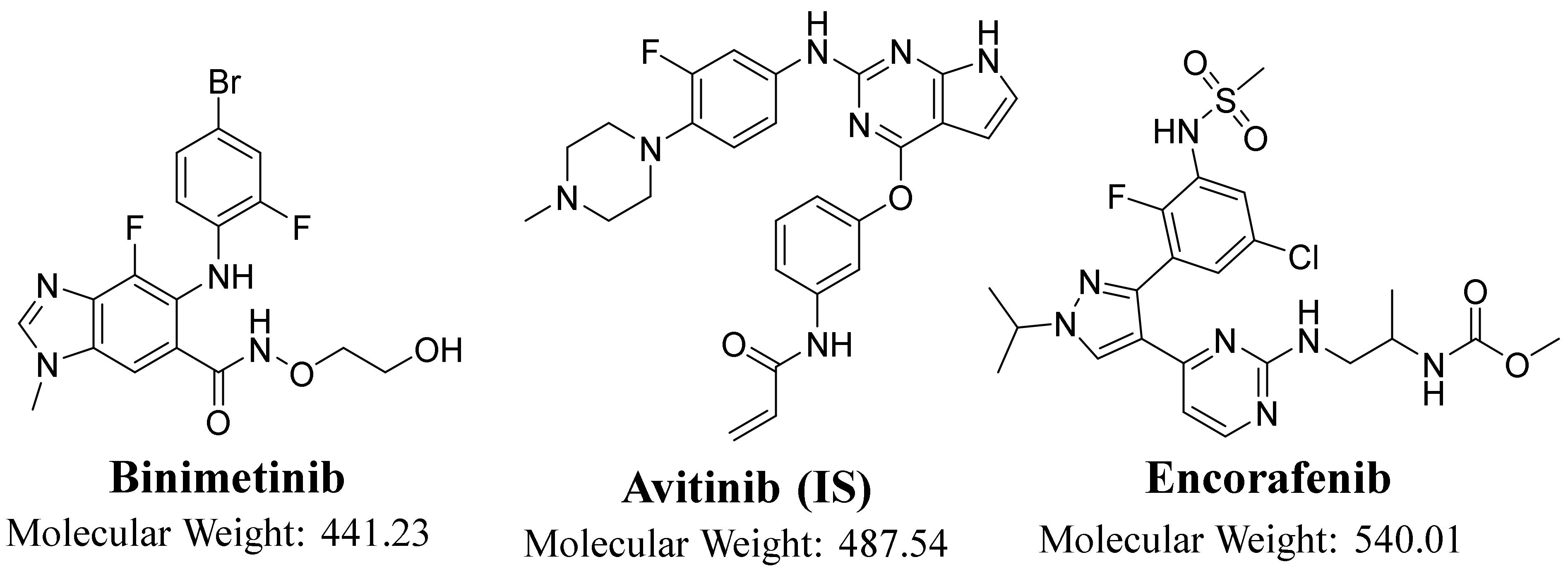
A Validated LC–MS/MS Assay for the Simultaneous Quantification of the FDA-Approved Anticancer Mixture (Encorafenib and Binimetinib): Metabolic Stability Estimation
Abstract
1. Introduction
2. Results and Discussion
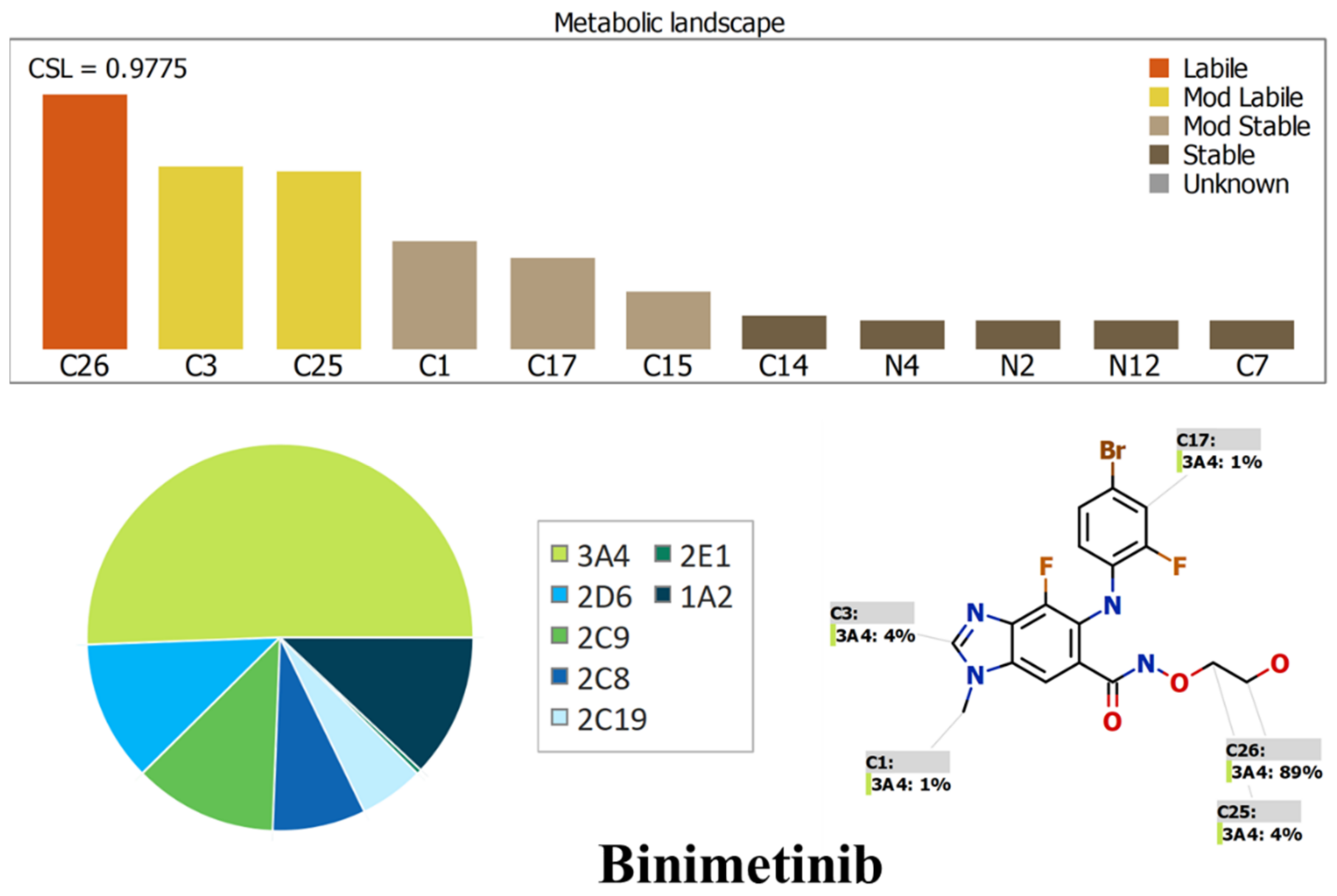
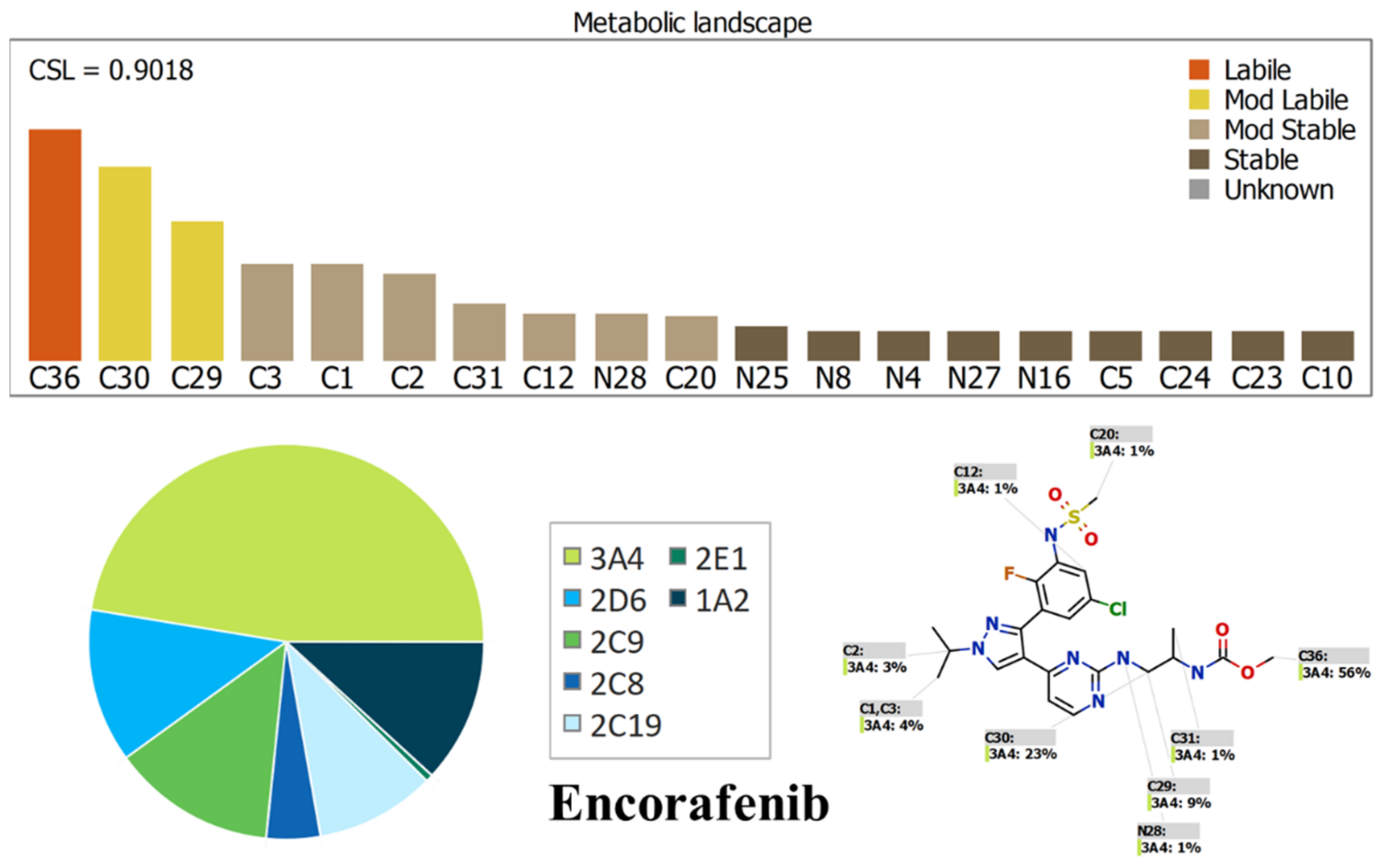
2.1. Results of In Silico ENF and BNB Metabolic Stability
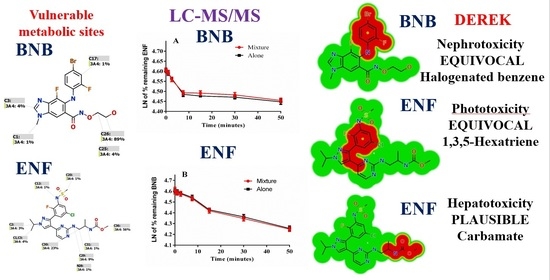
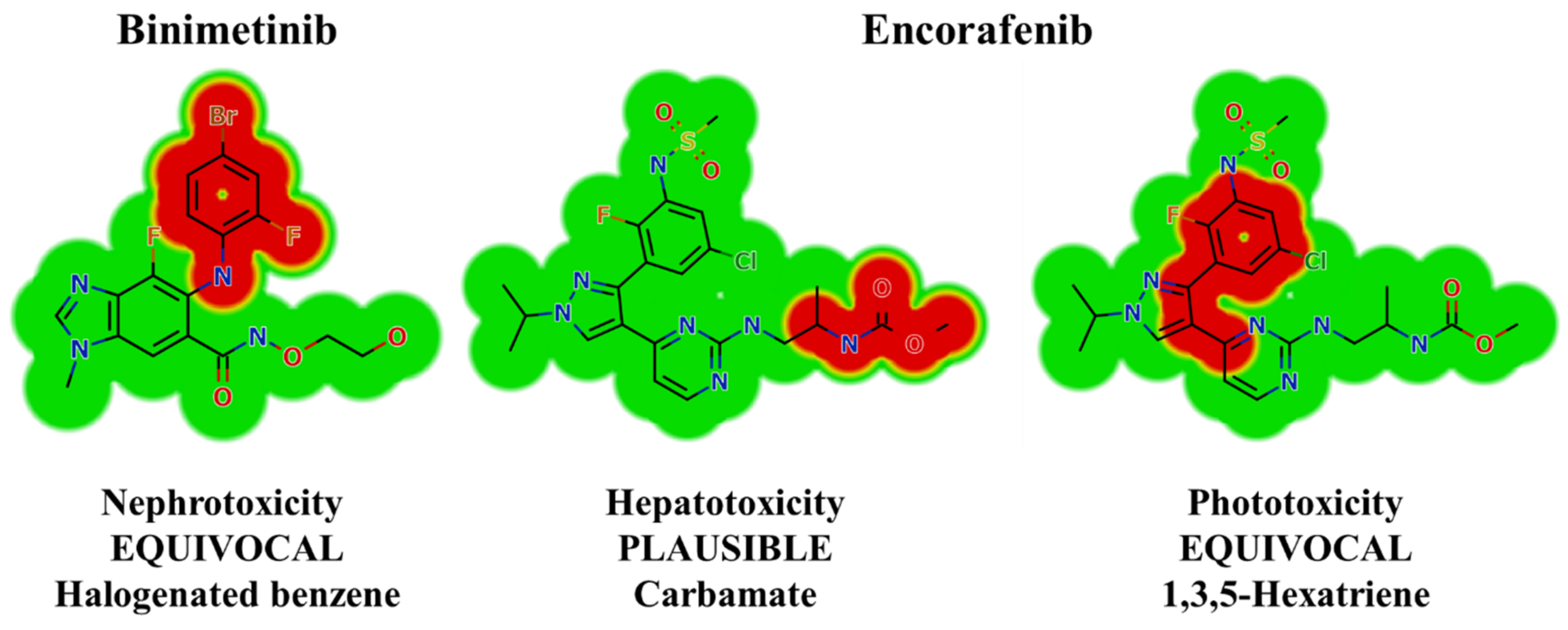
2.2. Results of In Silico BNB and ENF Structural Alert Sites and Toxicity Prediction
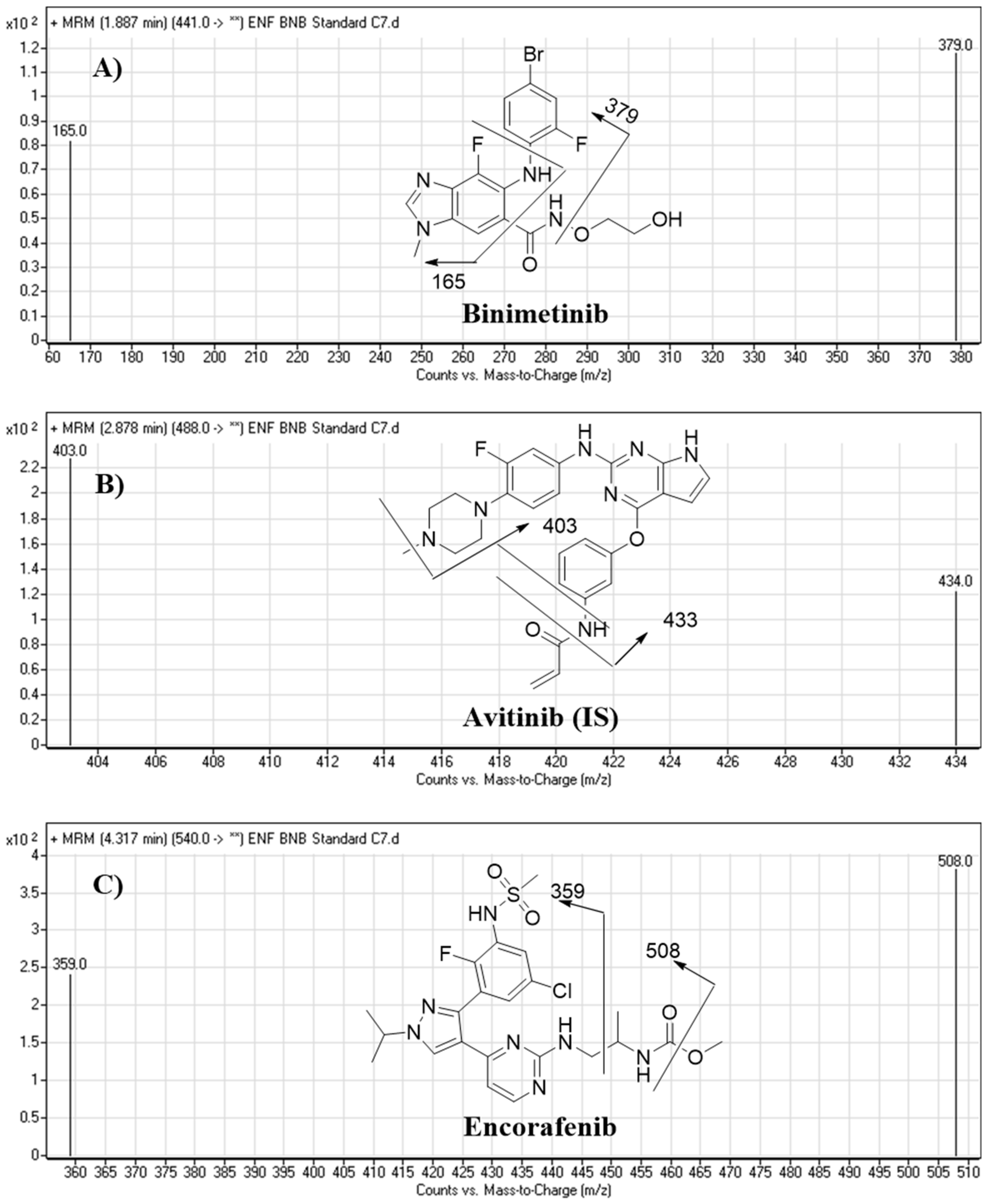
2.3. HPLC–MS/MS Methodology
2.4. LC–MS/MS Method Validation
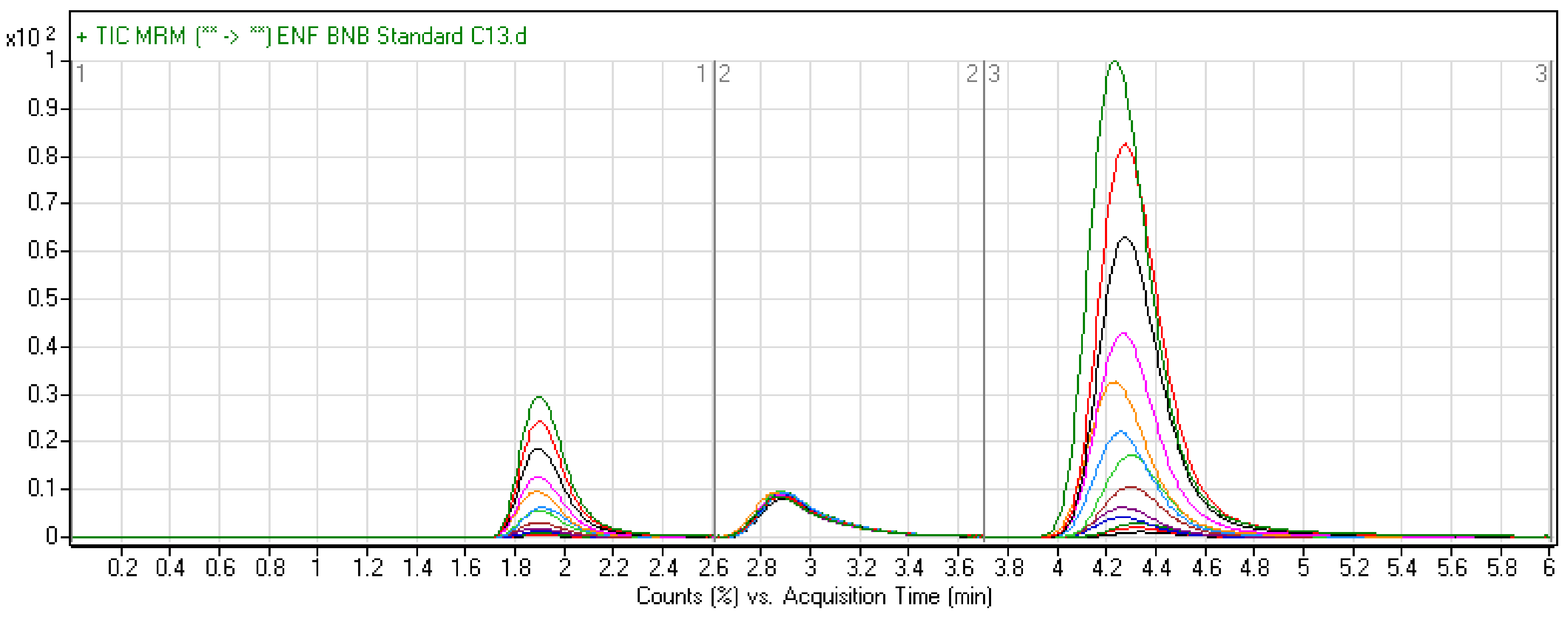
2.4.1. Specificity
2.4.2. Sensitivity and Linearity
2.4.3. Precision and Accuracy
2.4.4. Matrix Effects and Extraction Recovery
2.4.5. Stability
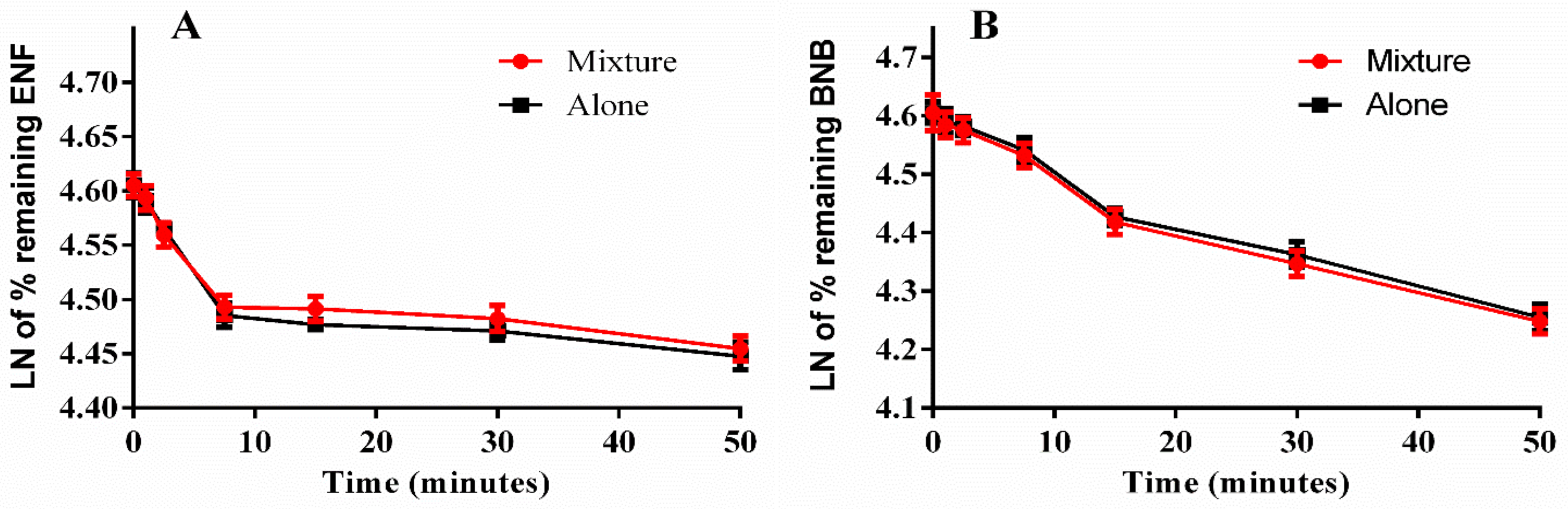
2.5. Metabolic Stability
3. Materials and Methods
3.1. Chemicals and Reagents
3.2. In Silico Prediction of ENF and BNB Metabolic Vulnerability and Toxicity Using StarDrop WhichP450 and DEREK Modules
3.3. LC–MS/MS Methodology
3.4. Standard Solutions of ENF and BNB
3.5. Preparation of Calibration Standards
3.6. BNB and ENF Extraction from HLM Matrix
3.7. Method Validation
3.7.1. Specificity, Linearity, and Sensitivity
3.7.2. Accuracy, Precision, and Stability
3.7.3. Matrix Effect and Extraction Recovery
3.8. Metabolic Stability Assessment of BNB and ENF
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Sample Availability
References
- Cummins, D.L.; Cummins, J.M.; Pantle, H.; Silverman, M.A.; Leonard, A.L.; Chanmugam, A. Cutaneous Malignant Melanoma. Mayo Clin. Proc. 2006, 81, 500–507. [Google Scholar] [CrossRef] [PubMed]
- Hall, C.S.; Ross, M.; Bauldry, J.B.B.; Upshaw, J.; Karhade, M.G.; Royal, R.; Patel, S.; Lucci, A. Circulating tumor cells in Stage IV melanoma patients. J. Am. Coll. Surg. 2018, 227, 116–124. [Google Scholar] [CrossRef] [PubMed]
- Raedler, L.A. Braftovi (Encorafenib) Plus Mektovi (Binimetinib) Third BRAF/MEK Inhibition Combination Approved for Metastatic Melanoma with BRAF Mutation. Available online: https://www.ahdbonline.com/select-drug-profiles/2756-braftovi-encorafenib-plus-mektovi-binimetinib-third-braf-mek-inhibition-combination-approved-for-metastatic-melanoma-with-braf-mutation (accessed on 15 June 2020).
- Meier, F.; Schittek, B.; Busch, S.; Garbe, C.; Smalley, K.; Satyamoorthy, K.; Li, G.; Herlyn, M. The RAS/RAF/MEK/ERK and PI3K/AKT signaling pathways present molecular targets for the effective treatment of advanced melanoma. Front. Biosci. 2005, 10, 2986–3001. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Zhou, F.; Dijke, P. Signaling interplay between transforming growth factor-β receptor and PI3K/AKT pathways in cancer. Trends Biochem. Sci. 2013, 38, 612–620. [Google Scholar] [CrossRef] [PubMed]
- Trojaniello, C.; Festino, L.; Vanella, V.; Ascierto, P.A. Encorafenib in combination with binimetinib for unresectable or metastatic melanoma with BRAF mutations. Expert Rev. Clin. Pharmacol. 2019, 12, 259–266. [Google Scholar] [CrossRef] [PubMed]
- Sun, J.; Zager, J.S.; Eroglu, Z. Encorafenib/binimetinib for the treatment of BRAF-mutant advanced, unresectable, or metastatic melanoma: Design, development, and potential place in therapy. OncoTargets Ther. 2018, 11, 9081–9089. [Google Scholar] [CrossRef] [PubMed]
- Shirley, M. Encorafenib and Binimetinib: First global approvals. Drugs 2018, 78, 1277–1284. [Google Scholar] [CrossRef] [PubMed]
- Turner, M.C.; Rossfeld, K.; Salama, A.K.; Tyler, D.; Beasley, G. Can binimetinib, encorafenib and masitinib be more efficacious than currently available mutation-based targeted therapies for melanoma treatment? Expert Opin. Pharmacother. 2017, 18, 487–495. [Google Scholar] [CrossRef]
- Dummer, R.; Ascierto, P.A.; Gogas, H.J.; Arance, A.; Mandala, M.; Liszkay, G.; Garbe, C.; Schadendorf, D.; Krajsova, I.; Gutzmer, R.; et al. Encorafenib plus binimetinib versus vemurafenib or encorafenib in patients with BRAF-mutant melanoma (COLUMBUS): A multicentre, open-label, randomised phase 3 trial. Lancet Oncol. 2018, 19, 603–615. [Google Scholar] [CrossRef]
- Di, L. The role of drug metabolizing enzymes in clearance. Expert Opin. Drug Metab. Toxicol. 2014, 10, 379–393. [Google Scholar] [CrossRef]
- Wienkers, L.C.; Heath, T.G. Predicting in vivo drug interactions from in vitro drug discovery data. Nat. Rev. Drug Discov. 2005, 4, 825–833. [Google Scholar] [CrossRef]
- Pelkonen, O.; Turpeinen, M.; Uusitalo, J.; Rautio, A.; Raunio, H. Prediction of drug metabolism and interactions on the basis of in vitro investigations. Basic Clin. Pharmacol. Toxicol. 2005, 96, 167–175. [Google Scholar] [CrossRef]
- Coecke, S.; Ahr, H.; Blaauboer, B.J.; Bremer, S.; Casati, S.; Castell, J.; Combes, R.; Corvi, R.; Crespi, C.L.; Cunningham, M.L. Metabolism: A bottleneck in in vitro toxicological test development: The report and recommendations of ECVAM workshop 54. Altern. Lab. Anim. 2006, 34, 49–84. [Google Scholar] [CrossRef]
- Eddershaw, P.J.; Beresford, A.P.; Bayliss, M.K. ADME/PK as part of a rational approach to drug discovery. Drug Discov. Today 2000, 5, 409–414. [Google Scholar] [CrossRef]
- Kola, I.; Landis, J. Can the pharmaceutical industry reduce attrition rates? Nat. Rev. Drug Discov. 2004, 3, 711–716. [Google Scholar] [CrossRef]
- Khanna, I. Drug discovery in pharmaceutical industry: Productivity challenges and trends. Drug Discov. Today 2012, 17, 1088–1102. [Google Scholar] [CrossRef]
- Ulenberg, S.; Bączek, T. Metabolic stability studies of lead compounds supported by separation techniques and chemometrics analysis. J. Sep. Sci. 2021, 44, 373–386. [Google Scholar] [CrossRef]
- Attwa, M.W.; Kadi, A.A.; Abdelhameed, A.S. Phase I metabolic profiling and unexpected reactive metabolites in human liver microsome incubations of X-376 using LC-MS/MS: Bioactivation pathway elucidation and in silico toxicity studies of its metabolites. RSC Adv. 2020, 10, 5412–5427. [Google Scholar] [CrossRef]
- Attwa, M.W.; Kadi, A.A.; Abdelhameed, A.S. Detection and characterization of olmutinib reactive metabolites by LC–MS/MS: Elucidation of bioactivation pathways. J. Sep. Sci. 2020, 43, 708–718. [Google Scholar] [CrossRef]
- Meirson, T.; Asher, N.; Bomze, D.; Markel, G. Safety of BRAF + MEK Inhibitor Combinations: Severe Adverse Event Evaluation. Cancers 2020, 12, 1650. [Google Scholar] [CrossRef]
- Matsudate, Y. Case of inflammatory arthritis induced by encorafenib and binimetinib in a patient with melanoma. J. Dermatol. 2020, 47. [Google Scholar] [CrossRef]
- Lièvre, A.; de la Fouchardière, C.; Samalin, E.; Benoist, S.; Phelip, J.M.; André, T.; Lledo, G. BRAF V600E-mutant colorectal cancers: Where are we? Bull. Cancer 2020, 107, 881–895. [Google Scholar] [CrossRef]
- Sullivan, R.J.; Weber, J.S.; Patel, S.P.; Dummer, R.; Carlino, M.S.; Tan, D.S.; Lebbe, C.; Siena, S.; Élez, E.; Wollenberg, L.; et al. A phase 1b/2 study of the BRAF inhibitor encorafenib plus the MEK inhibitor binimetinib in patients with BRAF V600E/K-mutant solid tumors. Clin. Cancer Res. 2020, 26, 5102–5112. [Google Scholar] [CrossRef]
- Baranczewski, P.; Stańczak, A.; Sundberg, K.; Svensson, R.; Wallin, A.; Jansson, J.; Garberg, P.; Postlind, H. Introduction to in vitro estimation of metabolic stability and drug interactions of new chemical entities in drug discovery and development. Pharmacol. Rep. 2006, 58, 453–472. [Google Scholar]
- Rousset, M.; Titier, K.; Bouchet, S.; Dutriaux, C.; Pham-Ledard, A.; Prey, S.; Canal-Raffin, M.; Molimard, M. An UPLC-MS/MS method for the quantification of BRAF inhibitors (vemurafenib, dabrafenib) and MEK inhibitors (cobimetinib, trametinib, binimetinib) in human plasma. Application to treated melanoma patients. Clin. Chim. Acta 2017, 470, 8–13. [Google Scholar] [CrossRef]
- Sparidans, R.W.; Rosing, H.; Rood, J.J.M.; Schellens, J.H.M.; Beijnen, J.H. Liquid chromatography-tandem mass spectrometric assay for therapeutic drug monitoring of the B-Raf inhibitor encorafenib, the EGFR inhibitors afatinib, erlotinib and gefitinib and the O-desmethyl metabolites of erlotinib and gefitinib in human plasma. J. Chromatogr. B Anal. Technol. Biomed. Life Sci. 2016, 1033–1034, 390–398. [Google Scholar] [CrossRef]
- Kadav, A.A.; Vora, D.N. Stability indicating UPLC method for simultaneous determination of atorvastatin, fenofibrate and their degradation products in tablets. J. Pharm. Biomed. Anal. 2008, 48, 120–126. [Google Scholar] [CrossRef]
- Al Bratty, M. Stability-indicating UPLC Method for simultaneous analysis of protein kinase inhibitors, binimetinib, and encorafenib in pure active pharmaceutical ingredient (API) and formulation. Lat. Am. J. Pharm. 2020, 39, 1428–1436. [Google Scholar]
- Huynh, H.H.; Pressiat, C.; Sauvageon, H.; Madelaine, I.; Maslanka, P.; Lebbé, C.; Thieblemont, C.; Goldwirt, L.; Mourah, S. Development and Validation of a Simultaneous Quantification Method of 14 Tyrosine Kinase Inhibitors in Human Plasma Using LC-MS/MS. Ther. Drug Monit. 2017, 39, 43–54. [Google Scholar] [CrossRef]
- Jiang, H.; Zeng, J.; Titsch, C.; Voronin, K.; Akinsanya, B.; Luo, L.; Shen, H.; Desai, D.D.; Allentoff, A.; Aubry, A.F.; et al. Fully validated LC-MS/MS assay for the simultaneous quantitation of coadministered therapeutic antibodies in cynomolgus monkey serum. Anal. Chem. 2013, 85, 9859–9867. [Google Scholar] [CrossRef]
- Van Erp, N.P.; de Wit, D.; Guchelaar, H.J.; Gelderblom, H.; Hessing, T.J.; Hartigh, J. A validated assay for the simultaneous quantification of six tyrosine kinase inhibitors and two active metabolites in human serum using liquid chromatography coupled with tandem mass spectrometry. J. Chromatogr. B Anal. Technol. Biomed. Life Sci. 2013, 937, 33–43. [Google Scholar] [CrossRef] [PubMed]
- Attwa, M.W.; Kadi, A.A.; Alrabiah, H.; Darwish, H.W. LC-MS/MS reveals the formation of iminium and quinone methide reactive intermediates in entrectinib metabolism: In vivo and in vitro metabolic investigation. J. Pharm. Biomed. Anal. 2018, 160, 19–30. [Google Scholar] [CrossRef] [PubMed]
- Rowland, M.; Benet, L.Z.; Graham, G.G. Clearance concepts in pharmacokinetics. J. Pharm. Biopharm. 1973, 1, 123–136. [Google Scholar] [CrossRef] [PubMed]
- Wilkinson, G.R.; Shand, D.G. Commentary: A physiological approach to hepatic drug clearance. Clin. Pharm. 1975, 18, 377–390. [Google Scholar]
- Houston, J.B. Utility of in vitro drug metabolism data in predicting in vivo metabolic clearance. Biochem. Pharm. 1994, 47, 1469–1479. [Google Scholar] [CrossRef]
- Obach, R.S.; Baxter, J.G.; Liston, T.E.; Silber, B.M.; Jones, B.C.; MacIntyre, F.; Rance, D.J.; Wastall, P. The prediction of human pharmacokinetic parameters from preclinical and in vitro metabolism data. J. Pharm. Exp. 1997, 283, 46–58. [Google Scholar]
- Inoue, K.; Yoshimi, Y.; Hino, T.; Oka, H. Simultaneous determination of avermectins in bovine tissues by LC-MS/MS. J. Sep. Sci. 2009, 32, 3596–3602. [Google Scholar] [CrossRef]
- Surowiec, I.; Koc, M.; Antti, H.; Wikström, P.; Moritz, T. LC-MS/MS profiling for detection of endogenous steroids and prostaglandins in tissue samples. J. Sep. Sci. 2011, 34, 2650–2658. [Google Scholar] [CrossRef]
- Shah, H.J.; Kundlik, M.L.; Patel, N.K.; Subbaiah, G.; Patel, D.M.; Suhagia, B.N.; Patel, C.N. Rapid determination of losartan and losartan acid in human plasma by multiplexed LC–MS/MS. J. Sep. Sci. 2009, 32, 3388–3394. [Google Scholar] [CrossRef]
- Attwa, M.W.; Darwish, H.W.; Alhazmi, H.A.; Kadi, A.A. Investigation of metabolic degradation of new ALK inhibitor: Entrectinib by LC-MS/MS. Clin. Chim. Acta 2018, 485, 298–304. [Google Scholar]
- Attwa, M.W.; Kadi, A.A.; Darwish, H.W.; Amer, S.M.; AlRabiah, H. LC-ESI-MS/MS identification and characterization of ponatinib in vivo phase I and phase II metabolites. Clin. Chim. Acta 2018, 485, 144–151. [Google Scholar] [CrossRef]
- Attwa, M.W.; Kadi, A.A.; Abdelhameed, A.S. Reactive intermediates and bioactivation pathways characterization of avitinib by LC–MS/MS: In vitro metabolic investigation. J. Pharm. Biomed. Anal. 2019, 164, 659–667. [Google Scholar] [CrossRef]
- Attwa, M.W.; Kadi, A.A.; Abdelhameed, A.S. Characterization of reactive intermediates formation in dacomitinib metabolism and bioactivation pathways elucidation by LC-MS/MS: In vitro phase I metabolic investigation. RSC Adv. 2018, 8, 38733–38744. [Google Scholar] [CrossRef]
- Bioanalytical Method Validation Guidance for Industry. Available online: https://www.fda.gov/regulatory-information/search-fda-guidance-documents/bioanalytical-method-validation-guidance-industry (accessed on 12 June 2020).
- Attwa, M.W.; Kadi, A.A.; Darwish, H.W.; Amer, S.M.; Alrabiah, H. A reliable and stable method for the determination of foretinib in human plasma by LC-MS/MS: Application to metabolic stability investigation and excretion rate. Eur. J. Mass Spectrom. 2018, 24, 344–351. [Google Scholar] [CrossRef]
- Darwish, H.W.; Kadi, A.A.; Attwa, M.W.; Almutairi, H.S. Investigation of metabolic stability of the novel ALK inhibitor brigatinib by liquid chromatography tandem mass spectrometry. Clin. Chim. Acta 2018, 480, 180–185. [Google Scholar] [CrossRef]
- Smith, G. European Medicines Agency guideline on bioanalytical method validation: What more is there to say? Bioanalysis 2012, 4, 865–868. [Google Scholar] [CrossRef]
- Manzo, A.; Montanino, A.; Costanzo, R.; Sandomenico, C.; Palumbo, G.; Schettino, C.; Daniele, G.; Morabito, A.; Perrone, F.; Piccirillo, M.C. Chapter 33—EGFR Mutations: Best Results from Second- and Third-Generation Tyrosine Kinase Inhibitors. In Oncogenomics; Dammacco, F., Silvestris, F., Eds.; Academic Press: Cambridge, MA, USA, 2019; pp. 477–486. [Google Scholar]
- Amer, S.M.; Kadi, A.A.; Darwish, H.W.; Attwa, M.W. Liquid chromatography tandem mass spectrometry method for the quantification of vandetanib in human plasma and rat liver microsomes matrices: Metabolic stability investigation. Chem. Cent. J. 2017, 11, 45. [Google Scholar] [CrossRef]
- Caldwell, G.; Yan, Z. Optimization in Drug Discovery: In Vitro Methods; Springer Science & Business Media: Berlin/Heidelberg, Germany, 2014. [Google Scholar]
- Leahy, D.E. Integrating in vitro ADMET data through generic physiologically based pharmacokinetic models. Expert Opin. Drug Metab. Toxicol. 2006, 2, 619–628. [Google Scholar] [CrossRef]
| Concentration in ng/mL | BNB | ENF | ||||||
|---|---|---|---|---|---|---|---|---|
| Mean a | SD | RSD % | Accuracy % | Mean a | SD | RSD % | Accuracy % | |
| 5 | 5.72 | 0.13 | 2.27 | 114.35 | 5.70 | 0.04 | 0.63 | 113.94 |
| 10 | 10.09 | 0.36 | 3.58 | 100.93 | 10.56 | 0.06 | 0.57 | 105.57 |
| 15 (LQC) | 14.98 | 0.24 | 1.60 | 99.84 | 15.24 | 0.14 | 0.94 | 101.63 |
| 20 | 19.46 | 0.36 | 1.86 | 97.29 | 20.17 | 0.35 | 1.75 | 100.86 |
| 30 | 29.62 | 0.32 | 1.08 | 98.75 | 30.80 | 0.18 | 0.57 | 102.66 |
| 50 | 49.48 | 1.41 | 2.85 | 98.96 | 49.87 | 0.76 | 1.52 | 99.75 |
| 80 | 82.15 | 2.40 | 2.92 | 102.69 | 79.06 | 1.30 | 1.65 | 98.83 |
| 100 | 98.21 | 1.36 | 1.39 | 98.21 | 97.41 | 0.77 | 0.79 | 97.41 |
| 150 (MQC) | 148.71 | 1.51 | 1.02 | 99.14 | 149.30 | 0.83 | 0.55 | 99.53 |
| 200 | 200.18 | 1.42 | 0.71 | 100.09 | 199.68 | 1.47 | 0.74 | 99.84 |
| 300 | 303.91 | 5.47 | 1.80 | 101.30 | 302.84 | 1.00 | 0.33 | 100.95 |
| 400 (HQC) | 396.97 | 2.22 | 0.56 | 99.24 | 400.59 | 3.46 | 0.86 | 100.15 |
| 500 | 500.44 | 2.89 | 0.58 | 100.09 | 498.75 | 2.08 | 0.42 | 99.75 |
| % Recovery | 100.84 | 101.61 | ||||||
| SD | 4.29 | 4.20 | ||||||
| HLM Matrix | Mean | SD | % RSD | % Accuracy | |||
|---|---|---|---|---|---|---|---|
| BNB | Conc. in ng/mL | 15.00 (LQ) | Intra-day assay * | 14.98 | 0.20 | 1.31 | 99.8 |
| Inter-day assay ** | 15.02 | 0.37 | 2.48 | 104 | |||
| 150.00 (MQ) | Intra-day assay | 148.71 | 1.24 | 0.83 | 99.1 | ||
| Inter-day assay | 148.68 | 1.37 | 0.92 | 100 | |||
| 400.00 (HQ) | Intra-day assay | 396.97 | 1.81 | 0.46 | 99.2 | ||
| Inter-day assay | 397.78 | 3.52 | 0.88 | 101 | |||
| ENF | 15.00 (LQ) | Intra-day assay | 15.24 | 0.12 | 0.77 | 101 | |
| Inter-day assay | 15.16 | 0.39 | 2.60 | 96.1 | |||
| 150.00 (MQ) | Intra-day assay * | 149.30 | 0.67 | 0.45 | 99.5 | ||
| Inter-day assay | 148.20 | 1.88 | 1.27 | 96.7 | |||
| 400.00 (HQ) | Intra-day assay | 400.59 | 2.83 | 0.71 | 100 | ||
| Inter-day assay | 399.80 | 2.92 | 0.73 | 100 | |||
| Concentration (ng/mL) | HLM Matrix | |||||
|---|---|---|---|---|---|---|
| BNB | ENF | |||||
| 15 | 150 | 400 | 15 | 150 | 400 | |
| Mean a | 15.09 | 148.56 | 398.35 | 15.15 | 148.40 | 400.35 |
| Recovery (%) | 100 | 99.1 | 99.6 | 101 | 98.9 | 100 |
| SD | 0.38 | 1.30 | 3.55 | 0.36 | 1.79 | 3.03 |
| Precision (RSD %) | 2.51 | 0.87 | 0.89 | 2.38 | 1.21 | 0.76 |
| Nominal Concentration (ng/mL) | Mean (ng/mL) | Recovery % | Precision (RSD %) | |
|---|---|---|---|---|
| BNB | Room temperature for 8 h | 14.89 ± 0.2 | 99.25 | 3.07 |
| 15 | ||||
| 150 | 147.38 ± 2.74 | 98.25 | 1.70 | |
| 400 | 395.31 ± 4.14 | 98.85 | 1.23 | |
| Three freeze–thaw cycles | 14.75 ± 0.27 | 98.32 | 0.46 | |
| 15 | ||||
| 150 | 145.18 ± 2.04 | 96.79 | 2.52 | |
| 400 | 389.91 ± 3.19 | 97.48 | 4.87 | |
| Stored at 4 °C for 24 h | 15.01 ± 0.42 | 100.05 | 2.80 | |
| 15 | ||||
| 150 | 146.38 ± 3.99 | 97.59 | 2.73 | |
| 400 | 397.31 ± 4.89 | 99.33 | 1.23 | |
| Stored at −20 °C for 30 days | ||||
| 15 | 14.51 ± 0.67 | 96.72 | 4.59 | |
| 150 | 146.58 ± 2.39 | 97.72 | 1.63 | |
| 400 | 395.11 ± 4.57 | 98.78 | 1.16 | |
| ENF | Room temperature for 8 h | |||
| 15 | 15.25 ± 0.3 | 101.64 | 2.00 | |
| 150 | 150.83 ± 4.17 | 100.55 | 2.76 | |
| 400 | 405.84 ± 4.45 | 101.46 | 1.10 | |
| Three freeze–thaw cycles | ||||
| 15 | 14.61 ± 0.26 | 97.37 | 1.80 | |
| 150 | 145.83 ± 3.93 | 97.22 | 2.70 | |
| 400 | 398.64 ± 3.95 | 99.66 | 0.99 | |
| Stored at 4 °C for 24 h | ||||
| 15 | 14.51 ± 0.35 | 97.37 | 2.37 | |
| 150 | 143.43 ± 2.85 | 96.95 | 1.96 | |
| 400 | 392.84 ± 3.16 | 98.21 | 0.81 | |
| Stored at −20 °C for 30 days | ||||
| 15 | 15.03 ± 0.28 | 100.18 | 1.84 | |
| 150 | 144.83 ± 3.90 | 96.55 | 2.69 | |
| 400 | 393.64 ± 3.05 | 98.41 | 0.77 | |
| Parameters | ENF | BNB | ||
|---|---|---|---|---|
| ENF Alone | ENF with BNB | BNB Alone | BNB with ENF | |
| Regression equation a | y = −0.0161x + 4.6059 | y = −0.0151x + 4.6042 | y = −0.0115x + 4.6096 | y = −0.0119x + 4.605 |
| Slope | 0.0161 | 0.0151 | 0.0115 | 0.0119 |
| t1/2 b | 43.1 min | 45.9 min | 60.3 min | 58.2 min |
| CLint c | 16.09 µL/min/mg | 15.09 µL/min/mg | 11.49 µL/min/mg | 11.89 µL/min/mg |
| R2 d | 0.9972 | 0.9906 | 0.9757 | 0.9817 |
| Agilent 1200 HPLC | Triple Quadrupole 6410 QqQ | ||
|---|---|---|---|
| Isocratic mobile phase | ACN (38%) | ESI source | Positive mode |
| 10 mM ammonium formate in H2O (62%) adjusted with formic acid to pH 3.8 | Drying gas: N2 gas Pressure (60 psi) Flow rate (12 L/min) | ||
| Injection volume: 2 μL | |||
| Flow rate: 0.2 mL/min | |||
| Agilent Hypersil BDS-C18 | Length 125 mm, fully porous particle size (3.0 μm) and internal diameter (2.0 mm) | Source temperature: 350 °C | |
| Capillary voltage: 4000 V | |||
| Mode | MRM mode | ||
| Collision cell gas | Nitrogen with high purity | ||
| Analytes | Binimetinib (BNB) | BNB MRM transitions | m/z 441 → m/z 379, FVa: 140 V, CEb: of 22 eV |
| m/z 441 → m/z 165, FV: 140 V, CE: of 20 eV | |||
| Encorafenib (ENF) | ENF MRM Transitions | m/z 540 → m/z 508, FV: 135 V, CE: 18 eV | |
| m/z 540 → m/z 359, FV: 140 V, CE: 20 eV | |||
| IS | Avitinib (AVB) | AVB MRM transitions | m/z 488 → m/z 433, FV: 145 V, CE: of 15 eV |
| m/z 488 → m/z 403, FV: 145 V, CE: of 16 eV | |||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Attwa, M.W.; Darwish, H.W.; Al-Shakliah, N.S.; Kadi, A.A. A Validated LC–MS/MS Assay for the Simultaneous Quantification of the FDA-Approved Anticancer Mixture (Encorafenib and Binimetinib): Metabolic Stability Estimation. Molecules 2021, 26, 2717. https://doi.org/10.3390/molecules26092717
Attwa MW, Darwish HW, Al-Shakliah NS, Kadi AA. A Validated LC–MS/MS Assay for the Simultaneous Quantification of the FDA-Approved Anticancer Mixture (Encorafenib and Binimetinib): Metabolic Stability Estimation. Molecules. 2021; 26(9):2717. https://doi.org/10.3390/molecules26092717
Chicago/Turabian StyleAttwa, Mohamed W., Hany W. Darwish, Nasser S. Al-Shakliah, and Adnan A. Kadi. 2021. "A Validated LC–MS/MS Assay for the Simultaneous Quantification of the FDA-Approved Anticancer Mixture (Encorafenib and Binimetinib): Metabolic Stability Estimation" Molecules 26, no. 9: 2717. https://doi.org/10.3390/molecules26092717
APA StyleAttwa, M. W., Darwish, H. W., Al-Shakliah, N. S., & Kadi, A. A. (2021). A Validated LC–MS/MS Assay for the Simultaneous Quantification of the FDA-Approved Anticancer Mixture (Encorafenib and Binimetinib): Metabolic Stability Estimation. Molecules, 26(9), 2717. https://doi.org/10.3390/molecules26092717