Investigation of the New Inhibitors by Sulfadiazine and Modified Derivatives of α-D-glucopyranoside for White Spot Syndrome Virus Disease of Shrimp by In Silico: Quantum Calculations, Molecular Docking, ADMET and Molecular Dynamics Study
Abstract
1. Introduction
2. Computational Details of Procedure
2.1. Optimization and Ligand Preparation
2.2. Protein Preparation and Collection
2.3. Molecular Docking and Visualization of Docking
2.4. Pharmacokinetics and ADMET Studies
2.5. Lipinski Rule and Pharmacokinetics
2.6. Molecular Dynamic
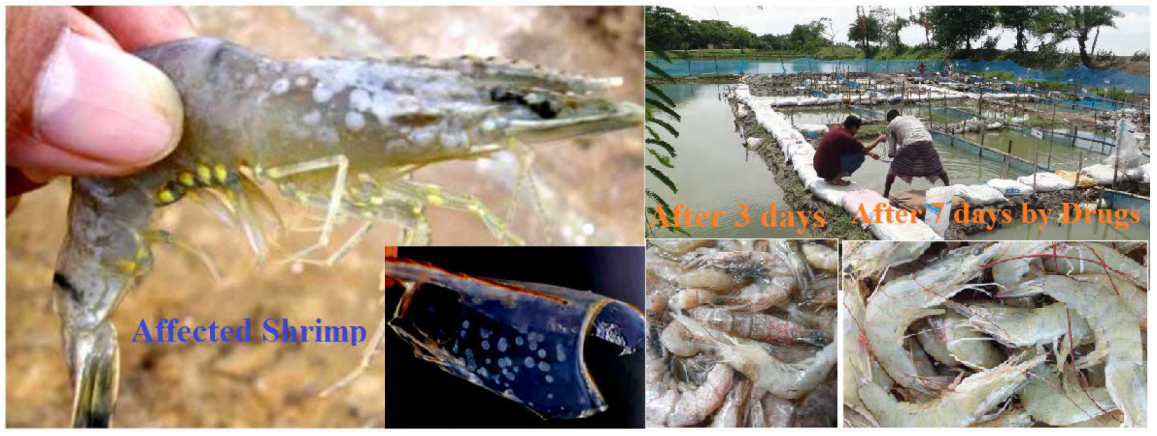
2.7. White Spot Trial Procedure
3. Results and Discussions
3.1. Optimized Structure
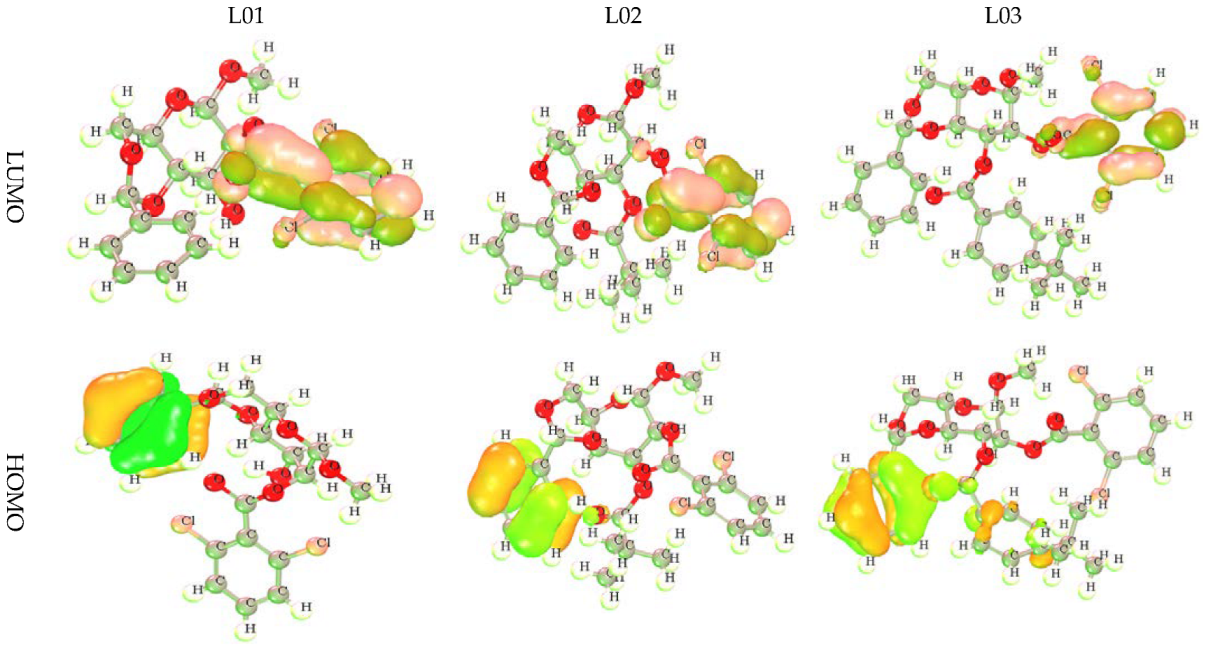
3.2. HOMO, LUMO, and Chemical Reactivity Descriptors
3.3. Frontier Molecular Orbital: HOMO and LUMO
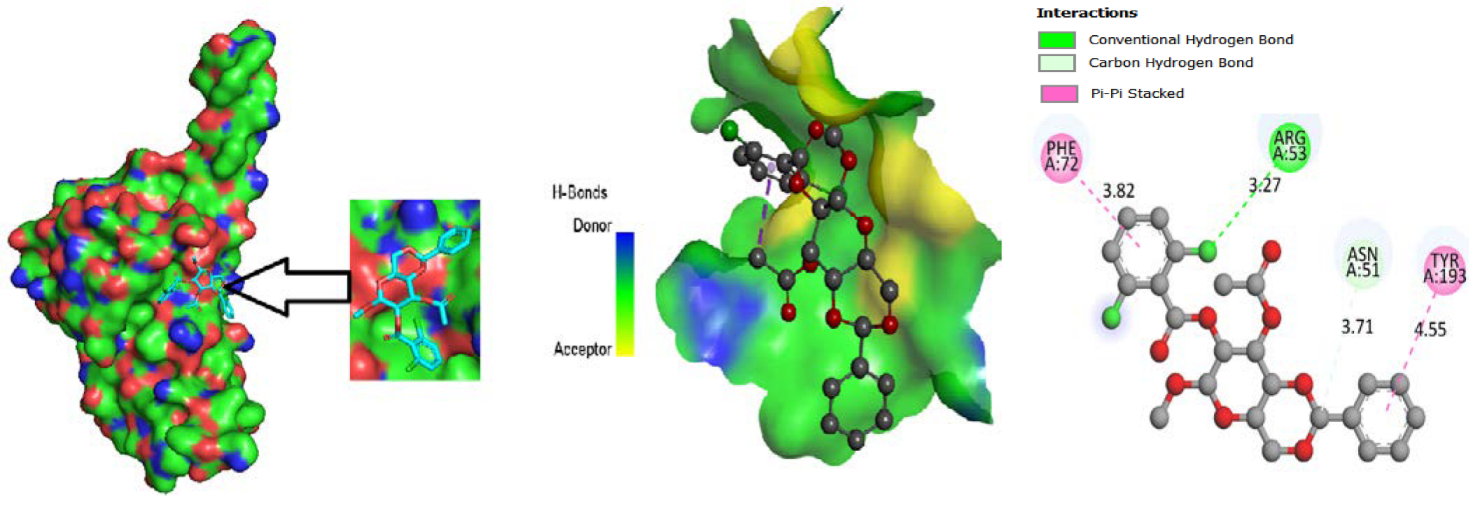
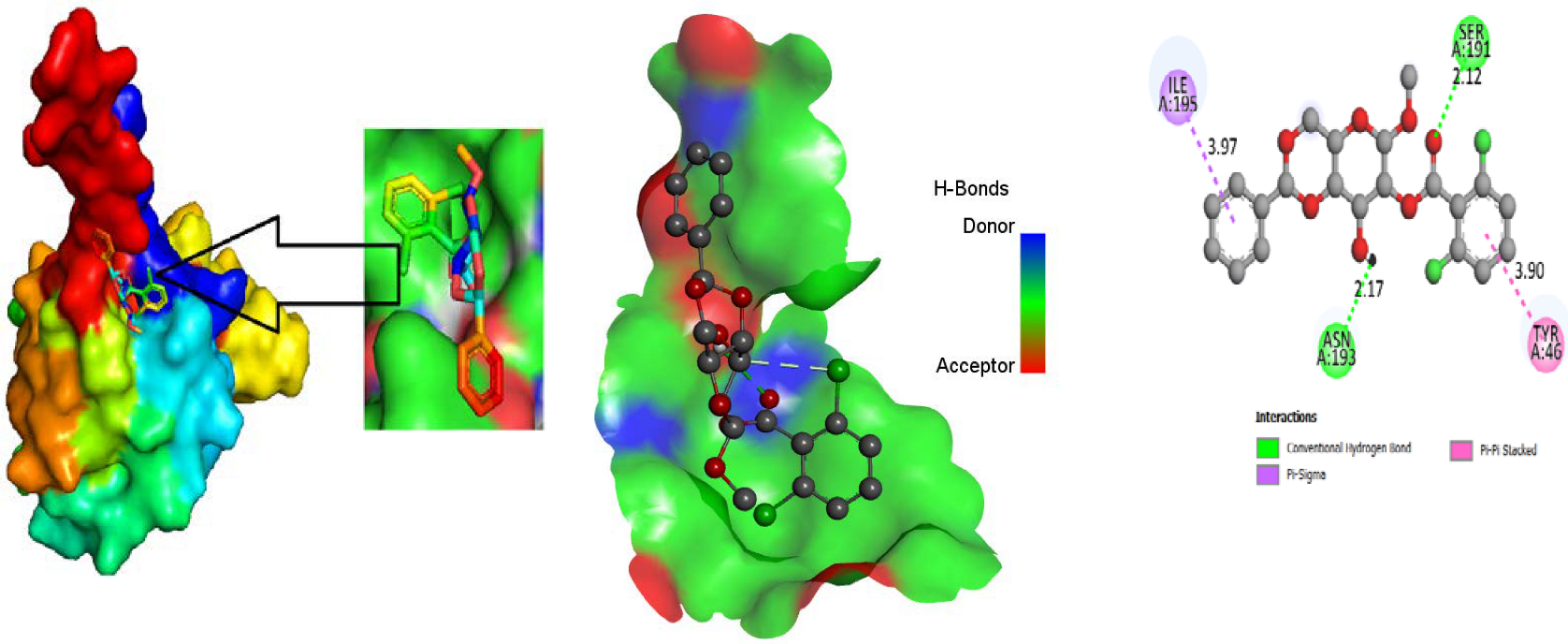
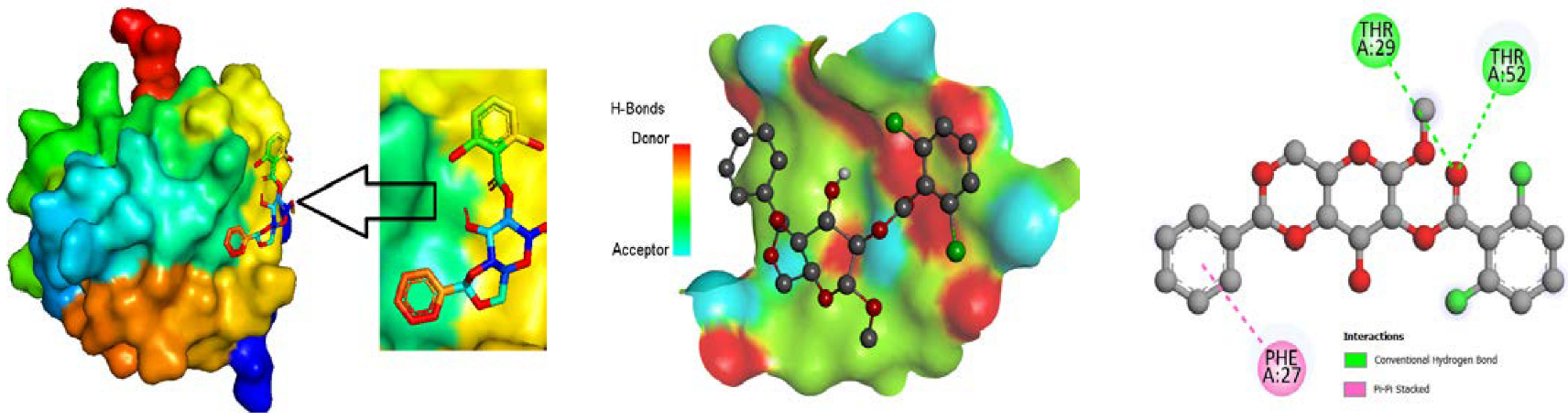
3.4. Molecular Docking
3.5. Protein-Ligand Interaction
3.6. Pharmacokinetics and Drug-Likeness Study
3.7. Pharmacokinetics and ADMET Studies
3.8. Aquatic and Non-Aquatic Toxicity
3.9. Protein-Ligand Interaction
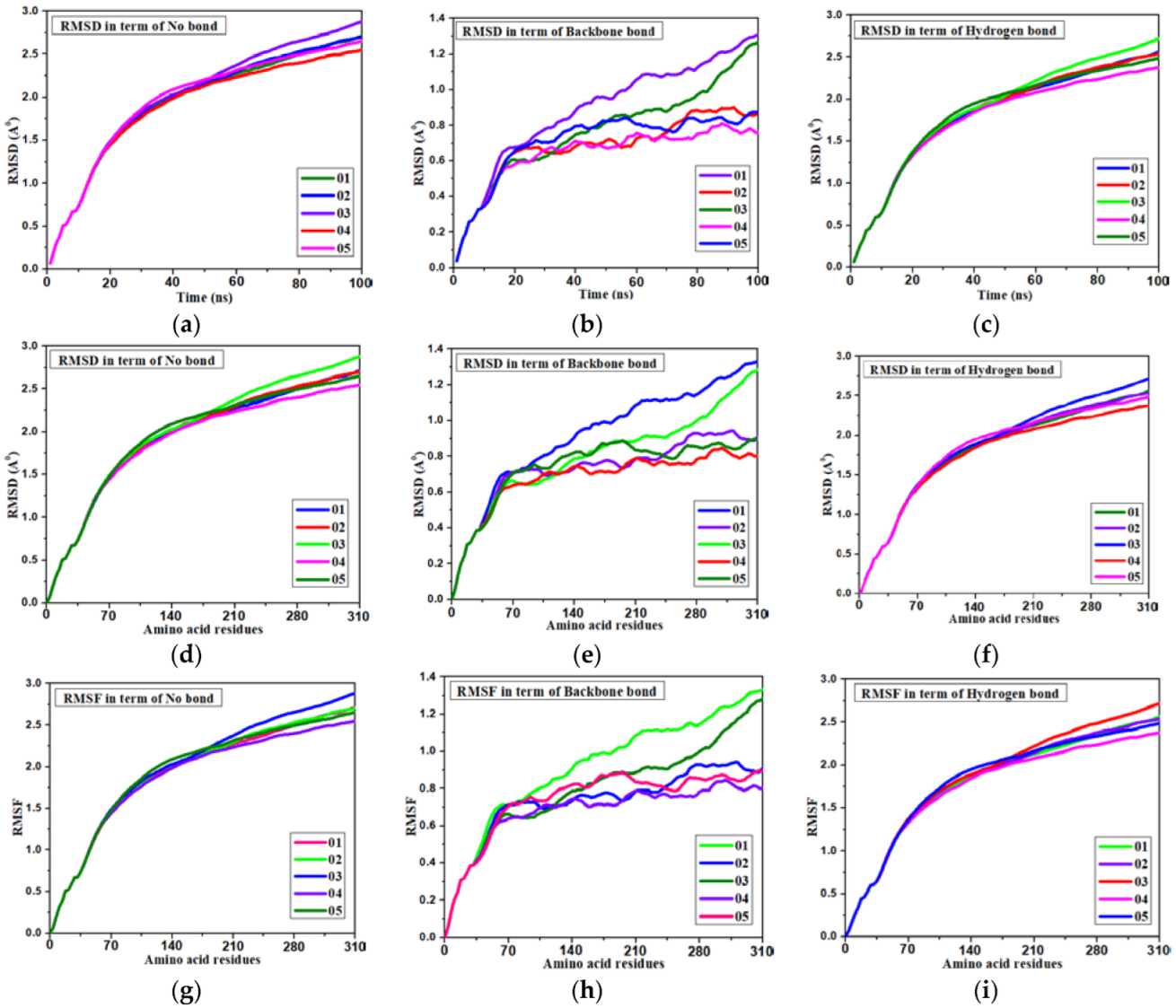
3.10. Molecular Dynamics
3.11. Trial Results
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Shamsuzzaman, M.; Mozumder, M.M.H.; Mitu, S.J.; Ahamad, A.F.; Bhyuian, S. The economic contribution of fish and fish trade in Bangladesh. Aquac. Fish. 2020, 5, 174–181. [Google Scholar] [CrossRef]
- Muir, J.F.J.A.F. Fish, feeds, and food security. Anim. Front. 2013, 3, 28–34. [Google Scholar] [CrossRef][Green Version]
- Thorpe, A.; Andrew, N.L.; Allison, E.H. Fisheries and poverty reduction. Nutr. Nat. Resour. 2007, 2, 85. [Google Scholar] [CrossRef]
- Hossain, M.A.R. An overview of Fisheries sector of Bangladesh. Res. Agric. Livest. Fish. 2014, 1, 109–126. [Google Scholar] [CrossRef]
- Craig, J.H.; Barr, A.S.; Bean, J.J.F. The Bangladesh floodplain fisheries. Fish. Res. 2004, 66, 271–286. [Google Scholar] [CrossRef]
- Islam, M.S. Perspectives of the coastal and marine fisheries of the Bay of Bengal, Bangladesh. Ocean. Coast. Manag. 2003, 46, 763–796. [Google Scholar] [CrossRef]
- Shamsuzzaman, M.M.; Islam, M.M.; Tania, N.J.; Abdullah Al-Mamun, M.; Barman, P.P.; Xu, X. Fisheries resources of Bangladesh: Present status and future direction. Aquac. Fish. 2017, 2, 145–156. [Google Scholar] [CrossRef]
- Dey, S. Relationship between rice production, fisheries production and gross domestic product (gdp) in bangladesh: Co integrating Regression Analysis (1971–2017). Int. J. Econ. Financ. Issues 2020, 1, 201–216. [Google Scholar]
- Chakraborty, B.K. Status of Fish Diversity and Production in Bangladesh. Integr. Biol. Resour. Prosper. 2021, 1, 85–99. [Google Scholar]
- Protom Alo Desk Report. Bangladesh attains self-sufficiency in fish production in terms of consumption against demand: Minister. Prothom. Alo. 2021. Available online: https://en.prothomalo.com/bangladesh/parliament/bangladesh-attains-self-sufficiency-in-fish-production-in-terms-of-consumption-against-demand-minister (accessed on 15 September 2021).
- Hussain, M.G. Freshwater fishes of Bangladesh: Fisheries, biodiversity and habitat. Aquat. Ecosyst. Heal. Manag. 2010, 13, 85–93. [Google Scholar] [CrossRef]
- Hossain, M.M.I.; Aminul, M.; Ridgway, S.; Matsuishi, T. Management of inland open water fisheries resources of Bangladesh: Issues and options. Fish. Res. 2006, 77, 275–284. [Google Scholar] [CrossRef]
- Alam, M.F.T.; Kenneth, J. Current constraints and future possibilities for Bangladesh fisheries. Food Policy 2001, 26, 297–313. [Google Scholar] [CrossRef]
- Habib, T.B. Bangladesh sees significant rise in fish production in a decade. Financ. Express 2021, 7. Available online: https://thefinancialexpress.com.bd/trade/bangladesh-sees-significant-rise-in-fish-production-in-a-decade-1636256725 (accessed on 7 November 2021).
- Toufique, K.A.; Belton, B. Is aquaculture pro-poor? Empirical evidence of impacts on fish consumption in Bangladesh. Wolrd Dev. 2014, 64, 609–620. [Google Scholar] [CrossRef]
- Ahmed, N.; Diana, J.S.J.O.; Management, C. Threatening “white gold”: Impacts of climate change on shrimp farming in coastal Bangladesh. Ocean. Coast. Manag. 2015, 114, 42–52. [Google Scholar] [CrossRef]
- Ahmed, N. Linking prawn and shrimp farming towards a green economy in Bangladesh: Confronting climate change. Ocean Coast. Manag. 2013, 75, 33–42. [Google Scholar] [CrossRef]
- Swapan, M.; Gavin, M. A desert in the delta: Participatory assessment of changing livelihoods induced by commercial shrimp farming in Southwest Bangladesh. Ocean Coast. Manag. 2011, 54, 45–54. [Google Scholar] [CrossRef]
- Department of Fisheries, Ministry of Fisheries: Banglades. Yearbook of Fisheries Statistics of Bangladesh (July 2017–June 2018); Fisheries Resources Survey System (FRSS): Dhaka, Bangladesh, 2018; Volume 35, pp. 1–129. [Google Scholar]
- Rahman, M.; Hossain, M. Production and Export of Shrimp of Bangladesh: Problems and Prospects. Progress. Agric. 2013, 20, 163–171. [Google Scholar] [CrossRef]
- Paul, B.G.; Vogl, C.R. Impacts of shrimp farming in Bangladesh: Challenges and alternatives. Ocean. Coast. Manag. 2009, 54, 201–211. [Google Scholar] [CrossRef]
- Alam, S.M.; Pokrant, B.; Yakupitiyage, A.; Phillips, M.J. Economic returns of disease-affected extensive shrimp farming in southwest Bangladesh. Aquac. Int. 2007, 15, 363–370. [Google Scholar] [CrossRef]
- Hossain, M.S.; Karunasagar, I.; Karunasagar, I. Detection of white spot syndrome virus (WSSV) in wild captured shrimp and in non-cultured crustaceans from shrimp ponds in Bangladesh by polymerase chain reaction. Fish Pathol. 2001, 36, 93–95. [Google Scholar] [CrossRef][Green Version]
- Talukder, A.S.; Punom, N.J.; Eshik, M.E.; Begum, M.K.; Islam, H.R.; Hossain, Z.; Rahman, M.S. Molecular identification of white spot syndrome virus (WSSV) and associated risk factors for white spot disease (WSD) prevalence in shrimp (Penaeus monodon) aquaculture in Bangladesh. J. Invertebr. Pathol. 2021, 179, 107535. [Google Scholar] [CrossRef]
- Hasan, N.A.; Haque, M.M.; Hinchliffe, S.J.; Guilder, J. A sequential assessment of WSD risk factors of shrimp farming in Bangladesh: Looking for a sustainable farming system. Aquaculture 2020, 526, 735348. [Google Scholar] [CrossRef]
- Colorni, A.; Burgess, P. Cryptocaryon irritans Brown 1951, the cause of ‘white spot disease’in marine fish: An update. Aquar. Sci. Conserv. 1997, 1, 217–238. [Google Scholar] [CrossRef]
- Singh, I.S.B.; Manjusha, M.; Pai, S.; Philip, R. Fenneropenaeus indicus is protected from white spot disease by oral administration of inactivated white spot syndrome virus. Dis. Aquat. Org. 2005, 66, 265–270. [Google Scholar] [CrossRef] [PubMed]
- van Hulten, M.C.; Witteveldt, J.; Peters, S.; Kloosterboer, N.; Tarchini, R.; Fiers, M.; Sandbrink, H.; Lankhorst, R.K.; Vlak, J.M. The white spot syndrome virus DNA genome sequence. Virology 2001, 286, 7–22. [Google Scholar] [CrossRef] [PubMed]
- Escobedo-Bonilla, C.M.; Alday-Sanz, V.; Wille, M.; Sorgeloos, P.; Pensaert, M.B.; Nauwynck, H.J. A review on the morphology, molecular characterization, morphogenesis and pathogenesis of white spot syndrome virus. J. Fish Dis. 2008, 31, 1–18. [Google Scholar] [CrossRef]
- Graslund, S.; Bengtsson, E.B. Chemicals and biological products used in south-east Asian shrimp farming, and their potential impact on the environment—A review. Sci. Total Environ. 2001, 280, 93–131. [Google Scholar] [CrossRef]
- Primavera, J.H. Socio-economic impacts of shrimp culture. Aquac. Res. 1997, 28, 815–827. [Google Scholar] [CrossRef]
- Rajasegar, M.; Srinivasan, M.; Rajaram, R. Phytoplankton diversity associated with the shrimp farm development in Vellar estuary, south India. Seaweed Res. Utiln 2000, 22, 125–213. [Google Scholar]
- Long, P.H. Sulfadiazine: The 2-Sulfanilamidopyrimidine Analogue of Sulfanilamide. J. Am. Med. Assoc. 1941, 116, 2399–2401. [Google Scholar] [CrossRef]
- Wheeler, C.; Plummer, N. Sulfadiazine and sodium sulfadiazine: A comparison of certain of their clinical and pharmacologic values. Ann. Intern. Med. 1942, 16, 269. [Google Scholar] [CrossRef]
- Kohn, W.; Becke, A.D.; Parr, R.G. Density Functional Theory of Electronic Structure. J. Phys. Chem. 1996, 100, 12974–12980. [Google Scholar] [CrossRef]
- Parr, R.G. Density functional theory of atoms and molecules. In Horizons of Quantum Chemistry; Springer: Berlin/Heidelberg, Germany, 1980; pp. 5–15. [Google Scholar] [CrossRef]
- Parr, R.G.; Yang, W. Density functional approach to the frontier-electron theory of chemical reactivity. J. Am. Chem. Soc. 1984, 106, 4049–4050. [Google Scholar] [CrossRef]
- Tang, X.; Wu, J.; Sivaraman, J.; Hew, C.L. Crystal Structures of Major Envelope Proteins VP26 and VP28 from White Spot Syndrome Virus Shed Light on Their Evolutionary Relationship. J. Virol. 2007, 81, 6709–6717. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Wu, J.; Song, J.; Sivaraman, J.; Hew, C.L. Identification of a novel nonstructural protein, VP9, from white spot syndrome virus: Its structure reveals a ferredoxin fold with specific metal binding sites. J. Virol. 2006, 80, 10419–10427. [Google Scholar] [CrossRef]
- Accelrys Software Inc. Discovery Studio Modeling Environment. 2013. Release 4., San Diego, CA. Available online: https://research.csc.fi/documents/48467/72092/Discovery_Studio_4.0_Product_Release_Document.pdf/0e006e19-2e8f-45e8-aad5-e10c744ee285 (accessed on 10 November 2021).
- Tsaioun, K.; Blaauboer, B.J.; Hartung, T. Evidence-based absorption, distribution, metabolism, excretion (ADME) and its interplay with alternative toxicity methods. Altern. Anim. Exp. ALTEX 2016, 33, 343–358. [Google Scholar] [CrossRef]
- Chandrasekaran, B.; Abed, S.N.; Al-Attraqchi, O.; Kuche, K.; Tekade, R.K. Computer-aided prediction of pharmacokinetic (ADMET) properties. In Dosage Form Design Parameters; Elsevier: Amsterdam, The Netherlands, 2018; pp. 731–755. [Google Scholar]
- Pires, D.E.V.; Blundell, T.L.; Ascher, D.B. pkCSM: Predicting Small-Molecule Pharmacokinetic and Toxicity Properties Using Graph-Based Signatures. J. Med. Chem. 2015, 58, 4066–4072. [Google Scholar] [CrossRef]
- Daina, A.; Michielin, O.; Zoete, V. SwissADME: A free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci. Rep. 2017, 7, 42717. [Google Scholar] [CrossRef]
- Phillips, J.C.; Hardy, D.J.; Maia, J.D.C.; Stone, J.E.; Ribeiro, J.V.; Bernardi, R.C.; Buch, R.; Fiorin, G.; Hénin, J.; Jiang, W.; et al. Scalable molecular dynamics on CPU and GPU architectures with NAMD. J. Chem. Phys. 2020, 153, 044130. [Google Scholar] [CrossRef]
- Skjevik, A.; Madej, B.D.; Dickson, C.J.; Teigen, K.; Walker, R.C.; Gould, I.R. All-atom lipid bilayer self-assembly with the AMBER and CHARMM lipid force fields. Chem. Commun. 2015, 51, 4402–4405. [Google Scholar] [CrossRef]
- López, J.; Anitescu, C.; Valizadeh, N.; Rabczuk, T.; Alajlan, N. Structural shape optimization using Bézier triangles and a CAD-compatible boundary representation. Eng. Comput. 2020, 36, 1657–1672. [Google Scholar] [CrossRef]
- Kumer, A.; Ahmed, B.; Sharif, A.; Al-Mamun, A. A Theoretical Study of Aniline and Nitrobenzene by Computational Overview. Asian J. Phys. Chem. Sci. 2017, 4, 1–12. [Google Scholar] [CrossRef]
- Kumer, A.; Paul, S.; Sarker, N.; Islam, M.J. The Prediction of Thermo Physical, Vibrational Spectroscopy, Chemical Reactivity, Biological Properties of Morpholinium Borate, Phosphate, Chloride and Bromide Ionic Liquid: A DFT Study. Int. J. New Chem. 2019, 6, 236–253. [Google Scholar] [CrossRef]
- Kumer, A.; Sarker, N.; Paul, S. The theoretical investigation of HOMO, LUMO, thermophysical properties and QSAR study of some aromatic carboxylic acids using HyperChem programming. Int. J. Chem. Technol. 2019, 3, 26–37. [Google Scholar] [CrossRef]
- Kumer, A.; Sarker, N.; Paul, S.; Zannat, A. The Theoretical Prediction of Thermophysical properties, HOMO, LUMO, QSAR and Biological Indics of Cannabinoids (CBD) and Tetrahhdrocannabinol (THC) by Computational Chemistry. Adv. J. Chem. A 2019, 2, 190–202. [Google Scholar] [CrossRef]
- Kumer, A.; Sarker, M.N.; Paul, S. The thermo physical, HOMO, LUMO, Vibrational spectroscopy and QSAR study of morphonium formate and acetate Ionic Liquid Salts using computational method. Turk. Comput. Theor. Chem. 2019, 3, 59–68. [Google Scholar] [CrossRef]
- Kumer, A.; Sarker, M.N.; Sunanda, P. The Simulating Study of HOMO, LUMO, thermo physical and Quantitative Structure of Activity Relationship (QSAR) of Some Anticancer Active Ionic Liquids. Eurasian J. Environ. Res. 2019, 3, 1–10. [Google Scholar]
- Nath, A.; Kumer, A.; Zaben, F.; Khan, M.W. Investigating the binding affinity, molecular dynamics, and ADMET properties of 2, 3-dihydrobenzofuran derivatives as an inhibitor of fungi, bacteria, and virus protein. Beni-Suef. Univ. J. Basic Appl. Sci. 2021, 10, 36. [Google Scholar] [CrossRef]
- Kawsar, S.M.A.; Kumer, A. Computational investigation of methyl α-D-glucopyranoside derivatives as inhibitor against bacteria, fungi and COVID-19 (SARS-2). J. Chil. Chem. Soc. 2021, 66, 5206–5214. [Google Scholar] [CrossRef]
- Kumer, A.; Khan, M.W. The effect of alkyl chain and electronegative atoms in anion on biological activity of anilinium carboxylate bioactive ionic liquids and computational approaches by DFT functional and molecular docking. Heliyon 2021, 7, e07509. [Google Scholar] [CrossRef]
- Kumer, A.; Khan, M.W. Synthesis, characterization, antimicrobial activity and computational exploration of ortho toludinium carboxylate ionic liquids. J. Mol. Struct. 2021, 1245, 131087. [Google Scholar] [CrossRef]
- Hornig, H.; Woolley, P.; Lührmann, R. Decoding at the ribosomal A site: Antibiotics, misreading and energy of aminoacyl-tRNA binding. Biochimie 1987, 69, 803–813. [Google Scholar] [CrossRef]
- Babahedari, A.K.; Soureshjani, E.H.; Shamsabadi, M.K.; Kabiri, H. The Comprehensive Evaluation Docking of Methicillin Drug Containing Isoxazole Derivatives, as Targeted Antibiotics to Staphylococcus Aureus. J. Bionanosci. 2013, 7, 288–291. [Google Scholar] [CrossRef]
- Cheng, K.; Zheng, Q.Z.; Qian, Y.; Shi, L.; Zhao, J.; Zhu, H.L. Synthesis, antibacterial activities and molecular docking studies of peptide and Schiff bases as targeted antibiotics. Bioorganic Med. Chem. 2009, 17, 7861–7871. [Google Scholar] [CrossRef]
- Hermann, T.; Westhof, E. Docking of Cationic Antibiotics to Negatively Charged Pockets in RNA Folds. J. Med. Chem. 1999, 42, 1250–1261. [Google Scholar] [CrossRef]
- Athar, M.; Sona, A.N.; Bekono, B.D.; Ntie-Kang, F. Fundamental physical and chemical concepts behind “drug-likeness” and “natural product-likeness”. In Fundamental Concepts; De Gruyter: Berlin/Boston, Germany, 2020; pp. 55–80. [Google Scholar]
- Lipinski, C.A.; Lombardo, F.; Dominy, B.W.; Feeney, P.J. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv. Drug Deliv. Rev. 1997, 23, 3–25. [Google Scholar] [CrossRef]
- van Breemen, R.B.; Li, Y. Caco-2 cell permeability assays to measure drug absorption. Expert Opin. Drug Metab. Toxicol. 2005, 1, 175–185. [Google Scholar] [CrossRef]
- Kumer, A.; Chakma, U.; Matin, M.M.; Akash, S.; Howlader, D.; Chandro, A. The computational screening of inhibitor for black fungus and white fungus by D-glucofuranose derivatives using in silico and SAR study. Org. Commun. 2021, 14, 305–322. [Google Scholar] [CrossRef]
- Patel, H.M.; Noolvi, M.N.; Sharma, P.; Jaiswal, V.; Bansal, S.; Lohan, S.; Kumar, S.S.; Abbot, V.; Dhiman, S.; Bhardwaj, V. Quantitative structure–activity relationship (QSAR) studies as strategic approach in drug discovery. Med. Chem. Res. 2014, 23, 4991–5007. [Google Scholar] [CrossRef]
- Trott, O.; Olson, A.J. AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 2010, 31, 455–461. [Google Scholar] [CrossRef]
- Rahman, M.A.; Matin, M.M.; Kumer, A.; Chakma, U.; Rahman, M. Modified D-Glucofuranoses as New Black Fungus Protease Inhibitors: Computational Screening, Docking, Dynamics, and QSAR Study. Phys. Chem. Res. 2022, 10, 195–209. [Google Scholar]
- Kumer, A.; Chakma, U.; Kawsar, S.M.A. The Inhibitory Effect of Some Natural Bioactive D-Glucopyranoside Derivatives against SARS-CoV-2 Main Protease (Mpro) and Spike Protease (Spro). ASM Sci. J. 2021, 16, 1–18. [Google Scholar] [CrossRef]


| Title | PDB ID: 2ED6 | PDB ID: 2GJ2 | PDB ID: 2GJI | PDB ID: 2EDM |
|---|---|---|---|---|
| Organism | Shrimp white spot syndrome virus | Shrimp white spot syndrome virus | Shrimp white spot syndrome virus | Shrimp white spot syndrome virus |
| Resolution | 2.00 Å | 2.35 Å | N/A | 2.20 Å |
| R-Value Free | 0.281 | 0.275 | N/A | 0.278 |
| References | [38] | [39] | [39] | [38] |
| Protein Name with PDB ID | Grid Box Size | |
|---|---|---|
| Center | Dimension (Å) | |
| Envelope Protein WSSV (PDB 2ED6) | X = 28.2583 | X = 38.9011 |
| Y = 106.048 | Y = 67.0482 | |
| Z = 92.9776 | Z = 45.525 | |
| White Spot Syndrome Virus (PDB 2GJI) | X = −8.6514 | X = 32.888 |
| Y = 15.6227 | Y = 33.828 | |
| Z = −5.5754 | Z = 43.396 | |
| White Spot Syndrome Virus (PDB 2EDM) | X= 37.1819 | X= 39.3455 |
| Y= 35.3181 | Y= 44.655 | |
| Z= 92.9466 | Z= 61.178 | |
| White Spot Syndrome Virus (PDB 2GJ2) | X = 36.2550 | X = 34.8480 |
| Y = 1.4367 | Y = 37.8846 | |
| Z = −6.1508 | Z= 28.8952 | |
| Ligand. | LUMO | HOMO | A = −LUMO | I = −HOMO | Energy Gap = I − A | Hardness | Electrophilicity | |||
|---|---|---|---|---|---|---|---|---|---|---|
| L01 | −1.685 | −8.445 | 1.685 | 8.445 | 6.760 | −5.065 | 3.38 | 5.065 | 0.2959 | 3.7950 |
| L02 | −1.550 | −8.528 | 1.550 | 8.528 | 6.978 | −5.039 | 3.489 | 5.039 | 0.2866 | 3.6388 |
| L03 | −1.413 | −8.159 | 1.413 | 8.159 | 6.746 | −4.786 | 3.373 | 4.786 | 0.3965 | 3.3955 |
| L04 | −1.647 | −8.896 | 1.647 | 8.896 | 7.249 | −5.2715 | 3.6245 | 5.271 | 0.2759 | 3.8335 |
| L05 | −1.594 | −8.837 | 1.594 | 8.837 | 7.243 | −5.2155 | 3.6215 | 5.215 | 0.2761 | 3.7555 |
| L06 | −1.701 | −8.605 | 1.701 | 8.605 | 6.904 | −5.1530 | 3.452 | 5.153 | 0.2897 | 3.8461 |
| L07 | −1.580 | −8.573 | 1.580 | 8.573 | 6.993 | −5.0765 | 3.4965 | 5.076 | 0.2860 | 3.6852 |
| L08 | −1.624 | −8.909 | 1.624 | 8.909 | 7.285 | −5.2665 | 3.6425 | 5.266 | 0.2745 | 3.8073 |
| L09 | −0.68 | −8.673 | 0.68 | 8.673 | 7.993 | −4.6765 | 3.9965 | 4.6765 | 0.2503 | 2.7361 |
| L10 | −1.240 | −7.877 | 1.240 | 7.877 | 6.637 | −4.5585 | 3.3185 | 4.5585 | 0.3013 | 3.1309 |
| L11 | −2.163 | −9.146 | 2.163 | 9.146 | 6.983 | −5.6545 | 3.4915 | 5.6545 | 0.2864 | 4.5787 |
| L12 | −1.745 | −8.464 | 1.745 | 8.464 | 6.719 | −5.1045 | 3.3595 | 5.1045 | 0.2977 | 3.8779 |
| Ligands | Envelope Protein WSSV (PDB ID: 2ED6) | Main Protease of WSSV (PDB ID: 2GJ2) | Main Protease of WSSV (PDB ID: 2GJI) | Main Protease of WSSV (PDB ID: 2EDM) |
|---|---|---|---|---|
| L01 | −6.4 | −6.20 | −6.2 | −7.0 |
| L02 | −5.6 | −6.30 | −6.0 | −6.4 |
| L03 | −6.5 | −6.90 | −6.1 | −6.6 |
| L04 | −6.6 | −6.20 | −5.7 | −6.3 |
| L05 | −5.6 | −6.20 | −5.7 | −6.0 |
| L06 | −6.2 | −5.80 | −5.5 | −6.1 |
| L07 | −5.6 | −5.80 | −5.0 | −5.7 |
| L08 | −5.5 | −5.10 | −4.7 | −5.5 |
| L09 | −5.6 | −6.54 | −5.9 | −5.7 |
| L10 | −5.7 | −6.74 | −5.6 | −5.4 |
| L11 | −6.1 | −6.4 | −6.4 | −6.4 |
| L12 | −5.5 | −5.4 | −5.1 | −5.8 |
| Ligand | Inhibitor Constant (µM) | Ligand Efficiency (kcal/mol) | Internal Energy (kcal/mol) | Electrostatic Energy (kcal/mol) | Total Internal Energy (kcal/mol) | Torsional Energy (kcal/mol) | Unbound Energy (kcal/mol) |
|---|---|---|---|---|---|---|---|
| L01 | 40.00 | −0.19 | −7.37 | −0.15 | −2.60 | 1.79 | −2.60 |
| L02 | 36.00 | −0.17 | −8.44 | −0.24 | −2.73 | 2.39 | −2.73 |
| L03 | 10.28 | −0.15 | −9.06 | −0.09 | −4.49 | 2.68 | −4.49 |
| L04 | 16.21 | −0.38 | −7.43 | −1.40 | −1.58 | 0.89 | −1.58 |
| L05 | 11.43 | −0.40 | −7.64 | −1.37 | −0.27 | 0.89 | −0.27 |
| L06 | 17.18 | −0.43 | −6.88 | −0.46 | −3.71 | 2.45 | −2.08 |
| L07 | 17.25 | −0.36 | −6.69 | −0.64 | −4.10 | 2.32 | −1.94 |
| L08 | 18.66 | −0.32 | −6.51 | −0.69 | −3.98 | 2.11 | −1.96 |
| L09 | 22.23 | −0.22 | −6.34 | −0.71 | −3.78 | 2.67 | −1.76 |
| L10 | 21.23 | −0.24 | −6.10 | −0.88 | −3.67 | 3.20 | −2.44 |
| L11 | 20.01 | −0.26 | −5.90 | −0.81 | −4.20 | 2.45 | −2.87 |
| L12 | 18.56 | −0.28 | −6.10 | −0.96 | −4.36 | 2.98 | 2.62 |
| Ligands | NBR | HBA | HBD | TPSA, Ų | Consensus Log Po/w | Log Kp (Skin Permeation), cm/s | Lipinski Rule | MW | Bioavailability Score | GI Absorption | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Result | Violation | ||||||||||
| L01 | 05 | 07 | 01 | 83.45 | 3.16 | −6.69 | Yes | 00 | 455.29 | 0.55 | High |
| L02 | 08 | 08 | 0 | 89.52 | 4.23 | −5.81 | No | 01 | 539.40 | 0.55 | High |
| L03 | 09 | 08 | 0 | 89.52 | 5.64 | −4.90 | No | 02 | 615.50 | 0.17 | Low |
| L04 | 07 | 08 | 00 | 89.52 | 3.37 | −6.54 | Yes | 00 | 497.32 | 0.55 | High |
| L05 | 10 | 08 | 00 | 89.52 | 4.47 | −5.83 | Yes | 01 | 539.40 | 0.55 | High |
| L06 | 11 | 08 | 00 | 89.52 | 4.75 | −5.53 | Yes | 01 | 553.43 | 0.55 | High |
| L07 | 06 | 07 | 00 | 80.29 | 7.79 | −2.93 | No | 02 | 649.65 | 0.17 | Low |
| L08 | 19 | 08 | 00 | 89.52 | 7.56 | −3.13 | No | 02 | 665.64 | 0.17 | Low |
| L09 | 03 | 06 | 02 | 109.47 | −0.26 | −8.16 | Yes | 0 | 260.36 | 0.55 | High |
| L10 | 03 | 04 | 01 | 119.13 | 1.18 | −7.41 | Yes | 0 | 267.33 | 0.55 | High |
| L11 | 02 | 10 | 07 | 201.85 | −1.04 | −9.62 | No | 02 | 460.43 | 0.11 | Low |
| L12 | N/A | N/A | N/A | N/A | N/A | N/A | N/A | N/A | 528.16 | N/A | N/A |
| Ligands. | Caco-2 Permeability | Blood Brain Barrier Permeant | P-I Glycoprotein Inhibitor | P-Glycoprotein Substrate | Total Clearance | CYP2C9 Inhibitor | CYP 1A2 Inhibitor |
|---|---|---|---|---|---|---|---|
| L01 | 1.47 | No | Yes | No | 0.595 | No | No |
| L02 | 1.807 | No | Yes | No | 0.431 | No | No |
| L03 | 1.878 | No | Yes | No | 0.424 | Yes | No |
| L04 | 1.70 | No | Yes | No | 0.561 | No | No |
| L05 | 0.747 | No | No | No | 0.711 | No | Yes |
| L06 | 1.778 | No | Yes | No | 0.627 | Yes | Yes |
| L07 | 1.758 | No | Yes | No | 0.705 | No | No |
| L08 | 1.59 | No | Yes | No | 0.873 | No | No |
| L09 | −0.018 | No | No | No | 0.642 | No | No |
| L10 | 1.296 | No | No | No | −0.112 | No | No |
| L11 | −0.538 | No | No | Yes | 0.456 | No | No |
| L12 | −0.595 | N/A | No | Yes | 0.225 | No | No |
| Ligands | Max Tolerated Dose (mg/kg/day) | Oral Rat Chronic Toxicity ((LOAEL) | Hepatotoxicity | AMES Toxicity | Water Solubility, Log S | Oral Rat Acute Toxicity (LD50) (mol/kg) | T. Pyriformis Toxicity (log μg/L) |
|---|---|---|---|---|---|---|---|
| L01 | 0.581 | 1.556 | No | No | −4.658 | 2.746 | 0.285 |
| L02 | 0.674 | 1.530 | No | No | −4.219 | 3.034 | 0.285 |
| L03 | 0.590 | 1.113 | No | No | −3.698 | 2.910 | 0.285 |
| L04 | 0.822 | 1.522 | No | No | −4.674 | 3.264 | 0.285 |
| L05 | 0.438 | 10.30 | No | Yes | −2.892 | 2.482 | 0.285 |
| L06 | 0.763 | 1.524 | No | No | −5.509 | 3.148 | 0.285 |
| L07 | 0.525 | 1.396 | No | No | −4.321 | 2.302 | 0.285 |
| L08 | 0.700 | 1.497 | No | No | −5.021 | 2.621 | 0.285 |
| L09 | 1.156 | 1.97 | Yes | No | −2.954 | 2.234 | 0.285 |
| L10 | 1.014 | 1.838 | Yes | No | −3.076 | 2.348 | 0.285 |
| L11 | 1.136 | 5.156 | No | No | −2.528 | 5.156 | 0.285 |
| L12 | 1.045 | 4.524 | No | No | −2.497 | 2.456 | 0.285 |
| S.L. No. | Name of the Drugs | Dose | Cure Rate |
|---|---|---|---|
| 1 | Oxytetracycline (OTC) 50% with vitamin C | OTC-10 g/kg feed for 7 days and VC 10 g/kg feed for 10 days | 40–45% |
| 2 | Sulfadiazine (SFD) | 10 g/kg feed for 7 days | 5–7% |
| 3 | Oxytetracycline dehydrate (OTCD) | 10 g/kg feed for 7 days | 10–15% |
| 4 | p-Mercapto-Sulfadiazine (p-M-SFD) | 10 g/kg feed for 7 days | 00% |
| 5 | Oxytetracycline 50%+ sulfadiazine | OTC-5 g and SFD-5 g/kg feed for 7 days | 30–35% |
| 6 | Sulfadiazine with vitamin | SFD-10 g/kg feed and VC 10 g/kg feed for 7 days | 3–5% |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kumer, A.; Chakma, U.; Rana, M.M.; Chandro, A.; Akash, S.; Elseehy, M.M.; Albogami, S.; El-Shehawi, A.M. Investigation of the New Inhibitors by Sulfadiazine and Modified Derivatives of α-D-glucopyranoside for White Spot Syndrome Virus Disease of Shrimp by In Silico: Quantum Calculations, Molecular Docking, ADMET and Molecular Dynamics Study. Molecules 2022, 27, 3694. https://doi.org/10.3390/molecules27123694
Kumer A, Chakma U, Rana MM, Chandro A, Akash S, Elseehy MM, Albogami S, El-Shehawi AM. Investigation of the New Inhibitors by Sulfadiazine and Modified Derivatives of α-D-glucopyranoside for White Spot Syndrome Virus Disease of Shrimp by In Silico: Quantum Calculations, Molecular Docking, ADMET and Molecular Dynamics Study. Molecules. 2022; 27(12):3694. https://doi.org/10.3390/molecules27123694
Chicago/Turabian StyleKumer, Ajoy, Unesco Chakma, Md Masud Rana, Akhel Chandro, Shopnil Akash, Mona M. Elseehy, Sarah Albogami, and Ahmed M. El-Shehawi. 2022. "Investigation of the New Inhibitors by Sulfadiazine and Modified Derivatives of α-D-glucopyranoside for White Spot Syndrome Virus Disease of Shrimp by In Silico: Quantum Calculations, Molecular Docking, ADMET and Molecular Dynamics Study" Molecules 27, no. 12: 3694. https://doi.org/10.3390/molecules27123694
APA StyleKumer, A., Chakma, U., Rana, M. M., Chandro, A., Akash, S., Elseehy, M. M., Albogami, S., & El-Shehawi, A. M. (2022). Investigation of the New Inhibitors by Sulfadiazine and Modified Derivatives of α-D-glucopyranoside for White Spot Syndrome Virus Disease of Shrimp by In Silico: Quantum Calculations, Molecular Docking, ADMET and Molecular Dynamics Study. Molecules, 27(12), 3694. https://doi.org/10.3390/molecules27123694







