Micro-Raman Characterization of Structural Features of High-k Stack Layer of SOI Nanowire Chip, Designed to Detect Circular RNA Associated with the Development of Glioma
Abstract
1. Introduction
2. Results
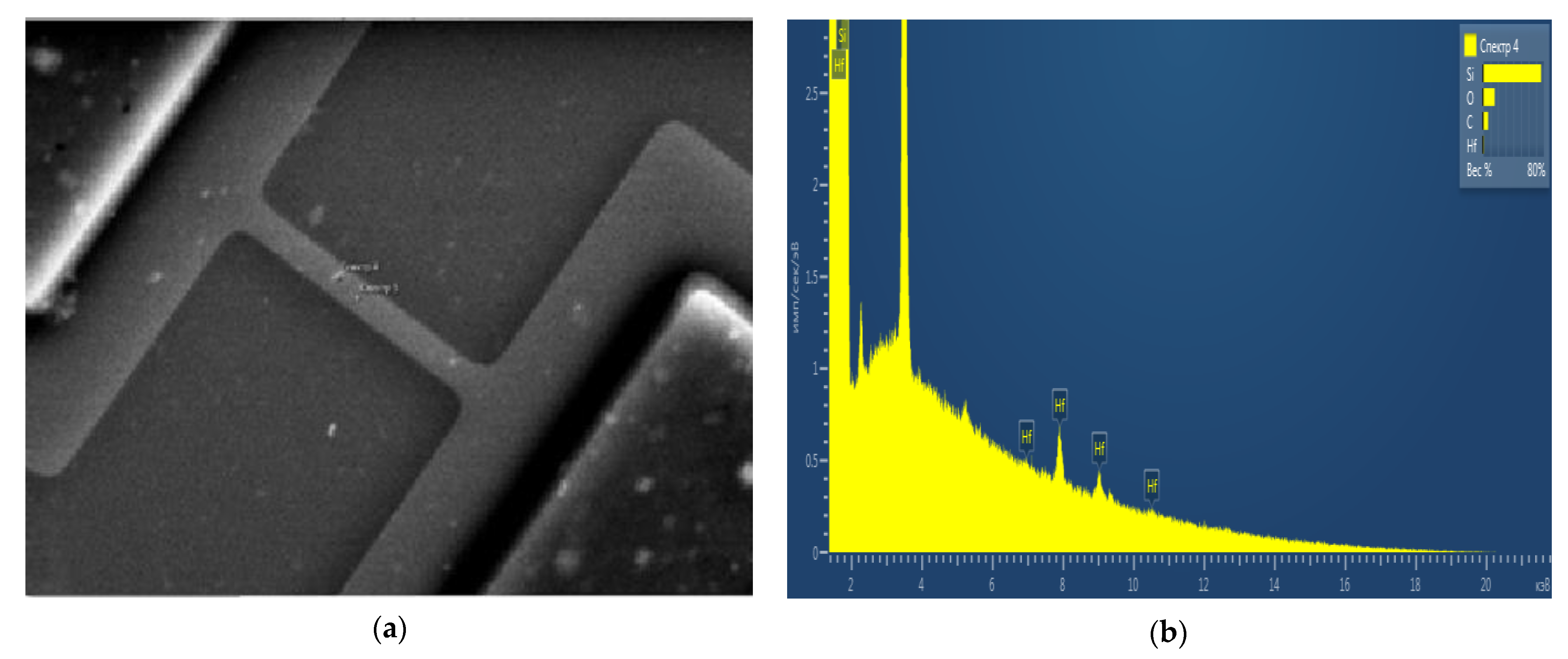
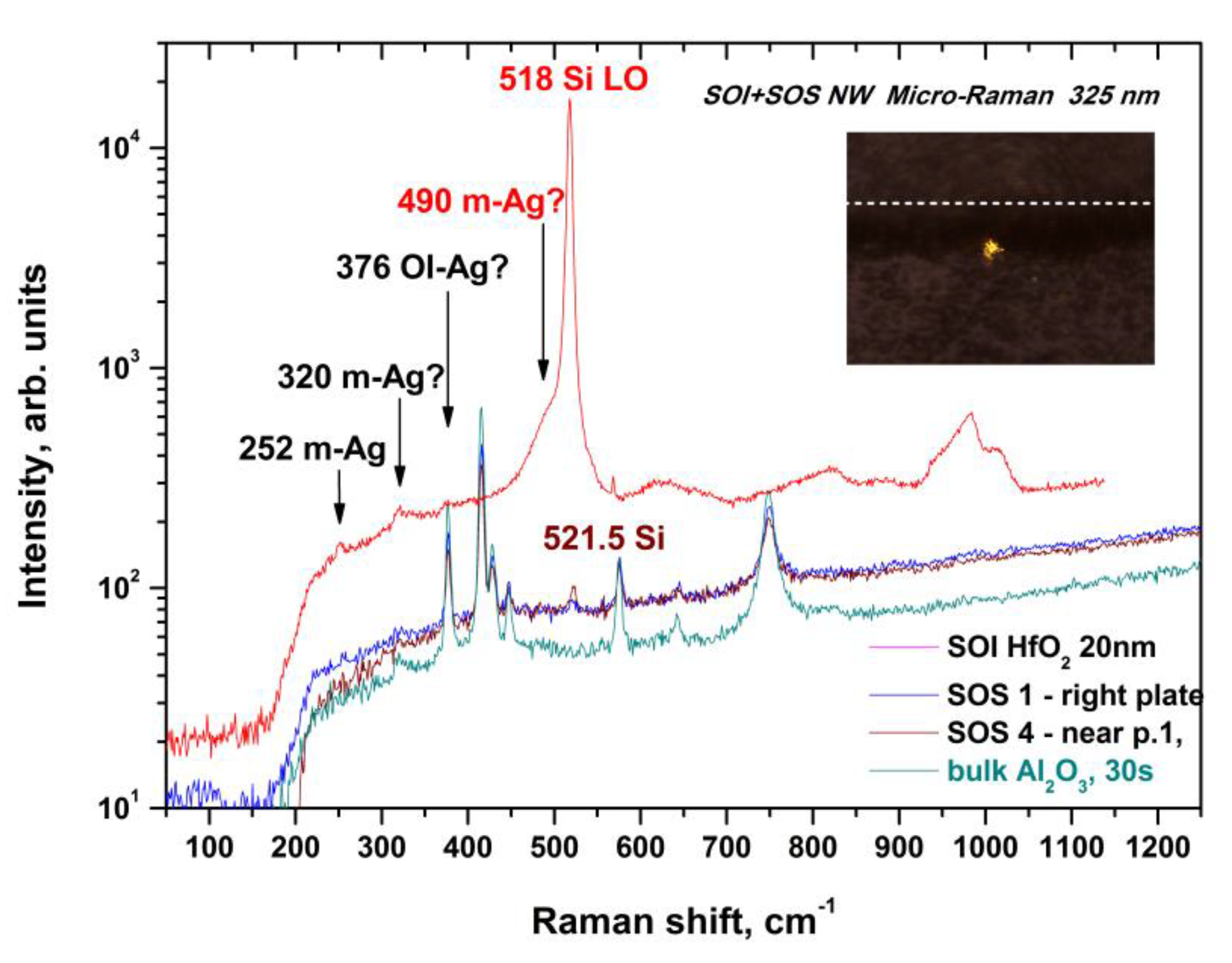
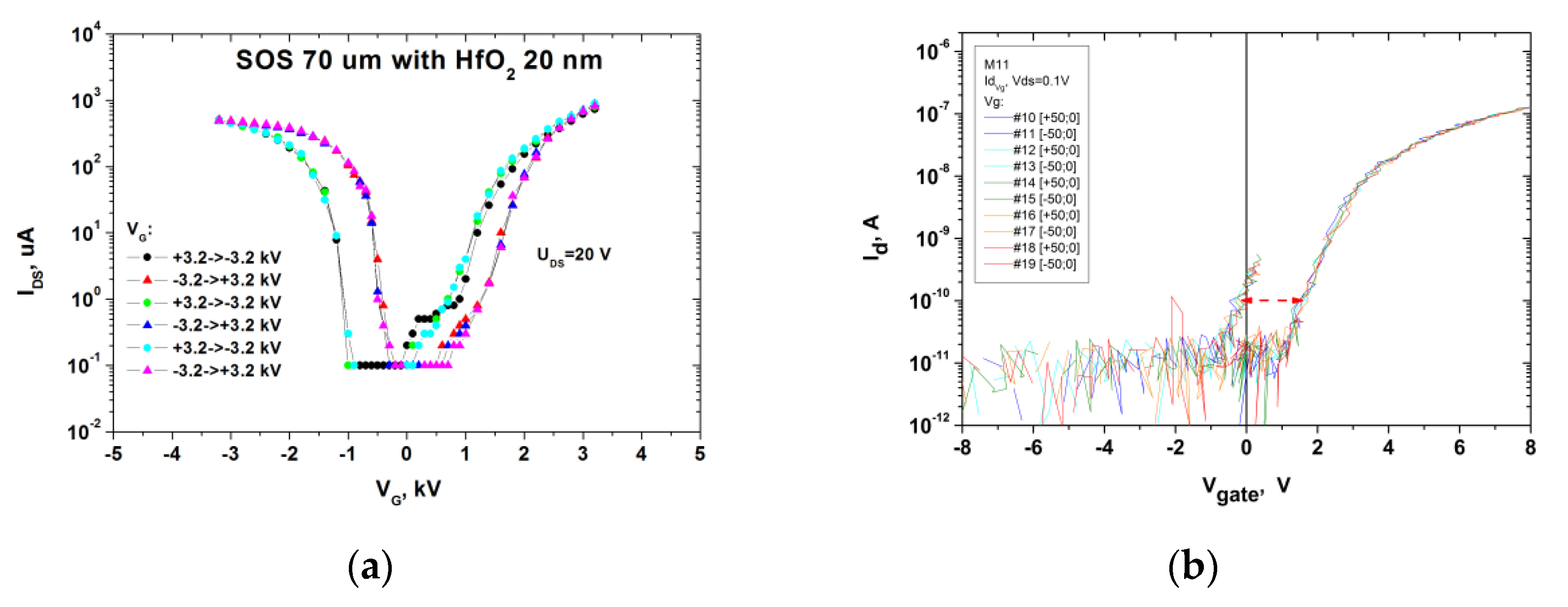
2.1. Monitoring of the Quality of h-k-SOI-NW Chip with High-k Cover Dielectrics of Different Composition by Micro-Raman Spectroscopy
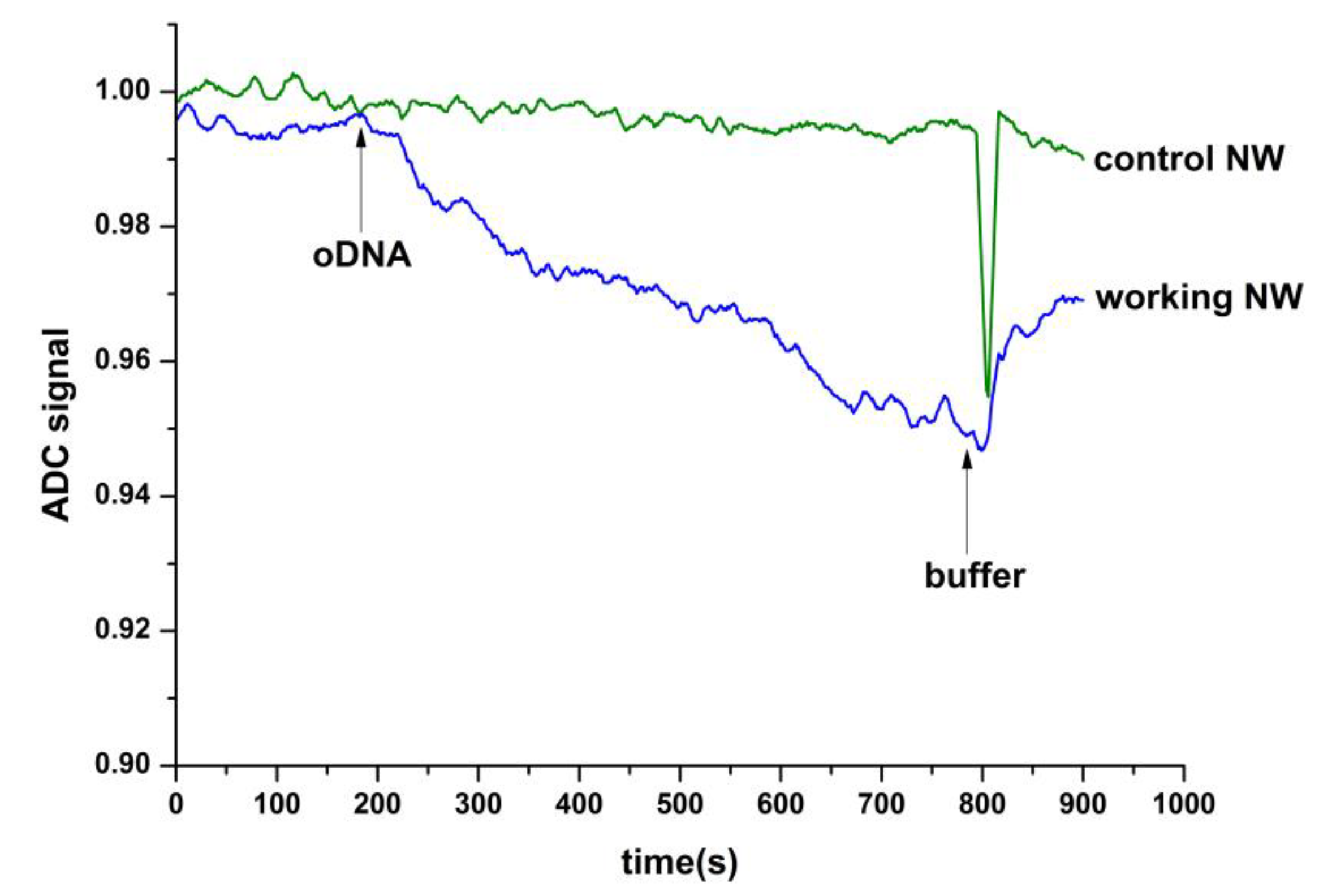
2.2. Demonstration of the Detection Capabilities of Synthetic Analogues of circRNA-oDNA, Associated with Glioma by h-k-SOI-NW Chip
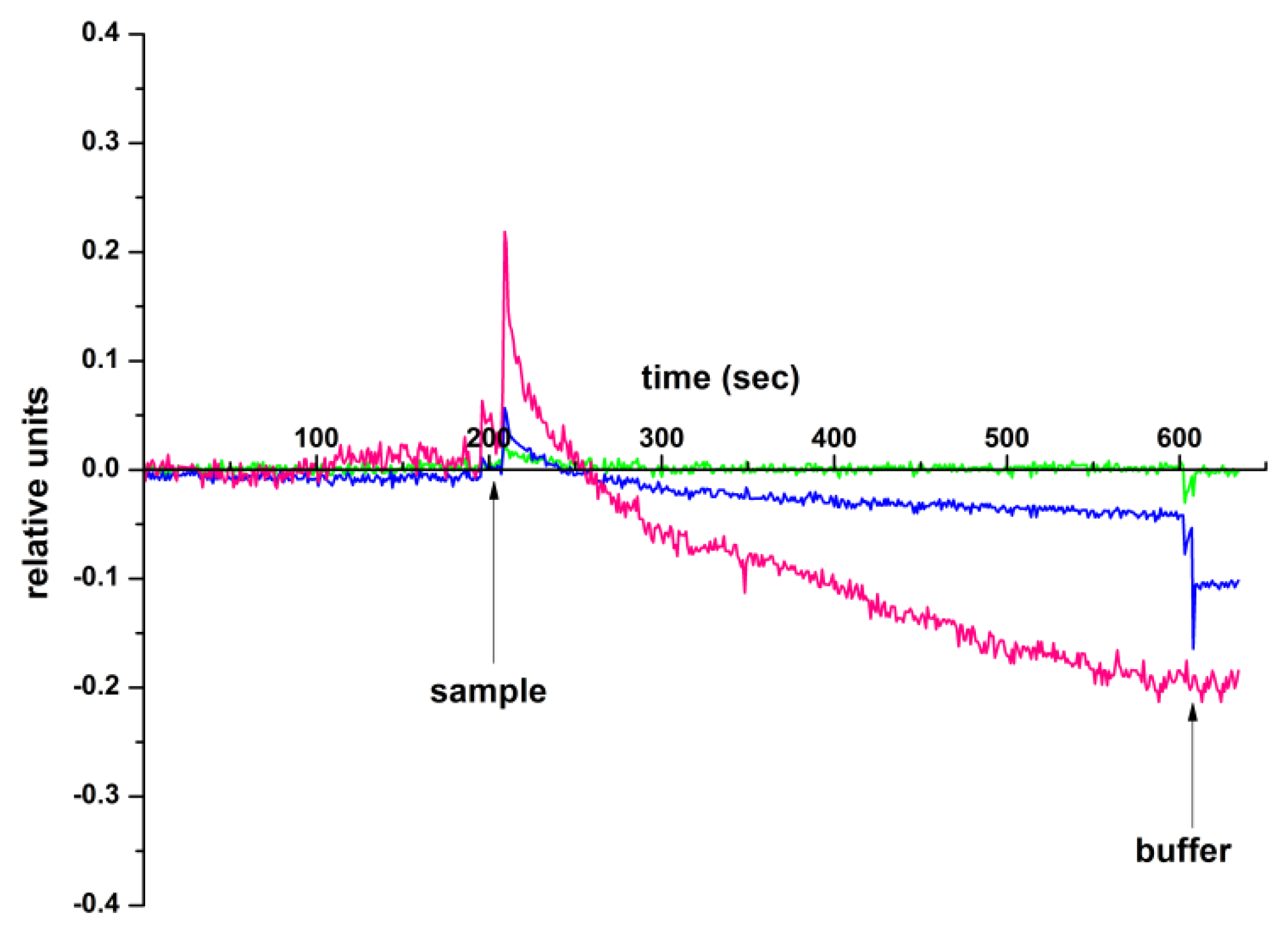
2.3. Demonstration of the Detection Capabilities of circ-SHKBP1, Associated with Glioma in Blood Plasma by h-k-SOI-NW Chip
3. Discussion
4. Materials and Methods
4.1. Chemicals
4.2. Oligonucleotides
4.3. Equipment
4.4. Fabrication of h-k-SOI-NW Chip
4.5. Modification of h-k-SOI-NW Chip Surface
4.6. Covalent Immobilization of Oligonucleotide Probes
4.7. Target oDNAs in Buffer Solutions Preparation
4.8. Plasma Samples
4.9. Electrical Measurements
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Chao, J.; Cao, W.; Su, S.; Weng, L.; Song, S.; Fan, C.; Wang, L.-H. Nanostructure-based surface-enhanced Raman scattering biosensors for nucleic acids and proteins. J. Mater. Chem. B 2016, 4, 1757–1769. [Google Scholar] [CrossRef]
- Žukovskaja, O.; Jahn, I.J.; Weber, K.; Cialla-May, D.; Popp, J. Detection of Pseudomonas aeruginosa Metabolite Pyocyanin in Water and Saliva by Employing the SERS Technique. Sensors 2017, 17, 1704. [Google Scholar] [CrossRef] [PubMed]
- Guo, J.; Zeng, F.; Guo, J.; Ma, X. Preparation and application of microfluidic SERS substrate: Challenges and future perspectives. J. Mater. Sci. Technol. 2020, 37, 96–103. [Google Scholar] [CrossRef]
- Rissin, D.M.; Kan, C.W.; Campbell, T.G.; Howes, S.C.; Fournier, D.R.; Song, L.; Piech, T.; Patel, P.P.; Chang, L.; Rivnak, A.J.; et al. Single-molecule enzyme-linked immunosorbent assay detects serum proteins at subfemtomolar concentrations. Nat. Biotechnol. 2010, 28, 595–599. [Google Scholar] [CrossRef]
- Malsagova, K.A.; Pleshakova, T.O.; Galiullin, R.A.; Shumov, I.D.; Kozlov, A.F.; Romanova, T.S.; Popov, V.P.; Glukhov, A.V.; Konev, V.A.; Archakov, A.I.; et al. Nanowire Aptamer-Sensitized Biosensor Chips with Gas Plasma-Treated Surface for the Detection of Hepatitis C Virus Core Antigen. Coatings 2020, 10, 753. [Google Scholar] [CrossRef]
- Yang, F.; Zhang, G.-J. Silicon nanowire-transistor biosensor for study of molecule-molecule interactions. Rev. Anal. Chem. 2014, 33. [Google Scholar] [CrossRef]
- Patolsky, F.; Zheng, G.; Hayden, O.; Lakadamyali, M.; Zhuang, X.; Lieber, C.M. Electrical detection of single viruses. Proc. Natl. Acad. Sci. USA 2004, 101, 14017–14022. [Google Scholar] [CrossRef]
- Naumova, O.V.; Fomin, B.; Nasimov, D.; Dudchenko, N.V.; Devyatova, S.F.; Zhanaev, E.D.; Popov, V.; Latyshev, A.V.; Aseev, A.L.; Ivanov, Y.D.; et al. SOI nanowires as sensors for charge detection. Semicond. Sci. Technol. 2010, 25. [Google Scholar] [CrossRef]
- Popov, V.P.; Antonova, A.I.; Frantsuzov, A.A.; Safronov, L.N.; Feofanov, G.N.; Naumova, O.V.; Kilanov, D.V. Properties of silicon-on-insulator structures and devices. Semiconductors 2001, 35, 1030–1037. [Google Scholar] [CrossRef]
- Gao, X.P.A.; Zheng, G.; Lieber, C.M. Subthreshold Regime has the Optimal Sensitivity for Nanowire FET Biosensors. Nano Lett. 2010, 10, 547–552. [Google Scholar] [CrossRef]
- Wang, H.; Han, X.; Ou, X.; Lee, C.-S.; Zhang, X.; Lee, S.-T. Silicon nanowire based single-molecule SERS sensor. Nanoscale 2013, 5, 8172–8176. [Google Scholar] [CrossRef]
- Elfström, N.; Juhasz, R.; Sychugov, I.; Engfeldt, T.; Karlström, A.E.; Linnros, J. Surface Charge Sensitivity of Silicon Nanowires: Size Dependence. Nano Lett. 2007, 7, 2608–2612. [Google Scholar] [CrossRef]
- Hahm, J.-I.; Lieber, C.M. Direct Ultrasensitive Electrical Detection of DNA and DNA Sequence Variations Using Nanowire Nanosensors. Nano Lett. 2004, 4, 51–54. [Google Scholar] [CrossRef]
- Malsagova, K.A.; Pleshakova, T.O.; Kozlov, A.F.; Shumov, I.D.; Ilnitskii, M.A.; Miakonkikh, A.V.; Popov, V.P.; Rudenko, K.V.; Glukhov, A.V.; Kupriyanov, I.N.; et al. Micro-Raman Spectroscopy for Monitoring of Deposition Quality of High-k Stack Protective Layer onto Nanowire FET Chips for Highly Sensitive miRNA Detection. Biosensors 2018, 8, 72. [Google Scholar] [CrossRef] [PubMed]
- Popov, V.P.; Ilnitskii, M.A.; Zhanaev, E.D.; Myakon’Kich, A.V.; Rudenko, K.V.; Glukhov, A.V. Biosensor properties of SOI nanowire transistors with a PEALD Al2O3 dielectric protective layer. Semiconductors 2016, 50, 632–638. [Google Scholar] [CrossRef]
- He, Q.; Zhao, L.; Liu, Y.; Liu, X.; Zheng, J.; Yu, H.; Cai, H.; Ma, J.; Liu, L.; Wang, P.; et al. Circ-SHKBP1 Regulates the Angiogenesis of U87 Glioma-Exposed Endothelial Cells through miR-544a/FOXP1 and miR-379/FOXP2 Pathways. Mol. Ther. Nucleic Acids 2018, 10, 331–348. [Google Scholar] [CrossRef] [PubMed]
- Ohgaki, H.; Kleihues, P. Genetic Pathways to Primary and Secondary Glioblastoma. Am. J. Pathol. 2007, 170, 1445–1453. [Google Scholar] [CrossRef] [PubMed]
- Ohgaki, H.; Dessen, P.; Jourde, B.; Horstmann, S.; Nishikawa, T.; Di Patre, P.-L.; Burkhard, C.; Schüler, D.; Probst-Hensch, N.M.; Maiorka, P.C.; et al. Genetic Pathways to Glioblastoma. Cancer Res. 2004, 64, 6892–6899. [Google Scholar] [CrossRef] [PubMed]
- Miller, A.M.; Shah, R.; Pentsova, E.I.; Pourmaleki, M.; Briggs, S.; Distefano, N.; Zheng, Y.; Skakodub, A.; Mehta, S.A.; Campos, C.; et al. Tracking tumour evolution in glioma through liquid biopsies of cerebrospinal fluid. Nat. Cell Biol. 2019, 565, 654–658. [Google Scholar] [CrossRef]
- Zachariah, M.; Oliveira-Costa, J.P.; Carter, B.S.; Stott, S.L.; Nahed, B.V. Blood-based biomarkers for the diagnosis and monitoring of gliomas. Neuro Oncol. 2018, 20, 1155–1161. [Google Scholar] [CrossRef]
- Chen, L.-C.; Zhang, H.-Y.; Qin, Z.-Y.; Wang, Y.; Mao, Y.; Yao, Y.; Zhou, L.-F. Serological Identification of URGCP as a Potential Biomarker for Glioma. CNS Neurosci. Ther. 2013, 20, 301–307. [Google Scholar] [CrossRef]
- Ludwig, K.; Kornblum, H.I. Molecular markers in glioma. J. Neuro Oncol. 2017, 134, 505–512. [Google Scholar] [CrossRef]
- Lisitsa, A.V.; Ponomarenko, E.A.; Lokhov, P.G.; Archakov, A.I. Postgenomic Medicine: Alternative to Biomarkers. Ann. Russ. Acad. Med. Sci. 2016, 71. [Google Scholar] [CrossRef]
- Kros, J.M.; Mustafa, D.M.; Dekker, L.J.; Smitt, P.A.S.; Luider, T.M.; Zheng, P.-P. Circulating glioma biomarkers. Neuro Oncol. 2014, 17, 343–360. [Google Scholar] [CrossRef] [PubMed]
- Archakov, A.I.; Ivanov, Y.D.; Lisitsa, A.V.; Zgoda, V.G. AFM fishing nanotechnology is the way to reverse the Avogadro number in proteomics. Proteomics 2007, 7, 4–9. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Ridzon, D.A.; Broomer, A.J.; Zhou, Z.; Lee, D.H.; Nguyen, J.T.; Barbisin, M.; Xu, N.L.; Mahuvakar, V.R.; Andersen, M.R.; et al. Real-time quantification of microRNAs by stem-loop RT-PCR. Nucleic Acids Res. 2005, 33, e179. [Google Scholar] [CrossRef] [PubMed]
- Hansen, T.B.; Jensen, T.I.; Clausen, B.H.; Bramsen, J.B.; Finsen, B.; Damgaard, C.K.; Kjems, J. Natural RNA circles function as efficient microRNA sponges. Nat. Cell Biol. 2013, 495, 384–388. [Google Scholar] [CrossRef]
- Jeck, W.R.; Sorrentino, J.A.; Wang, K.; Slevin, M.K.; Burd, C.E.; Liu, J.; Marzluff, W.F.; Sharpless, N. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA 2012, 19, 141–157. [Google Scholar] [CrossRef] [PubMed]
- Memczak, S.; Jens, M.; Elefsinioti, A.; Torti, F.; Krueger, J.; Rybak-Wolf, A.; Maier, L.; Mackowiak, S.; Gregersen, L.H.; Munschauer, M.; et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nat. Cell Biol. 2013, 495, 333–338. [Google Scholar] [CrossRef]
- Greene, J.; Baird, A.-M.; Brady, L.; Lim, M.; Gray, S.; McDermott, R.; Finn, S.P. Circular RNAs: Biogenesis, Function and Role in Human Diseases. Front. Mol. Biosci. 2017, 4, 38. [Google Scholar] [CrossRef]
- Li, Y.; Zheng, Q.; Bao, C.; Li, S.; Guo, W.; Zhao, J.; Chen, D.; Gu, J.; He, X.; Huang, S. Circular RNA is enriched and stable in exosomes: A promising biomarker for cancer diagnosis. Cell Res. 2015, 25, 981–984. [Google Scholar] [CrossRef] [PubMed]
- Holdt, L.M.; Kohlmaier, A.; Teupser, D. Molecular roles and function of circular RNAs in eukaryotic cells. Cell. Mol. Life Sci. 2018, 75, 1071–1098. [Google Scholar] [CrossRef]
- Quintard, P.E.; Barbéris, P.; Mirgorodsky, A.P.; Merle-Méjean, T. Comparative Lattice-Dynamical Study of the Raman Spectra of Monoclinic and Tetragonal Phases of Zirconia and Hafnia. J. Am. Ceram. Soc. 2004, 85, 1745–1749. [Google Scholar] [CrossRef]
- Naumova, O.; Fomin, B.; Popov, V.; Strelchuk, V.; Nikolenko, A.; Nazarov, A. An Experimental Study of Properties of Ultrathin Si Layer with Bonded Si/SiO2 Interface. Adv. Mater. Res. 2013, 854, 3–10. [Google Scholar] [CrossRef]
- Ivanov, Y.D.; Pleshakova, T.O.; Kozlov, A.F.; Malsagova, K.A.; Krohin, N.V.; Shumyantseva, V.V.; Shumov, I.D.; Popov, V.P.; Naumova, O.V.; Fomin, B.I.; et al. SOI nanowire for the high-sensitive detection of HBsAg and α-fetoprotein. Lab Chip 2012, 12, 5104–5111. [Google Scholar] [CrossRef] [PubMed]
- Malsagova, K.A.; Ivanov, Y.D.; Pleshakova, T.O.; Kaysheva, A.L.; Shumov, I.D.; Kozlov, A.F.; Archakov, A.I.; Popov, V.; Fomin, B.I.; Latyshev, A.V. A SOI-nanowire biosensor for the multiple detection of D-NFATc1 protein in the serum. Anal. Methods 2015, 7, 8078–8085. [Google Scholar] [CrossRef]

| Plasma Sample No. | Age | Sex | Diagnosis | Morphological Characteristic |
|---|---|---|---|---|
| 005 | 54 | Female | Neoplasm of an uncertain nature of the brain and central nervous system | Brain tissue with a marginal zone of glioma in one of the fragments, within the studied biopsy |
| 96 | 70 | Male | Prostatic hyperplasia | - |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ivanov, Y.D.; Malsagova, K.A.; Popov, V.P.; Kupriyanov, I.N.; Pleshakova, T.O.; Galiullin, R.A.; Ziborov, V.S.; Dolgoborodov, A.Y.; Petrov, O.F.; Miakonkikh, A.V.; et al. Micro-Raman Characterization of Structural Features of High-k Stack Layer of SOI Nanowire Chip, Designed to Detect Circular RNA Associated with the Development of Glioma. Molecules 2021, 26, 3715. https://doi.org/10.3390/molecules26123715
Ivanov YD, Malsagova KA, Popov VP, Kupriyanov IN, Pleshakova TO, Galiullin RA, Ziborov VS, Dolgoborodov AY, Petrov OF, Miakonkikh AV, et al. Micro-Raman Characterization of Structural Features of High-k Stack Layer of SOI Nanowire Chip, Designed to Detect Circular RNA Associated with the Development of Glioma. Molecules. 2021; 26(12):3715. https://doi.org/10.3390/molecules26123715
Chicago/Turabian StyleIvanov, Yuri D., Kristina A. Malsagova, Vladimir P. Popov, Igor N. Kupriyanov, Tatyana O. Pleshakova, Rafael A. Galiullin, Vadim S. Ziborov, Alexander Yu. Dolgoborodov, Oleg F. Petrov, Andrey V. Miakonkikh, and et al. 2021. "Micro-Raman Characterization of Structural Features of High-k Stack Layer of SOI Nanowire Chip, Designed to Detect Circular RNA Associated with the Development of Glioma" Molecules 26, no. 12: 3715. https://doi.org/10.3390/molecules26123715
APA StyleIvanov, Y. D., Malsagova, K. A., Popov, V. P., Kupriyanov, I. N., Pleshakova, T. O., Galiullin, R. A., Ziborov, V. S., Dolgoborodov, A. Y., Petrov, O. F., Miakonkikh, A. V., Rudenko, K. V., Glukhov, A. V., Smirnov, A. Y., Usachev, D. Y., Gadzhieva, O. A., Bashiryan, B. A., Shimansky, V. N., Enikeev, D. V., Potoldykova, N. V., & Archakov, A. I. (2021). Micro-Raman Characterization of Structural Features of High-k Stack Layer of SOI Nanowire Chip, Designed to Detect Circular RNA Associated with the Development of Glioma. Molecules, 26(12), 3715. https://doi.org/10.3390/molecules26123715









