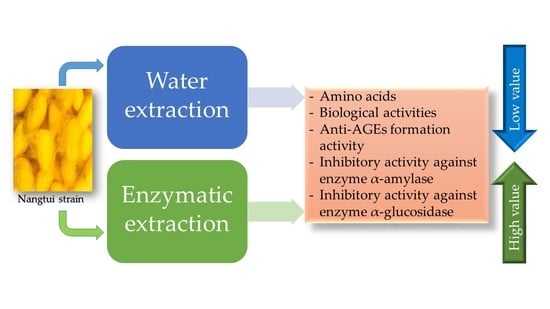
Amino Acid Profile and Biological Properties of Silk Cocoon as Affected by Water and Enzyme Extraction
Abstract
1. Introduction
2. Results and Discussion
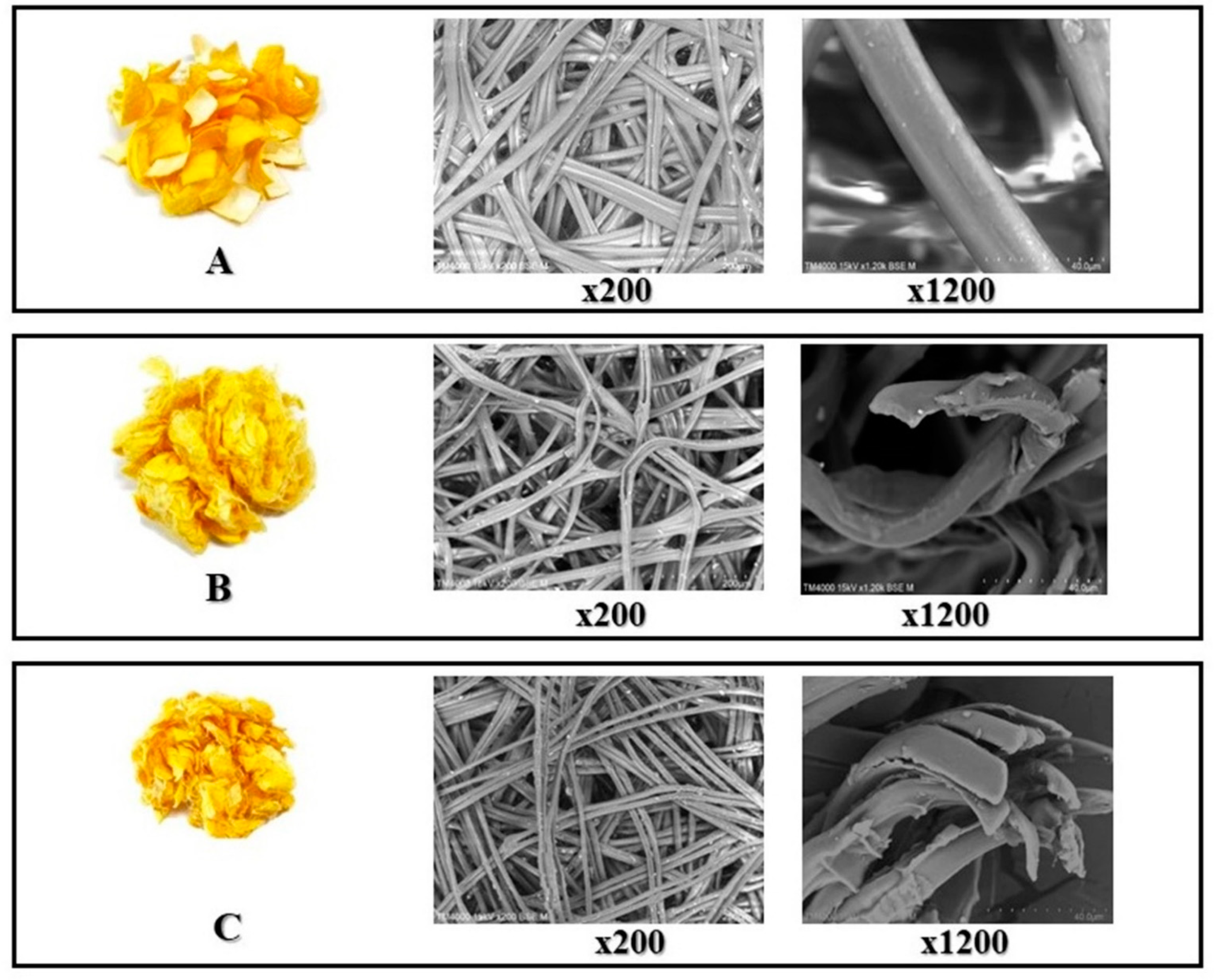
2.1. Microstructure of Silk Fibers
2.2. The Difference Due to Extraction Method on the Amino-Acid Content of Silk Protein Extract (SPE) and Protein Content
2.3. The Effect of Extraction Method on Amino-Acid Contributions to Taste
2.4. The Effect of Extraction Method on Total Phenolic Content and Total Flavonoid Content
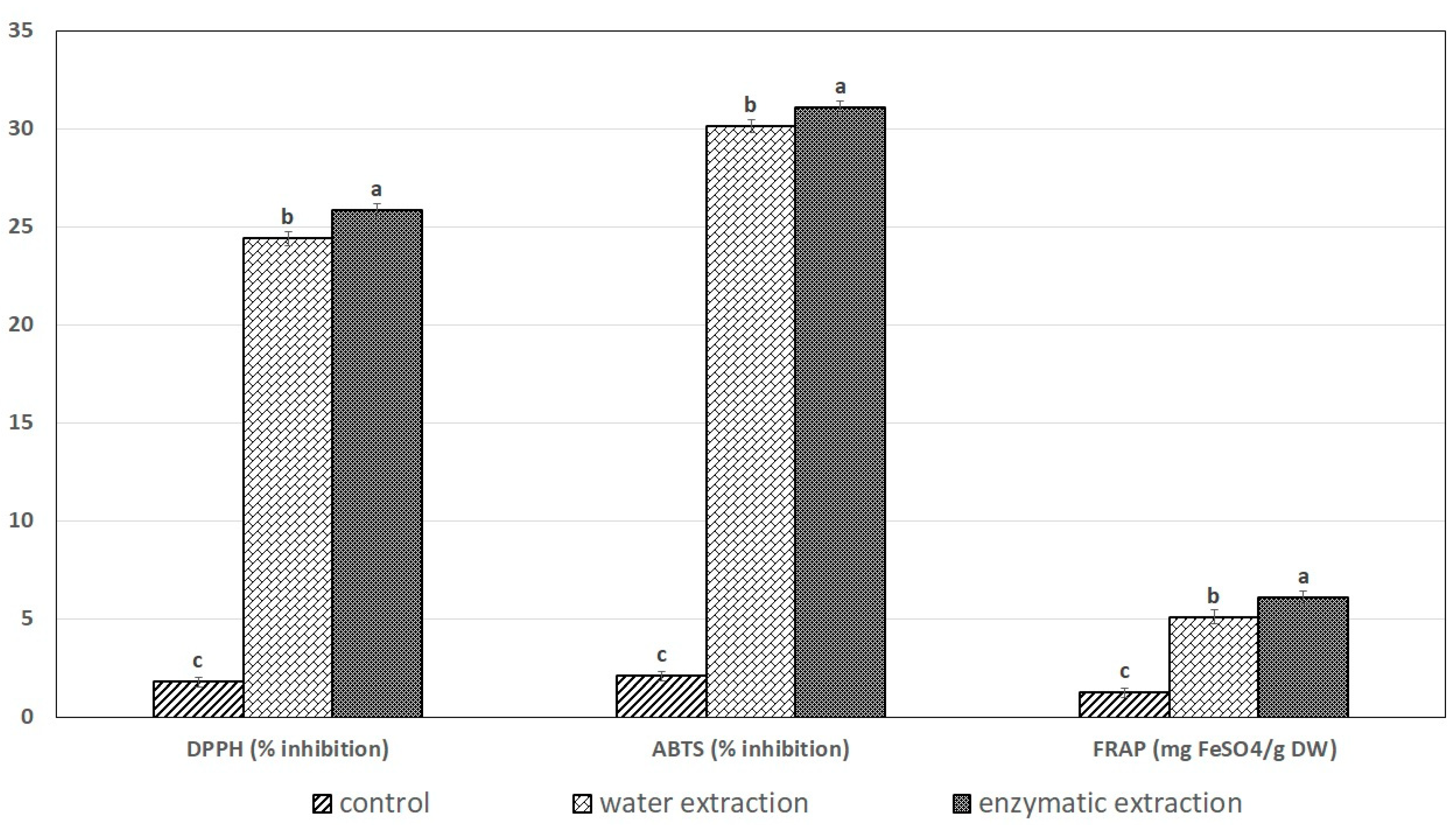
2.5. The Effects of Exteaction Method on DPPH Radical-Scavenging Activity, ABTS+• and the FRAP Assay
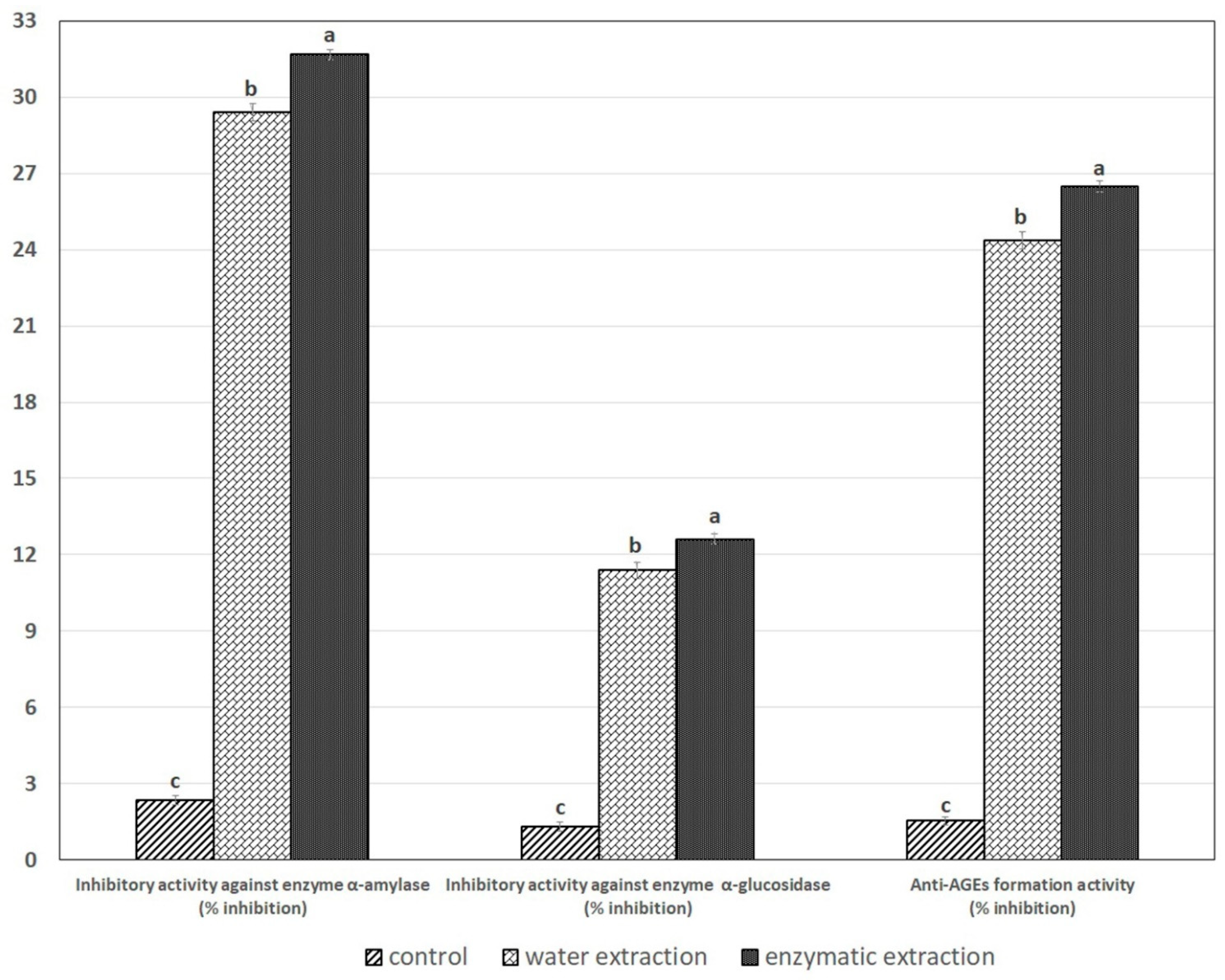
2.6. Inhibitory Activity against the α-Amylase, α-Glucosidase and Anti-AGEs Formation Activity in SPE
3. Materials and Methods
3.1. Silk Cocoons
3.2. Chemicals and Reagents
3.3. Extraction Methods
3.4. Microstructure of Silk Fibers
3.5. Protein Determination
3.6. Amino-Acid Content by LCMS/MS
3.7. Total Phenolic Content (TPC)
3.8. Total Flavonoid Content (TFC)
3.9. DPPH Radical Scavenging Activity
3.10. Antioxidant Activity by ABTS Assay
3.11. Ferric Reducing/Antioxidant Power Assay (FRAP)
3.12. Inhibitory Activity against Enzyme α-Amylase
3.13. Inhibitory Activity against Enzyme α-Glucosidase
3.14. Evaluation of Anti-AGEs Formation Activity
3.15. Statistical Analysis
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Sample Availability
References
- Koh, L.D.; Cheng, Y.; Teng, C.P.; Khin, Y.W.; Loh, X.J.; Tee, S.Y.; Low, M.; Ye, E.; Yu, H.-D.; Zhang, Y.-W.; et al. Structures, mechanical properties and applications of silk fibroin materials. Prog. Polym. Sci. 2015, 46, 86–110. [Google Scholar] [CrossRef]
- Aghaz, F.; Hajarian, H.; Shabankareh, H.K.; Abdolmohammadi, A. Effect of sericin supplementation in maturation medium on cumulus cell expansion, oocyte nuclear maturation, and subsequent embryo development in Sanjabi ewes during the breeding season. Theriogenology 2015, 84, 1631–1635. [Google Scholar] [CrossRef] [PubMed]
- Sothornvit, R.; Chollakup, R.; Suwanruji, P. Extracted sericin from silk waste for film formation. Songklanakarin J. Sci. Technol. 2010, 32, 17–22. [Google Scholar]
- Züge, L.C.B.; Silva, V.R.; Hamerski, F.; Ribani, M.; Gimenes, M.L.; Scheer, A.P. Emulsifying properties of sericin obtained from hot water degumming process. J. Food Process Eng. 2017, 40, 1–10. [Google Scholar] [CrossRef]
- Cao, T.-T.; Zhang, Y.-Q. Processing and characterization of silk sericin from Bombyx mori and its application in biomaterials and biomedicines. Mater. Sci. Eng. 2016, 61, 940–952. [Google Scholar] [CrossRef] [PubMed]
- Lamboni, L.; Gauthier, M.; Yang, G.; Wang, Q. Silk sericin: A versatile material for tissue engineering and drug delivery. Biotechnol. Adv. 2015, 33, 1855–1867. [Google Scholar] [CrossRef]
- Srnivas, N.; Kumar, R.; Merchant, M. Extraction & characterization of sericin and its immobilization on hydroxylapatite nanoparticles for tissue engineering applications. Int. J. Chemtech. Res. 2014, 7, 2017–2124. [Google Scholar]
- Suktham, K.; Koobkokkruad, T.; Wutikhun, T.; Surassmo, S. Efficiency of resveratrol-loaded sericin nanoparticles: Promising bionanocarriers for drug delivery. Int. J. Pharm. 2018, 537, 48–56. [Google Scholar] [CrossRef]
- Puangphet, A.; Tiyaboonchai, W.; Thongsook, T. Inhibitory effect of sericin hydrolysate on polyphenol oxidase and browning of fresh-cut products. Int. Food Res. J. 2015, 22, 1623–1630. [Google Scholar]
- Bungthong, C.; Siriamornpun, S. Changes in amino acid profiles and bioactive compounds of Thai silk cocoons as affected by water extraction. Molecules 2021, 26, 2033. [Google Scholar] [CrossRef]
- Yang, Y.; Lee, S.M.; Lee, H.S.; Lee, K.H. Recovery of silk sericin from soap: Alkaline degumming solution. Int. J. Ind. Entomol. 2013, 27, 203–208. [Google Scholar] [CrossRef]
- Zhao, Z.L.; Li, W.W.; Wang, F.; Zhang, Y.Q. Using of hydrated lime water as a novel degumming agent of silk and sericin recycling from wastewater. J. Clean. Prod. 2018, 172, 2090–2096. [Google Scholar] [CrossRef]
- Mahmoodi, N.M.; Arami, M.; Mazaheri, F.; Rahimi, S. Degradation of sericin (degumming) of Persian silk by ultrasound and enzymes as a cleaner and environmentally friendly process. J. Clean. Prod. 2010, 18, 146–151. [Google Scholar] [CrossRef]
- Silva, V.R.; Ribani, M.; Gimenes, M.L.; Scheer, A.P. High molecular weight sericin obtained by high temperature and ultrafiltration process. Procedia Eng. 2012, 42, 833–841. [Google Scholar] [CrossRef]
- Rocha, L.K.H.; Favaro, L.I.L.; Rios, A.C.; Silva, E.C.; Silva, W.F.; Stigliani, T.P.; Guilger, M.; Lima, R.; Oliveira Junior, J.M.; Aranha, N.; et al. Sericin from Bombyx mori cocoons. Part I: Extraction and physicochemical biological characterization for biopharmaceutical applications. Process Biochem. 2017, 61, 163–177. [Google Scholar] [CrossRef]
- Muniglia, L.; Claisse, N.; Baudelet, P.H.; Ricochon, G. Enzymatic aqueous extraction (EAE). Green Chem. Sustain. Technol. 2014, 167–204. [Google Scholar] [CrossRef]
- Sangwong, G.; Sumida, M.; Sutthikhum, V. Antioxidant activity of chemically and enzymatically modified sericin extracted from cocoons of Bombyx mori. Biocatal. Agric. Biotechnol. 2016, 5, 155–161. [Google Scholar] [CrossRef]
- Freddi, G.; Mossotti, R.; Innocenti, R. Degumming of silk fabric with several proteases. J. Biotechnol. 2003, 106, 101–112. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.; Zhu, Y.; Shi, Z.; Jiang, W.; Liu, X.; Ni, Q.Q. Degumming of raw silk via steam treatment. J. Clean. Prod. 2018, 203, 492–497. [Google Scholar] [CrossRef]
- Yakul, K.; Takenaka, S.; Nakamura, K.; Techapun, C.; Leksawasdi, N.; Seesuriyachan, P.; Watanabe, M.; Chaiyaso, T. Characterization of thermostable alkaline protease from Bacillus halodurans SE5 and its application in degumming coupled with sericin hydrolysate production from yellow cocoon. Process Biochem. 2019, 78, 63–70. [Google Scholar] [CrossRef]
- Wongpinyochit, T.; Totten, J.D.; Johnston, B.F.; Seib, F.P. Microfluidic-assisted silk nanoparticle tuning. Nanoscale Adv. 2019. [Google Scholar] [CrossRef]
- Singh, L.R.; Devi, Y.R.; Devi, S.K. Enzymological characterization of pineapple extract for potential application in oak tasar (Antheraea proylei J.) silk cocoon cooking and reeling. Electron. J. Biotechnol. 2003, 6, 198–207. [Google Scholar] [CrossRef]
- Nakpathom, M.; Somboon, B.; Narumol, N. Papain enzymatic degumming of Thai Bombyx mori silk fibers. Microsc. Microanal. Res. 2009, 23, 142–146. [Google Scholar]
- Pandey, J.P.; Mishra, P.K.; Dinesh, K.; Sinha, A.K.; Prasad, B.C.; Singh, B.M.; Paul, T.K. Possible efficacy of 26 kDa Antheraea mylitta Cocoonase in Cocoon-cooking. Int. J. Biol. Chem. 2011, 5, 215–226. [Google Scholar] [CrossRef]
- Kristinsson, H.G.; Rasco, B.A. Biochemical and functional properties of Atlantic salmon (Salmo salar) muscle proteins hydrolyzed with various alkaline proteases. J. Agric. Food Chem. 2000, 48, 657–666. [Google Scholar] [CrossRef] [PubMed]
- Adler-Nissen, J. Relationship of structure to taste of peptides and peptide mixtures. In Protein Tailoring for Food and Medical Uses; Freeney, R.E., Whitaker, J.R., Eds.; Marcel Dekker: New York, NY, USA, 1986; pp. 97–112. [Google Scholar]
- Ness, A.R.; Powles, J.W. Fruit and vegetables, and cardiovascular disease: A review. Int. J. Epidemiol. 1997, 26, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Halliwell, B. Antioxidant defense mechanisms: From the beginning to the end. Free Radic. Res. 1999, 31, 261–272. [Google Scholar] [CrossRef]
- Chaovanalikit, A.; Mingmuang, A.; Kitbunluewit, T.; Choldumrongkool, N.; Sondee, J.; Chupratum, S. Anthocyanin and total phenolics content of mangosteen and effect of processing on the quality of mangosteen products. Int. Food Res. J. 2012, 19, 1047–1053. [Google Scholar]
- Kumar, J.P.; Mandal, B.B. Antioxidant potential of mulberry and non-mulberry silk sericin and its implications in biomedicine. Free Radic. Biol. Med. 2017, 108, 803–818. [Google Scholar] [CrossRef]
- Li, B.B.; Smith, B.; Hossain, M.M. Extraction of phenolics from citrus peels: II. Enzyme-assisted extraction method. Sep. Purif. Technol. 2006, 48, 189–196. [Google Scholar] [CrossRef]
- Wanyo, P.; Meeso, N.; Siriamornpun, S. Effects of different treatments on the antioxidant properties and phenolic compounds of rice bran and rice husk. Food Chem. 2014, 157, 457–463. [Google Scholar] [CrossRef] [PubMed]
- Zambrowicz, A.; Pokora, M.; Eckert, E.; Szołtysik, M.; Dąbrowska, A.; Chrzanowska, J.; Trziszka, T. Antioxidant and antimicrobial activity of lecithin free egg yolk protein preparation hydrolysates obtained with digestive enzymes. Funct. Foods Health Dis. 2012, 2, 487–500. [Google Scholar] [CrossRef]
- Suzuki, Y.; Aoki, R.; Hayashi, H. Assignment of a p-nitrophenylα-D-glucopyranoside-hydrolyzing α-glucosidase of Bacillus cereus ATCC 7064 to an exo-oligo-1,6-glucosidase. Proteins 1982, 704, 476–483. [Google Scholar] [CrossRef]
- Hara, Y.; Honda, M. Inhibition of rat small intestinal sucrase and α-glucosidase activities by tea polyphenol. Biosci. Biotechnol. Biochem. 1992, 57, 123–124. [Google Scholar] [CrossRef]
- Kim, J.S.; Kwon, C.S.; Son, K.H. Inhibition of α-glucosidase and amylase by luteolin, a flavonoid. Biosci. Biotechnol. Biochem. 2000, 64, 2458–2461. [Google Scholar] [CrossRef] [PubMed]
- Matsui, T.; Ueda, T.; Oki, T.; Sugita, K.; Terahara, N.; Matsumoto, K. α-Glucosidase inhibitory action of natural acylated anthocyanins. 1. Survey of natural pigments with potent inhibitory activity. J. Agric. Food Chem. 2001, 49, 1948–1951. [Google Scholar] [CrossRef]
- Ardestani, A.; Yazdanparast, R. Cyperus rotundus suppresses AGE formation and protein oxidation in a model of fructose-mediated protein glycoxidation. Int. J. Biol. Macromol. 2007, 41, 572–578. [Google Scholar] [CrossRef]
- Peng, X.; Zheng, Z.; Cheung, K.W.; Shan, F.; Ren, G.X.; Chen, S.F.; Wang, M. Inhibitory effect of mung bean extract and its constituents vitexin and isovitexin on the formation of advanced glycation endproducts. Food Chem. 2008, 106, 457–481. [Google Scholar] [CrossRef]
- Gugliucci, A.; Bastos, D.H.; Schulze, J.; Souza, M.F. Caffeic and chlorogenic acids in Ilex paraguariensis extracts are the main inhibitors of AGE generation by methylglyoxal in model proteins. Fitoterapia 2009, 80, 339–344. [Google Scholar] [CrossRef]
- Landis-Piwowar, K.R.; Huo, C.; Chen, D.; Milacic, V.; Shi, G.; Chan, T.H.; Dou, Q.P. A novel prodrug of the green tea polyphenol (−)-epigallocatechin-3-gallate as a potential anticancer agent. Cancer Res. 2007, 67, 4303–4310. [Google Scholar] [CrossRef]
- Mandel, S.; Weinreb, O.; Amit, T.; Youdim, B.H. Cell signaling pathways in the neuroprotective actions of the green tea polyphenol (-)-epigallocatechin-3-gallate: Implications for neurodegenerative diseases. J. Neurochem. 2004, 88, 1555–1569. [Google Scholar] [CrossRef]
- Kowluru, R.A.; Kanwar, M. Effects of curcumin on retinal oxidative stress and inflammation in diabetes. Nutr. Metab. 2007, 4, 8. [Google Scholar] [CrossRef] [PubMed]
- Saiga, A.; Tanabe, S.; Nishimura, T. Antioxidant activity of peptides obtained from porcine myofibrillar proteins by protease treatment. J. Agric. Food Chem. 2003, 51, 3661–3667. [Google Scholar] [CrossRef]
- Zainol, M.K.; Abdhamid, A.; Yusof, S.; Muse, R. Antioxidative activity and total phenolic compounds of leaf, root and petiole of four accessions of Centella asiatica (L.) Urban. Food Chem. 2003, 81, 575–581. [Google Scholar] [CrossRef]
- Chen, H.M.; Muramoto, K.; Yamauchi, F.; Nokihara, K. Antioxidant Activity of Designed Peptides Based on the Antioxidative Peptide Isolated from Digests of a Soybean Protein. J. Agric. Food Chem. 2014, 44, 2619–2623. [Google Scholar] [CrossRef]
- Vaithanomsat, P.; Punyasawon, C. Process optimization for the production of Philosamia ricini (Eri Silk) pupae hydrolysate. Kasetsart J. (Nat. Sci.) 2008, 42, 341–352. [Google Scholar]
- Chumroenphat, T.; Somboonwatthanakul, I.; Saensouk, S.; Siriamornpun, S. The diversity of biologically active compounds in the rhizomes of recently discovered zingiberaceae plants native to north eastern Thailand. Pharmacogn. J. 2019, 11, 1014–1022. [Google Scholar] [CrossRef]
- Bradford, M.M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 1976, 72, 248–254. [Google Scholar] [CrossRef]
- Chumroenphat, T.; Somboonwatthanakul, I.; Saensouk, S.; Siriamornpun, S. Changes in curcuminoids and chemical components of turmeric (Curcuma longa L.) under freeze-drying and low-temperature drying methods. Food Chem. 2021, 339, 128121. [Google Scholar] [CrossRef]
- Kubola, J.; Siriamornpun, S. Phenolic contents and antioxidant activities of bitter gourd (Momordica charantia L.) leaf, stem and fruit fraction extracts in vitro. Food Chem. 2008, 110, 881–890. [Google Scholar] [CrossRef] [PubMed]
- Brand-Williams, W.; Cuvelier, M.; Berset, C. Use of a free radical method to evaluate antioxidant activity. LWT 1995, 28, 25–30. [Google Scholar] [CrossRef]
- Re, R.; Pellegrini, N.; Proteggente, A.; Pannala, A.; Yang, M.; Rice-Evans, C. Antioxidant activity applying an improved ABTS radical cation decolorization assay. Free Radic. Biol. Med. 1999, 26, 1231–1237. [Google Scholar] [CrossRef]
- Benzie, I.F.F.; Strain, J.J. The ferric reducing ability of plasma (FRAP) as a measure of “antioxidant power”: The FRAP assay. Anal. Biochem. 1996, 239, 70–76. [Google Scholar] [CrossRef] [PubMed]
- Xiao, Z.; Storms, R.; Tsang, A. A quantitative starch–iodine method for measuring alpha-amylase and glucoamylase activities. Anal. Biochem. 2006, 351, 146–148. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.-J.; Zhao, J.-L. Corn silk (Zea mays L.), a source of natural antioxidants with α-amylase, α-glucosidase, advanced glycation and diabetic nephropathy inhibitory activities. Biomed. Pharmacother. 2019, 110, 510–517. [Google Scholar] [CrossRef]
- Vinson, J.A.; Howard, T.B. Inhibition of protein glycation and advanced glycation end products by ascorbic acid and other vitamins and nutrients. J. Nutr. Biochem. 1996, 7, 659–663. [Google Scholar] [CrossRef]



| Parameters | Extraction Method | |||
|---|---|---|---|---|
| Control | Water | Enzyme | ||
| Amino acid content (µg/g DW) Essential amino acids | Phenylalanine | 1.39 ± 0.06 c | 10.37 ± 0.17 b | 35.11 ± 0.12 a |
| Valine | 2.04 ± 0.09 c | 5.31 ± 0.08 b | 30.60 ± 0.16 a | |
| Tryptophan | 1.73 ± 0.15 c | 3.34 ± 0.27 b | 28.28 ± 0.28 a | |
| Threonine | 1.44 ± 0.09 c | 24.41 ± 0.17 b | 32.21 ± 0.21 a | |
| Isoleucine | 0.88 ± 0.02 c | 2.86 ± 0.11 b | 18.80 ± 0.13 a | |
| Methionine | 0.53 ± 0.07 c | 2.72 ± 0.08 b | 6.96 ± 0.12 a | |
| Histidine | 0.17 ± 0.06 c | 10.60 ± 0.13 b | 15.57 ± 0.33 a | |
| Arginine | 0.60 ± 0.07 c | 5.39 ± 0.15 b | 43.47 ± 0.55 a | |
| Lysine | 1.39 ± 0.17 c | 3.42 ± 0.19 b | 21.16 ± 0.07 a | |
| Leucine | 0.46 ± 0.09 c | 2.53 ± 0.11 b | 12.67 ± 0.10 a | |
| Non-essential amino acids | Glycine | 2.93 ± 0.06 c | 36.45 ± 0.34 b | 54.55 ± 0.36 a |
| Glutamic acid | 0.92 ± 0.04 c | 22.66 ± 0.23 b | 30.68 ± 0.18 a | |
| Aspartic acid | 1.86 ± 0.07 c | 26.10 ± 0.18 b | 31.72 ± 0.25 a | |
| Glutamine | 1.90 ± 0.04 c | 22.28 ± 0.05 b | 39.70 ± 0.14 a | |
| Serine | 1.98 ± 0.06 c | 10.15 ± 0.11 b | 31.17 ± 0.16 a | |
| Tyrosine | 0.73 ± 0.05 c | 3.26 ± 0.19 b | 38.18 ± 0.12 a | |
| Alanine | 0.83 ± 0.05 c | 5.27 ± 0.08 b | 28.77 ± 0.18 a | |
| Asparagine | 1.14 ± 0.09 c | 5.81 ± 0.17 b | 16.98 ± 0.09 a | |
| Total amino acids | 22.87 ± 0.11 c | 202.96 ± 0.21 b | 516.61 ± 0.32 a | |
| Protein content (mg/g) | 0.84 ± 0.07 c | 2.15 ± 0.05 b | 3.26 ± 0.08 a | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bungthong, C.; Wrigley, C.; Sonteera, T.; Siriamornpun, S. Amino Acid Profile and Biological Properties of Silk Cocoon as Affected by Water and Enzyme Extraction. Molecules 2021, 26, 3455. https://doi.org/10.3390/molecules26113455
Bungthong C, Wrigley C, Sonteera T, Siriamornpun S. Amino Acid Profile and Biological Properties of Silk Cocoon as Affected by Water and Enzyme Extraction. Molecules. 2021; 26(11):3455. https://doi.org/10.3390/molecules26113455
Chicago/Turabian StyleBungthong, Chuleeporn, Colin Wrigley, Thanathat Sonteera, and Sirithon Siriamornpun. 2021. "Amino Acid Profile and Biological Properties of Silk Cocoon as Affected by Water and Enzyme Extraction" Molecules 26, no. 11: 3455. https://doi.org/10.3390/molecules26113455
APA StyleBungthong, C., Wrigley, C., Sonteera, T., & Siriamornpun, S. (2021). Amino Acid Profile and Biological Properties of Silk Cocoon as Affected by Water and Enzyme Extraction. Molecules, 26(11), 3455. https://doi.org/10.3390/molecules26113455