tuberculosis. Structure 2006, 14, 275–285. [Google Scholar] [CrossRef] [PubMed]
- Bellale, E.; Naik, M.; VB, V.; Ambady, A.; Narayan, A.; Ravishankar, S.; Ramachandran, V.; Kaur, P.; McLaughlin, R.; Whiteaker, J.; et al. Diarylthiazole: An Antimycobacterial Scaffold Potentially Targeting PrrB-PrrA Two-Component System. J. Med. Chem. 2014, 57, 6572–6582. [Google Scholar] [CrossRef] [PubMed]
- Ewann, F.; Jackson, M.; Pethe, K.; Cooper, A.; Mielcarek, N.; Ensergueix, D.; Gicquel, B.; Locht, C.; Supply, P. Transient Requirement of the PrrA-PrrB Two-Component System for Early Intracellular Multiplication of Mycobacterium tuberculosis. Infect. Immun. 2002, 70, 2256–2263. [Google Scholar] [CrossRef] [PubMed]
- Villellas, C.; Coeck, N.; Meehan, C.J.; Lounis, N.; de Jong, B.; Rigouts, L.; Andries, K. Unexpected High Prevalence of Resistance-Associated Rv0678 Variants in MDR-TB Patients without Documented Prior Use of Clofazimine or Bedaquiline. J. Antimicrob. Chemother. 2017, 72, 684–690. [Google Scholar]
- Hartkoorn, R.C.; Uplekar, S.; Cole, S.T. Cross-Resistance between Clofazimine and Bedaquiline through Upregulation of MmpL5 in Mycobacterium tuberculosis. Antimicrob. Agents Chemother. 2014, 58, 2979–2981. [Google Scholar] [CrossRef]
- Radhakrishnan, A.; Kumar, N.; Wright, C.C.; Chou, T.-H.; Tringides, M.L.; Bolla, J.R.; Lei, H.-T.; Rajashankar, K.R.; Su, C.-C.; Purdy, G.E.; et al. Crystal Structure of the Transcriptional Regulator Rv0678 of Mycobacterium tuberculosis. J. Biol. Chem. 2014, 289, 16526–16540. [Google Scholar] [CrossRef]
- Omar, S.V.; Ismail, F.; Ndjeka, N.; Kaniga, K.; Ismail, N.A. Bedaquiline-Resistant Tuberculosis Associated with Rv0678 Mutations. N. Engl. J. Med. 2022, 386, 93–94. [Google Scholar] [CrossRef]
- Ioerger, T.R.; O’Malley, T.; Liao, R.; Guinn, K.M.; Hickey, M.J.; Mohaideen, N.; Murphy, K.C.; Boshoff, H.I.M.; Mizrahi, V.; Rubin, E.J.; et al. Identification of New Drug Targets and Resistance Mechanisms in Mycobacterium tuberculosis. PLoS ONE 2013, 8, e75245. [Google Scholar] [CrossRef]
- Duckworth, B.P.; Wilson, D.J.; Nelson, K.M.; Boshoff, H.I.; Barry, C.E., 3rd; Aldrich, C.C. Development of a Selective Activity-Based Probe for Adenylating Enzymes: Profiling MbtA Involved in Siderophore Biosynthesis from Mycobacterium tuberculosis. ACS Chem. Biol. 2012, 7, 1653–1658. [Google Scholar] [CrossRef]
- Ferguson, L.; Wells, G.; Bhakta, S.; Johnson, J.; Guzman, J.; Parish, T.; Prentice, R.A.; Brucoli, F. Integrated Target-Based and Phenotypic Screening Approaches for the Identification of Anti-Tubercular Agents That Bind to the Mycobacterial Adenylating Enzyme MbtA. Chem. Med. Chem. 2019, 14, 1735–1741. [Google Scholar] [CrossRef]
- Nelson, K.M.; Viswanathan, K.; Dawadi, S.; Duckworth, B.P.; Boshoff, H.I.; Barry, C.E., 3rd; Aldrich, C.C. Synthesis and Pharmacokinetic Evaluation of Siderophore Biosynthesis Inhibitors for Mycobacterium tuberculosis. J. Med. Chem. 2015, 58, 5459–5475. [Google Scholar] [CrossRef] [PubMed]
- Lun, S.; Guo, H.; Adamson, J.; Cisar, J.S.; Davis, T.D.; Chavadi, S.S.; Warren, J.D.; Quadri, L.E.N.; Tan, D.S.; Bishai, W.R. Pharmacokinetic and in Vivo Efficacy Studies of the Mycobactin Biosynthesis Inhibitor Salicyl-AMS in Mice. Antimicrob. Agents Chemother. 2013, 57, 5138–5140. [Google Scholar] [CrossRef] [PubMed]
- Luo, J.; Li, X.; Song, Y.; Liu, H.; Zheng, K.; Xia, X.; Zhang, A.-M. Detection of Mycobacterium tuberculosis in Clinical Sputum by a Unique Gene in MTB Strains Called Conserved Protein TB18.5 (TB18.5). J. Clin. Lab. Anal. 2021, 35, e24033. [Google Scholar] [CrossRef] [PubMed]
- Malhotra, V.; Agrawal, R.; Duncan, T.R.; Saini, D.K.; Clark-Curtiss, J.E. Mycobacterium tuberculosis Response Regulators, DevR and NarL, Interact in Vivo and Co-Regulate Gene Expression during Aerobic Nitrate Metabolism. J. Biol. Chem. 2015, 290, 8294–8309. [Google Scholar] [CrossRef]
- Kumar, N.; Srivastava, R.; Prakash, A.; Lynn, A.M. Structure-Based Virtual Screening, Molecular Dynamics Simulation and MM-PBSA toward Identifying the Inhibitors for Two-Component Regulatory System Protein NarL of Mycobacterium tuberculosis. J. Biomol. Struct. Dyn. 2019, 38, 3396–3410. [Google Scholar] [CrossRef]
- Kumar, M.; Khan, F.G.; Sharma, S.; Kumar, R.; Faujdar, J.; Sharma, R.; Chauhan, D.S.; Singh, R.; Magotra, S.K.; Khan, I.A. Identification of Mycobacterium tuberculosis Genes Preferentially Expressed during Human Infection. Microb. Pathog. 2011, 50, 31–38. [Google Scholar] [CrossRef]
- Botella, H.; Peyron, P.; Levillain, F.; Poincloux, R.; Poquet, Y.; Brandli, I.; Wang, C.; Tailleux, L.; Tilleul, S.; Charrière, G.M.; et al. Mycobacterial p(1)-Type ATPases Mediate Resistance to Zinc Poisoning in Human Macrophages. Cell Host Microbe 2011, 10, 248–259. [Google Scholar] [CrossRef]
- Padilla-Benavides, T.; Long, J.E.; Raimunda, D.; Sassetti, C.M.; Argüello, J.M. A Novel P(1B)-Type Mn2+-Transporting ATPase Is Required for Secreted Protein Metallation in Mycobacteria. J. Biol. Chem. 2013, 288, 11334–11347. [Google Scholar] [CrossRef]
- Cimino, M.; Thomas, C.; Namouchi, A.; Dubrac, S.; Gicquel, B.; Gopaul, D.N. Identification of DNA Binding Motifs of the Mycobacterium Tuberculosis PhoP/PhoR Two-Component Signal Transduction System. PLoS ONE 2012, 7, e42876. [Google Scholar] [CrossRef]
- Xing, D.; Ryndak, M.B.; Wang, L.; Kolesnikova, I.; Smith, I.; Wang, S. Asymmetric Structure of the Dimerization Domain of PhoR, a Sensor Kinase Important for the Virulence of Mycobacterium tuberculosis. ACS Omega 2017, 2, 3509–3517. [Google Scholar] [CrossRef]
- Broset, E.; Martín, C.; Gonzalo-Asensio, J. Evolutionary Landscape of the Mycobacterium tuberculosis Complex from the Viewpoint of PhoPR: Implications for Virulence Regulation and Application to Vaccine Development. MBio 2015, 6, e01289-15. [Google Scholar] [CrossRef] [PubMed]
- Fujiwara, M.; Kawasaki, M.; Hariguchi, N.; Liu, Y.; Matsumoto, M. Mechanisms of Resistance to Delamanid, a Drug for Mycobacterium tuberculosis. Tuberculosis 2018, 108, 186–194. [Google Scholar] [CrossRef] [PubMed]
- Kumar, P.; Kumar, D.; Parikh, A.; Rananaware, D.; Gupta, M.; Singh, Y.; Nandicoori, V.K. The Mycobacterium tuberculosis Protein Kinase K Modulates Activation of Transcription from the Promoter of Mycobacterial Monooxygenase Operon through Phosphorylation of the Transcriptional Regulator VirS. J. Biol. Chem. 2009, 284, 11090–11099. [Google Scholar] [CrossRef] [PubMed]
- Singh, S.; Goswami, N.; Tyagi, A.K.; Khare, G. Unraveling the Role of the Transcriptional Regulator VirS in Low pH-Induced Responses of Mycobacterium tuberculosis and Identification of VirS Inhibitors. J. Biol. Chem. 2019, 294, 10055–10075. [Google Scholar] [CrossRef]
- Baños-Mateos, S.; van Roon, A.-M.M.; Lang, U.F.; Maslen, S.L.; Skehel, J.M.; Lamers, M.H. High-Fidelity DNA Replication in Mycobacterium tuberculosis Relies on a Trinuclear Zinc Center. Nat. Commun. 2017, 8, 855. [Google Scholar] [CrossRef]
- Nasir, N.; Kisker, C. Mechanistic Insights into the Enzymatic Activity and Inhibition of the Replicative Polymerase Exonuclease Domain from Mycobacterium tuberculosis. DNA Repair 2019, 74, 17–25. [Google Scholar] [CrossRef]
- Alland, D.; Steyn, A.J.; Weisbrod, T.; Aldrich, K.; Jacobs, W.R., Jr. Characterization of the Mycobacterium tuberculosis iniBAC Promoter, a Promoter That Responds to Cell Wall Biosynthesis Inhibition. J. Bacteriol. 2000, 182, 1802–1811. [Google Scholar] [CrossRef]
- Williams, M.; Mizrahi, V.; Kana, B.D. Molybdenum Cofactor: A Key Component of Mycobacterium Tuberculosis Pathogenesis? Crit. Rev. Microbiol. 2014, 40, 18–29. [Google Scholar] [CrossRef]
- Titgemeyer, F.; Amon, J.; Parche, S.; Mahfoud, M.; Bail, J.; Schlicht, M.; Rehm, N.; Hillmann, D.; Stephan, J.; Walter, B.; et al. A Genomic View of Sugar Transport in Mycobacterium smegmatis and Mycobacterium tuberculosis. J. Bacteriol. 2007, 189, 5903–5915. [Google Scholar] [CrossRef]
- Gygli, S.M.; Borrell, S.; Trauner, A.; Gagneux, S. Antimicrobial Resistance in Mycobacterium tuberculosis: Mechanistic and Evolutionary Perspectives. FEMS Microbiol. Rev. 2017, 41, 354–373. [Google Scholar] [CrossRef]
- Graham, F.; Hatfull, W.R.J. Molecular Genetics of Mycobacteria; American Society of Microbiology: Washington, DC, USA, 2014; ISBN 9781555818838. [Google Scholar]
- Tsui, C.K.M.; Wong, D.; Narula, G.; Gardy, J.L.; Hsiao, W.W.H.; Av-Gay, Y. Genome Sequences of the Mycobacterium tuberculosis H37Rv-ptkA Deletion Mutant and Its Parental Strain. Genome Announc. 2017, 5, e01156-17. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Durbin, R. Fast and Accurate Long-Read Alignment with Burrows-Wheeler Transform. Bioinformatics 2010, 26, 589–595. [Google Scholar] [CrossRef]
- McKenna, A.; Hanna, M.; Banks, E.; Sivachenko, A.; Cibulskis, K.; Kernytsky, A.; Garimella, K.; Altshuler, D.; Gabriel, S.; Daly, M.; et al. The Genome Analysis Toolkit: A MapReduce Framework for Analyzing next-Generation DNA Sequencing Data. Genome Res. 2010, 20, 1297–1303. [Google Scholar] [CrossRef]
- Cingolani, P.; Platts, A.; Wang, L.L.; Coon, M.; Nguyen, T.; Wang, L.; Land, S.J.; Lu, X.; Ruden, D.M. A Program for Annotating and Predicting the Effects of Single Nucleotide Polymorphisms, SnpEff: SNPs in the Genome of Drosophila Melanogaster Strain w1118; Iso-2; Iso-3. Fly 2012, 6, 80–92. [Google Scholar] [CrossRef] [PubMed]
- Cole, S.T.; Brosch, R.; Parkhill, J.; Garnier, T.; Churcher, C.; Harris, D.; Gordon, S.V.; Eiglmeier, K.; Gas, S.; Barry, C.E., 3rd; et al. Deciphering the Biology of Mycobacterium tuberculosis from the Complete Genome Sequence. Nature 1998, 393, 537–544. [Google Scholar] [CrossRef] [PubMed]
- Milne, I.; Bayer, M.; Stephen, G.; Cardle, L.; Marshall, D. Tablet: Visualizing Next-Generation Sequence Assemblies and Mappings. Methods Mol. Biol. 2016, 1374, 253–268. [Google Scholar] [PubMed]
- Wood, D.E.; Salzberg, S.L. Kraken: Ultrafast Metagenomic Sequence Classification Using Exact Alignments. Genome Biol. 2014, 15, R46. [Google Scholar] [CrossRef]
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D.; et al. SPAdes: A New Genome Assembly Algorithm and Its Applications to Single-Cell Sequencing. J. Comput. Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef]
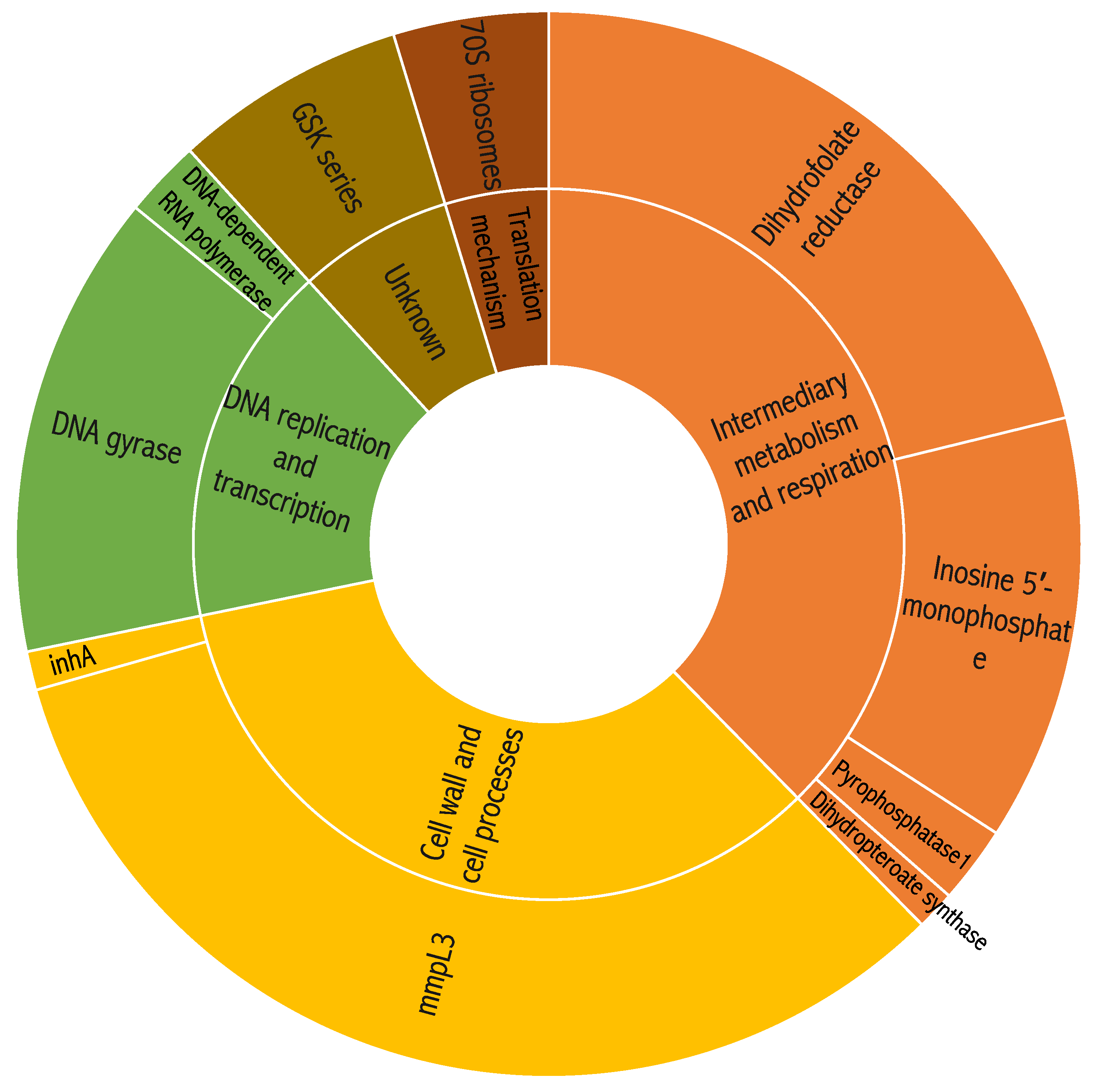
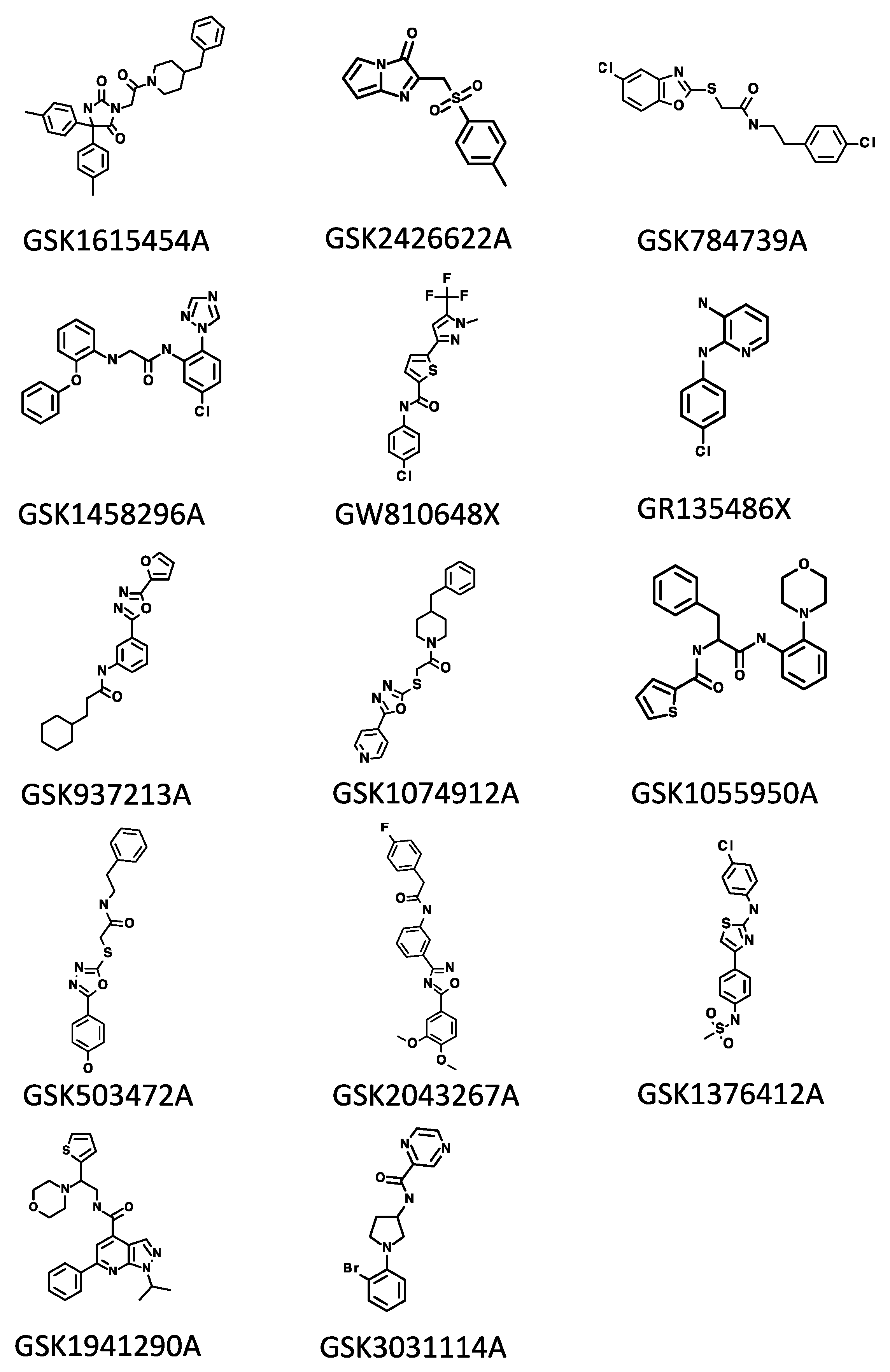

| ID | Molecular Weight | Intracellular MIC90 (μM) | Resistant Mutant Selection Media | Number of Resistant Colonies |
|---|---|---|---|---|
| 213A | 365 | 0.6 | 5× MIC glucose | 5 |
| 267A | 433 | 0.16 | 2× MIC90 ADC | 3 |
| 290A | 476 | 0.6 | 2× MIC90 ADS | 2 |
| 950A | 436 | 1.26 | 2× MIC90 ADC | 4 |
| 739A | 381 | 2 | 2× MIC90 acetate | 3 |
| 472A | 355 | 2 | 2× MIC90 acetate | 4 |
| 412A | 380 | 0.32 | 2× MIC90 ADC | 3 |
| 412A | 380 | 0.32 | 2× MIC90 glucose | 6 |
| 296A | 420 | 1 | 2× MIC90 acetate | 6 |
| 648X | 386 | 1.58 | 2× MIC90 ADC | 2 |
| 454A | 496 | 2 | 2× MIC90 ADC | 3 |
| 1114A | 347 | 2.51 | 5× MIC90 glucose | 2 |
| 486X | 220 | 0.16 | 5× MIC90 ADC | 3 |
| 912A | 394 | 1.58 | 2× MIC90 acetate | 4 |
| 622A | 288 | 2 | 5× MIC90 glucose | 3 |
| 705A | 310.8 | 1.26 | 2× MIC acetate | 1 |
| Compound | Number of Mutants | Gene/ ORFs | Genetic Polymorphisms (Frequency) | Relevant Codon Change (Frequency) | Product |
|---|---|---|---|---|---|
| 213A | 5 | mmpL3 | 755A > G; 758G > A; 875T > C; 1985C > A; 2051T > C | Tyr252Cys; Gly253Glu; Ile292Thr; Ala662Glu; Val684Ala | Conserved membrane transport protein |
| sugl | 16C > T | Gln6*, gained stop codon (2) | Involved in transport of sugar across the membrane. Translocation of the substrate. | ||
| 267A | 3 | mmpL3 | 763T > C; 765C > G; 1932 C > A | Phe255Leu (2); Phe644Leu | Conserved membrane transport protein |
| 290A | 2 | mmpL3 | 1909C > A | Leu637Ile (2) | Conserved membrane transport protein |
| Rv0370c | 474G > T | Val158Val (2) | Unknown. possible Oxidoreductase | ||
| 950A | 4 | rpoB | 1720G > A | Asp574Asn (4) | Transcription of DNA into RNA |
| Rv3629 | 641G > A | Gly214Glu (4) | Probable conserved integral membrane protein | ||
| 739A | 3 | ethA | 611T > C | Met204Thr (2) | Monooxygenase that activates the pro-drug ethionamide (ETH) |
| rpsO | 157C > T | Arg53Trp | 30S ribosomal protein S15 | ||
| Rv1024 | 154C > T | Pro52Ser | Possible conserved membrane protein | ||
| Rv3220c | 746C > CA (2) | indels, frameshift variant | Probable two-component sensor kinase | ||
| 472A | 4 | ethA | 611T > C | Met204Thr (3) | Monooxygenase that activates the pro-drug ETH |
| rpsO | 157C > T | Arg53Trp | 30S ribosomal protein S15 | ||
| Rv3220c | 746C > CA (3) | indels, frameshift variant | Probable two-component sensor kinase | ||
| 412A | 9 | prrB | 452T > C (7); 548C > T; 875A > G | Leu151Pro (7); Thr183Ile; Gln292Arg; | Two-component regulatory system PRRA/PRRB |
| moaC3 | 392A > G | Asp131Gly | Probable molybdenum cofactor biosynthesis protein | ||
| iniB | 290C > T | Thr97Ile | Isoniazid inductible gene protein. | ||
| narL | 298G > C | Ala100Pro | Possible nitrate/nitrite response transcriptional regulatory protein | ||
| ctpl | 3113TGCGAG > T | Indels, frameshift variant | Probable cation-transporter ATPase I | ||
| 296A | 6 | TB18.5 | 145G > C; 236A > G; 243C > A; 277A > G | Val49Leu (2); Tyr79Cys; His81Gln; Thr93Ala (2) | Conserved protein |
| Rv1948 | 122C > A | Ala41Glu | Hypothetical protein | ||
| pks6 | 667G > A | Val223Ile (2) | Probable membrane-bound polyketide synthase | ||
| 648X | 2 | ctpC | 1511C > T | Ser504Phe | Probable metal cation-transporting P-type ATPase C |
| 454A | 3 | Rv0678 | 466 G > GC (3) | indels, frameshift variation | Conserved protein |
| mbtA | 1369CT > C (3) | Indels, frameshift variation | Bifunctional enzyme MbtA: salicyl-AMP ligase (SAL-AMP ligase) + salicyl-S-ArCP synthetase | ||
| 1114A | 2 | dnaE1 | 2215A > G | Met739Val | Probable DNA polymerase III (alpha chain) DnaE1 (DNA nucleotidyltransferase) |
| virS | 983C > A | Pro328His | Virulence-regulating transcriptional regulator VirS (AraC/XylS family) | ||
| Rv0585c | 1202A > C | Asp401Ala | Conserved integral membrane protein | ||
| sugl | 16C > T | Gln6* (2), gained stop codon | Involved in transport of sugar across the membrane. Responsible for the translocation of the substrate across the membrane. | ||
| 486X | 3 | phoR | 661G > C | Ala221Pro, | Possible two-component system response sensor kinase membrane associated PhoR |
| fbiC | 1082C > A | Thr361Lys, | Probable F420 biosynthesis protein FbiC | ||
| fbiA | 866T > A | Leu289Gln (2) | Probable F420 biosynthesis protein FbiA | ||
| Rv3327 | 296C > G | Pro100Ala | Probable transposase fusion protein | ||
| 912A | 3 | ethA | 205T > C; 190T > C | Trp69Arg; Phe64Ile | Monooxygenase that activates the pro-drug ETH |
| Rv2542 | 1042G > A | Ala348Thr (2) | Conserved hypothetical protein | ||
| Rv3083 | 783G > A; 806T > C | Trp261*, gained stop codon; Leu269Pro | Probable monooxygenase (hydroxylase) | ||
| 622A | 3 | ftsK | 1192T > C | Ser398Pro | Possible cell division transmembrane protein |
| virS | 712G > T | Val238Phe | Virulence-regulating transcriptional regulator VirS (AraC/XylS family) | ||
| sugl | 16C > T | Gln6* (3), gained stop codon | Involved in transport of sugar across the membrane. Responsible for the translocation of the substrate across the membrane. | ||
| 705A | 1 | Rv3083 | 380_381 G > GA, indels, frame shift variant | Glu127_Thr128fs | Probable monooxygenase (hydroxylase) |
| sugl | 16C > T | Gln6*, gained stop codon | Involved in transport of sugar across the membrane. Responsible for the translocation of the substrate across the membrane. |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tsui, C.K.M.; Sorrentino, F.; Narula, G.; Mendoza-Losana, A.; del Rio, R.G.; Herrán, E.P.; Lopez, A.; Bojang, A.; Zheng, X.; Remuiñán-Blanco, M.J.; et al. Hit Compounds and Associated Targets in Intracellular Mycobacterium tuberculosis. Molecules 2022, 27, 4446. https://doi.org/10.3390/molecules27144446
Tsui CKM, Sorrentino F, Narula G, Mendoza-Losana A, del Rio RG, Herrán EP, Lopez A, Bojang A, Zheng X, Remuiñán-Blanco MJ, et al. Hit Compounds and Associated Targets in Intracellular Mycobacterium tuberculosis. Molecules. 2022; 27(14):4446. https://doi.org/10.3390/molecules27144446
Chicago/Turabian StyleTsui, Clement K. M., Flavia Sorrentino, Gagandeep Narula, Alfonso Mendoza-Losana, Ruben Gonzalez del Rio, Esther Pérez Herrán, Abraham Lopez, Adama Bojang, Xingji Zheng, Modesto Jesus Remuiñán-Blanco, and et al. 2022. "Hit Compounds and Associated Targets in Intracellular Mycobacterium tuberculosis" Molecules 27, no. 14: 4446. https://doi.org/10.3390/molecules27144446
APA StyleTsui, C. K. M., Sorrentino, F., Narula, G., Mendoza-Losana, A., del Rio, R. G., Herrán, E. P., Lopez, A., Bojang, A., Zheng, X., Remuiñán-Blanco, M. J., & Av-Gay, Y. (2022). Hit Compounds and Associated Targets in Intracellular Mycobacterium tuberculosis. Molecules, 27(14), 4446. https://doi.org/10.3390/molecules27144446






