EGCG, a Green Tea Catechin, as a Potential Therapeutic Agent for Symptomatic and Asymptomatic SARS-CoV-2 Infection
Abstract
1. Introduction
2. Results
2.1. Inflammation Associated with COVID-19 Pathobiology and Possible Targeting by Catechins
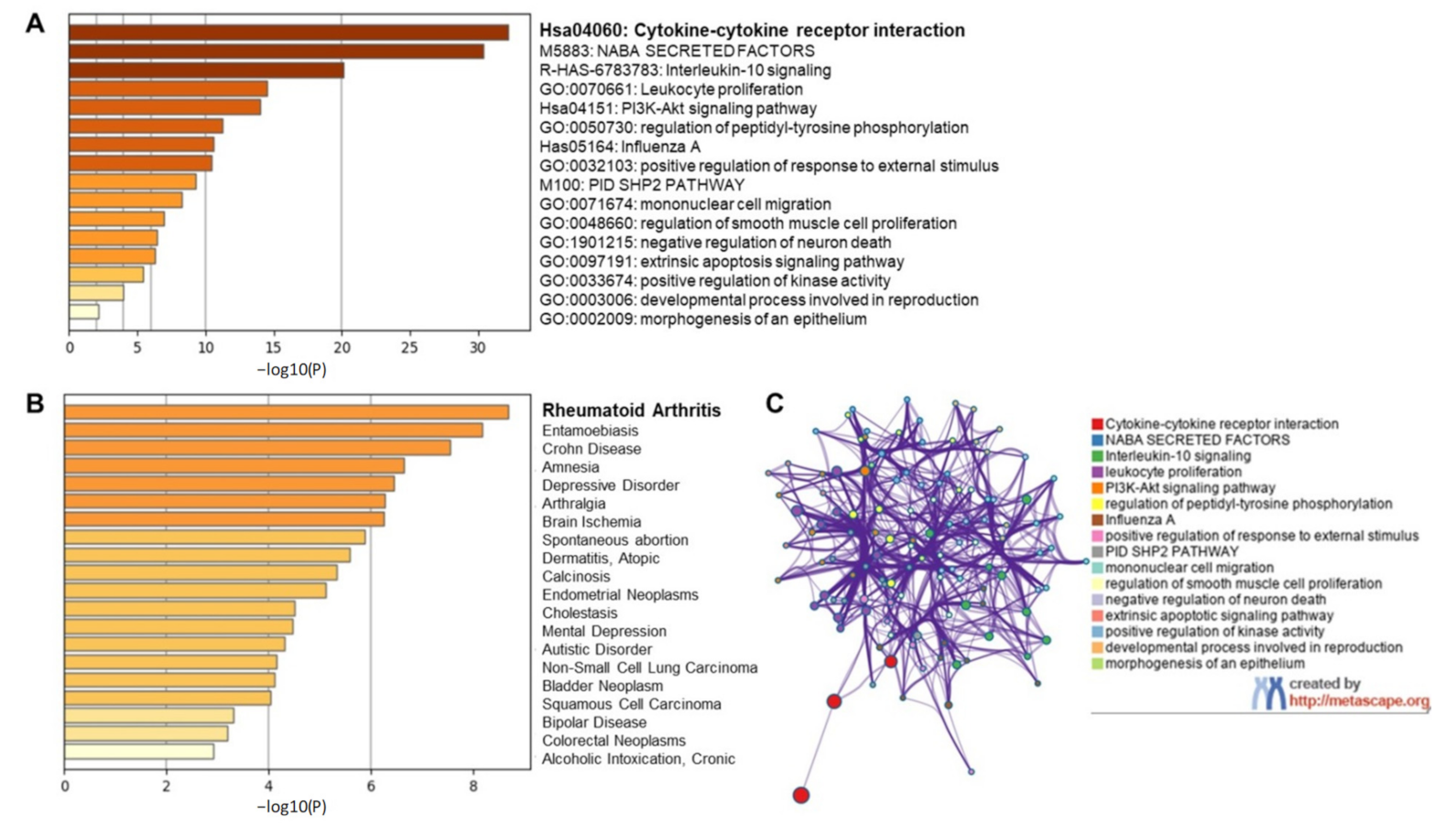
2.1.1. Inflammatory Response in COVID-19 Infection
2.1.2. Targeting Individual Cytokines Might Not Be a Viable Option for COVID-19 Amelioration
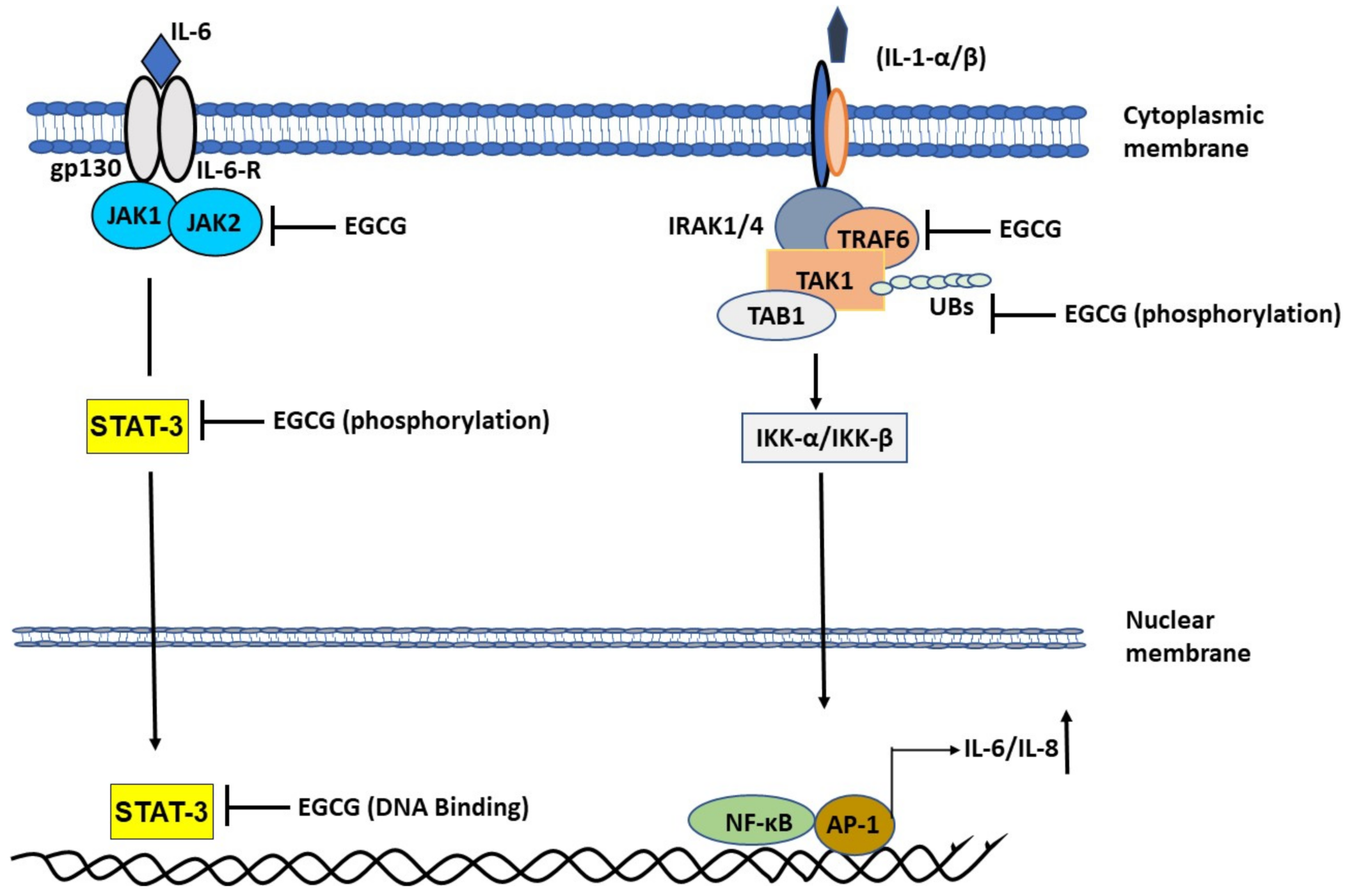
2.1.3. EGCG Ameliorates Interleukin-1 (IL-1)-Induced IL-6 Expression
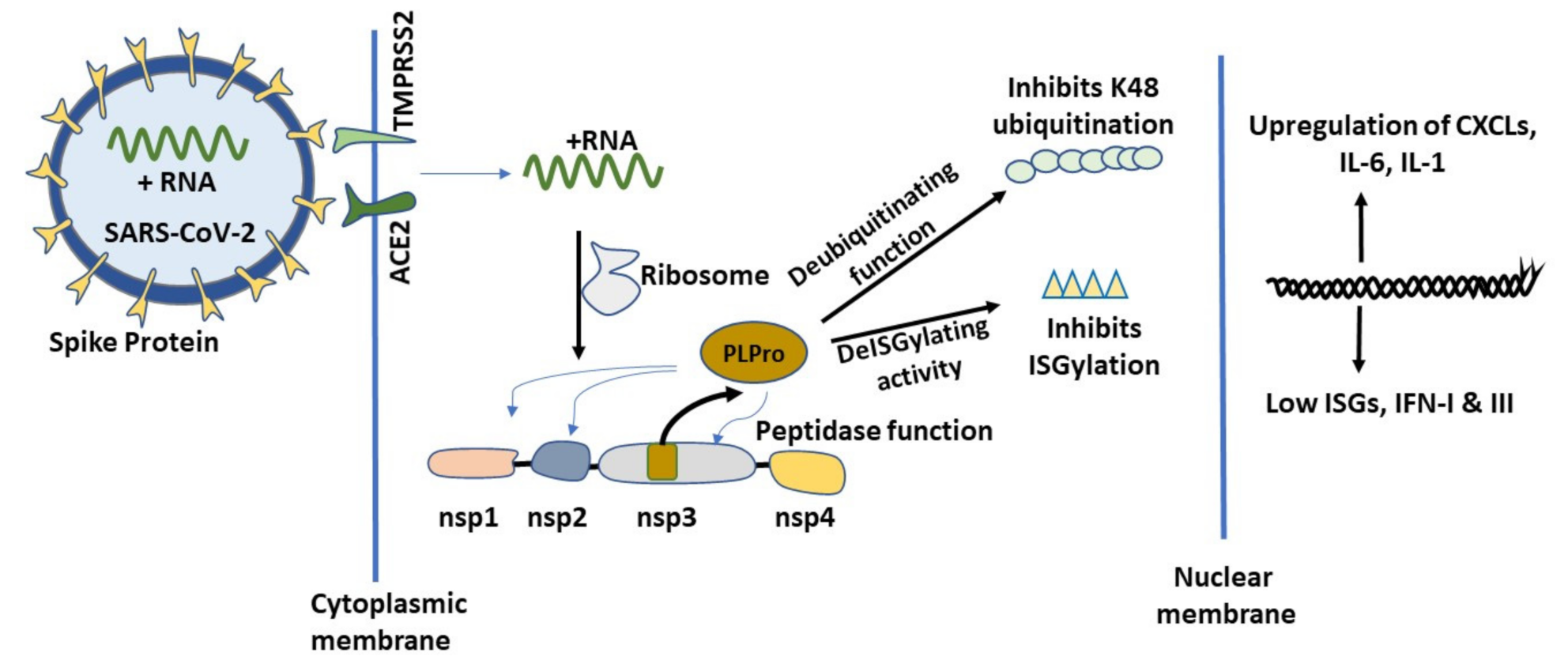
2.2. Initial Cellular Events of COVID-19 and the Role of PLPro
2.2.1. Chronic Inflammatory Response and Cytokine Storm in COVID-19
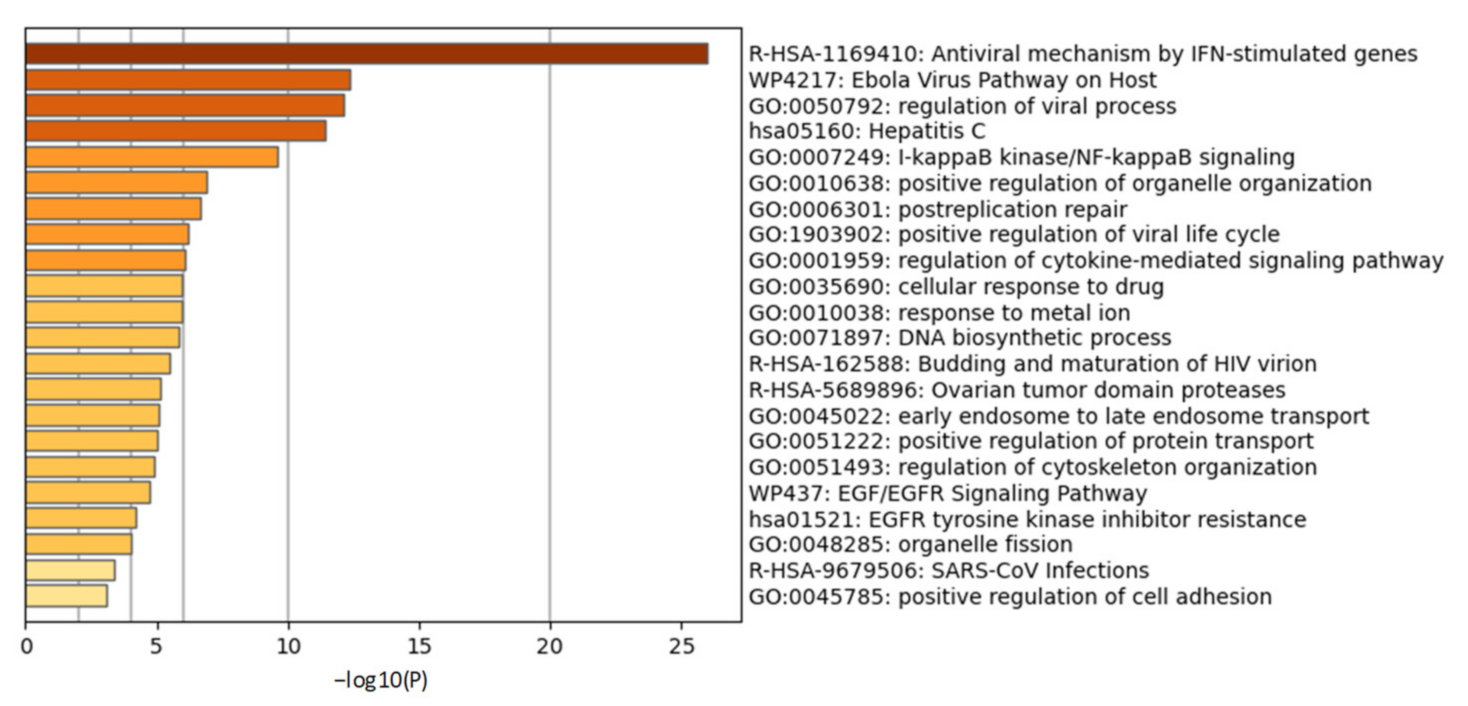
2.2.2. Cellular Ubiquitination and ISGylation Processes in Infection
2.2.3. PLPro Targets Cellular Ubiquitination and ISGylation Processes
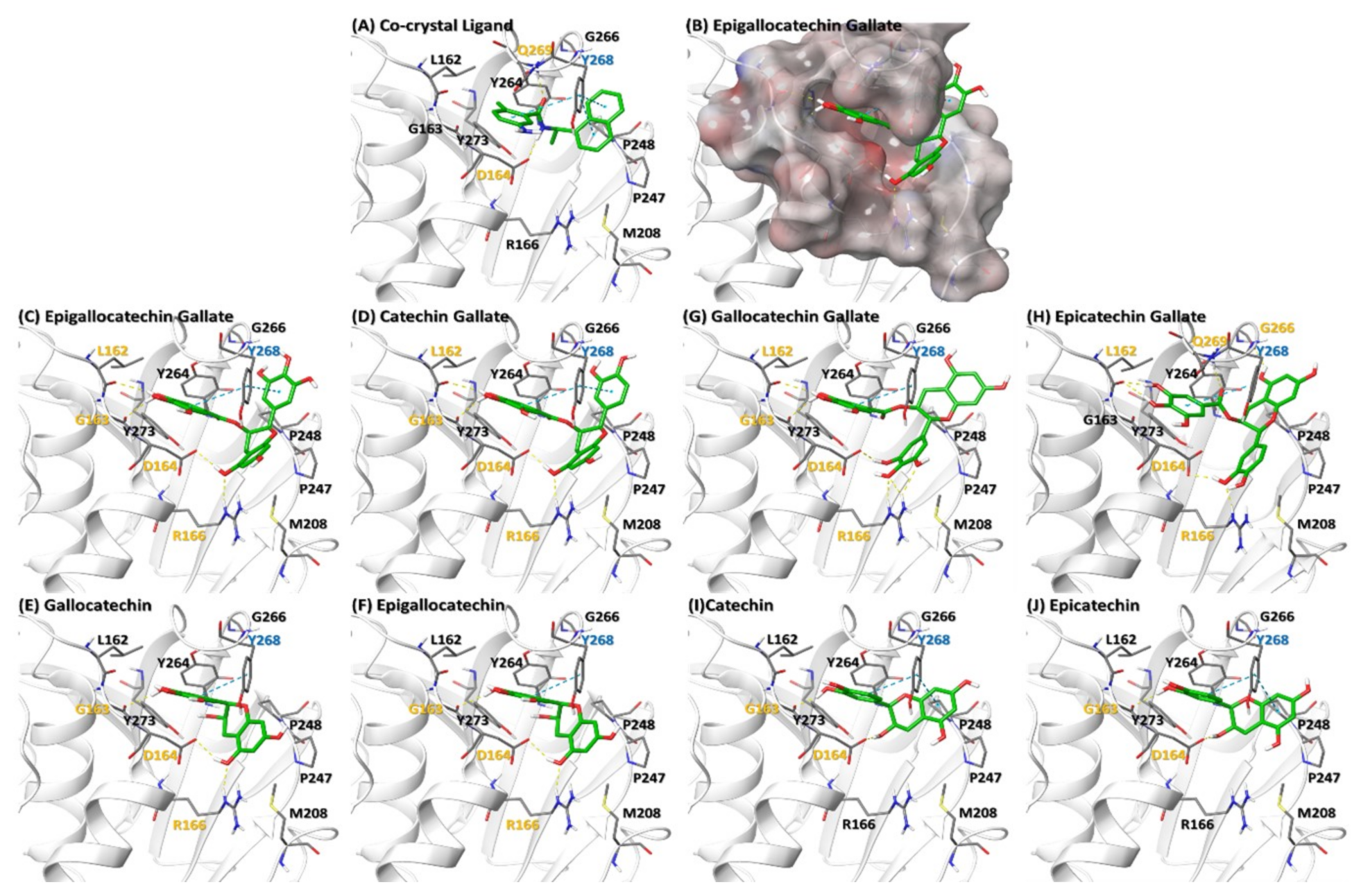
2.2.4. Catechins May Counteract Inhibition of ISGylation/Ubiquitination by PLPro
3. Methods
3.1. Molecular Docking of Catechins on PLPro
3.2. Gene Ontology Studies
3.3. Keywords Searched
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
Disclaimer
Abbreviations
| COVID-19 | Coronavirus disease 2019 |
| SARS-CoV-2 | severe acute respiratory syndrome coronavirus 2 |
| EGCG | epigallocatechin gallate |
| PLPro | Papain-like protease |
| S protein | SARS-CoV-2 spike glycoprotein |
| EC | (−)-epicatechin |
| EGC | (−)-epigallocatechin |
| Mpro | Main proteinase |
| TRAIL | TNF-related apoptosis-inducing ligand |
| TNF | Tumor necrosis factor |
| M-CSF | macrophage colony-stimulating factor |
| GRO-α | Grown regulated oncogene-α |
| G-CSF | Granulocyte colony stimulating factor |
| IL | Interleukin |
| β-NGF | β-Nerve growth factor |
| MCP-1 | Monocyte chemoattractant protein-1 |
| SCF | Skp1-cullin 1-F-box |
| IP-10 | Interferon gamma-induced protein 10 (also known as CXCL10) |
| PDGF-BB | Platelet derived growth factor-BB |
| IFN | Interferon |
| PI3K | Phosphoinositide 3-kinase |
| Akt | Protein kinase B |
| TNF | Tumor necrosis factor |
| CRP | C-Reactive protein |
| TIMP1 | Tissue inhibitor of metalloproteinase |
| CKM | Creatine Kinase M |
| JAK | Janus kinase |
| STAT | Signal transducer and activator of transcription protein |
| RDS | respiratory distress syndrome |
| ACE2 | Angiotensin I Converting Enzyme 2 |
| MAPK | Mitogen-activated protein kinase |
| TRAF6 | TNF Receptor Associated Factor 6 |
| TAK1 | Mitogen-Activated Protein Kinase Kinase Kinase 7 |
| NF-κB | Nuclear factor kappa B |
| TAM | Tumor-associated macrophage |
| MT1-MMP | Membrane type 1-matrix metalloproteinase |
| CSF | Colony stimulating factor |
| MSCs | Mesenchymal stromal cells |
| MIF | macrophage migration inhibitory factor |
| AD | Atopic Dermatitis |
| SEB | staphylococcal enterotoxin B |
| TAC | Transverse aortic constriction |
| CXCL | chemokine C-X-C ligand |
| NLRP3 | NOD-, LRR- and pyrin domain-containing protein 3 |
| SEB | Staphylococcal enterotoxin B |
| TLR | Toll-like receptor |
| LPS+Aβ | lipopolysaccahride+amyloid β protein |
| AT cells | Alveolar type 1 and 2 cells |
| DCs | Dendritic cells |
| TMPRSS2 | Transmembrane Serine Protease 2 |
| ATP | Adenosine triphosphate |
| DNA | Deoxyribonucleic acid |
| RNA | Ribonucleic acid |
| ASC | Apoptosis-associated speck-like protein |
| DAMPs | Damage associated molecular patterns |
| MIP1α | Macrophage inflammatory protein 1α |
| NK cells | Natural killer cells |
| UB | Ubiquitin |
| ISG | Interferon stimulated gene |
| UPS | Ubiquitin proteasome signaling |
| RIG-I | Retinoic acid-inducible gene I |
| TRIM25 | Tripartite motif-containing protein 25 |
| IRF | Interferon regulatory factor |
| STING | Stimulator of interferon genes |
| TSG101 | Tumor Susceptibility Gene 101 Protein |
| PKR | Protein kinase RNA-activated |
| CHMP5 | Charged Multivesicular Body Protein 5 |
| BECN1 | Beclin 1 |
| ERK1 | Mitogen-activated protein kinase 3 |
| IFIT1 | Interferon induced protein with tetratricopeptide repeats 1 |
| MxA | Myxovirus resistance protein 1 |
| p53 | Tumor protein 53 |
| PCNA | Proliferating cell nuclear antigen |
| HIF1α | Hypoxia-inducible factor 1-alpha |
| IQGAP1 | IQ Motif Containing GTPase Activating Protein 1 |
| UBC13 | Ubiquitin-conjugating enzyme E2 13 |
| UBCH6 | Ubiquitin-conjugating Enzyme H6 |
| PP2Cβ | Protein phosphatase 2Cβ |
| TRIM25 | Tripartite motif containing 25 |
| 4EHP | Eukaryotic Translation Initiation Factor 4E Family Member 2 |
| PML–RARα | Fusion protein of PML nuclear body scaffold protein and retinoic acid α receptor |
| δnp63α | Tumor protein 63 isoform |
| Nsp | Non-structural protein |
| GCG | (−)-gallocatechingallate |
| CG | (−)-catechingallate |
| GC | (−)-gallocatechin |
| C | (−)-catechin |
References
- Baden, L.R.; El Sahly, H.M.; Essink, B.; Kotloff, K.; Frey, S.; Novak, R.; Diemert, D.; Spector, S.A.; Rouphael, N.; Creech, C.B.; et al. Efficacy and safety of the mrna-1273 sars-cov-2 vaccine. N. Engl. J. Med. 2020, 384, 403–416. [Google Scholar] [CrossRef]
- Xia, X. Domains and functions of spike protein in sars-cov-2 in the context of vaccine design. Viruses 2021, 13, 109. [Google Scholar] [CrossRef] [PubMed]
- CDC. Emerging Sars-Cov-2 Variants. Available online: https://www.cdc.gov/coronavirus/2019-ncov/more/science-and-research/scientific-brief-emerging-variants.html# (accessed on 28 January 2021).
- Liu, W.; Dong, M.; Bo, L.; Li, C.; Liu, Q.; Li, Y.; Ma, L.; Xie, Y.; Fu, E.; Mu, D.; et al. Epigallocatechin-3-gallate ameliorates seawater aspiration-induced acute lung injury via regulating inflammatory cytokines and inhibiting jak/stat1 pathway in rats. Mediat. Inflamm. 2014, 2014, 612593. [Google Scholar] [CrossRef] [PubMed]
- Sano, M.; Tabata, M.; Suzuki, M.; Degawa, M.; Miyase, T.; Maeda-Yamamoto, M. Simultaneous determination of twelve tea catechins by high-performance liquid chromatography with electrochemical detection. Analyst 2001, 126, 816–820. [Google Scholar] [CrossRef] [PubMed]
- Jeong, J.H.; Kim, H.J.; Lee, T.J.; Kim, M.K.; Park, E.S.; Choi, B.S. Epigallocatechin 3-gallate attenuates neuronal damage induced by 3-hydroxykynurenine. Toxicology 2004, 195, 53–60. [Google Scholar] [CrossRef] [PubMed]
- Roomi, M.W.; Ivanov, V.; Kalinovsky, T.; Niedzwiecki, A.; Rath, M. In Vitro and In Vivo antitumorigenic activity of a mixture of lysine, proline, ascorbic acid, and green tea extract on human breast cancer lines mda-mb-231 and mcf-7. Med. Oncol. 2005, 22, 129–138. [Google Scholar] [CrossRef] [PubMed]
- Babu, P.V.; Sabitha, K.E.; Shyamaladevi, C.S. Therapeutic effect of green tea extract on oxidative stress in aorta and heart of streptozotocin diabetic rats. Chem. Biol. Interact. 2006, 162, 114–120. [Google Scholar] [CrossRef] [PubMed]
- Tsuneki, H.; Ishizuka, M.; Terasawa, M.; Wu, J.B.; Sasaoka, T.; Kimura, I. Effect of green tea on blood glucose levels and serum proteomic patterns in diabetic (db/db) mice and on glucose metabolism in healthy humans. BMC Pharmacol. 2004, 4, 18. [Google Scholar] [CrossRef] [PubMed]
- Mukhtar, H.; Wang, Z.Y.; Katiyar, S.K.; Agarwal, R. Tea components: Antimutagenic and anticarcinogenic effects. Prev. Med. 1992, 21, 351–360. [Google Scholar] [CrossRef]
- Takabayashi, F.; Harada, N.; Yamada, M.; Murohisa, B.; Oguni, I. Inhibitory effect of green tea catechins in combination with sucralfate on helicobacter pylori infection in mongolian gerbils. J. Gastroenterol. 2004, 39, 61–63. [Google Scholar] [CrossRef]
- Mukoyama, A.; Ushijima, H.; Nishimura, S.; Koike, H.; Toda, M.; Hara, Y.; Shimamura, T. Inhibition of rotavirus and enterovirus infections by tea extracts. Jpn. J. Med. Sci. Biol. 1991, 44, 181–186. [Google Scholar] [CrossRef] [PubMed]
- Fechtner, S.; Singh, A.; Chourasia, M.; Ahmed, S. Molecular insights into the differences in anti-inflammatory activities of green tea catechins on il-1beta signaling in rheumatoid arthritis synovial fibroblasts. Toxicol. Appl. Pharmacol. 2017, 329, 112–120. [Google Scholar] [CrossRef] [PubMed]
- Singh, A.K.; Umar, S.; Riegsecker, S.; Chourasia, M.; Ahmed, S. Regulation of transforming growth factor beta-activated kinase activation by epigallocatechin-3-gallate in rheumatoid arthritis synovial fibroblasts: Suppression of k(63)-linked autoubiquitination of tumor necrosis factor receptor-associated factor 6. Arthritis Rheumatol. 2016, 68, 347–358. [Google Scholar] [CrossRef] [PubMed]
- Ge, M.; Xiao, Y.; Chen, H.; Luo, F.; Du, G.; Zeng, F. Multiple antiviral approaches of (−)-epigallocatechin-3-gallate (EGCG) against porcine reproductive and respiratory syndrome virus infection in vitro. Antiviral. Res. 2018, 158, 52–62. [Google Scholar] [CrossRef] [PubMed]
- Carneiro, B.M.; Batista, M.N.; Braga, A.C.S.; Nogueira, M.L.; Rahal, P. The green tea molecule EGCG inhibits zika virus entry. Virology 2016, 496, 215–218. [Google Scholar] [CrossRef]
- Lu, J.W.; Hsieh, P.S.; Lin, C.C.; Hu, M.K.; Huang, S.M.; Wang, Y.M.; Liang, C.Y.; Gong, Z.; Ho, Y.J. Synergistic effects of combination treatment using EGCG and suramin against the chikungunya virus. Biochem. Biophys. Res. Commun. 2017, 491, 595–602. [Google Scholar] [CrossRef] [PubMed]
- Raekiansyah, M.; Buerano, C.C.; Luz, M.A.D.; Morita, K. Inhibitory effect of the green tea molecule EGCG against dengue virus infection. Arch. Virol. 2018, 163, 1649–1655. [Google Scholar] [CrossRef] [PubMed]
- Mou, Q.; Jiang, Y.; Zhu, L.; Zhu, Z.; Ren, T. EGCG induces beta-defensin 3 against influenza a virus h1n1 by the mapk signaling pathway. Exp. Ther. Med. 2020, 20, 3017–3024. [Google Scholar] [PubMed]
- Yamaguchi, K.; Honda, M.; Ikigai, H.; Hara, Y.; Shimamura, T. Inhibitory effects of (−)-epigallocatechin gallate on the life cycle of human immunodeficiency virus type 1 (hiv-1). Antiviral. Res. 2002, 53, 19–34. [Google Scholar] [CrossRef]
- Reid, S.P.; Shurtleff, A.C.; Costantino, J.A.; Tritsch, S.R.; Retterer, C.; Spurgers, K.B.; Bavari, S. Hspa5 is an essential host factor for ebola virus infection. Antiviral. Res. 2014, 109, 171–174. [Google Scholar] [CrossRef]
- Tallei, T.E.; Tumilaar, S.G.; Niode, N.J.; Fatimawali; Kepel, B.J.; Idroes, R.; Effendi, Y.; Sakib, S.A.; Emran, T.B. Potential of plant bioactive compounds as sars-cov-2 main protease (m(pro)) and spike (s) glycoprotein inhibitors: A molecular docking study. Scientifica 2020, 2020, 6307457. [Google Scholar] [CrossRef] [PubMed]
- Long, Q.X.; Tang, X.J.; Shi, Q.L.; Li, Q.; Deng, H.J.; Yuan, J.; Hu, J.L.; Xu, W.; Zhang, Y.; Lv, F.J.; et al. Clinical and immunological assessment of asymptomatic sars-cov-2 infections. Nat. Med. 2020, 26, 1200–1204. [Google Scholar] [CrossRef] [PubMed]
- Schett, G.; Manger, B.; Simon, D.; Caporali, R. Covid-19 revisiting inflammatory pathways of arthritis. Nat. Rev. Rheumatol. 2020, 16, 465–470. [Google Scholar] [CrossRef] [PubMed]
- Sanyaolu, A.; Okorie, C.; Marinkovic, A.; Patidar, R.; Younis, K.; Desai, P.; Hosein, Z.; Padda, I.; Mangat, J.; Altaf, M. Comorbidity and its impact on patients with covid-19. SN Compr. Clin. Med. 2020, 2, 1069–1076. [Google Scholar] [CrossRef] [PubMed]
- Tang, Y.; Liu, J.; Zhang, D.; Xu, Z.; Ji, J.; Wen, C. Cytokine storm in covid-19: The current evidence and treatment strategies. Front. Immunol. 2020, 11, 1708. [Google Scholar] [CrossRef] [PubMed]
- D’Alessandro, A.; Thomas, T.; Dzieciatkowska, M.; Hill, R.C.; Francis, R.O.; Hudson, K.E.; Zimring, J.C.; Hod, E.A.; Spitalnik, S.L.; Hansen, K.C. Serum proteomics in covid-19 patients: Altered coagulation and complement status as a function of il-6 level. J. Proteome Res. 2020, 19, 4417–4427. [Google Scholar] [CrossRef]
- Luo, M.; Liu, J.; Jiang, W.; Yue, S.; Liu, H.; Wei, S. Il-6 and cd8+ t cell counts combined are an early predictor of in-hospital mortality of patients with covid-19. JCI Insight 2020, 5, e139024. [Google Scholar] [CrossRef] [PubMed]
- Stone, J.H.; Frigault, M.J.; Serling-Boyd, N.J.; Fernandes, A.D.; Harvey, L.; Foulkes, A.S.; Horick, N.K.; Healy, B.C.; Shah, R.; Bensaci, A.M.; et al. Efficacy of tocilizumab in patients hospitalized with covid-19. N. Engl. J. Med. 2020, 383, 2333–2344. [Google Scholar] [CrossRef] [PubMed]
- Campochiaro, C.; Della-Torre, E.; Cavalli, G.; De Luca, G.; Ripa, M.; Boffini, N.; Tomelleri, A.; Baldissera, E.; Rovere-Querini, P.; Ruggeri, A.; et al. Efficacy and safety of tocilizumab in severe covid-19 patients: A single-centre retrospective cohort study. Eur. J. Intern. Med. 2020, 76, 43–49. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Zhang, Y.; Qiao, W.; Zhang, J.; Qi, Z. Baricitinib, a drug with potential effect to prevent sars-cov-2 from entering target cells and control cytokine storm induced by covid-19. Int. Immunopharmacol. 2020, 86, 106749. [Google Scholar] [CrossRef]
- Mehta, P.; McAuley, D.F.; Brown, M.; Sanchez, E.; Tattersall, R.S.; Manson, J.J.; The HLH across Speciality Collaboration, UK. Covid-19: Consider cytokine storm syndromes and immunosuppression. Lancet 2020, 395, 1033–1034. [Google Scholar] [CrossRef]
- Bozzi, G.; Mangioni, D.; Minoia, F.; Aliberti, S.; Grasselli, G.; Barbetta, L.; Castelli, V.; Palomba, E.; Alagna, L.; Lombardi, A.; et al. Anakinra combined with methylprednisolone in patients with severe covid-19 pneumonia and hyperinflammation: An observational cohort study. J. Allergy Clin. Immunol. 2020, 147, 561–566.e4. [Google Scholar] [CrossRef] [PubMed]
- Singh, A.K.; Haque, M.; O’Sullivan, K.; Chourasia, M.; Ouseph, M.M.; Ahmed, S. Suppression of monosodium urate crystal-induced inflammation by inhibiting tgf-beta-activated kinase 1-dependent signaling: Role of the ubiquitin proteasome system. Cell Mol. Immunol. 2021, 18, 162–170. [Google Scholar] [CrossRef]
- Singh, A.K.; Fechtner, S.; Chourasia, M.; Sicalo, J.; Ahmed, S. Critical role of il-1alpha in il-1beta-induced inflammatory responses: Cooperation with nf-kappabp65 in transcriptional regulation. FASEB J. 2019, 33, 2526–2536. [Google Scholar] [CrossRef] [PubMed]
- Ning, W.; Wang, S.; Dong, X.; Liu, D.; Fu, L.; Jin, R.; Xu, A. Epigallocatechin-3-gallate (EGCG) suppresses the trafficking of lymphocytes to epidermal melanocytes via inhibition of jak2: Its implication for vitiligo treatment. Biol. Pharm. Bull. 2015, 38, 1700–1706. [Google Scholar] [CrossRef]
- Lee, S.Y.; Jung, Y.O.; Ryu, J.G.; Oh, H.J.; Son, H.J.; Lee, S.H.; Kwon, J.E.; Kim, E.K.; Park, M.K.; Park, S.H.; et al. Epigallocatechin-3-gallate ameliorates autoimmune arthritis by reciprocal regulation of t helper-17 regulatory t cells and inhibition of osteoclastogenesis by inhibiting stat3 signaling. J. Leukoc. Biol. 2016, 100, 559–568. [Google Scholar] [CrossRef] [PubMed]
- Singh, A.K. Epigallocatechin-3-gallate (EGCG) suppresses systemic inflammation by inhibiting il-6-induced stat3 activation in cultured hepatocytes and in liver tissue of adjuvant-induced arthritis (AIA) rats. In Proceedings of the ACR/ARHP Annual Meeting, Chicago, IL, USA, 19–24 October 2018. [Google Scholar]
- Li, J. Neuroprotective effect of (−)-epigallocatechin-3-gallate on autoimmune thyroiditis in a rat model by an anti-inflammation effect, anti-apoptosis and inhibition of trail signaling pathway. Exp. Ther. Med. 2018, 15, 1087–1092. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.S.; Michael, A.; Butler-Manuel, S.A. Advances in the treatment of ovarian cancer: A potential role of antiinflammatory phytochemicals. Discov. Med. 2012, 13, 7–17. [Google Scholar]
- Jang, J.Y.; Lee, J.K.; Jeon, Y.K.; Kim, C.W. Exosome derived from epigallocatechin gallate treated breast cancer cells suppresses tumor growth by inhibiting tumor-associated macrophage infiltration and m2 polarization. BMC Cancer 2013, 13, 421. [Google Scholar] [CrossRef] [PubMed]
- Akhtar, N.; Haqqi, T.M. Epigallocatechin-3-gallate suppresses the global interleukin-1beta-induced inflammatory response in human chondrocytes. Arthritis Res. Ther. 2011, 13, R93. [Google Scholar] [CrossRef] [PubMed]
- Zgheib, A.; Lamy, S.; Annabi, B. Epigallocatechin gallate targeting of membrane type 1 matrix metalloproteinase-mediated src and janus kinase/signal transducers and activators of transcription 3 signaling inhibits transcription of colony-stimulating factors 2 and 3 in mesenchymal stromal cells. J. Biol. Chem. 2013, 288, 13378–13386. [Google Scholar]
- Noh, S.U.; Cho, E.A.; Kim, H.O.; Park, Y.M. Epigallocatechin-3-gallate improves dermatophagoides pteronissinus extract-induced atopic dermatitis-like skin lesions in nc/nga mice by suppressing macrophage migration inhibitory factor. Int. Immunopharmacol. 2008, 8, 1172–1182. [Google Scholar] [CrossRef]
- Hisano, M.; Yamaguchi, K.; Inoue, Y.; Ikeda, Y.; Iijima, M.; Adachi, M.; Shimamura, T. Inhibitory effect of catechin against the superantigen staphylococcal enterotoxin b (SEB). Arch. Dermatol. Res. 2003, 295, 183–189. [Google Scholar] [CrossRef] [PubMed]
- Koh, S.H.; Kim, S.H.; Kwon, H.; Park, Y.; Kim, K.S.; Song, C.W.; Kim, J.; Kim, M.H.; Yu, H.J.; Henkel, J.S.; et al. Epigallocatechin gallate protects nerve growth factor differentiated pc12 cells from oxidative-radical-stress-induced apoptosis through its effect on phosphoinositide 3-kinase/akt and glycogen synthase kinase-3. Brain Res. Mol. Brain Res. 2003, 118, 72–81. [Google Scholar] [CrossRef]
- Gundimeda, U.; McNeill, T.H.; Schiffman, J.E.; Hinton, D.R.; Gopalakrishna, R. Green tea polyphenols potentiate the action of nerve growth factor to induce neuritogenesis: Possible role of reactive oxygen species. J. Neurosci. Res. 2010, 88, 3644–3655. [Google Scholar] [CrossRef] [PubMed]
- Yang, N.; Shang, Y.X. Epigallocatechin gallate ameliorates airway inflammation by regulating treg/th17 imbalance in an asthmatic mouse model. Int. Immunopharmacol. 2019, 72, 422–428. [Google Scholar] [CrossRef]
- Han, Y.; Wang, Q.; Fan, X.; Chu, J.; Peng, J.; Zhu, Y.; Li, Y.; Li, X.; Shen, L.; Asenso, J.; et al. Epigallocatechin gallate attenuates overloadinduced cardiac ecm remodeling via restoring t cell homeostasis. Mol. Med. Rep. 2017, 16, 3542–3550. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Wang, J.; Pae, M.; Meydani, S.N.; Wu, D. Epigallocatechin-3-gallate inhibits expression of receptors for T cell regulatory cytokines and their downstream signaling in mouse CD4+ T cells. J. Nutr. 2012, 142, 566–571. [Google Scholar] [CrossRef]
- Qin, S.; Alcorn, J.F.; Craigo, J.K.; Tjoeng, C.; Tarwater, P.M.; Kolls, J.K.; Reinhart, T.A. Epigallocatechin-3-gallate reduces airway inflammation in mice through binding to proinflammatory chemokines and inhibiting inflammatory cell recruitment. J. Immunol. 2011, 186, 3693–3700. [Google Scholar] [CrossRef] [PubMed]
- Weber, A.A.; Neuhaus, T.; Skach, R.A.; Hescheler, J.; Ahn, H.Y.; Schror, K.; Ko, Y.; Sachinidis, A. Mechanisms of the inhibitory effects of epigallocatechin-3 gallate on platelet-derived growth factor-bb-induced cell signaling and mitogenesis. FASEB J. 2004, 18, 128–130. [Google Scholar] [CrossRef]
- Zhong, X.; Liu, M.; Yao, W.; Du, K.; He, M.; Jin, X.; Jiao, L.; Ma, G.; Wei, B.; Wei, M. Epigallocatechin-3-gallate attenuates microglial inflammation and neurotoxicity by suppressing the activation of canonical and noncanonical inflammasome via tlr4/nf-kappab pathway. Mol. Nutr. Food Res. 2019, 63, e1801230. [Google Scholar] [CrossRef]
- Ahn, H.Y.; Hadizadeh, K.R.; Seul, C.; Yun, Y.P.; Vetter, H.; Sachinidis, A. Epigallocathechin-3 gallate selectively inhibits the pdgf-bb-induced intracellular signaling transduction pathway in vascular smooth muscle cells and inhibits transformation of sis-transfected nih 3t3 fibroblasts and human glioblastoma cells (a172). Mol. Biol. Cell 1999, 10, 1093–1104. [Google Scholar] [CrossRef] [PubMed]
- Koch, E. Previfenon® as Chemoprophylaxis of Covid-19 in Health Workers (Herd). Available online: https://clinicaltrials.gov/ct2/show/NCT04446065 (accessed on 31 January 2021).
- Laxminarayan, R.; Wahl, B.; Dudala, S.R.; Gopal, K.; Mohan, B.C.; Neelima, S.; Reddy, J.K.S.; Radhakrishnan, J.; Lewnard, J.A. Epidemiology and transmission dynamics of Covid-19 in two indian states. Science 2020, 370, 691–697. [Google Scholar] [CrossRef]
- Rao, V.U.S.; Arakeri, G.; Subash, A.; Rao, J.; Jadhav, S.; Suhail Sayeed, M.; Rao, G.; Brennan, P.A. Covid-19: Loss of bridging between innate and adaptive immunity? Med. Hypotheses 2020, 144, 109861. [Google Scholar] [CrossRef] [PubMed]
- Tay, M.Z.; Poh, C.M.; Renia, L.; MacAry, P.A.; Ng, L.F.P. The trinity of covid-19: Immunity, inflammation and intervention. Nat. Rev. Immunol. 2020, 20, 363–374. [Google Scholar] [CrossRef]
- Zhao, C.; Zhao, W. Nlrp3 inflammasome—A key player in antiviral responses. Front. Immunol. 2020, 11, 211. [Google Scholar] [CrossRef]
- Roh, J.S.; Sohn, D.H. Damage-associated molecular patterns in inflammatory diseases. Immune Netw. 2018, 18, e27. [Google Scholar] [CrossRef]
- Fajgenbaum, D.C.; June, C.H. Cytokine storm. N. Engl. J. Med. 2020, 383, 2255–2273. [Google Scholar] [CrossRef]
- Nagashima, S.; Mendes, M.C.; Martins, C.A.P.; Borges, N.H.; Godoy, T.M.; Miggiolaro, A.; da Deziderio, S.F.; Machado-Souza, C.; de Noronha, L. Endothelial dysfunction and thrombosis in patients with covid-19-brief report. Arter. Thromb. Vasc. Biol. 2020, 40, 2404–2407. [Google Scholar] [CrossRef] [PubMed]
- Couzin-Frankel, J. The long haul. Science 2020, 369, 614–617. [Google Scholar] [CrossRef]
- Ackermann, M.; Verleden, S.E.; Kuehnel, M.; Haverich, A.; Welte, T.; Laenger, F.; Vanstapel, A.; Werlein, C.; Stark, H.; Tzankov, A.; et al. Pulmonary vascular endothelialitis, thrombosis, and angiogenesis in covid-19. N. Engl. J. Med. 2020, 383, 120–128. [Google Scholar] [CrossRef] [PubMed]
- Merad, M.; Martin, J.C. Pathological inflammation in patients with covid-19: A key role for monocytes and macrophages. Nat. Rev. Immunol. 2020, 20, 355–362. [Google Scholar] [CrossRef] [PubMed]
- Perng, Y.C.; Lenschow, D.J. Isg15 in antiviral immunity and beyond. Nat. Rev. Microbiol. 2018, 16, 423–439. [Google Scholar]
- Oshiumi, H.; Matsumoto, M.; Seya, T. Ubiquitin-mediated modulation of the cytoplasmic viral rna sensor rig-i. J. Biochem. 2012, 151, 5–11. [Google Scholar] [CrossRef]
- Chang, C.Y.; Liu, H.M.; Chang, M.F.; Chang, S.C. Middle East respiratory syndrome coronavirus nucleocapsid protein suppresses type I and type III interferon induction by targeting RIG-I signaling. J. Virol. 2020, 94. [Google Scholar] [CrossRef] [PubMed]
- Schubert, K.; Karousis, E.D.; Jomaa, A.; Scaiola, A.; Echeverria, B.; Gurzeler, L.A.; Leibundgut, M.; Thiel, V.; Muhlemann, O.; Ban, N. Sars-cov-2 nsp1 binds the ribosomal mrna channel to inhibit translation. Nat. Struct. Mol. Biol. 2020, 27, 959–966. [Google Scholar] [CrossRef]
- Spiegel, M.; Pichlmair, A.; Martinez-Sobrido, L.; Cros, J.; Garcia-Sastre, A.; Haller, O.; Weber, F. Inhibition of beta interferon induction by severe acute respiratory syndrome coronavirus suggests a two-step model for activation of interferon regulatory factor 3. J. Virol. 2005, 79, 2079–2086. [Google Scholar] [CrossRef] [PubMed]
- Clemente, V.; D’Arcy, P.; Bazzaro, M. Deubiquitinating enzymes in coronaviruses and possible therapeutic opportunities for covid-19. Int. J. Mol. Sci. 2020, 21, 3492. [Google Scholar] [CrossRef]
- Han, Y.S.; Chang, G.G.; Juo, C.G.; Lee, H.J.; Yeh, S.H.; Hsu, J.T.; Chen, X. Papain-like protease 2 (PLP2) from severe acute respiratory syndrome coronavirus (sars-cov): Expression, purification, characterization, and inhibition. Biochemistry 2005, 44, 10349–10359. [Google Scholar] [CrossRef]
- Lindner, H.A.; Fotouhi-Ardakani, N.; Lytvyn, V.; Lachance, P.; Sulea, T.; Menard, R. The papain-like protease from the severe acute respiratory syndrome coronavirus is a deubiquitinating enzyme. J. Virol. 2005, 79, 15199–15208. [Google Scholar] [CrossRef] [PubMed]
- Bekes, M.; Rut, W.; Kasperkiewicz, P.; Mulder, M.P.; Ovaa, H.; Drag, M.; Lima, C.D.; Huang, T.T. Sars hcov papain-like protease is a unique lys48 linkage-specific di-distributive deubiquitinating enzyme. Biochem. J. 2015, 468, 215–226. [Google Scholar] [CrossRef]
- Bekes, M.; van der Heden van Noort, G.J.; Ekkebus, R.; Ovaa, H.; Huang, T.T.; Lima, C.D. Recognition of lys48-linked di-ubiquitin and deubiquitinating activities of the sars coronavirus papain-like protease. Mol. Cell 2016, 62, 572–585. [Google Scholar] [CrossRef] [PubMed]
- Lindner, H.A.; Lytvyn, V.; Qi, H.; Lachance, P.; Ziomek, E.; Menard, R. Selectivity in isg15 and ubiquitin recognition by the sars coronavirus papain-like protease. Arch. Biochem. Biophys. 2007, 466, 8–14. [Google Scholar] [CrossRef]
- Shin, D.; Mukherjee, R.; Grewe, D.; Bojkova, D.; Baek, K.; Bhattacharya, A.; Schulz, L.; Widera, M.; Mehdipour, A.R.; Tascher, G.; et al. Papain-like protease regulates sars-cov-2 viral spread and innate immunity. Nature 2020, 587, 657–662. [Google Scholar] [CrossRef] [PubMed]
- Klemm, T.; Ebert, G.; Calleja, D.J.; Allison, C.C.; Richardson, L.W.; Bernardini, J.P.; Lu, B.G.; Kuchel, N.W.; Grohmann, C.; Shibata, Y.; et al. Mechanism and inhibition of the papain-like protease, plpro, of sars-cov-2. EMBO J. 2020, 39, e106275. [Google Scholar] [CrossRef]
- Fu, Z.; Huang, B.; Tang, J.; Liu, S.; Liu, M.; Ye, Y.; Liu, Z.; Xiong, Y.; Zhu, W.; Cao, D.; et al. The complex structure of grl0617 and sars-cov-2 plpro reveals a hot spot for antiviral drug discovery. Nat. Commun. 2021, 12, 488. [Google Scholar] [CrossRef] [PubMed]
- Harder, E.; Damm, W.; Maple, J.; Wu, C.; Reboul, M.; Xiang, J.Y.; Wang, L.; Lupyan, D.; Dahlgren, M.K.; Knight, J.L.; et al. Opls3: A force field providing broad coverage of drug-like small molecules and proteins. J. Chem. Theory Comput. 2016, 12, 281–296. [Google Scholar] [CrossRef] [PubMed]
- Friesner, R.A.; Murphy, R.B.; Repasky, M.P.; Frye, L.L.; Greenwood, J.R.; Halgren, T.A.; Sanschagrin, P.C.; Mainz, D.T. Extra precision glide: Docking and scoring incorporating a model of hydrophobic enclosure for protein-ligand complexes. J. Med. Chem. 2006, 49, 6177–6196. [Google Scholar] [CrossRef]
- Hirano, T.; Akira, S.; Taga, T.; Kishimoto, T. Biological and clinical aspects of interleukin 6. Immunol. Today 1990, 11, 443–449. [Google Scholar] [CrossRef]
- Ouyang, J.; Zhu, K.; Liu, Z.; Huang, J. Prooxidant effects of epigallocatechin-3-gallate in health benefits and potential adverse effect. Oxid. Med. Cell Longev. 2020, 2020, 9723686. [Google Scholar] [CrossRef] [PubMed]
- Molinari, M.; Watt, K.D.; Kruszyna, T.; Nelson, R.; Walsh, M.; Huang, W.Y.; Nashan, B.; Peltekian, K. Acute liver failure induced by green tea extracts: Case report and review of the literature. Liver Transpl. 2006, 12, 1892–1895. [Google Scholar] [CrossRef] [PubMed]
- Bedrood, Z.; Rameshrad, M.; Hosseinzadeh, H. Toxicological effects of camellia sinensis (green tea): A review. Phytother. Res. 2018, 32, 1163–1180. [Google Scholar] [PubMed]
- Cabrera, C.; Artacho, R.; Gimenez, R. Beneficial effects of green tea—A review. J. Am. Coll. Nutr. 2006, 25, 79–99. [Google Scholar] [CrossRef] [PubMed]
- Eng, Q.Y.; Thanikachalam, P.V.; Ramamurthy, S. Molecular understanding of epigallocatechin gallate (EGCG) in cardiovascular and metabolic diseases. J. Ethnopharmacol. 2018, 210, 296–310. [Google Scholar] [PubMed]
| Cytokine | Inhibitory Effect of EGCG | Reference |
|---|---|---|
| TRAIL | Suppresses TRAIL expression levels in auto immune thyroiditis. | [39] |
| Sensitizes TRAIL induced apoptosis in ovarian cancer cells, via repression of nuclear factor kappa B (NF-κB) and inhibition of TNF-α and IL-6. | [40] | |
| M-CSF | Down regulates M-CSF via miR-16 mediated inhibition of Tumor-associated macrophage (TAM) infiltration and M2 polarization in breast cancer cells. | [41] |
| GRO-α | Inhibits GRO-α expression levels by blocking IL-1β mediated stimulation in human Chondrocytes. | [42] |
| G-CSF | EGCG is chemopreventive through membrane type 1-matrix metalloproteinase (MT1-MMP) intracellular mediated signaling and sequential activation of the Src and JAK/STAT pathways thus antagonizing concavalin-A-induced colony stimulating factor (CSF)-2 and CSF-3 gene expression in mesenchymal stromal cells (MSCs). | [43] |
| IL-6 | Inhibits IL-6 in epithelial ovarian cancer cells. | [44] |
| Inhibits IL-6 expression levels by blocking IL-1β mediated stimulation in human chondrocytes. | [42] | |
| IL-2 | EGCG exerts anti-inflammatory effect by inhibiting macrophage migration inhibitory factor (MIF) in the pathogenesis of atopic dermatitis (AD). | [44] |
| EGCG binds to staphylococcal enterotoxin B (SEB) and neutralizes it in a dose dependent manner and inhibits SEB-induced IL-2 in AD. | [45] | |
| β-NGF | EGCG via the PI3K/Akt, glycogen synthase kinase-3 pathway and downstream signaling through cytochrome c and caspase-3 pathways could exert trophic factor effect in neurodegenerative diseases associated with oxidative injury. | [46] |
| EGCG significantly exalted NGF-induced neurite outgrowth by increasing the expression levels of mRNA and proteins for the neuronal markers neurofilament-L and growth associated protein-43. | [47] | |
| IL-10 | EGCG ameliorated airway inflammation and eosinophil infiltrations in asthmatic mice by increasing the production of IL-10, the number of CD4+CD25+ Foxp3+ Treg cells and expression of Foxp3 mRNA in the lung tissue. | [48] |
| MCP-1 | EGCG inhibits MCP-1, by suppressing IL-1β mediated stimulation in human Chondrocytes. | [42] |
| SCF | No articles found | |
| IL-15 | EGCG regulates effector T cells and naïve T cell population and restores the balance of T helper (Th) cell 17/regulatory T cells, via STAT3 and STAT5. EGCG rescued IL‑7 production and decreased the levels of IL‑15 in transverse aortic constriction (TAC) rats with a therapeutic potential in inhibiting cardiac extracellular matrix remodeling. | [49] |
| IL-8 | Inhibits IL-6 expression levels by blocking IL-1β mediated stimulation in human chondrocytes. | [42] |
| IL-7 | EGCG suppresses IL-7 signaling by inhibiting the expression of IL-7R and IL-2R receptor subunit (common γ chain) involved in IL-2 mediated T-cell regulation. | [50] |
| IP-10 | EGCG reduces airway inflammation via anti-inflammatory mechanism in the airway cells by binding directly to chemokines C-X-C ligand (CXCL)9, CXCL10, and CXCL11, thus limiting their biological activities. | [51] |
| PDGF-BB | EGCG, either by plasma membrane incorporated or soluble form, interacts directly with PDGF-BB via the galloyl group in the third position and interferes with PDGF-BB-induced mitogenic signaling pathway by inhibiting tyrosine phosphorylation of the PDGF-Rβ thereby preventing specific receptor binding, inhibiting downstream signal transduction pathways and cell proliferation. | [52] |
| IFN-γ | Reduced IFN-γ levels by suppressing NF-κB pathway. | [39] |
| EGCG binds to staphylococcal enterotoxin B (SEB) and neutralizes it in a dose dependent manner and inhibits SEB-induced IFN-γ production and IL-2. | [45] | |
| IL-18 | EGCG attenuates microglial inflammation and neurotoxicity by suppressing NLRP3 and caspase-11-dependent inflammasome via toll-like receptor (TLR)4/NF-κB pathway in lipopolysaccahride+amyloid β protein (LPS+Aβ)-induced rat primary microglia and hippocampus of APP/PS1 mice by inhibiting the expression of ionised calcium-binding adapter molecule-1, cleaved IL-1β, and cleaved IL-18 induced by LPS+Aβ. | [53] |
| EGCG exhibits anti-cancer and anti-atherosclerotic activity by a selective inhibition of the tyrosine phosphorylation of PDGF-Rβ and its downstream signaling pathway. | [54] | |
| IL-2rγ | EGCG impacts T-cell regulation by inhibiting IL-2 proprietary α chains resulting in impaired signaling. | [50] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chourasia, M.; Koppula, P.R.; Battu, A.; Ouseph, M.M.; Singh, A.K. EGCG, a Green Tea Catechin, as a Potential Therapeutic Agent for Symptomatic and Asymptomatic SARS-CoV-2 Infection. Molecules 2021, 26, 1200. https://doi.org/10.3390/molecules26051200
Chourasia M, Koppula PR, Battu A, Ouseph MM, Singh AK. EGCG, a Green Tea Catechin, as a Potential Therapeutic Agent for Symptomatic and Asymptomatic SARS-CoV-2 Infection. Molecules. 2021; 26(5):1200. https://doi.org/10.3390/molecules26051200
Chicago/Turabian StyleChourasia, Mukesh, Purushotham Reddy Koppula, Aruna Battu, Madhu M. Ouseph, and Anil K. Singh. 2021. "EGCG, a Green Tea Catechin, as a Potential Therapeutic Agent for Symptomatic and Asymptomatic SARS-CoV-2 Infection" Molecules 26, no. 5: 1200. https://doi.org/10.3390/molecules26051200
APA StyleChourasia, M., Koppula, P. R., Battu, A., Ouseph, M. M., & Singh, A. K. (2021). EGCG, a Green Tea Catechin, as a Potential Therapeutic Agent for Symptomatic and Asymptomatic SARS-CoV-2 Infection. Molecules, 26(5), 1200. https://doi.org/10.3390/molecules26051200






