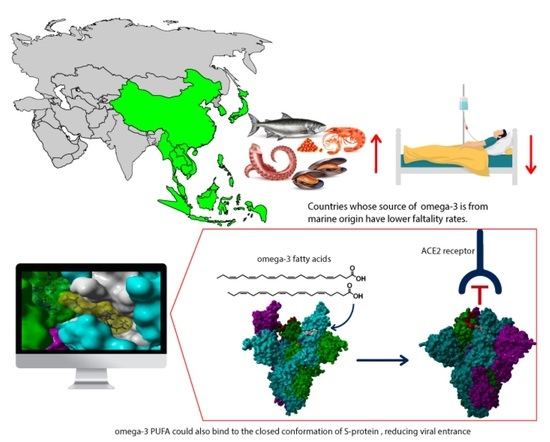
In Silico Study of Polyunsaturated Fatty Acids as Potential SARS-CoV-2 Spike Protein Closed Conformation Stabilizers: Epidemiological and Computational Approaches
Abstract
1. Introduction
2. Results and Discussion
2.1. Epidemiological-Based Analysis
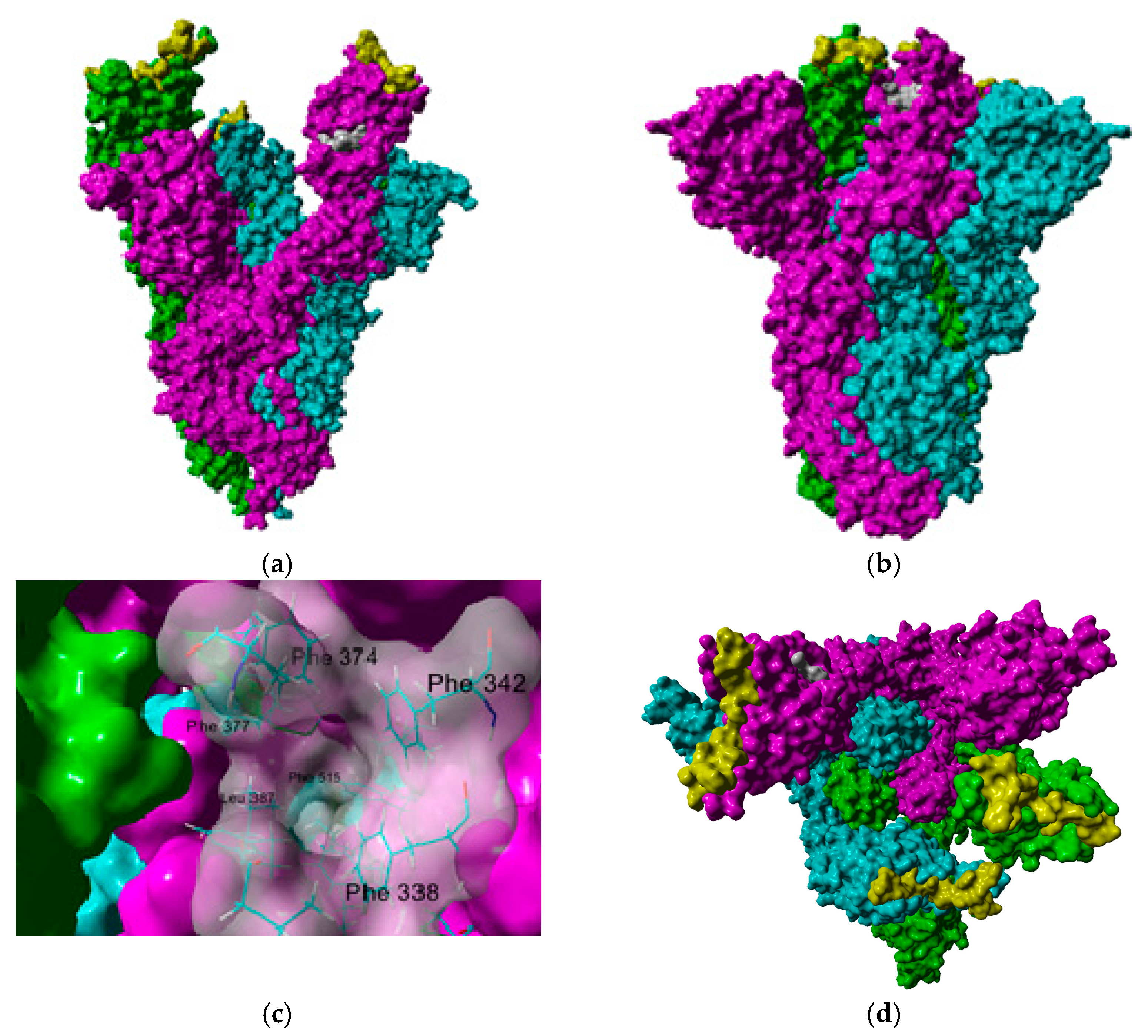
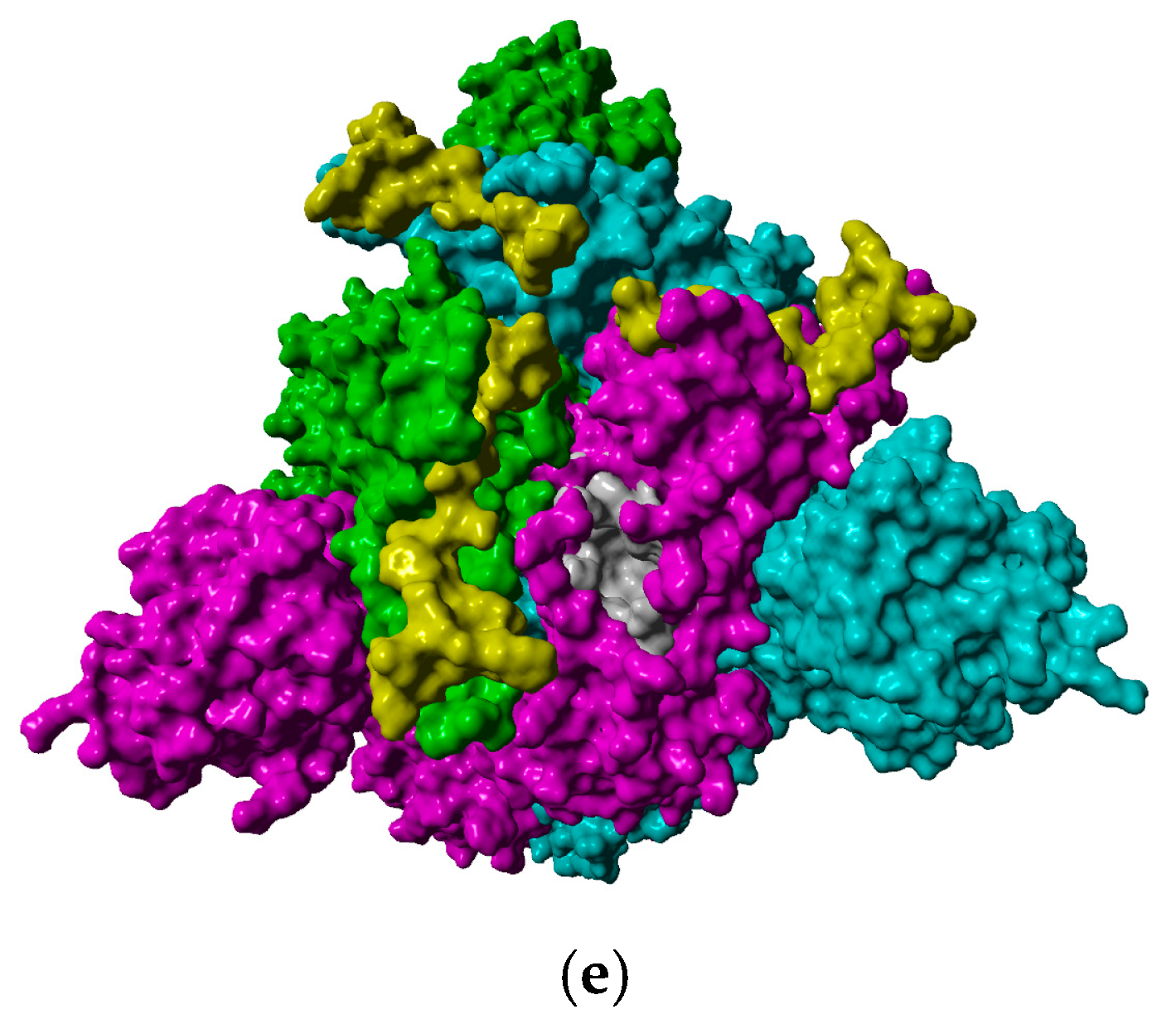
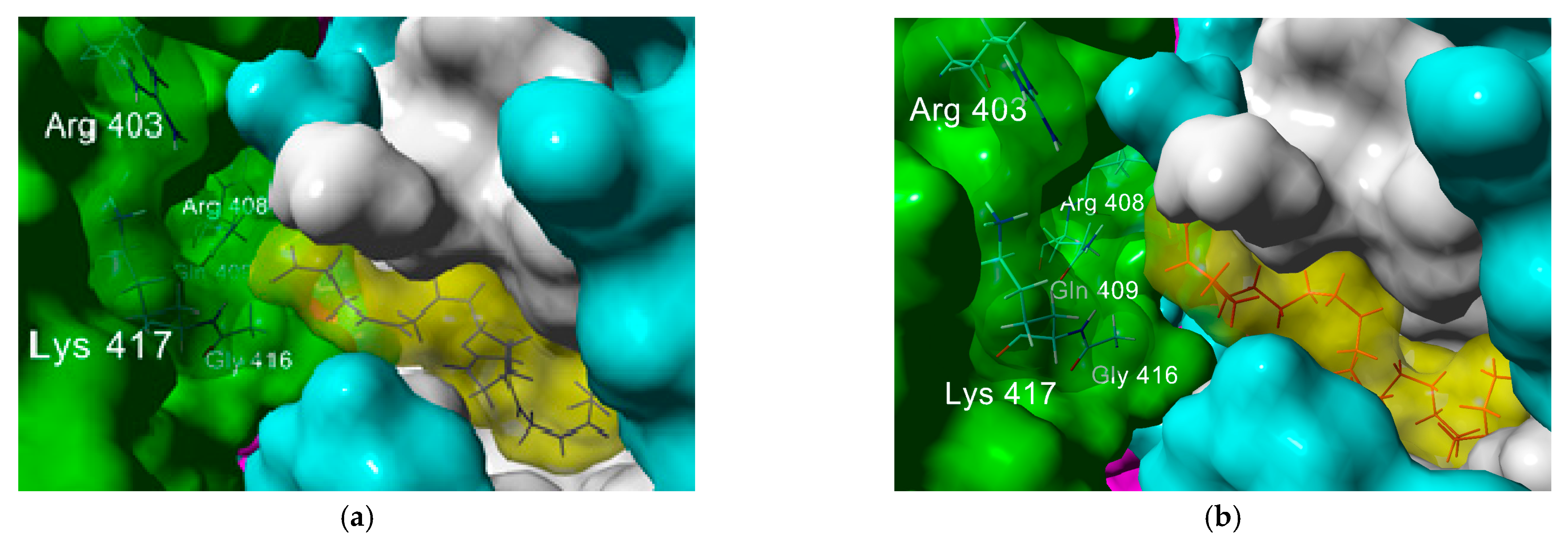
2.2. In Silico Studies
3. Materials and Methods
3.1. Epidemiology Analysis
3.2. Molecular Docking Studies
3.3. Molecular Dynamics Simulations
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Zumla, A.; Chan, J.F.W.; Azhar, E.I.; Hui, D.S.C.; Yuen, K.Y. Coronaviruses—drug discovery and therapeutic options. Nat. Rev. Drug Discov. 2016, 15, 327–347. [Google Scholar] [CrossRef] [PubMed]
- Letko, M.; Marzi, A.; Munster, V. Functional assessment of cell entry and receptor usage for SARS-CoV-2 and other lineage B betacoronaviruses. Nat. Microbiol. 2020. [Google Scholar] [CrossRef] [PubMed]
- Hoffmann, M.; Kleine-Weber, H.; Schroeder, S.; Krüger, N.; Herrler, T.; Erichsen, S.; Schiergens, T.S.; Herrler, G.; Wu, N.H.; Nitsche, A.; et al. SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and Is Blocked by a Clinically Proven Protease Inhibitor. Cell 2020, 181. [Google Scholar] [CrossRef] [PubMed]
- Boopathi, S.; Poma, A.B.; Kolandaivel, P. Novel 2019 coronavirus structure, mechanism of action, antiviral drug promises and rule out against its treatment. J. Biomol. Struct. Dyn. 2020. [Google Scholar] [CrossRef]
- Tay, M.Z.; Poh, C.M.; Rénia, L.; MacAry, P.A.; Ng, L.F.P. The trinity of COVID-19: Immunity, inflammation and intervention. Nat. Rev. Immunol. 2020, 20, 363–374. [Google Scholar] [CrossRef]
- Zhou, P.; Yang, X.L.; Wang, X.G.; Hu, B.; Zhang, L.; Zhang, W.; Si, H.R.; Zhu, Y.; Li, B.; Huang, C.L.; et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 2020, 579, 270–273. [Google Scholar] [CrossRef]
- Udugama, B.; Kadhiresan, P.; Kozlowski, H.N.; Malekjahani, A.; Osborne, M.; Li, V.Y.C.; Chen, H.; Mubareka, S.; Gubbay, J.B.; Chan, W.C.W. Diagnosing COVID-19: The Disease and Tools for Detection. ACS Nano 2020, 14, 3822–3835. [Google Scholar] [CrossRef]
- Walls, A.C.; Park, Y.J.; Tortorici, M.A.; Wall, A.; McGuire, A.T.; Veesler, D. Structure, Function, and Antigenicity of the SARS-CoV-2 Spike Glycoprotein. Cell 2020. [Google Scholar] [CrossRef]
- Henderson, R.; Edwards, R.J.; Mansouri, K.; Janowska, K.; Stalls, V.; Gobeil, S.M.C.; Kopp, M.; Li, D.; Parks, R.; Hsu, A.L.; et al. Controlling the SARS-CoV-2 spike glycoprotein conformation. Nat. Struct. Mol. Biol. 2020, 27, 925–933. [Google Scholar] [CrossRef]
- Cai, Y.; Zhang, J.; Xiao, T.; Peng, H.; Sterling, S.M.; Walsh, R.M.; Rawson, S.; Rits-Volloch, S.; Chen, B. Distinct conformational states of SARS-CoV-2 spike protein. Science 2020, 369, 1586–1592. [Google Scholar] [CrossRef]
- Toelzer, C.; Gupta, K.; Yadav, S.K.N.; Borucu, U.; Davidson, A.D.; Kavanagh Williamson, M.; Shoemark, D.K.; Garzoni, F.; Staufer, O.; Milligan, R.; et al. Free fatty acid binding pocket in the locked structure of SARS-CoV-2 spike protein. Science 2020, 370, 725–730. [Google Scholar] [CrossRef] [PubMed]
- Badger, J.; Minor, I.; Kremer, M.J.; Oliveira, M.A.; Smith, T.J.; Griffith, J.P.; Guerin, D.M.A.; Krishnaswamy, S.; Luo, M.; Rossmann, M.G.; et al. Structural analysis of a series of antiviral agents complexed with human rhinovirus 14. Proc. Natl. Acad. Sci. USA 1988, 85, 3304–3308. [Google Scholar] [CrossRef] [PubMed]
- Oliveira, M.A.; Zhao, R.; Lee, W.M.; Kremer, M.J.; Minor, I.; Rueckert, R.R.; Diana, G.D.; Pevear, D.C.; Dutko, F.J.; McKinlay, M.A.; et al. The structure of human rhinovirus 16. Structure 1993, 1, 51–68. [Google Scholar] [CrossRef]
- Casanova, V.; Sousa, F.H.; Stevens, C.; Barlow, P.G. Antiviral therapeutic approaches for human rhinovirus infections. Future Virol. 2018, 13, 505–518. [Google Scholar] [CrossRef] [PubMed]
- Beigel, J.H.; Tomashek, K.M.; Dodd, L.E.; Mehta, A.K.; Zingman, B.S.; Kalil, A.C.; Hohmann, E.; Chu, H.Y.; Luetkemeyer, A.; Kline, S.; et al. Remdesivir for the Treatment of Covid-19–Preliminary Report. N. Engl. J. Med. 2020. [Google Scholar] [CrossRef] [PubMed]
- Calder, P.C.; Grimble, R.F. Polyunsaturated fatty acids, inflammation and immunity. Eur. J. Clin. Nutr. 2002, 56. [Google Scholar] [CrossRef]
- Saini, R.K.; Keum, Y.S. Omega-3 and omega-6 polyunsaturated fatty acids: Dietary sources, metabolism, and significance–A review. Life Sci. 2018, 203, 255–267. [Google Scholar] [CrossRef]
- Micha, R.; Khatibzadeh, S.; Shi, P.; Fahimi, S.; Lim, S.; Andrews, K.G.; Engell, R.E.; Powles, J.; Ezzati, M.; Mozaffarian, D. Global, regional, and national consumption levels of dietary fats and oils in 1990 and 2010: A systematic analysis including 266 country-specific nutrition surveys. BMJ Br. Med. J. 2014, 348. [Google Scholar] [CrossRef]
- Diet Low in Seafood Omega-3 Fatty Acids–Level 3 Risk; Institute for Health Metrics and Evaluation. Available online: http://www.healthdata.org/results/gbd_summaries/2019/diet-low-in-seafood-omega-3-fatty-acids-level-3-risk (accessed on 12 January 2021).
- Weill, P.; Plissonneau, C.; Legrand, P.; Rioux, V.; Thibault, R. May omega-3 fatty acid dietary supplementation help reduce severe complications in Covid-19 patients? Biochimie 2020, 179, 275–280. [Google Scholar] [CrossRef]
- Calder, P.C. n-3 Polyunsaturated fatty acids, inflammation, and inflammatory diseases. Am. J. Clin. Nutr. 2006, 83, 1505S–1519S. [Google Scholar] [CrossRef]
- Vardhana, S.A.; Wolchok, J.D. The many faces of the anti-COVID immune response. J. Exp. Med. 2020, 217. [Google Scholar] [CrossRef] [PubMed]
- Frieman, M.; Heise, M.; Baric, R. SARS coronavirus and innate immunity. Virus Res. 2008, 133, 101–112. [Google Scholar] [CrossRef] [PubMed]
- Calder, P.C. Marine omega-3 fatty acids and inflammatory processes: Effects, mechanisms and clinical relevance. Biochim. Biophys. Acta—Mol. Cell Biol. Lipids 2015, 1851, 469–484. [Google Scholar] [CrossRef] [PubMed]
- Browning, L.M.; Walker, C.G.; Mander, A.P.; West, A.L.; Madden, J.; Gambell, J.M.; Young, S.; Wang, L.; Jebb, S.A.; Calder, P.C. Incorporation of eicosapentaenoic and docosahexaenoic acids into lipid pools when given as supplements providing doses equivalent to typical intakes of oily fish1-4. Am. J. Clin. Nutr. 2012, 96, 748–758. [Google Scholar] [CrossRef] [PubMed]
- Sorci, G.; Faivre, B.; Morand, S. Explaining among-country variation in COVID-19 case fatality rate. Sci. Rep. 2020, 10, 18909. [Google Scholar] [CrossRef]
- García De Acilu, M.; Leal, S.; Caralt, B.; Roca, O.; Sabater, J.; Masclans, J.R. The Role of Omega-3 Polyunsaturated Fatty Acids in the Treatment of Patients with Acute Respiratory Distress Syndrome: A Clinical Review. Biomed Res. Int. 2015. [Google Scholar] [CrossRef]
- Susser, M. The logic in ecological: I. The logic of analysis. Am. J. Public Health 1994, 84, 825–829. [Google Scholar] [CrossRef]
- Hernández-Rodríguez, M.; C Rosales-Hernández, M.; E Mendieta-Wejebe, J.; Martínez-Archundia, M.; Correa Basurto, J. Current Tools and Methods in Molecular Dynamics (MD) Simulations for Drug Design. Curr. Med. Chem. 2016, 23, 3909–3924. [Google Scholar] [CrossRef]
- Kindler, E.; Thiel, V. SARS-CoV and IFN: Too Little, Too Late. Cell Host Microbe 2016, 19, 139–141. [Google Scholar] [CrossRef]
- Ye, Q.; Wang, B.; Mao, J. The pathogenesis and treatment of the “Cytokine Storm” in COVID-19. J. Infect. 2020, 80, 607–613. [Google Scholar] [CrossRef]
- World Health Organization. WHO’s Presence in Countries, Territories and Areas; WHO: Geneva, Switzerland, 2017. [Google Scholar]
- Berman, H.M.; Westbrook, J.; Feng, Z.; Gilliland, G.; Bhat, T.N.; Weissig, H.; Shindyalov, I.N.; Bourne, P.E. The protein data bank. Nucleic Acids Res. 2000, 28, 235–242. [Google Scholar] [CrossRef] [PubMed]
- Thomsen, R.; Christensen, M.H. MolDock: A new technique for high-accuracy molecular docking. J. Med. Chem. 2006, 49, 3315–3321. [Google Scholar] [CrossRef] [PubMed]
- Krieger, E.; Vriend, G. YASARA View—molecular graphics for all devices—from smartphones to workstations. Bioinformatics 2014, 30, 2981–2982. [Google Scholar] [CrossRef] [PubMed]
- Krieger, E.; Vriend, G. New ways to boost molecular dynamics simulations. J. Comput. Chem. 2015, 36, 996–1007. [Google Scholar] [CrossRef] [PubMed]
- Trott, O.; Olson, A.J. AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 2009, 31. [Google Scholar] [CrossRef] [PubMed]
- Duan, Y.; Wu, C.; Chowdhury, S.; Lee, M.C.; Xiong, G.; Zhang, W.; Yang, R.; Cieplak, P.; Luo, R.; Lee, T.; et al. A point-charge force field for molecular mechanics simulations of proteins based on condensed-phase quantum mechanical calculations. J. Comput. Chem. 2003, 24, 1999–2012. [Google Scholar] [CrossRef]

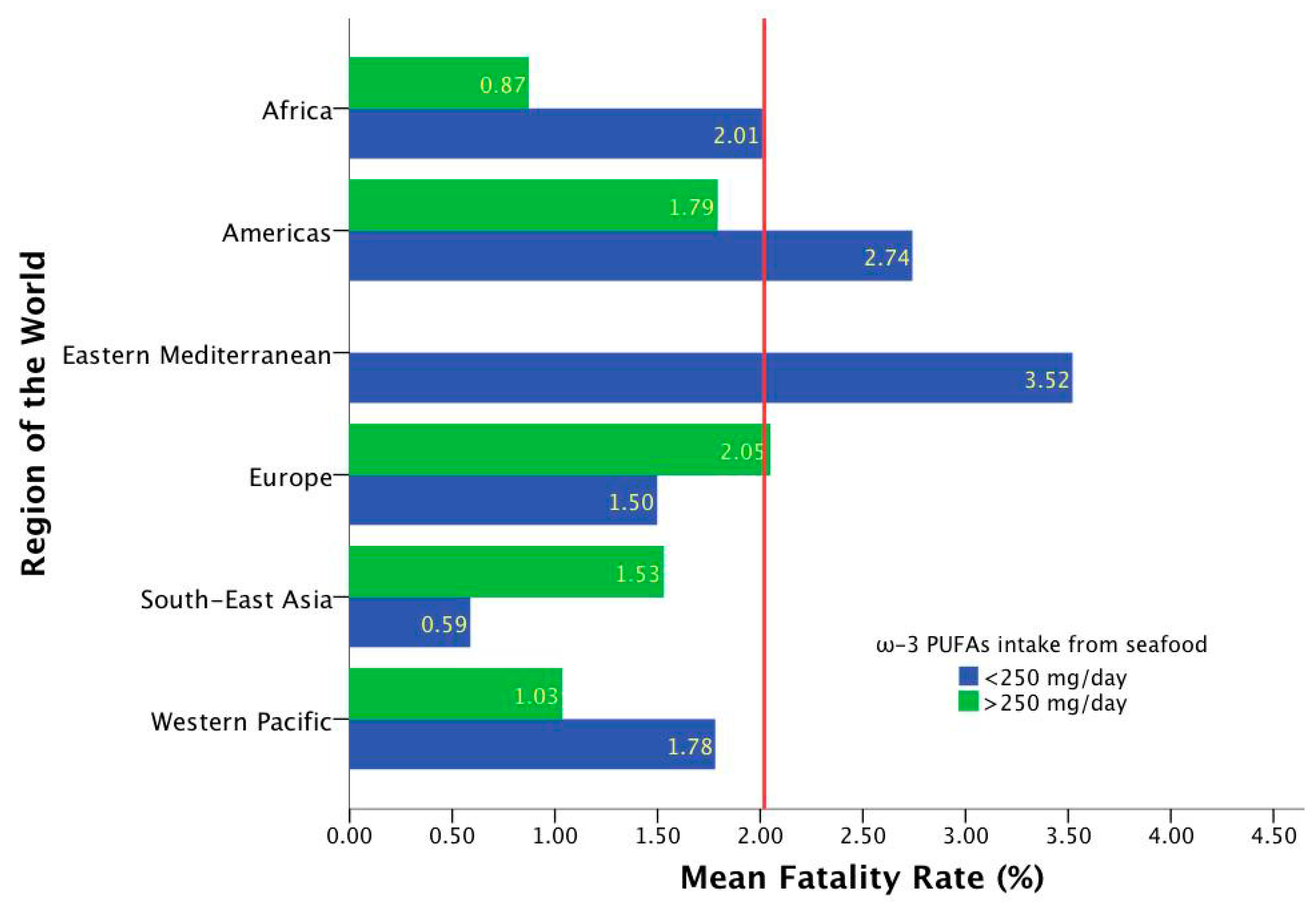

| Region According to WHO (Number of Included Countries) | Total Cumulative Cases 1 n = 186 | Total Cumulative Cases per 1 Million Population 1 n = 186 | Fatality Rates 1 n = 186 | ω-3 Marine Source Intake (mg/day) n = 186 | ω-3 Plant Source Intake (mg/day) n = 186 | ω-3 Marine + Plants Intake (mg/day) n = 186 |
|---|---|---|---|---|---|---|
| Global (186) | 336,937.59 ± 116,734.903 | 11,334.15 ± 14,635.058 | 2.02 ± 2.494 | 216.05 ± 380.148 | 838.81 ± 636.988 | 1054.86 ± 692.244 |
| Africa (46) | 32,392.63 ± 116,734.903 | 1895.015 ± 3441.064 | 1.94 ± 1.297 | 118.41 ± 195.488 | 726.96 ± 523.41 | 847.369 ± 576.333 |
| Americas (34) | 781,579.21 ± 2,466,433.13 | 12,101.95 ± 11,971.255 | 2.60 ± 1.889 | 164.26 ± 334.524 | 1005.29 ± 938.574 | 1169.55 ± 942.427 |
| Eastern Mediterranean (22) | 186,972.68 ± 225,982.552 | 14,092.86 ± 14,666.749 | 3.52 ± 6.036 | 45.14 ± 23.77 | 1027.45 ± 504.376 | 1072.59 ± 510.274 |
| Europe (51) | 268,574.41 ± 564,798.905 | 24,162.05 ± 16,505.756 | 1.65 ± 0.835 | 232.27 ± 254.529 | 904.27 ± 477.182 | 1136.55 ± 556.833 |
| South-East Asia (11) | 984,590.45 ± 2,818,704.55 | 4280.50 ± 7106.939 | 1.01 ± 1.017 | 634.00 ± 1132.984 | 517.27 ± 485.418 | 1151.27 ± 1094.051 |
| Western Pacific (22) | 39,338.55 ± 95,821.332 | 914.59 ± 2221.254 | 1.17 ± 1.565 | 424.59 ± 236.084 | 631.59 ± 690.112 | 1056.18 ± 639.115 |
| p2 | 0.058 | 0.000 ***,a,b,f,h,i,j,k,l,m | 0.008 *,a,b | 0.000 ***,b,c,d,e,f,g | 0.044 * | 0.307 |
| ω-3 Source Intake | Total Cumulative Cases | Total Cumulative Cases per 1 Million Population | Fatality Rates |
|---|---|---|---|
| ω-3 marine and plants | 0.219 ** | 0.257 ** | 0.041 |
| ω-3 marine source | −0.087 | 0.020 | −0.141 |
| ω-3 plants source | 0.321 *** | 0.329 *** | 0.165 * |
| Yasara Structure (Kcal/mol) | Molegro Virtual Docker | ||||
|---|---|---|---|---|---|
| Common Name | Fatty Acid Type | Closed | Open | Closed | Open |
| Tetracosahexaenoic acid (Nisinic acid) | 24:6 (ω-3) | 9.43 | 4.78 | −126.7 | −91.2 |
| Tetracosapentaenoic acid | 24:5 (ω-6) | 8.58 | 4.53 | −124.3 | −97.0 |
| Nervonic acid | 24:1 (ω-9) | 7.91 | 4.41 | −115.9 | −73.7 |
| Docosahexaenoic acid (DHA) | 22:6 (ω-3) | 8.87 | 4.72 | −120.1 | −85.3 |
| Docosapentaenoic acid (Osbond acid) | 22:5 (ω-6) | 8.69 | 4.84 | −117.9 | −77.9 |
| Erucic acid | 22:1 (ω-9) | 7.72 | 4.49 | −104.9 | −71.5 |
| Eicosapentaenoic acid (EPA) | 20:5 (ω-3) | 8.58 | 5.55 | −110.8 | −88.2 |
| Arachidonic acid (AA) | 20:4 (ω-6) | 8.23 | 4.80 | −116.1 | −82.3 |
| Gondoic acid | 20:1 (ω-9) | 7.44 | 5.11 | −106.3 | −79.9 |
| Lignoceric acid | 24:0 | 8.01 | 4.10 | −110.2 | −72.3 |
| Behenic acid | 22:0 | 7.37 | 4.09 | −109.1 | −72.1 |
| Arachidic acid | 20:0 | 6.59 | 4.19 | −108.2 | −73.1 |
| Linoleic acid | 18:2 (ω-6) | 6.79 | 4.37 | −104.8 | −79.6 |
| Hydrogen Bond | Electrostatic | |||||
|---|---|---|---|---|---|---|
| Phe 374A | Arg 403C | Asp 405C | Gln 409C | Arg 403C | Arg 408C | |
| Erucic acid (ω-9) | n.a. | n.a. | n.a. | n.a. | n.a. | n.a. |
| Osbond acid (ω-6) | 3.21 | 2.94 | 2.06 | 2.88 | 4.85 | n.a. |
| Docoesahexaenoic acid (ω-3) | 3.53 | 3.11 | 1.94 | 2.09 | 4.85 | n.a. |
| Behenic acid | 1.89 | n.a. | n.a. | 2.03 | n.a. | 3.7 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Vivar-Sierra, A.; Araiza-Macías, M.J.; Hernández-Contreras, J.P.; Vergara-Castañeda, A.; Ramírez-Vélez, G.; Pinto-Almazán, R.; Salazar, J.R.; Loza-Mejía, M.A. In Silico Study of Polyunsaturated Fatty Acids as Potential SARS-CoV-2 Spike Protein Closed Conformation Stabilizers: Epidemiological and Computational Approaches. Molecules 2021, 26, 711. https://doi.org/10.3390/molecules26030711
Vivar-Sierra A, Araiza-Macías MJ, Hernández-Contreras JP, Vergara-Castañeda A, Ramírez-Vélez G, Pinto-Almazán R, Salazar JR, Loza-Mejía MA. In Silico Study of Polyunsaturated Fatty Acids as Potential SARS-CoV-2 Spike Protein Closed Conformation Stabilizers: Epidemiological and Computational Approaches. Molecules. 2021; 26(3):711. https://doi.org/10.3390/molecules26030711
Chicago/Turabian StyleVivar-Sierra, Alonso, María José Araiza-Macías, José Patricio Hernández-Contreras, Arely Vergara-Castañeda, Gabriela Ramírez-Vélez, Rodolfo Pinto-Almazán, Juan Rodrigo Salazar, and Marco A. Loza-Mejía. 2021. "In Silico Study of Polyunsaturated Fatty Acids as Potential SARS-CoV-2 Spike Protein Closed Conformation Stabilizers: Epidemiological and Computational Approaches" Molecules 26, no. 3: 711. https://doi.org/10.3390/molecules26030711
APA StyleVivar-Sierra, A., Araiza-Macías, M. J., Hernández-Contreras, J. P., Vergara-Castañeda, A., Ramírez-Vélez, G., Pinto-Almazán, R., Salazar, J. R., & Loza-Mejía, M. A. (2021). In Silico Study of Polyunsaturated Fatty Acids as Potential SARS-CoV-2 Spike Protein Closed Conformation Stabilizers: Epidemiological and Computational Approaches. Molecules, 26(3), 711. https://doi.org/10.3390/molecules26030711