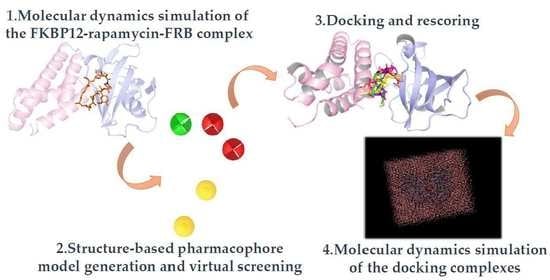
In Silico Strategy for Targeting the mTOR Kinase at Rapamycin Binding Site by Small Molecules
Abstract
1. Introduction
2. Results and Discussion
2.1. Molecular Dynamics Simulation
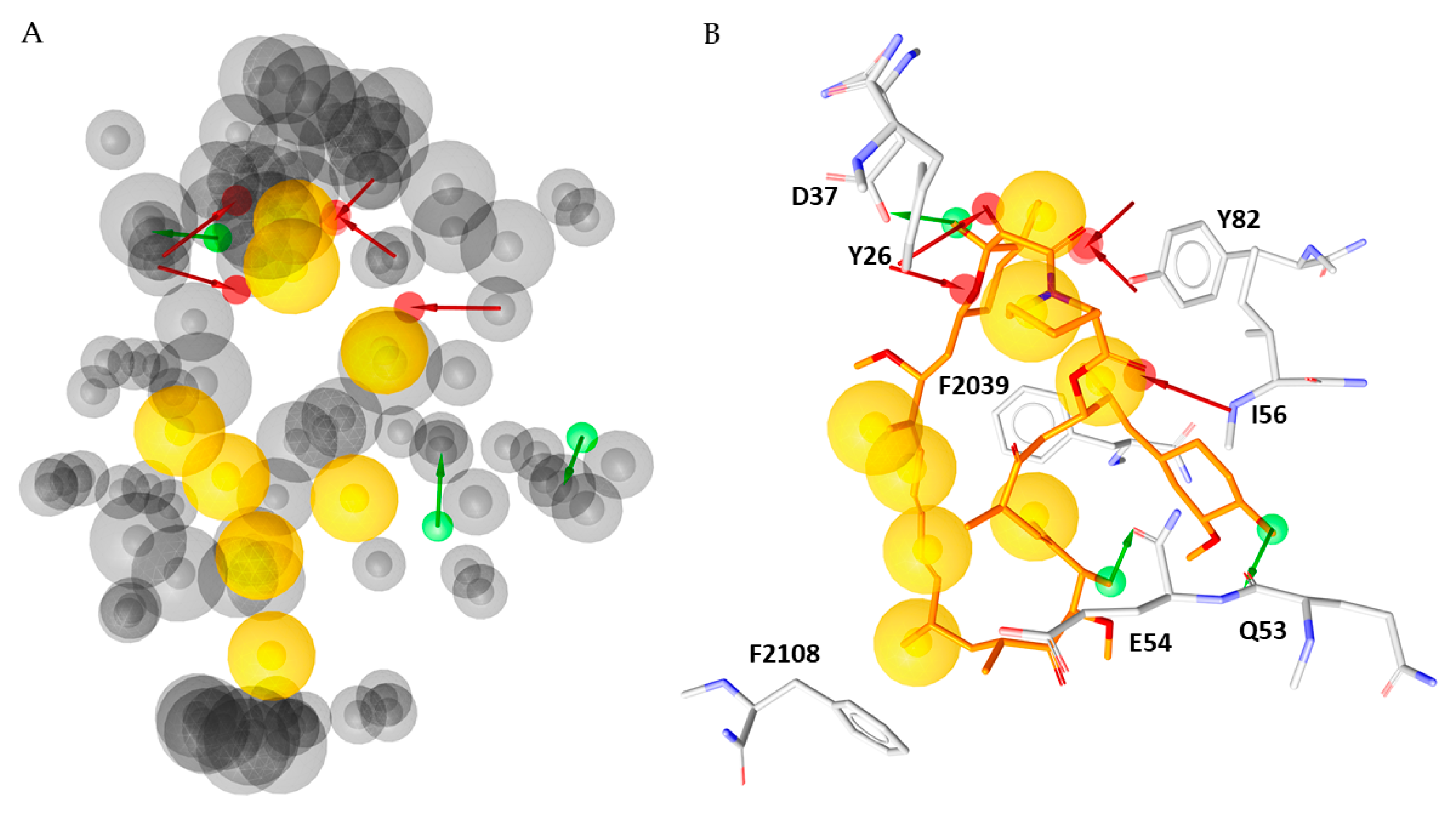
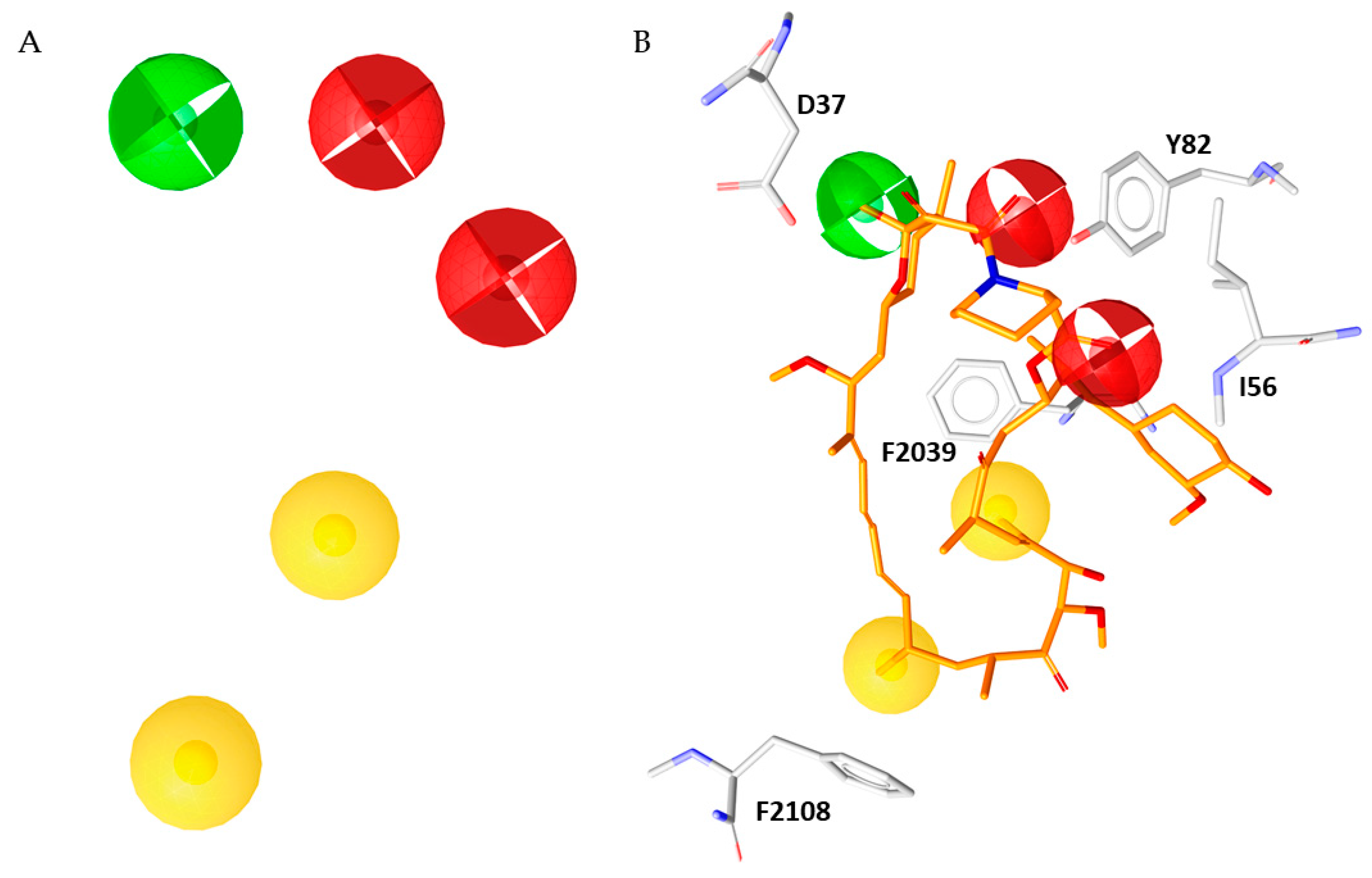
2.2. Pharmacophore Modelling
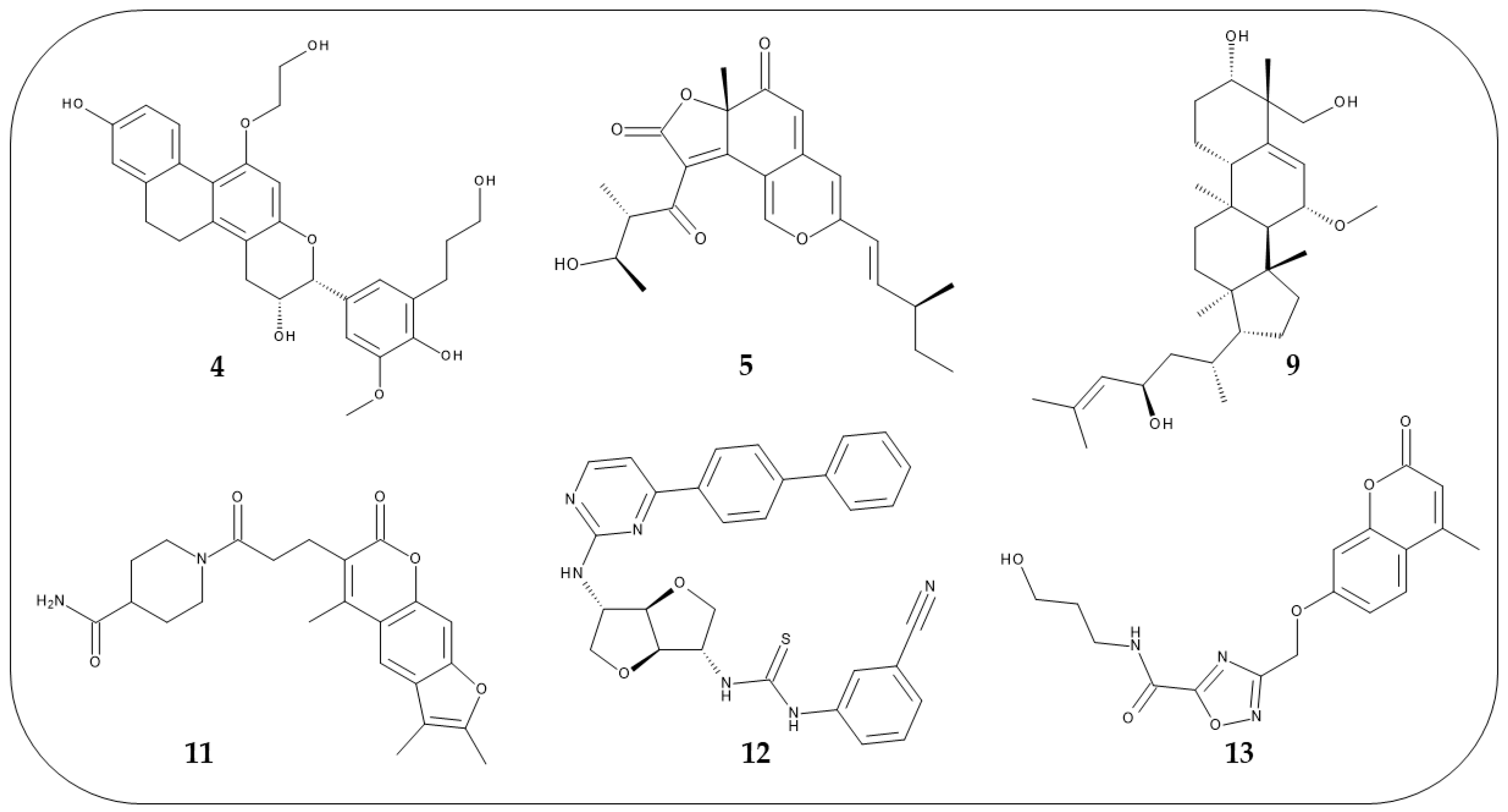
2.3. Virtual Screening
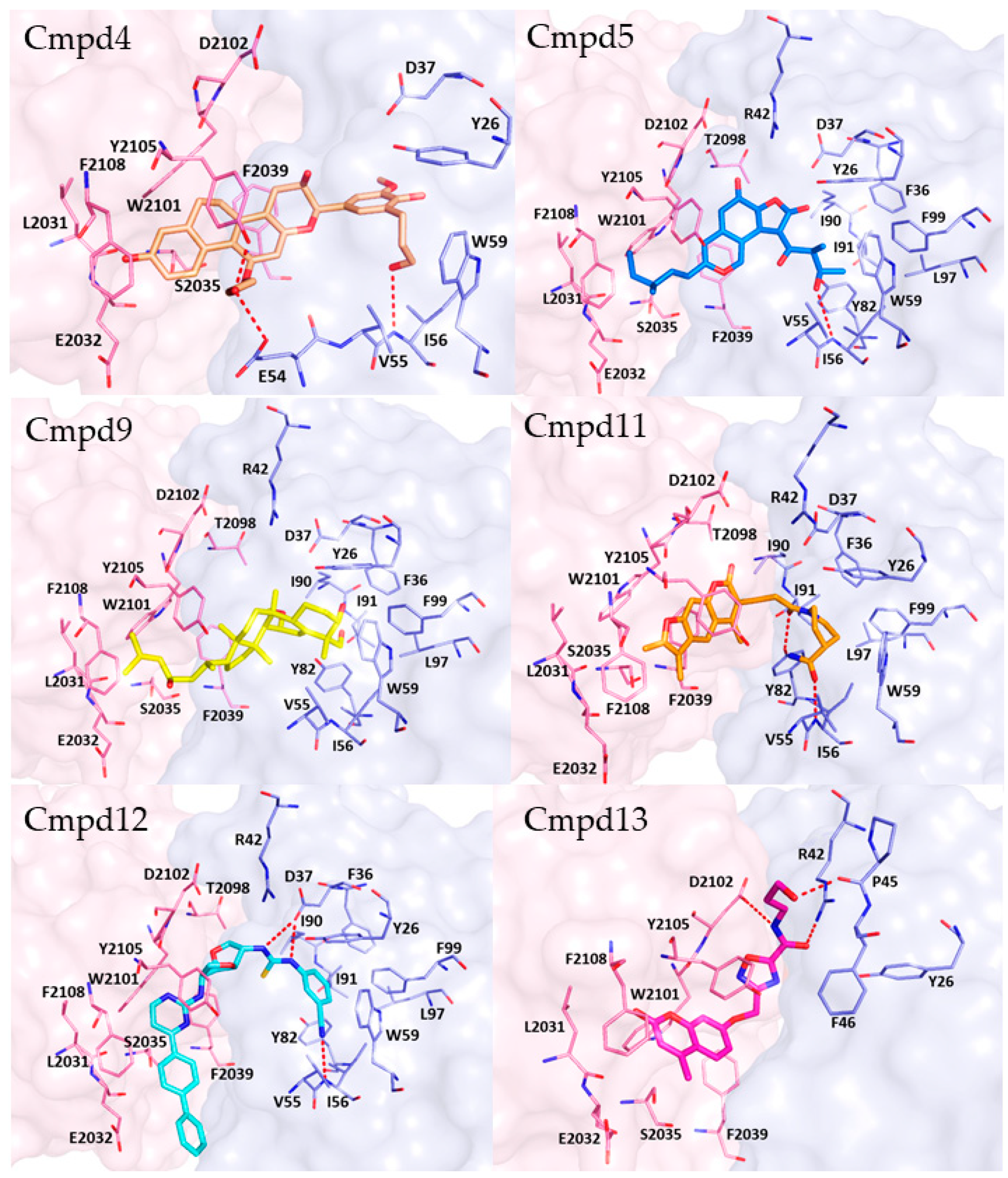
2.4. Molecular Docking, Rescoring and MD Simulation
3. Materials and Methods
3.1. Molecular Dynamics Simulation and Clustering
3.2. Pharmacophore Modelling and Virtual Screening
3.3. Docking and Rescoring
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Citraro, R.; Leo, A.; Constanti, A.; Russo, E.; De Sarro, G. mTOR pathway inhibition as a new therapeutic strategy in epilepsy and epileptogenesis. Pharmacol. Res. 2016, 107, 333–343. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Reeves, C.; Michalak, Z.; Coppola, A.; Diehl, B.; Sisodiya, S.M.; Thom, M. Evidence for mTOR pathway activation in a spectrum of epilepsy-associated pathologies. Acta Neuropathol. Commun. 2014, 2, 71. [Google Scholar] [CrossRef]
- Ostendorf, A.P.; Wong, M. mTOR inhibition in epilepsy: Rationale and clinical perspectives. CNS Drugs 2015, 29, 91–99. [Google Scholar] [CrossRef] [PubMed]
- Choi, J.; Chen, J.; Schreiber, S.L.; Clardy, J. Structure of the FKBP12-rapamycin complex interacting with the binding domain of human FRAP. Science 1996, 273, 239–242. [Google Scholar] [CrossRef]
- Chen, Y.; Zhou, X. Research progress of mTOR inhibitors. Eur. J. Med. Chem. 2020, 208, 112820. [Google Scholar] [CrossRef]
- Banaszynski, L.A.; Liu, C.W.; Wandless, T.J. Characterization of the FKBP.rapamycin.FRB ternary complex. J. Am. Chem. Soc. 2005, 127, 4715–4721. [Google Scholar] [CrossRef]
- Guduru, S.K.R.; Arya, P. Synthesis and biological evaluation of rapamycin-derived, next generation small molecules. Medchemcomm 2018, 9, 27–43. [Google Scholar] [CrossRef] [PubMed]
- Santulli, G.; Totary-Jain, H. Tailoring mTOR-based therapy: Molecular evidence and clinical challenges. Pharmacogenomics 2013, 14, 1517–1526. [Google Scholar] [CrossRef]
- Becher, O.J.; Gilheeney, S.W.; Khakoo, Y.; Lyden, D.C.; Haque, S.; De Braganca, K.C.; Kolesar, J.M.; Huse, J.T.; Modak, S.; Wexler, L.H.; et al. A phase I study of perifosine with temsirolimus for recurrent pediatric solid tumors. Pediatr. Blood Cancer 2017, 64. [Google Scholar] [CrossRef]
- Geiger, J.L.; Bauman, J.E.; Gibson, M.K.; Gooding, W.E.; Varadarajan, P.; Kotsakis, A.; Martin, D.; Gutkind, J.S.; Hedberg, M.L.; Grandis, J.R.; et al. Phase II trial of everolimus in patients with previously treated recurrent or metastatic head and neck squamous cell carcinoma. Head Neck 2016, 38, 1759–1764. [Google Scholar] [CrossRef] [PubMed]
- Meng, L.H.; Zheng, X.F. Toward rapamycin analog (rapalog)-based precision cancer therapy. Acta Pharmacol. Sin. 2015, 36, 1163–1169. [Google Scholar] [CrossRef]
- Oza, A.M.; Pignata, S.; Poveda, A.; McCormack, M.; Clamp, A.; Schwartz, B.; Cheng, J.; Li, X.; Campbell, K.; Dodion, P.; et al. Randomized phase II trial of ridaforolimus in advanced endometrial carcinoma. J. Clin. Oncol. 2015, 33, 3576–3582. [Google Scholar] [CrossRef]
- Park, H.; Garrido-Laguna, I.; Naing, A.; Fu, S.; Falchook, G.S.; Piha-Paul, S.A.; Wheler, J.J.; Hong, D.S.; Tsimberidou, A.M.; Subbiah, V.; et al. Phase I dose-escalation study of the mTOR inhibitor sirolimus and the HDAC inhibitor vorinostat in patients with advanced malignancy. Oncotarget 2016, 7, 67521–67531. [Google Scholar] [CrossRef]
- Luengo, J.I.; Yamashita, D.S.; Dunnington, D.; Beck, A.K.; Rozamus, L.W.; Yen, H.K.; Bossard, M.J.; Levy, M.A.; Hand, A.; Newman-Tarr, T.; et al. Structure-activity studies of rapamycin analogs: Evidence that the C-7 methoxy group is part of the effector domain and positioned at the FKBP12-FRAP interface. Chem. Biol. 1995, 2, 471–481. [Google Scholar] [CrossRef][Green Version]
- Nelson, F.C.; Stachel, S.J.; Eng, C.P.; Sehgal, S.N. Manipulation of the C(22)-C(27) region of rapamycin: Stability issues and biological implications. Bioorg. Med. Chem. Lett. 1999, 9, 295–300. [Google Scholar] [CrossRef]
- Ruan, B.; Pong, K.; Jow, F.; Bowlby, M.; Crozier, R.A.; Liu, D.; Liang, S.; Chen, Y.; Mercado, M.L.; Feng, X.; et al. Binding of rapamycin analogs to calcium channels and FKBP52 contributes to their neuroprotective activities. Proc. Natl. Acad. Sci. USA 2008, 105, 33–38. [Google Scholar] [CrossRef]
- Wieder, M.; Perricone, U.; Boresch, S.; Seidel, T.; Langer, T. Evaluating the stability of pharmacophore features using molecular dynamics simulations. Biochem. Biophys. Res. Commun. 2016, 470, 685–689. [Google Scholar] [CrossRef] [PubMed]
- Choudhury, C.; Priyakumar, U.D.; Sastry, G.N. Dynamics based pharmacophore models for screening potential inhibitors of mycobacterial cyclopropane synthase. J. Chem. Inf. Model. 2015, 55, 848–860. [Google Scholar] [CrossRef]
- Deng, J.; Lee, K.W.; Sanchez, T.; Cui, M.; Neamati, N.; Briggs, J.M. Dynamic receptor-based pharmacophore model development and its application in designing novel HIV-1 integrase inhibitors. J. Med. Chem. 2005, 48, 1496–1505. [Google Scholar] [CrossRef]
- Sohn, Y.S.; Park, C.; Lee, Y.; Kim, S.; Thangapandian, S.; Kim, Y.; Kim, H.H.; Suh, J.K.; Lee, K.W. Multi-conformation dynamic pharmacophore modeling of the peroxisome proliferator-activated receptor γ for the discovery of novel agonists. J. Mol. Graph. Model. 2013, 46, 1–9. [Google Scholar] [CrossRef]
- Perricone, U.; Wieder, M.; Seidel, T.; Langer, T.; Padova, A.; Almerico, A.M.; Tutone, M. A molecular dynamics-shared pharmacophore approach to boost early-enrichment virtual screening: A case study on peroxisome proliferator-activated receptor α. ChemMedChem 2017, 12, 1399–1407. [Google Scholar] [CrossRef]
- Crino, P.B. Mechanistic target of rapamycin (mTOR) signaling in status epilepticus. Epilepsy. Behav. 2019, 101, 106550. [Google Scholar] [CrossRef]
- Van Duyne, G.D.a.S.R.F.a.S.S.L.a.C.J. Atomic structure of the rapamycin human immunophilin FKBP-12 complex. J. Am. Chem. Soc. 1991, 113, 7433–7434. [Google Scholar] [CrossRef]
- Desmond Molecular Dynamics System. Maestro-Desmond Interoperability Tools; Schrödinger: New York, NY, USA, 2020. [Google Scholar]
- Wolber, G.; Langer, T. LigandScout: 3-D pharmacophores derived from protein-bound ligands and their use as virtual screening filters. J. Chem. Inf. Model. 2005, 45, 160–169. [Google Scholar] [CrossRef]
- Sterling, T.; Irwin, J.J. ZINC 15--Ligand Discovery for Everyone. J. Chem. Inf. Model. 2015, 55, 2324–2337. [Google Scholar] [CrossRef]
- Friesner, R.A.; Murphy, R.B.; Repasky, M.P.; Frye, L.L.; Greenwood, J.R.; Halgren, T.A.; Sanschagrin, P.C.; Mainz, D.T. Extra precision glide: Docking and scoring incorporating a model of hydrophobic enclosure for protein-ligand complexes. J. Med. Chem. 2006, 49, 6177–6196. [Google Scholar] [CrossRef] [PubMed]
- Friesner, R.A.; Banks, J.L.; Murphy, R.B.; Halgren, T.A.; Klicic, J.J.; Mainz, D.T.; Repasky, M.P.; Knoll, E.H.; Shelley, M.; Perry, J.K.; et al. Glide: A new approach for rapid, accurate docking and scoring. 1. Method and assessment of docking accuracy. J. Med. Chem. 2004, 47, 1739–1749. [Google Scholar] [CrossRef] [PubMed]
- Halgren, T.A.; Murphy, R.B.; Friesner, R.A.; Beard, H.S.; Frye, L.L.; Pollard, W.T.; Banks, J.L. Glide: A new approach for rapid, accurate docking and scoring. 2. Enrichment factors in database screening. J. Med. Chem. 2004, 47, 1750–1759. [Google Scholar] [CrossRef]
- Ramírez, D.; Caballero, J. Is it reliable to take the molecular docking top scoring position as the best solution without considering available structural data? Molecules 2018, 23, 1038. [Google Scholar] [CrossRef]
- Sastry, G.M.; Adzhigirey, M.; Day, T.; Annabhimoju, R.; Sherman, W. Protein and ligand preparation: Parameters, protocols, and influence on virtual screening enrichments. J. Comput. Aided. Mol. Des. 2013, 27, 221–234. [Google Scholar] [CrossRef]
- Schrödinger Release 2020-4: Maestro; Schrödinger, LLC: New York, NY, USA, 2020.
- Harder, E.; Damm, W.; Maple, J.; Wu, C.; Reboul, M.; Xiang, J.Y.; Wang, L.; Lupyan, D.; Dahlgren, M.K.; Knight, J.L.; et al. OPLS3: A force field providing broad coverage of drug-like small molecules and proteins. J. Chem. Theory Comput. 2016, 12, 281–296. [Google Scholar] [CrossRef] [PubMed]
- Schrödinger Release 2020-4: LigPrep; Schrödinger, LLC: New York, NY, USA, 2020.
- Shelley, J.C.; Cholleti, A.; Frye, L.L.; Greenwood, J.R.; Timlin, M.R.; Uchimaya, M. Epik: A software program for pK(a) prediction and protonation state generation for drug-like molecules. J. Comput. Aided. Mol. Des. 2007, 21, 681–691. [Google Scholar] [CrossRef] [PubMed]
- Greenwood, J.R.; Calkins, D.; Sullivan, A.P.; Shelley, J.C. Towards the comprehensive, rapid, and accurate prediction of the favorable tautomeric states of drug-like molecules in aqueous solution. J. Comput. Aided. Mol. Des. 2010, 24, 591–604. [Google Scholar] [CrossRef] [PubMed]

| Receptor | Native Ligand | Coordinates of Grid Center | Dimensions of the Outer Box (Å) | Dimensions of the Inner Box (Å) |
|---|---|---|---|---|
| FKBP12-FRB domain (PDB code 1FAP) | Rapamycin | X = −8.777 Y = 26.997 Z = 36.484 | X = 27.539 Y = 27.539 Z = 27.539 | X = 10 Y = 10 Z = 10 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Vittorio, S.; Gitto, R.; Adornato, I.; Russo, E.; De Luca, L. In Silico Strategy for Targeting the mTOR Kinase at Rapamycin Binding Site by Small Molecules. Molecules 2021, 26, 1103. https://doi.org/10.3390/molecules26041103
Vittorio S, Gitto R, Adornato I, Russo E, De Luca L. In Silico Strategy for Targeting the mTOR Kinase at Rapamycin Binding Site by Small Molecules. Molecules. 2021; 26(4):1103. https://doi.org/10.3390/molecules26041103
Chicago/Turabian StyleVittorio, Serena, Rosaria Gitto, Ilenia Adornato, Emilio Russo, and Laura De Luca. 2021. "In Silico Strategy for Targeting the mTOR Kinase at Rapamycin Binding Site by Small Molecules" Molecules 26, no. 4: 1103. https://doi.org/10.3390/molecules26041103
APA StyleVittorio, S., Gitto, R., Adornato, I., Russo, E., & De Luca, L. (2021). In Silico Strategy for Targeting the mTOR Kinase at Rapamycin Binding Site by Small Molecules. Molecules, 26(4), 1103. https://doi.org/10.3390/molecules26041103