Quinolone Complexes with Lanthanide Ions: An Insight into their Analytical Applications and Biological Activity
Abstract
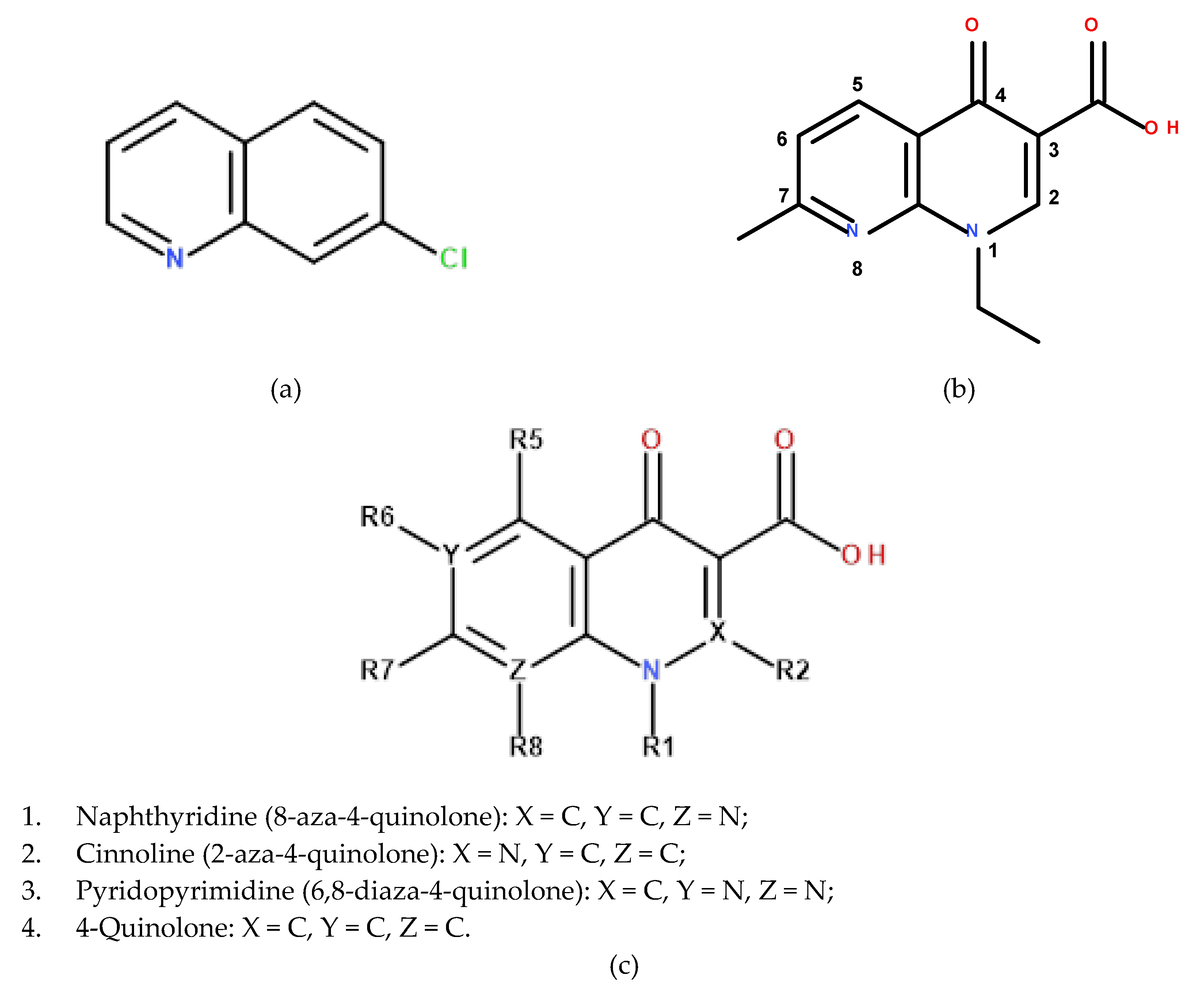
1. Introduction
2. Metal Complexes of Quinolones with Lanthanide Ions
3. Biological Activity of Metal Complexes of Quinolones with Lanthanide Ions
4. Analytical Applications of Metal Complexes of Quinolones with Lanthanide Ions
4.1. Quantitative Determination of Quinolones
| a. | FQ-Tb3+-Ce4+-SO3−-H2SO4 system: | |
| Ce4+ + HSO3- → Ce3+ + HSO3* 2HSO3* → S2O62- + 2H+ S2O62− → SO42- + SO2* | SO2* + Ln(FQ)23+ → SO2 + Ln(FQ*)23+ Ln(FQ*)23+ → Ln*(FQ)23+ Ln*(FQ)23+ → Ln(FQ)23+ + hν, FQ = levofloxacin, moxifloxacin, trovafloxacin [110]; | |
| b. | FQ-Ce4+-S2O42−-HNO3 system: | |
| S2O32−/S2O42− + H+ → HSO3− + H2O Ce4+ + HSO3− → HSO3∗ + Ce3+ 2HSO3∗ → S2O62− + 2H+ S2O62− → SO42− + SO2∗ | SO2∗ → SO2 + hν Ce4+ +FQ + H+ → Ce3+ + FQ oxide + H2O SO2∗ +FQ oxide → FQ oxide∗ + SO2 FQ oxide∗ → FQ oxide + hν, FQ = norfloxacin [109]; | |
| c. | Ln+3-FQ-Ce4+-S2O42− system: | |
| S2O42−/S2O32− + H+→ HSO3− Ce4+ + HSO3−→ HSO3∗ + Ce3+ 2HSO3∗ → S2O62− + 2H+ S2O62− → SO42− + SO2∗ | SO2* + [Ln–FQ]3+→SO2 + [Ln–FQ*]3+ [Ln–FQ∗]3+→ [Ln∗–FQ]3+ [Ln∗–FQ]3+→ [Ln–FQ]3++hν, FQ = gatifloxacin, enoxacin, Ln = Eu3+, Dy3+ [109]. |
4.2. Quantitative Determination of Lanthanide and Miscellaneous Ions
4.3. Determination of Nucleic Acids
4.4. Other Applications
5. Conclusions
Funding
Conflicts of Interest
References
- Zhang, G.-F.; Zhang, S.; Pan, B.; Liu, X.; Feng, L.-S. 4-Quinolone derivatives and their activities against Gram positive pathogens. Eur. J. Med. Chem. 2018, 143, 710–723. [Google Scholar] [CrossRef]
- Kocsis, B.; Domokos, J.; Szabo, D. Chemical structure and pharmacokinetics of novel quinolone agents represented by avarofloxacin, delafloxacin, finafloxacin, zabofloxacin and nemonoxacin. Ann. Clin. Microbiol. Antimicrob. 2016, 15, 34. [Google Scholar] [CrossRef] [PubMed]
- Dhiman, P.; Arora, N.; Thanikachalam, P.V.; Monga, V. Recent advances in the synthetic and medicinal perspective of quinolones: A review. Bioorg. Chem. 2019, 92, 103291. [Google Scholar] [CrossRef] [PubMed]
- Lesher, G.Y.; Froelich, E.J.; Gruett, M.D.; Bailey, J.H.; Brundage, R.P. 1,8-Naphthyridine derivatives. A new class of chemotherapeutic agents. J. Med. Pharm. Chem. 1962, 91, 1063–1065. [Google Scholar] [CrossRef] [PubMed]
- Ball, P.; Fernald, A.; Tillotson, G. Therapeutic advances of new fluoroquinolones. Expert Opin. Investig. Drugs 1998, 7, 761–783. [Google Scholar] [CrossRef] [PubMed]
- Pham, T.D.M.; Ziora, Z.M.; Blaskovich, M.A.T. Quinolone antibiotics. Medchemcomm 2019, 10, 1719–1739. [Google Scholar] [CrossRef] [PubMed]
- Andriole, V.T. The quinolones: Past, present, and future. Clin. Infect. Dis. 2005, 41, S113–S119. [Google Scholar] [CrossRef]
- Uivarosi, V. Metal Complexes of Quinolone Antibiotics and Their Applications: An Update. Molecules 2013, 18, 11153–11197. [Google Scholar] [CrossRef]
- Nagaraja, V.; Godbole, A.A.; Henderson, S.R.; Maxwell, A. DNA topoisomerase I and DNA gyrase as targets for TB therapy. Drug Discov. Today 2017, 22, 510–518. [Google Scholar] [CrossRef]
- Charrier, C.; Salisbury, A.-M.; Savage, V.J.; Duffy, T.; Moyo, E.; Chaffer-Malam, N.; Ooi, N.; Newman, R.; Cheung, J.; Metzger, R.; et al. Novel Bacterial Topoisomerase Inhibitors with Potent Broad-Spectrum Activity against Drug-Resistant Bacteria. Antimicrob. Agents Chemother. 2017, 61, e02100-16. [Google Scholar] [CrossRef]
- Kato, J.; Nishimura, Y.; Imamura, R.; Niki, H.; Hiraga, S.; Suzuki, H. New topoisomerase essential for chromosome segregation in E. coli. Cell 1990, 63, 393–404. [Google Scholar] [CrossRef]
- Idowu, T.; Schweizer, F. Ubiquitous Nature of Fluoroquinolones: The Oscillation between Antibacterial and Anticancer Activities. Antibiotics 2017, 6, 26. [Google Scholar] [CrossRef] [PubMed]
- Papillon, J.; Ménétret, J.-F.; Batisse, C.; Hélye, R.; Schultz, P.; Potier, N.; Lamour, V. Structural insight into negative DNA supercoiling by DNA gyrase, a bacterial type 2A DNA topoisomerase. Nucleic Acids Res. 2013, 41, 7815–7827. [Google Scholar] [CrossRef] [PubMed]
- Klostermeier, D. Why Two? On the Role of (A-)Symmetry in Negative Supercoiling of DNA by Gyrase. Int. J. Mol. Sci. 2018, 19, 1489. [Google Scholar] [CrossRef]
- Ashley, R.E.; Dittmore, A.; McPherson, S.A.; Turnbough, C.L., Jr.; Neuman, K.C.; Osheroff, N. Activities of gyrase and topoisomerase IV on positively supercoiled DNA. Nucleic Acids Res. 2017, 45, 9611–9624. [Google Scholar] [CrossRef]
- Wang, J.C. Cellular roles of DNA topoisomerases: A molecular perspective. Nat. Rev. Mol. Cell Biol. 2002, 3, 430–440. [Google Scholar] [CrossRef]
- Correia, S.; Poeta, P.; Hébraud, M.; Capelo, J.L.; Igrejas, G. Mechanisms of quinolone action and resistance: Where do we stand? J. Med. Microbiol. 2017, 66, 551–559. [Google Scholar] [CrossRef]
- Blower, T.R.; Williamson, B.H.; Kerns, R.J.; Berger, J.M. Crystal structure and stability of gyrase-fluoroquinolone cleaved complexes from Mycobacterium tuberculosis. Proc. Natl. Acad. Sci. USA 2016, 113, 1706–1713. [Google Scholar] [CrossRef]
- Aldred, K.J.; Kerns, R.J.; Osheroff, N. Mechanism of quinolone action and resistance. Biochemistry 2014, 53, 1565–1574. [Google Scholar] [CrossRef]
- Takács-Novák, K.; Noszál, B.; Hermecz, I.; Keresztúri, G.; Podányi, B.; Szász, G. Protonation equilibria of quinolone antibacterials. J. Pharm. Sci. 1990, 79, 1023–1028. [Google Scholar] [CrossRef]
- Regiel-Futyra, A.; Dąbrowski, J.M.; Mazuryk, O.; Śpiewak, K.; Kyzioł, A.; Pucelik, B.; Brindell, M.; Stochel, G. Bioinorganic antimicrobial strategies in the resistance era. Coord. Chem. Rev. 2017, 351, 76–117. [Google Scholar] [CrossRef]
- Debnath, A.; Mogha, N.K.; Masram, D.T. Metal complex of the first-generation quinolone antimicrobial drug nalidixic acid: Structure and its biological evaluation. Appl. Biochem. Biotechnol. 2015, 175, 2659–2667. [Google Scholar] [CrossRef] [PubMed]
- Kouris, E.; Kalogiannis, S.; Perdih, F.; Turel, I.; Psomas, G. Cobalt(II) complexes of sparfloxacin: Characterization, structure, antimicrobial activity and interaction with DNA and albumins. J. Inorg. Biochem. 2016, 163, 18–27. [Google Scholar] [CrossRef] [PubMed]
- Joshi, R.; Pandey, N.; Tilak, R.; Yadav, S.K.; Mishra, H.; Pokharia, S. New triorganotin(IV) complexes of quinolone antibacterial drug sparfloxacin: Synthesis, structural characterization, DFT studies and biological activity. Appl. Organomet. Chem. 2018, 32, e4324. [Google Scholar] [CrossRef]
- El-Megharbel, S.M.; Hegab, M.S.; Manaaa, E.-S.A.; Al-Humaidi, J.Y.; Refat, M.S. Synthesis and physicochemical characterizations of coordination between palladium(II) metal ions with floroquinolone drugs as medicinal model against cancer cells: Novel metallopharmaceuticals. New J. Chem. 2018, 42, 9709–9719. [Google Scholar] [CrossRef]
- Boughougal, A.; Cherchali, F.Z.; Messai, A.; Attik, N.; Decoret, D.; Hologne, M.; Sanglar, C.; Pilet, G.; Tommasino, J.B.; Luneau, D. New model of metalloantibiotic: Synthesis, structure and biological activity of a zinc(II) mononuclear complex carrying two enrofloxacin and sulfadiazine antibiotics. New J. Chem. 2018, 42, 15346–15352. [Google Scholar] [CrossRef]
- Elshafie, H.S.; Sakr, S.H.; Sadeek, S.A.; Camele, I. Biological Investigations and Spectroscopic Studies of New Moxifloxacin/Glycine-Metal Complexes. Chem. Biodivers. 2019, 16, e1800633. [Google Scholar] [CrossRef]
- Feio, M.J.; Sousa, I.; Ferreira, M.; Cunha-Silva, L.; Saraiva, R.G.; Queirós, C.; Alexandre, J.G.; Claro, V.; Mendes, A.; Ortiz, R.; et al. Fluoroquinolone–metal complexes: A route to counteract bacterial resistance? J. Inorg. Biochem. 2014, 138, 129–143. [Google Scholar] [CrossRef]
- Dudev, T.; Chang, L.-Y.; Lim, C. Factors governing the substitution of La3+ for Ca2+ and Mg2+ in metalloproteins: A DFT/CDM study. J. Am. Chem. Soc. 2005, 127, 4091–4103. [Google Scholar] [CrossRef]
- Hermann, P.; Kotek, J.; Kubíček, V.; Lukeš, I. Gadolinium(III) complexes as MRI contrast agents: Ligand design and properties of the complexes. Dalt. Trans. 2008, 3027–3047. [Google Scholar] [CrossRef]
- Xiao, Y.-D.; Paudel, R.; Liu, J.; Cong, M.; Zhang, Z.-S.; Zhou, S.-K. MRI contrast agents: Classification and application (Review). Int. J. Mol. Med. 2016, 38, 1319–1326. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.-K.; Lee, G.H.; Chang, Y. Gadolinium as an MRI contrast agent. Future Med. Chem. 2018, 10, 639–661. [Google Scholar] [CrossRef] [PubMed]
- Montgomery, C.P.; New, E.J.; Parker, D.; Peacock, R.D. Enantioselective regulation of a metal complex in reversible binding to serum albumin: dynamic helicity inversion signalled by circularly polarised luminescence. Chem. Commun. 2008, 4261–4263. [Google Scholar] [CrossRef]
- Singh, G. Chemistry of Lanthanides and Actinides; Singh, G., Ed.; Discovery Publishing House: New Delhi, India, 2007; ISBN 81-8356-241-8. [Google Scholar]
- Cotton, S. Coordination Chemistry of the Lanthanides. In Lanthanide and Actinide Chemistry; Wiley Online Books; John Wiley & Sons, Ltd.: Hoboken, NJ, USA, 2006; pp. 35–60. ISBN 9780470010082. [Google Scholar]
- Yakubovich, Y.Y.; Alekseev, V.G. Hydrolysis constants of tervalent lanthanum and lanthanide ions in 0.1 M KNO3 solution. Russ. J. Inorg. Chem. 2012, 57, 911–915. [Google Scholar] [CrossRef]
- Lawrance, G.A. Complexes and Commerce. In Introduction to Coordination Chemistry; Lawrance, G.A., Ed.; John Wiley & Sons (Asia) Pte Ltd.: Chichester, UK, 2009; pp. 251–267. ISBN 978-0-470-51930-1. [Google Scholar]
- Chen, Z.; Xu, H. Near-Infrared (NIR) Luminescence from Lanthanide(III) Complexes. In Rare Earth Coordination Chemistry; Huang, C., Ed.; John Wiley & Sons (Asia) Pte Ltd.: Chichester, UK, 2010; pp. 473–527. ISBN 978-0-470-82485-6. [Google Scholar]
- Yang, L.; Tao, D.; Yang, X.; Li, Y.; Guo, Y. Synthesis, characterization, and antibacterial activities of some rare Earth metal complexes of pipemidic acid. Chem. Pharm. Bull. (Tokyo) 2003, 51, 494–498. [Google Scholar] [CrossRef] [PubMed]
- Návia, M.; Gálico, D.; Caires, F.; Luiz, J.; Nunes, R. Synthesis, thermal behavior, and spectroscopic study of the solid nalidixate of selected light trivalent lanthanides. J. Therm. Anal. Calorim. 2018, 132, 1717–1727. [Google Scholar] [CrossRef]
- Li, J.-B.; Yang, P.; Gao, F.; Han, G.-Y.; Yu, K.-B. Novel Lanthanide Complexes of Ciprofloxacin: Synthesis, Characterization, Crystal Structure and in vitro Antibacterial Activity Studies. Chin. J. Chem. 2001, 19, 598–605. [Google Scholar] [CrossRef]
- Čurman, D.; Živec, P.; Leban, I.; Turel, I.; Polishchuk, A.; Klika, K.D.; Karaseva, E.; Karasev, V. Spectral properties of Eu(III) compound with antibacterial agent ciprofloxacin (cfqH). Crystal structure of [Eu(cfqH)(cfq)(H2O)4]Cl2·4.55H2O. Polyhedron 2008, 27, 1489–1496. [Google Scholar] [CrossRef]
- Shaban, S.; El-Kemary, M.; Samir, G.; Elbaradei, H. Synthesis, characterization, antibacterial activities testing and the interaction of DNA with ciprofloxacin and its La(III)-based complex. J. Chin. Adv. Mater. Soc. 2018, 6, 123–133. [Google Scholar] [CrossRef]
- Wang, Y.-J.; Hu, R.-D.; Jiang, D.-H.; Zhang, P.-H.; Lin, Q.-Y.; Wang, Y.-Y. Synthesis, crystal structure, interaction with BSA and antibacterial activity of La(III) and Sm(III) complexes with enrofloxacin. J. Fluoresc. 2011, 21, 813–823. [Google Scholar] [CrossRef]
- Tonghuan, L.; Lu, H.; Xi, P.; Liu, X.; Xu, Z.; Chen, F.; Zeng, Z. Synthesis, Characterization, and DNA-binding Properties of Ln(III) Complexes Containing Gatifloxacin. J. Biomol. Struct. Dyn. 2009, 26, 293–300. [Google Scholar]
- Chen, M.; Zheng, Y.; Gao, J.; Wang, Q. Novel europium (III)-gatifloxacin complex structure with dual functionality for pH sensing and metal recognition in aqueous environment. Opt. Mater. 2016, 60, 1–5. [Google Scholar] [CrossRef]
- Sadeek, S.; El-Hamid, S.; Elaasser, M. Synthesis, characterization, antimicrobial and cytotoxicity studies of some transition metal complexes with gemifloxacin. Mon. Chem. Chem. Mon. 2015, 146, 1967–1982. [Google Scholar] [CrossRef]
- Sadeek, S.; El-Hamid, S. Synthesis, spectroscopic, thermal analysis and in vitro biological properties of some new metal complexes with gemifloxacin and 1,10-phenanthroline. J. Therm. Anal. Calorim. 2015, 124, 547–562. [Google Scholar] [CrossRef]
- Sakr, S.H.; Elshafie, H.S.; Camele, I.; Sadeek, S.A. Synthesis, Spectroscopic, and Biological Studies of Mixed Ligand Complexes of Gemifloxacin and Glycine with Zn(II), Sn(II), and Ce(III). Molecules 2018, 23, 1182. [Google Scholar] [CrossRef]
- Sadeek, S.A.; El-Shwiniy, W.H. Preparation, structural characterization and biological studies of some new levofloxacin metal complexes. J. Iran. Chem. Soc. 2017, 14, 1711–1723. [Google Scholar] [CrossRef]
- Sadeek, S.; El-Shwiniy, W.; El-Attar, M. Synthesis, characterization and antimicrobial investigation of some moxifloxacin metal complexes. Spectrochim. Acta A Mol. Biomol. Spectrosc. 2011, 84, 99–110. [Google Scholar] [CrossRef]
- Li, S.; Wang, Y.; Lin, Q.; Liu, W.; Ding, J.; Wang, Y. Synthesis, crystal structures of novel complexes of rare earth with norfloxacin, interaction with DNA and BSA. J. Rare Earths 2012, 30, 460–466. [Google Scholar] [CrossRef]
- Refat, M.; El-Hawary, W.; Mohamed, M. Study of the chemical chelates and anti-microbial effect of some metal ions in nanostructural form on the efficiency of antibiotic therapy “norfloxacin drug”. J. Mol. Struct. 2012, 1013, 45–54. [Google Scholar] [CrossRef]
- Shaban, S.; El-Kemary, M.; Samir, G.; El-Baradei, G.; Puchta, R. Norfloxacin La(III)-based complex: Synthesis, characterization and DNA binding studies. J. Coord. Chem. 2015, 68, 3247–3258. [Google Scholar] [CrossRef]
- Xu, M.; Chen, F.-J.; Huang, L.; Xi, P.; Zeng, Z. Binding of rare earth metal complexes with an ofloxacin derivative to bovine serum albumin and its effect on the conformation of protein. J. Lumin. 2011, 131, 1557–1565. [Google Scholar] [CrossRef]
- Xu, M.; Ma, Z.R.; Huang, L.; Chen, F.J.; Zeng, Z. Spectroscopic studies on the interaction between Pr(III) complex of an ofloxacin derivative and bovine serum albumin or DNA. Spectrochim. Acta A Mol. Biomol. Spectrosc. 2011, 78, 503–511. [Google Scholar] [CrossRef] [PubMed]
- El-Gamel, N.E.A.; Zayed, M.A. Synthesis, structural characterization and antimicrobial activity evaluation of metal complexes of sparfloxacin. Spectrochim. Acta A Mol. Biomol. Spectrosc. 2011, 82, 414–423. [Google Scholar] [CrossRef] [PubMed]
- Xu, M.; Zhang, Y.; Xu, Z.; Zeng, Z. Crystal structure, biological studies of water-soluble rare earth metal complexes with an ofloxacin derivative. Inorg. Chim. Acta 2012, 384, 324–332. [Google Scholar] [CrossRef]
- Andreu, V.; Blasco, C.; Pico, Y. Analytical strategies to determine quinolone residues in food and the environment. TrAC Trends Anal. Chem. 2007, 26, 534–556. [Google Scholar] [CrossRef]
- Prabhakaran, D.; Sukul, P.; Lamshoft, M.; Maheswari, M.A.; Zuhlke, S.; Spiteller, M. Photolysis of difloxacin and sarafloxacin in aqueous systems. Chemosphere 2009, 77, 739–746. [Google Scholar] [CrossRef]
- Kotzerke, A.; Hammesfahr, U.; Kleineidam, K.; Lamshöft, M.; Sören, T.-B.; Schloter, M.; Wilke, B.-M. Influence of difloxacin-contaminated manure on microbial community structure and function in soils. Biol. Fertil. Soils 2010, 47, 177–186. [Google Scholar] [CrossRef]
- Lopez de Alda, M. Environmental Behavior and Analysis of Veterinary and Human Drugs in Soils, Sediments and Sludge. TrAC Trends Anal. Chem. 2003, 22, 340–351. [Google Scholar]
- Kummerer, K. Drugs in the environment: emission of drugs, diagnostic aids and disinfectants into wastewater by hospitals in relation to other sources—A review. Chemosphere 2001, 45, 957–969. [Google Scholar] [CrossRef]
- Tamtam, F.; Mercier, F.; Le Bot, B.; Eurin, J.; Tuc Dinh, Q.; Clement, M.; Chevreuil, M. Occurrence and fate of antibiotics in the Seine River in various hydrological conditions. Sci. Total Environ. 2008, 393, 84–95. [Google Scholar] [CrossRef]
- Li, Y.L.; Hao, X.L.; Ji, B.Q.; Xu, C.L.; Chen, W.; Shen, C.Y.; Ding, T. Rapid determination of 19 quinolone residues in spiked fish and pig muscle by high-performance liquid chromatography (HPLC) tandem mass spectrometry. Food Addit. Contam. Part A Chem. Anal. Control Exp. Risk Assess. 2009, 26, 306–313. [Google Scholar] [CrossRef] [PubMed]
- Espinosa-Mansilla, A.; de la Pena, A.M.; Gomez, D.G.; Salinas, F. HPLC determination of enoxacin, ciprofloxacin, norfloxacin and ofloxacin with photoinduced fluorimetric (PIF) detection and multiemission scanning: application to urine and serum. J. Chromatogr. B Anal. Technol. Biomed. Life Sci. 2005, 822, 185–193. [Google Scholar] [CrossRef] [PubMed]
- Watabe, S.; Yokoyama, Y.; Nakazawa, K.; Shinozaki, K.; Hiraoka, R.; Takeshita, K.; Suzuki, Y. Simultaneous measurement of pazufloxacin, ciprofloxacin, and levofloxacin in human serum by high-performance liquid chromatography with fluorescence detection. J. Chromatogr. B Anal. Technol. Biomed. Life Sci. 2010, 878, 1555–1561. [Google Scholar] [CrossRef] [PubMed]
- Horstkotter, C.; Blaschke, G. Stereoselective determination of ofloxacin and its metabolites in human urine by capillary electrophoresis using laser-induced fluorescence detection. J. Chromatogr. B Biomed. Sci. Appl. 2001, 754, 169–178. [Google Scholar] [CrossRef]
- Hernandez, M.; Aguilar, C.; Borrull, F.; Calull, M. Determination of ciprofloxacin, enrofloxacin and flumequine in pig plasma samples by capillary isotachophoresis—Capillary zone electrophoresis. J. Chromatogr. B Anal. Technol. Biomed. Life Sci. 2002, 772, 163–172. [Google Scholar] [CrossRef]
- D’Agostino, P.A.; Hancock, J.R.; Chenier, C.L.; Lepage, C.R.J. Liquid chromatography electrospray tandem mass spectrometric and desorption electrospray ionization tandem mass spectrometric analysis of chemical warfare agents in office media typically collected during a forensic investigation. J. Chromatogr. A 2006, 1110, 86–94. [Google Scholar] [CrossRef]
- Ghoneim, M.M.; Radi, A.; Beltagi, A.M. Determination of Norfloxacin by square-wave adsorptive voltammetry on a glassy carbon electrode. J. Pharm. Biomed. Anal. 2001, 25, 205–210. [Google Scholar] [CrossRef]
- Tamer, A. Adsorptive stripping voltammetric determination of ofloxacin. Anal. Chim. Acta 1990, 231, 129–131. [Google Scholar] [CrossRef]
- Jaber, A.M.Y.; Lounici, A. Polarographic behaviour and determination of norfloxacin in tablets. Anal. Chim. Acta 1994, 291, 53–64. [Google Scholar] [CrossRef]
- Kilic, E.; Koseoglu, F.; Akay, M.A. The non-aqueous titrimetric assay of selected antibiotics using tetra-N-butylammonium hydroxide as titrant. J. Pharm. Biomed. Anal. 1994, 12, 347–352. [Google Scholar] [CrossRef]
- El-Brashy, A.M.; El-Sayed Metwally, M.; El-Sepai, F.A. Spectrophotometric determination of some fluoroquinolone antibacterials by binary complex formation with xanthene dyes. Farmaco 2004, 59, 809–817. [Google Scholar] [CrossRef] [PubMed]
- Mostafa, S.; El-Sadek, M.; Alla, E.A. Spectrophotometric determination of ciprofloxacin, enrofloxacin and pefloxacin through charge transfer complex formation. J. Pharm. Biomed. Anal. 2002, 27, 133–142. [Google Scholar] [CrossRef]
- Kaur, K.; Singh, B.; Malik, A. Chemiluminescence and Spectrofluorimetric Methods for Determination of Fluoroquinolones: A Review. Anal. Lett. 2011, 44, 1602–1639. [Google Scholar] [CrossRef]
- Sun, C.; Ping, H.; Zhang, M.; Li, H.; Guan, F. Spectroscopic studies on the lanthanide sensitized luminescence and chemiluminescence properties of fluoroquinolone with different structure. Spectrochim. Acta A Mol. Biomol. Spectrosc. 2011, 82, 375–382. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Fan, R.; Wang, P.; Wang, X.; Gao, S.; Dong, Y.; Wang, Y.; Yang, Y. Structure variations of a series of lanthanide complexes constructed from quinoline carboxylate ligands: Photoluminescent properties and PMMA matrix doping. RSC Adv. 2015, 5, 38254–38263. [Google Scholar] [CrossRef]
- Georges, J. Lanthanide-sensitized luminescence and applications to the determination of organic analytes. A review. Analyst 1993, 118, 1481–1486. [Google Scholar] [CrossRef]
- Gómez-Hens, A.; Aguilar-Caballos, M. Terbium-sensitized luminescence: A selective and versatile analytical approach. TrAC Trends Anal. Chem. 2002, 21, 131–141. [Google Scholar] [CrossRef]
- Junker, A.K.R.; Sørensen, T.J. Illuminating the Intermolecular vs. Intramolecular Excited State Energy Transfer Quenching by Europium(III) Ions. Eur. J. Inorg. Chem. 2019, 2019, 1201–1206. [Google Scholar] [CrossRef]
- Izosimov, I.N. Time-resolved Laser-induced Luminescence/Chemiluminescence Laser Spectroscopy and Detection of Actinides/Lanthanides. In Environmental Radiochemical Analysis VI; The Royal Society of Chemistry: London, UK, 2020; pp. 115–130. ISBN 978-1-78801-735-0. [Google Scholar]
- Singha, S.; Ahn, K.H. Detection of Ciprofloxacin in Urine through Sensitized Lanthanide Luminescence. Sensors 2016, 16, 2065. [Google Scholar] [CrossRef]
- Veiopoulou, C.J.; Ioannou, P.C.; Lianidou, E.S. Application of terbium sensitized fluorescence for the determination of fluoroquinolone antibiotics pefloxacin, ciprofloxacin and norfloxacin in serum. J. Pharm. Biomed. Anal. 1997, 15, 1839–1844. [Google Scholar] [CrossRef]
- Rodríguez-Díaz, R.; Aguilar-Caballos, M.; Gómez-Hens, A. Sensitive Determination of Fluoroquinolone Antibiotics in Milk Samples Using Time-Resolved Methodology. Anal. Lett. 2011, 37, 1163–1175. [Google Scholar] [CrossRef]
- Ocana, J.A.; Callejon, M.; Barragan, F.J. Terbium-sensitized luminescence determination of levofloxacin in tablets and human urine and serum. Analyst 2000, 125, 1851–1854. [Google Scholar] [CrossRef] [PubMed]
- Ocana, J.A.; Callejon, M.; Barragan, F.J. Application of terbium-sensitized luminescence for the determination of grepafloxacin in human urine and serum. J. Pharm. Sci. 2001, 90, 1553–1557. [Google Scholar] [CrossRef] [PubMed]
- Ocaña, J.A.; Barragán, F.J.; Callejón, M. Fluorescence and terbium-sensitised luminescence determination of garenoxacin in human urine and serum. Talanta 2004, 63, 691–697. [Google Scholar] [CrossRef] [PubMed]
- Shtykov, S.N. Chemical Analysis in Nanoreactors: Main Concepts and Applications. J. Anal. Chem. 2002, 57, 859–868. [Google Scholar] [CrossRef]
- Shtykov, S.N.; Smirnova, T.D.; Bylinkin, Y.G.; Kalashnikova, N.V.; Zhemerichkin, D.A. Determination of ciprofloxacin and enrofloxacin by the sensitized fluorescence of europium in the presence of the second ligand and micelles of anionic surfactants. J. Anal. Chem. 2007, 62, 136–140. [Google Scholar] [CrossRef]
- Tan, H.; Zhang, L.; Ma, C.; Song, Y.; Xu, F.; Chen, S.; Wang, L. Terbium-based coordination polymer nanoparticles for detection of ciprofloxacin in tablets and biological fluids. ACS Appl. Mater. Interfaces 2013, 5, 11791–11796. [Google Scholar] [CrossRef]
- Xu, X.; Feng, L.; Li, J.; Yuan, P.; Feng, J.; Wei, L.; Cheng, X. Rapid screening detection of fluoroquinolone residues in milk based on turn-on fluorescence of terbium coordination polymer nanosheets. Chin. Chem. Lett. 2018, 30, 549–552. [Google Scholar] [CrossRef]
- Liu, B.; Huang, Y.; Shen, Q.; Zhu, X.; Hao, Y.; Qu, P.; Xu, M. Turn-on fluorescence detection of ciprofloxacin in tablets based on lanthanide coordination polymer nanoparticles. RSC Adv. 2016, 6, 100743–100747. [Google Scholar] [CrossRef]
- Kamruzzaman, M.; Alam, A.-M.; Lee, S.; Suh, Y.; Kim, Y.H.; Kim, G.M.; Kim, H. Method for determination of fluoroquinolones based on the plasmonic interaction between their fluorescent terbium complexes and silver nanoparticles. Microchim. Acta 2011, 174, 353–360. [Google Scholar] [CrossRef]
- Kaur, B.; Kumar, R.; Chand, S.; Singh, K.; Malik, A.K. Determination of norfloxacin in urine and pharmaceutical samples using terbium doped zinc sulphide nanomaterials-sensitized fluorescence method. Spectrochim. Acta A Mol. Biomol. Spectrosc. 2019, 214, 261–268. [Google Scholar] [CrossRef] [PubMed]
- Yanez-Jacome, G.S.; Aguilar-Caballos, M.P.; Gomez-Hens, A. Luminescent determination of quinolones in milk samples by liquid chromatography/post-column derivatization with terbium oxide nanoparticles. J. Chromatogr. A 2015, 1405, 126–132. [Google Scholar] [CrossRef] [PubMed]
- Ding, F.; Zhao, H.; Jin, L.; Zheng, D. Study of the influence of silver nanoparticles on the second-order scattering and the fluorescence of the complexes of Tb(III) with quinolones and determination of the quinolones. Anal. Chim. Acta 2006, 566, 136–143. [Google Scholar] [CrossRef]
- Lee, S.H.; Wabaidur, S.M.; Alothman, Z.A.; Alam, S.M. Gold nanoparticles-based fluorescence enhancement of the terbium-levofloxacin system and its application in pharmaceutical preparations. Luminescence 2011, 26, 768–773. [Google Scholar] [CrossRef]
- Wei, L.; Cheng, X.; Lin, J.-M.; Cai, H.; Huang, F. Chemiluminescence mechanisms of cerium-norfloxacin and its application in urine analysis. Chem. Pap. 2009, 63, 358–365. [Google Scholar] [CrossRef]
- Zhu, Y.; Shu, G.; Yang, Y.; Dong, Q.; Zou, L. Synthesis and electrochemiluminescence properties of a new ternary terbium complex and its application for the determination of norfloxacin. J. Electroanal. Chem. 2014, 727, 113–119. [Google Scholar] [CrossRef]
- Jeon, C.; Khan, M.; Lee, S.; Karim, M.; Lee, H.; Suh, Y.; Alam, S.; Chung, H. Optical Flow-Through Sensor for the Determination of Norfloxacin Based on Emission of KMnO4–Na2SO3–Tb3+ System. J. Fluoresc. 2008, 18, 843–851. [Google Scholar] [CrossRef]
- Chen, L.; Wang, X.; Zhao, H.; Wang, K.; Jin, L. Flow-injection chemiluminescence determination of ofloxacin using the Ru(bpy)2(CIP)2+-Ce(IV) system and its application. Luminescence 2008, 23, 309–315. [Google Scholar] [CrossRef]
- Sun, H.; Li, L.; Wu, Y. Dysprosium-sensitized chemiluminescence reactions: Their mechanism and application to the determination of synthetic quinolone antibiotics. J. Anal. Chem. 2011, 66, 720. [Google Scholar] [CrossRef]
- Ocana-Gonzalez, J.A.; Ramos-Payan, M.; Fernandez-Torres, R.; Callejon-Mochon, M.; Bello-Lopez, M.A. Hollow-fiber liquid-phase microextraction for the direct determination of flumequine in urban wastewaters by flow-injection analysis with terbium-sensitized chemiluminescence. J. Sep. Sci. 2014, 37, 2738–2744. [Google Scholar] [CrossRef]
- Chen, S.; Ding, F.; Liu, Y.; Zhao, H. Electrochemiluminescence of terbium (III)-two fluoroquinolones-sodium sulfite system in aqueous solution. Spectrochim. Acta A Mol. Biomol. Spectrosc. 2006, 64, 130–135. [Google Scholar] [CrossRef]
- Murillo Pulgarín, J.A.; Alañón Molina, A.; Jiménez García, E. Simplex optimization of the variables influencing the determination of pefloxacin by time-resolved chemiluminescence. Spectrochim. Acta Part A Mol. Biomol. Spectrosc. 2018, 193, 117–124. [Google Scholar] [CrossRef] [PubMed]
- Lozano, V.A.; Tauler, R.; Ibanez, G.A.; Olivieri, A.C. Standard addition analysis of fluoroquinolones in human serum in the presence of the interferent salicylate using lanthanide-sensitized excitation-time decay luminescence data and multivariate curve resolution. Talanta 2009, 77, 1715–1723. [Google Scholar] [CrossRef] [PubMed]
- Sun, H.; Li, L.; Chen, X. Investigation on sensitized chemiluminescence systems and their mechanism for five fluoroquinolones. Anal. Chim. Acta 2006, 576, 192–199. [Google Scholar] [CrossRef] [PubMed]
- Ocaña, J.A.; Barragán, F.J.; Callejón, M.; De la Rosa, F. Application of Lanthanide-Sensitised Chemiluminescence to the Determination of Levofloxacin, Moxifloxacin and Trovafloxacin in Tablets. Microchim. Acta 2004, 144, 207–213. [Google Scholar] [CrossRef]
- Wang, L.; Guo, C.; Chu, Z.; Jiang, W. Luminescence enhancement effect for the determination of balofloxacin with balofloxacin–europium (III)–sodium dodecylbenzene sulfonate system. J. Lumin. 2009, 129, 90–94. [Google Scholar] [CrossRef]
- Cai, H.; Zhao, F.; Si, H.; Zhang, S.; Wang, C.; Qi, P. Highly sensitive luminescence assessment of bile acid using a balofloxacin-europium(III) probe in micellar medium. Bull. Korean Chem. Soc. 2012, 33, 4145–4149. [Google Scholar] [CrossRef]
- Zhao, F.; Qi, Y.; Xiong, W. Chemiluminescence determination of balofloxacin based on europium (III)-sensitized KBrO3-Na2S2O4 reaction in micellar medium. Bull. Korean Chem. Soc. 2012, 33, 204–208. [Google Scholar] [CrossRef]
- Qi, Y.; Zhao, F.; Xie, X.; Xu, X.; Ma, Z. Study on the cofluorescence effect of europium (III)-yttrium (III)-balofloxacin-sodium dodecyl sulfate system and its analytical application. Spectrosc. Lett. 2015, 48, 311–316. [Google Scholar] [CrossRef]
- Hernández-Arteseros, J.A.; Compañó, R.; Ferrer, R.; Prat, M.D. Application of principal component regression to luminescence data for the screening of ciprofloxacin and enrofloxacin in animal tissues. Analyst 2000, 125, 1155–1158. [Google Scholar] [CrossRef]
- Rieutord, A.; Vazquez, L.; Soursac, M.; Prognon, P.; Blais, J.; Bourget, P.; Mahuzier, G. Fluoroquinolones as sensitizers of lanthanide fluorescence: Application to the liquid chromatographic determination of ciprofloxacin using terbium. Anal. Chim. Acta 1994, 290, 215–225. [Google Scholar] [CrossRef]
- Sierra-Rodero, M.; Fernández-Romero, J.M.; Gómez-Hens, A. Determination of fluoroquinolone antibiotics by microchip capillary electrophoresis along with time-resolved sensitized luminescence of their terbium(III) complexes. Microchim. Acta 2014, 181, 1897–1904. [Google Scholar] [CrossRef]
- Rodríguez-Díaz, R.C.; Aguilar-Caballos, M.; Gómez-Hens, A. Simultaneous determination of ciprofloxacin and tetracycline in biological fluids based on dual-lanthanide sensitised luminescence using dry reagent chemical technology. Anal. Chim. Acta 2003, 494, 55–62. [Google Scholar] [CrossRef]
- Lian, N.; Zhao, H.; Sun, C.; Chen, S.; Lu, Y.; Jin, L. A study on terbium sensitized chemiluminescence of ciprofloxacin and its application. Microchem. J. 2003, 74, 223–230. [Google Scholar] [CrossRef]
- Liu, J.; Chen, K.; Li, B.; Zhu, Y. A novel method to determine ciprofloxacin by enhanced electrochemiluminescence of Tb(bpy)23+–K2S2O8 system in aqueous solution. Anal. Methods 2012, 4, 2355–2360. [Google Scholar] [CrossRef]
- Rizk, M.; Habib, I.H.I.; Mohamed, D.; Mowaka, S. Lanthanide-DNA probe for spectrofluorimetric determination of some 6- fluoroquinolones in eye-ear pharmaceutical preparations. Microchem. J. 2019, 150, 104138. [Google Scholar] [CrossRef]
- Attia, M.; Essawy, A.; Youssef, A. Europium-sensitized and simultaneous pH-assisted spectrofluorimetric assessment of ciprofloxacin, norfloxacin and gatifloxacin in pharmaceutical and serum samples. J. Photochem. Photobiol. A Chem. 2012, 236, 26–34. [Google Scholar] [CrossRef]
- Kamruzzaman, M.; Alam, A.; Kim, K.M.; Lee, S.H.; Suh, Y.S.; Kim, Y.H.; Kim, S.H.; Oh, S.H. Enhanced Luminescence of Lanthanide Complexes by Silver Nanoparticles for Ciprofloxacin Determination. J. Nanosci. Nanotechnol. 2012, 12, 6125–6130. [Google Scholar] [CrossRef]
- Wang, B.; Yan, B. A turn-on fluorescence probe Eu3+ functionalized Ga-MOF integrated with logic gate operation for detecting ppm-level ciprofloxacin (CIP) in urine. Talanta 2020, 208, 120438. [Google Scholar] [CrossRef]
- Sun, H.; Wu, Y.; Li, L. Dysprosium-sensitized chemiluminescence system for the determination of enoxacin in pharmaceutical preparations and biological fluids with flow-injection sampling. Drug Test. Anal. 2009, 1, 128–134. [Google Scholar] [CrossRef]
- Yi, L.; Zhao, H.; Chen, S.; Jin, L.; Zheng, D.; Wu, Z. Flow-injection analysis of two fluoquinolones by the sensitizing effect of terbium(III) on chemiluminescence of the potassium permanganate-sodium sulfite system. Talanta 2003, 61, 403–409. [Google Scholar] [CrossRef]
- Karim, M.M.; Lee, S.H. Determination of enoxacin using Tb composite nanoparticles sensitized luminescence method. J. Fluoresc. 2008, 18, 827–833. [Google Scholar] [CrossRef] [PubMed]
- Attia, M.S.; Sabry, D.; Youssef, A.O. Enrofloxacin Assessment by the Enhancement of the Red Emission of Eu3+ Optical Sensor. Anal. Chem. Lett. 2014, 4, 65–72. [Google Scholar] [CrossRef]
- Zdunek, J.; Benito-Peña, E.; Linares, A.; Falcimaigne-Cordin, A.; Orellana, G.; Haupt, K.; Moreno-Bondi, M.C. Surface-imprinted nanofilaments for europium-amplified luminescent detection of fluoroquinolone antibiotics. Chem. A Eur. J. 2013, 19, 10209–10216. [Google Scholar] [CrossRef]
- Ershadi, S.; Jouyban, A.; Shayanfar, A. Determination of Enrofloxacin in Milk Samples Using Silver Nanoparticle Enhanced Terbium-Sensitized Fluorescence Method. Food Anal. Methods 2017, 10, 3607–3614. [Google Scholar] [CrossRef]
- Kaczmarek, M.; Staninski, K.; Stodolny, M. Oxidation of certain fluoroquinolones using the chemical sources of reactive oxygen species in the presence of Tb(III) ions and analytical application of these systems. J. Lumin. 2019, 215, 116654. [Google Scholar] [CrossRef]
- Schneider, M.J.; Yun, L.; Lehotay, S.J. Terbium-sensitised luminescence screening method for fluoroquinolones in beef serum. Food Addit. Contam. Part A Chem. Anal. Control Exp. Risk Assess. 2013, 30, 666–669. [Google Scholar] [CrossRef]
- Sun, H.; Li, L. Flow-Injection Chemiluminescence Determination of Fleroxacin in Pharmaceutical Preparations and Human Urine. J. Chromatogr. Sep. Tech. 2010, 1, 104. [Google Scholar] [CrossRef]
- Liu, T.Y.; Qu, X.L.; Yan, B. A highly sensitive and selective “turn-on” fluorescent probe for detection of fleroxacin in human serum and urine based on a lanthanide functionalized metal-organic framework. Dalt. Trans. 2019, 48, 17945–17952. [Google Scholar] [CrossRef]
- Yan, W.; Tieli, Z.; Huichun, Z.; Linpei, J. Photochemical fluorescence enhancement of terbium-fleroxacin system and determination of fleroxacin. Anal. Lett. 2000, 33, 3303–3314. [Google Scholar] [CrossRef]
- Rodriguez-Diaz, R.C.; Fernandez-Romero, J.M.; Aguilar-Caballos, M.P.; Gomez-Hens, A. Chromatographic determination of flumequine in food samples by post-column derivatisation with terbium(III). Anal. Chim. Acta 2006, 578, 220–226. [Google Scholar] [CrossRef] [PubMed]
- Smirnova, T.D.; Shtykov, S.N.; Nevryueva, N.V.; Zhemerichkin, D.A.; Parashchenko, I.I. Fluorimetric assay of flumequine using sensitized terbium fluorescence in organized media. Pharm. Chem. J. 2011, 44, 635–638. [Google Scholar] [CrossRef]
- Castillo-García, M.L.; Aguilar-Caballos, M.P.; Gómez-Hens, A. A europium- and terbium-coated magnetic nanocomposite as sorbent in dispersive solid phase extraction coupled with ultra-high performance liquid chromatography for antibiotic determination in meat samples. J. Chromatogr. A 2015, 1425, 73–80. [Google Scholar] [CrossRef] [PubMed]
- Guo, C.; Dong, P.; Chu, Z.; Wang, L.; Jiang, W. Rapid determination of gatifloxacin in biological samples and pharmaceutical products using europium-sensitized fluorescence spectrophotometry. Luminescence 2008, 23, 7–13. [Google Scholar] [CrossRef] [PubMed]
- Attia, M.; Youssef, A.; Essawy, A.; Abdel-Mottaleb, M. A highly luminescent complexes of Eu(III) and Tb(III) with norfloxacin and gatifloxacin doped in sol–gel matrix: A comparable approach of using silica doped Tb(III) and Eu(III) as optical sensor. J. Lumin. 2012, 132, 2741–2746. [Google Scholar] [CrossRef]
- Ocaña, J.A.; Callejón, M.; Barragán, F.J.; De la Rosa, F.F. Lanthanide sensitized chemiluminescence determination of grepafloxacin in tablets and human urine. Anal. Chim. Acta 2003, 482, 105–113. [Google Scholar] [CrossRef]
- El-Hamshary, M.S.; Fouad, M.A.; Hanafi, R.S.; Al-Easa, H.S.; El-Moghazy, S.M. Screening and optimization of samarium-assisted complexation for the determination of norfloxacin, levofloxacin and lomefloxacin in their corresponding dosage forms employing spectrofluorimetry. Spectrochim. Acta A Mol. Biomol. Spectrosc. 2019, 206, 578–587. [Google Scholar] [CrossRef]
- Wang, J.M.; Lian, X.; Yan, B. Eu3+-Functionalized Covalent Organic Framework Hybrid Material as a Sensitive Turn-On Fluorescent Switch for Levofloxacin Monitoring in Serum and Urine. Inorg. Chem. 2019, 58, 9956–9963. [Google Scholar] [CrossRef]
- Wang, L.; Liu, J.; Wang, Z.; Wang, Y. Fluorescence resonance energy transfer between cerium ion(III) and levofloxacin in micellar solution and its analytical application to the determination of levofloxacin. Spectrosc. Lett. 2019, 52, 313–320. [Google Scholar] [CrossRef]
- Nie, L.H.; Zhao, H.C.; Wang, X.; Yi, L.; Lu, Y.; Jin, L.P.; Ma, H.M. Determination of lomefloxacin by terbium sensitized chemiluminescence method. Anal. Bioanal. Chem. 2002, 374, 1187–1190. [Google Scholar] [CrossRef]
- Rodriguez-Diaz, R.C.; Fernandez-Romero, J.M.; Aguilar-Caballos, M.P.; Gomez-Hens, A. Determination of fluoroquinolones in milk samples by postcolumn derivatization liquid chromatography with luminescence detection. J. Agric. Food Chem. 2006, 54, 9670–9676. [Google Scholar] [CrossRef]
- Beltyukova, S.; Teslyuk, O.; Egorova, A.; Tselik, E. Solid-Phase Luminescence Determination of Ciprofloxacin and Norfloxacin in Biological Fluids. J. Fluoresc. 2002, 12, 269–272. [Google Scholar] [CrossRef]
- Chen, S.L.; Liu, Y.; Zhao, H.C.; Jin, L.P.; Zhang, Z.L.; Zheng, Y.Z. Determination of norfloxacin using a terbium-sensitized electrogenerated chemiluminescence method. Luminescence 2006, 21, 20–25. [Google Scholar] [CrossRef] [PubMed]
- Bel’tyukova, S.V.; Malinka, E.V.; Liventsova, E.O. Determination of ofloxacin and norfloxacin in wastewaters of pharmaceutical enterprises using sensitized luminescence of Tb(III) complexes. J. Water Chem. Technol. 2008, 30, 38–41. [Google Scholar]
- Llorent-Martínez, E.; Garcia-Reyes, J.F.; Ortega-Barrales, P.; Molina-Díaz, A. Terbium-sensitized luminescence optosensor for the determination of norfloxacin in biological fluids. Anal. Chim. Acta 2005, 532, 159–164. [Google Scholar] [CrossRef]
- Hu, G.; Sheng, W.; Zhang, Y.; Wu, X.; Wang, S. A novel and sensitive fluorescence immunoassay for the detection of fluoroquinolones in animal-derived foods using upconversion nanoparticles as labels. Anal. Bioanal. Chem. 2015, 407, 8487–8496. [Google Scholar] [CrossRef]
- Zhang, X.; Ouyang, J.; Zhai, S.; Huang, G.; Zhao, H. Flow-injection with enhanced chemiluminescence detection of ofloxacin in human plasma. Luminescence 2005, 20, 362–369. [Google Scholar] [CrossRef]
- Attia, M.S.; Essawy, A.A.; Youssef, A.O.; Mostafa, M.S. Determination of ofloxacin using a highly selective photo probe based on the enhancement of the luminescence intensity of Eu3+--ofloxacin complex in pharmaceutical and serum samples. J. Fluoresc. 2012, 22, 557–564. [Google Scholar] [CrossRef]
- Llorent-Martinez, E.J.; Ortega-Barrales, P.; Molina-Diaz, A.; Ruiz-Medina, A. Implementation of terbium-sensitized luminescence in sequential-injection analysis for automatic analysis of orbifloxacin. Anal. Bioanal. Chem. 2008, 392, 1397–1403. [Google Scholar] [CrossRef]
- Chen, S.; Ma, H.; Zhao, H.; Feng, R.; Jin, L. Terbium-sensitized fluorescence method for the determination of pazufloxacin mesilate and its application. Anal. Sci. 2004, 20, 1075–1078. [Google Scholar] [CrossRef]
- Wang, X.-L.; Chen, S.-L.; Zhao, H.-C.; Jin, L.-P.; Li, X. Europium Sensitized Chemiluminescence Determination of Pazufloxacin Mesylate in Urine and Serum. Anal. Lett. 2005, 38, 971–979. [Google Scholar] [CrossRef]
- Yang, C.; Zhang, Z.; Chen, S.; Yang, F. Molecularly imprinted on-line solid-phase extraction combined with chemiluminescence for the determination of pazufloxacin mesilate. Microchim. Acta 2007, 159, 299–304. [Google Scholar] [CrossRef]
- Li, A.; Song, Z. Study of silver nanoparticles sensitized fluorescence and second-order scattering of terbium(III)-pefloxacin mesylate complex and determination of pefloxacin mesylate. Sci. World J. 2014, 2014, 742935. [Google Scholar] [CrossRef] [PubMed]
- Llorent-Martinez, E.J.; Ortega-Barrales, P.; Molina-Diaz, A. Multicommuted optosensor for the determination of pipemidic acid in biological fluids. Anal. Biochem. 2005, 347, 330–332. [Google Scholar] [CrossRef]
- Wang, X.L.; Li, A.Y.; Zhao, H.C.; Jin, L.P. A lanthanide sensitized chemiluminescence method of flow-injection for the determination of ulifloxacin and prulifloxacin. J. Anal. Chem. 2009, 64, 75–81. [Google Scholar] [CrossRef]
- Yu, F.; Li, L.; Chen, F.; Liu, W. Terbium sensitized chemiluminescence of potassium permanganate-sodium thiosulfateprulifloxacin system and its application. Anal. Lett. 2010, 43, 357–366. [Google Scholar] [CrossRef]
- Dong, P.; Xu, N.; Fu, B.; Wang, L. Rapid europium-sensitized fluorescent determination of ulifloxacin, the active metabolite of prulifloxacin, in human serum and urine. J. Pharm. Anal. 2011, 1, 46–50. [Google Scholar] [CrossRef]
- Davydov, N.; Zairov, R.; Mustafina, A.; Syakayev, V.; Tatarinov, D.; Mironov, V.; Eremin, S.; Konovalov, A.; Mustafin, M. Determination of fluoroquinolone antibiotics through the fluorescent response of Eu(III) based nanoparticles fabricated by layer-by-layer technique. Anal. Chim. Acta 2013, 784, 65–71. [Google Scholar] [CrossRef]
- Wang, N.-X.; Wang, L.; Jiang, W.; Ren, Y.-Z.; Si, Z.-K.; Qiu, X.-X.; Du, G.-Y.; Qi, P. Determination of neodymium, holmium and erbium in mixed rare earths by norfloxacin. Fresenius’ J. Anal. Chem. 1998, 361, 821–824. [Google Scholar] [CrossRef]
- Guo, C.; Lang, A.; Wang, L.; Jiang, W. The co-luminescence effect of a europium (III)–lanthanum (III)–gatifloxacin–sodium dodecylbenzene sulfonate system and its application for the determination of trace amount of europium (III). J. Lumin. 2010, 130, 591–597. [Google Scholar] [CrossRef]
- Turel, M.; Duerkop, A.; Yegorova, A.; Scripinets, Y.; Lobnik, A.; Samec, N. Detection of nanomolar concentrations of copper(II) with a Tb-quinoline-2-one probe using luminescence quenching or luminescence decay time. Anal. Chim. Acta 2009, 644, 53–60. [Google Scholar] [CrossRef] [PubMed]
- Abbasi, Y.; Ali, A.A.; Khan, M.; Saeed, M.; Naeem, K. Liquid-liquid extraction of scandium with nalidixic acid in dichloromethane. J. Radioanal. Nucl. Chem. 2012, 292, 277–283. [Google Scholar] [CrossRef]
- Turel, M.; Duerkop, A.; Yegorova, A.; Karasyov, A.; Scripinets, Y.; Lobnik, A. Microtiterplate phosphate assay based on luminescence quenching of a terbium complex amenable to decay time detection. Anal. Chim. Acta 2010, 675, 42–48. [Google Scholar] [CrossRef] [PubMed]
- Le Pecq, J.B.; Paoletti, C. A new fluorometric method for RNA and DNA determination. Anal. Biochem. 1966, 17, 100–107. [Google Scholar] [CrossRef]
- Rye, H.S.; Yue, S.; Wemmer, D.E.; Quesada, M.A.; Haugland, R.P.; Mathies, R.A.; Glazer, A.N. Stable fluorescent complexes of double-stranded DNA with bis-intercalating asymmetric cyanine dyes: Properties and applications. Nucleic Acids Res. 1992, 20, 2803–2812. [Google Scholar] [CrossRef] [PubMed]
- Wu, T.; Fang, B.; Chang, L.; Liu, M.; Chen, F. Sensitive determination of DNA based on the interaction between prulifloxacin-terbium(III) complex and DNA. Luminescence 2013, 28, 894–899. [Google Scholar] [CrossRef]
- Yegorova, A.; Karasyov, A.; Dürkop, A.; Ukrainets, I.; Antonovich, V. New luminescent terbium complex for the determination of DNA. Spectrochim. Acta A Mol. Biomol. Spectrosc. 2005, 61, 109–116. [Google Scholar] [CrossRef]
- Jenkins, Y.; Friedman, A.E.; Turro, N.J.; Barton, J.K. Characterization of dipyridophenazine complexes of ruthenium(II): The light switch effect as a function of nucleic acid sequence and conformation. Biochemistry 1992, 31, 10809–10816. [Google Scholar] [CrossRef]
- Wang, L.; Guo, C.; Fu, B.; Wang, L. Fluorescence determination of DNA using the gatifloxacin-europium(III) complex. J. Agric. Food Chem. 2011, 59, 1607–1611. [Google Scholar] [CrossRef]
- Yegorova, A.V.; Scripinets, Y.V.; Duerkop, A.; Karasyov, A.A.; Antonovich, V.P.; Wolfbeis, O.S. Sensitive luminescent determination of DNA using the terbium(III)-difloxacin complex. Anal. Chim. Acta 2007, 584, 260–267. [Google Scholar] [CrossRef]
- Chen, Y.; Chi, Y.; Wen, H.; Lu, Z. Sensitized luminescent terbium nanoparticles: preparation and time-resolved fluorescence assay for DNA. Anal. Chem. 2007, 79, 960–965. [Google Scholar] [CrossRef] [PubMed]
- Krasnoperov, L.N.; Marras, S.A.E.; Kozlov, M.; Wirpsza, L.; Mustaev, A. Luminescent Probes for Ultrasensitive Detection of Nucleic Acids. Bioconjug. Chem. 2010, 21, 319–327. [Google Scholar] [CrossRef] [PubMed]
- Juskowiak, B. Nucleic acid-based fluorescent probes and their analytical potential. Anal. Bioanal. Chem. 2011, 399, 3157–3176. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Selvin, P.R. Luminescent Polyaminocarboxylate Chelates of Terbium and Europium: The Effect of Chelate Structure. J. Am. Chem. Soc. 1995, 117, 8132–8138. [Google Scholar] [CrossRef]
- Soltani, N.; Manzoori, J.L.; Amjadi, M.; Lotfipour, F.; Jouyban, A. Development and validation of a spectrofluorimetric determination of calf thymus DNA using a terbium-danofloxacin probe. Pharm. Sci. 2016, 22, 2–8. [Google Scholar] [CrossRef][Green Version]
- Jakovljevic, I.; Petrovic, D.; Joksovic, L.; Lazarevic, I.; Jelikic-Stankov, M.; Djurdjevic, P. Complex formation equilibria between aluminum(III), gadolinium(III) and yttrium(III) ions and some fluoroquinolone ligands. Potentiometric and spectroscopic study. J. Coord. Chem. 2015, 68, 4272–4295. [Google Scholar] [CrossRef]
- Su, Y.; Xu, Y.; Yang, L.; Yang, J.; Weng, S.; Yu, Z.; Wu, J. New, rapid fluorescence stain method for histologic sections using lanthanide complexes. Anal. Biochem. 2005, 347, 89–93. [Google Scholar] [CrossRef]
- Lin, J.; Zheng, Y.; Wang, Q.; Zeng, Z.; Zhang, C.C. Novel lanthanide pH fluorescent probes based on multiple emissions and its visible-light-sensitized feature. Anal. Chim. Acta 2014, 839, 51–58. [Google Scholar] [CrossRef]
- Leyva, S.; Hernández, H. Synthesis of norfloxacin analogues catalyzed by Lewis and Brönsted acids: An alternative pathway. J. Fluor. Chem. 2010, 131, 982–988. [Google Scholar] [CrossRef]
- Polishchuk, A.; Karaseva, É.; Korpela, T.; Karasev, V.E. New luminophor-activators based on (fluoro)quinolone antibacterials. J. Lumin. 2008, 128, 1753–1757. [Google Scholar] [CrossRef]
| Ligand | Complex | Material/Ligand:Ln Molar Ratio/pH Adjustment | Mixing Mode | Heating/Cooling | Ref |
|---|---|---|---|---|---|
| Pipemidic acid (PPA) | [M(PPA)4Cl]Cl2 where M = La3+; [M(PPA)4]Cl3 where M = Ce3+, Pr3+, Nd3+, Sm3+, Tb3+, Dy3+, Y3+. | -ligand salt dissolved in ethanol mixed with PPA dissolved in acetic acid; -4:1; -no pH adjustment. | -refluxed on water bath 3–4 h; -cooling to room temperature. | [39] | |
| Nalidixic Acid (NAL) | [Ln(NAL)3]⋅nH2O n = 5 for La3+ n = 6 for Ce3+, Pr3+, Sm3+. | -aqueous solution of nalidixic salt with Na+, pH = 8.5; -aqueous solution of LnCl3, -pH = 5. | -nalidixate solution added slowly and under continuous stirring. | -heated until boiling, then kept in refrigerator 12 h; -deionized water added and repeated. | [40] |
| Ciprofloxacin (CPX) | [Ln(CPX)2]Cl⋅nH2O, n = 7–9, Ln = Pr3+, Nd3+, Sm3+, Eu3+, Gd3+, Tb3+, Dy3+, Ho3+, Er3+, Tm3+, Yb3+. | -ciprofloxacin HCl, lanthanide oxides; -2:1. | -CPFX dissolved in HCl solution and pyridine; -Ln(III) oxides dissolved in conc. HCl, heated to dryness and dissolved in water; -lanthanide solution dropped into CPFX, continuous stirring (2 h–5 days). | [41] | |
| [Ce(CPX)2(H2O)4]Cl⋅ (H2O)3.25⋅(C2H5OH)0.25 | -ciprofloxacin HCl, CeCl3⋅7H2O; -2:1. | -lanthanide dissolved in water, added to CPFX solution. | -stirring for 7 days at room temp; -20 mL ethanol added, left standing at room temp. | [41] | |
| [Eu(CPXH)(CPX)(H2O)4]Cl2⋅ 4.55H2O | -EuCl3⋅6H2O, ciprofloxacin HCl hydrate, 1,3-bis (dimethylamino)- 2 propranolol; -2:1. | -solvothermal method: substances in mixture of water and ethanol -placed in stainless steel autoclave. | -heating at 150 °C for 48 h, cooling; -isothermal evaporation in the air. | [42] | |
| [La(H2O)4(CPX)2]Cl | -LaCl3, ciprofloxacin; -2:1. | -both substances suspended in methanol, stirred for 10h. | -reflux 8 h. | [43] | |
| Enrofloxacin (EF) | [La2(EF)6(H2O)2]⋅14H2O, [Sm2(EF)6(H2O)2]⋅14H2O | -La2O3, Sm2O3 enrofloxacin, distilled water; -2:1. | -components sealed in 25ml Teflon- lined stainless vessel. | -heated at 435 K, 3 days; -cooled at room temp. | [44] |
| Gatifloxacin (GFLX) | [Ln(GFLX)3Cl3]⋅2H2O, Where Ln = La3+, Nd3+, Eu3+, Tb3+. | -GFLX, LnCl3⋅6H2O; -3:1. | -HGA dissolved in acetone added dropwise into solution of LnCl3⋅6H2O dissolved in ethanol. | -stirred and refluxed for 2 h. | [45] |
| [Eu(GFLX)3⋅2H2O] | -gatifloxacin, EuCl3⋅6H2O; -3:1; -pH = 9 with ammonium hydroxide. | -gatifloxacin and Eu3+ salt dissolved in deionized water, while stirring; -yellow solution obtained; | -stirring 2 h at room temp. | [46] | |
| Gemifloxacin (GMFX) | [La(GMFX)2(H2O)2]Cl3⋅3H2O [Ce(GMFX)2(H2O)2](SO4)2⋅2H2O. | -gemifloxacin, LaCl3⋅7H2O, Ce(SO4)2; -2:1. | -substances dissolved in acetone, metal salt solution added dropwise to the solution of gemifloxacin. | -stirring for 15 h at room temp. | [47] |
| [La(GMFX)(phen)(H2O)2]Cl3⋅6H2O [Ce(GMFX)(phen)(H2O)2] (SO4)2⋅3H2O. | -gemifloxacin, 1,10- phenanthroline, LaCl3⋅7H2O, Ce(SO4)2; -GMFX: metal: phen = 1:1:1. | -mixing hot saturated solution on GMFX in acetone with phen and metal salt. | -reflux for 3 h. | [48] | |
| [Ce(GMFX)(Gly)(H2O)2]Cl2⋅ H2O | -gemifloxacin, glycine, CeCl3⋅7H2O; -GMFX: metal: Gly = 1:1:1. | -metal salt dissolved in ethanol added dropwise to a stirred solution of GMFX and Gly in ethanol; | -reflux and stirring 5–6 h at 100 °C in water bath; | [49] | |
| Levofloxacin (LEVO) | [Ce(LEVO)2(H2O)2]SO4⋅5H2O | -levofloxacin, Ce(SO4)2; -2:1. | -metal salt added to magnetically stirred solution of LEVO and NaOH in methanol and acetone. | -stirring at room temp for 1 day. | [50] |
| Moxifloxacin (MOXI) | [Ce(MOXI)2](SO4)2⋅2H2O | -moxifloxacin, (NH4)4Ce(SO4)4⋅2H2O -2:1. | -ethanolic solution of metal salt added dropwise to a stirred ethanolic solution of moxifloxacin. | -24 h stirring at 50 °C on water bath. | [51] |
| Norfloxacin (NOR) | [N(CH3)4][Ln(NOR)4]⋅6H2O, where Ln = Nd3+, Sm3+, Ho3+; [N(CH3)4] = tetramethylammonium. | -norfloxacin, Ln(NO3)3⋅6H2O, imidazole; -3:1. | -norfloxacin, metallic salt and imidazole mixed with distilled water and sealed in a 25ml Teflon-lined stainless vessel. | -heating at 453 K for 3 days; -cooled slowly to room temp. | [52] |
| [La(NOR)3]⋅3H2O, [Ce(NOR)3]⋅2H2O | -norfloxacin, CeCl3⋅7H2O, LaCl3⋅6H2O; -3:1; -basic pH with ammonia solution 5% v/v. | a) -NOR suspended in methanol mixed with metal salt suspended in distilled water; -solution basified by adding ammonia solution; b) complexes obtained as nanoparticles by the following transformations: metal chlorides→metal carbonates→nano-oxides→metal chlorides→above described method; | a) -reflux at 80–90 °C for 4–5 h; b) -reflux at 80–90 °C for 4–5 h. | [53] | |
| [La(NOR)2Cl2]Cl | -2:1. | -substances suspended in methanol. | -gently stirring for 10 h and reflux for 8 h. | [54] | |
| Ofloxacin (OFLO) | [Pr(L-OFLO)(NO3)2(CH3OH)](NO3), [NdOFLO(NO3)2(CH3OH)](NO3), where L-OFLO = derivative obtained by esterification of ofloxacin. | -ofloxacin, NaOH, Pr(NO3)3⋅6H2O, Nd(NO3)3⋅6H2O; -ligand: NaOH: metal = 0.25: 0.25: 0.3. | -ligand (L) obtained by esterification of ofloxacin with ethanol, treatment with N2H4⋅H2O and refluxed with 2-hydroxybenz-aldehyde; -ligand and NaOH mixed in methanol; metal salt added. | -refluxed and stirred for 3 h; -cooling to room temperature. | [55] |
| [Pr(L-OFLO)2(NO3)](NO3)2, where L-OFLO = ofloxacin derivative. | -ofloxacin, NaOH, Pr(NO3)3⋅6H2O; -ligand: NaOH: metal = 0.25: 0.25: 0.3. | -ligand (L) obtained by esterification of ofloxacin with ethanol, treatment with N2H4⋅H2O and refluxed with 2-hydroxybenz-aldehyde; -ligand and NaOH mixed in methanol; metal salt added. | -stirring 3 hat room temp; | [56] | |
| Sparfloxacin (SPAR) | [La(SPAR)2NO3⋅H2O]⋅2H2O (1), [La(SPAR)(HL)NO3⋅H2O]⋅H2O (2), where L = DL-alanine. | -sparfoxacin, La(NO3)3⋅6H2O, DL-alanine; -sparfloxacin: metal = 2; 1; -sparfloxacin: DL-alanine: metal = 1: 1: 1. | -metal salt dissolved in hot water -ethanolic solution added to a hot ethanolic solution of sparfloxacin and DL-alanin. | -mixture stirred under reflux for 2 h; left to cool. | [57] |
| Ligand | Complex | Biological Activity Test | Results | Ref |
|---|---|---|---|---|
| Pipemidic acid (PPA) | [La(PPA)4Cl]Cl2 | -antibacterial activity on E. coli, S. aureus, P. aeruginosa similar to PPA; | -antibacterial activity on S. pneumoniae much greater than PPA; -NO activity on S. aureus | [39] |
| [M(PPA)4]Cl3 where M = Ce3+, Pr3+, Nd3+, Sm3+, Tb3+, Dy3+, Y3+. | -Pr, Sm, Y complexes have similar activity to PPA against E. coli, S. aureus, B. subtilis, S. pneumoniae; weaker activity on P. aeruginosa. | [39] | ||
| Ciprofloxacin (CPX) | [Er(CPX)2(H2O)8]Cl [Ce(CPX)2(H2O)4]Cl·(H2O)3.25(C2H5OH)0.25 | -MIC determined by broth tube dilution method; E. coli, P. aeruginosa, S. aureus; | -for Ce, the activity is 2.5, 2.5,1.25 fold higher than CPX; -for Er, the activity is 2.5, 1.25 fold higher, resp. 3.0 fold lower than CPX. | [41] |
| [La(H2O)4(CPX)2]Cl | -antibacterial activity against E. coli strains, through flat-filter paper method; | -the complex is less active than ciprofloxacin. | [43] | |
| Enrofloxacin (EF) | [La2(EF)6(H2O)2]⋅14H2O [Sm2(EF)6(H2O)2]⋅14H2O | -antibacterial activity tested against B. subtilis, S. aureus, E. coli through the diffusimetric method; | -both complexes have bactericidal properties greater than the ligand; -the Sm3+ complex is more active than the La3+ complex. | [44] |
| Gemifloxacin (GMFX) | [La(GMFX)2(H2O)2]Cl3⋅3H2O [Ce(GMFX)2(H2O)2] (SO4)2⋅2H2O | -antibacterial activity tested against S. aureus, B. subtilis, E. coli, P. aeruginosa, by diffusimetric method; -antifungal activity tested against C. albicans, A. awamori, Altenaria sp. by diffusimetric method; -cytotoxic activity tested against human breast carcinoma cell line (MCF-7 cells), human colon carcinoma cell line (HCT-116 cells), through crystal violet colorimetric viability assay; | -the activity of the La3+ complex is comparable to gemifloxacin, but the one of the Ce4+ complex is slightly higher; -only the Ce4+ complex is active and only against C. albicans; -results were compared to the activity of doxorubicin; both complexes were found to be active on both cell lines, but have IC50 higher than doxorubicin; Ce4+ complex more active than La3+ complex against the breast carcinoma cell line; La3+ complex more active than the Ce4+ complex against the colon carcinoma cell line. | [47] |
| [La(GMFX)(phen)(H2O)2] Cl3⋅6H2O [Ce(GMFX)(phen)(H2O)2] (SO4)2⋅3H2O | -antibacterial activity against S. aureus, B. subtilis, E. coli, P. aeruginosa, by diffusimetric method; -antifungal activity tested against C. albicans, A. awamori, Alternaria sp. by diffusimetric method; -cytotoxic activity tested against human breast carcinoma cell line (MCF-7 cells) and human colon carcinoma cell line (HCT-116 cells) by crystal violet staining viability assay; doxorubicin used as positive control; | -activity against S. aureus is comparable to GMFX; higher activities against the others; -comparable activity against C. albicans to GFX; no activity against other fungi strains; -complexes show cytotoxic activity, but lower than GMFX; phen also shows cytotoxic activity, but higher than GMFX. | [48] | |
| [Ce(GMFX)(Gly)(H2O)2] Cl2⋅H2O | -antibacterial activity tested against Xanthomonas campestris, Bacillus megeterium, E. coli, Clavibacter michiganesis; -antifungal activity tested against phytopathogenic fungi: Rhizoctonia solani, Sclerotinia sclerotium, Aspergillus niger, Botrytis cinerea, Penicillium digitatum; -antioxidant activity tested through DPPH and ABTS methods; | -GMFX and complex proved to be active against all strains, the weakest activity being against E. coli and the highest against C. michiganensis; -complex shows lower activity than GMFX against all strains; -complex activity comparable to that of GMFX. | [49] | |
| Levofloxacin (LEVO) | [Ce(LEVO)2(H2O)2]SO4⋅ 5H2O | -antibacterial activity tested against S. aureus, B. subtilis, B. otitidis, E. coli, P. aeruginosa, K. oxytoca, by cup-diffusion technique; -antifungal activity tested against A. flaurus, A. fumigatus, using the disc diffusion sensitivity method; | -the Ce4+ proves to be more active on B. subtilis and B. otitidis; -no antifungal activity noted; | [50] |
| Moxifloxacin (MOXI) | [Ce(MOXI)2](SO4)2⋅2H2O | -antibacterial activity tested against S. aureus, B. subtilis, B. otitidis, E. coli, P. aeruginosa, K. oxytoca by cup-diffusion method; | -the complex shows similar activity against E.coli; -no activity against P. aeruginosa and K. oxytoca; -higher activity against B. subtilis, B. otitidis and S. aureus. | [51] |
| Norfloxacin (NOR) | [La(NOR)3]⋅3H2O [Ce(NOR)3]⋅2H2O | -antibacterial activities tested using modified Kirby-Bauer disk diffusion method, against S. aureus, B. subtilis, E. coli, P. aeruginosa, C. albicans, A. flavus; positive controls used: tetracycline and amphotericin; | -complexes in nanoparticle form displayed greater activities than those in normal- particle form, but lower than the positive controls; -La3+ nanocomplex is the most active. | [53] |
| Ofloxacin (OFLO) | [Pr(L-OFLO)(NO3)2(CH3OH)] (NO3) [Nd(L-OFLO)(NO3)2(CH3OH)] (NO3), where L-OFLO = ofloxacin derivative. | -antioxidant activity tested through hydroxyl radical scavenging activity through the Fenton reaction; | -complexes show better activity than the ligand. | [55,58] |
| Sparfloxacin (SPAR) | [La(SPAR)2NO3⋅H2O]⋅2H2O (1) [La(SPAR)(HL)NO3⋅H2O]⋅ H2O (2), where L = DL-alanin. | -antibacterial activity tested against S. aureus, E. coli using modified Kirby-Bauer disc diffusion method; tetracycline used as control; -antifungal activity tested against A. flavus, C. albicans using modified Kirby-Bauer disc diffusion method; amphotericin B used as control; | -the complexes show the same activity as the free ligand, which is higher than the control, against both bacteria; -complexes and free ligand showed no antifungal activity. | [57] |
| Ligand | Complex | Binding of Bovine Serum Albumin/CT-DNA | Results | Ref |
|---|---|---|---|---|
| Ciprofloxacin (CPX) | [La(H2O)4(CPX)2]Cl | -CT-DNA binding properties investigated through UV-VIS spectroscopy and fluorescence quenching methods; | -binding ability of CPX and complex is the highest in basic medium and the lowest in acidic medium. | [43] |
| Enrofloxacin (CF) | [La2(EF)6(H2O)2]⋅14H2O [Sm2(EF)6(H2O)2]⋅14H2O | -BSA binding properties investigated through UV-VIS spectroscopy and fluorescence quenching methods; | -both complexes have the ability to quench the fluorescence of BSA, the Sm3+ complex more than the La3+; -the mechanism is mainly a static one. | [44] |
| Levofloxacin (LEVO) | [Ce(LEVO)2(H2O)2]SO4⋅5H2O | -CT-DNA degradation by testing electrophoretic mobility; | -the complex degrades the DNA completely compared to the metal salt. | [50] |
| Norfloxacin (NOR) | [N(CH3)4][Ln(NOR)4]⋅ 6H2O, where Ln = Nd3+, Sm3+, Ho3+; | -CT-DNA binding investigated by UV absorbance of complex in the presence of increasing amount of CT-DNA and by emission spectra of EB-DNA; | -the complex binds to CT-DNA stronger than norfloxacin; -interaction with BSA tested by measuring fluorescence quenching spectra; -complexes have strong binding ability; -the quenching process is a static one. | [52] |
| [La(NOR)2Cl2]Cl | -binding to CT-DNA, using UV-VIS absorption spectroscopy and time-resolved fluorescence spectroscopy; | -the complex shows moderate interaction with CT-DNA, by partial or non-intercalative binding modes. | [54] | |
| Ofloxacin (OFLO) | [Pr(L-OFLO)(NO3)2(CH3OH)] (NO3) [Nd(L-OFLO)(NO3)2(CH3OH)] (NO3), where L-OFLO = ofloxacin derivative. | -binding to BSA using fluorescence quenching and UV-VIS spectroscopy; -DNA binding tested through viscosity measurement, electronic absorption titration, ethidium bromide (EB) displacement experiments, CD spectra and cyclic voltammetry experiments; -DNA cleavage tested by electrophoresis; | -tests confirm the intercalative binding mode of the complexes; interaction between complexes and DNA is stronger than of the free ligand; -the complexes show a concentration dependent activity through hydrolytic pathways. | [55,58] |
| [Pr(L-OFLO)2(NO3)](NO3)2, where L-OFLO = ligand obtained from ofloxacin | -BSA binding tested through fluorescence quenching experiments; -CT-DNA binding tested through UV spectroscopy, UV titration experiments, competitive studies with EB, iodide quenching; -DNA cleavage tested through electrophoresis | -the complex binds to BSA with high affinity which induces a conformational change of BSA; -the complex binds to DNA more strongly than the free ligand; -the complex shows such activity. | [56] |
| Quinolone | Method (System, Optimum pH, Optimum Buffer Conditions) | λexitation/λemission | Limit of Detection/Limit ofQuantification (LOD/LOQ) | Matrix | Ref |
|---|---|---|---|---|---|
| Balofloxacin (BLFX) | BLFX-Eu3+-SDBS system; pH = 7.5-tris HCl buffer; | 335 nm/618 nm | LOD = 5 nM | Serum and urine samples | [111] |
| BLFX-Eu3+-SDBS system; pH = 7.5-tris HCl buffer; | 335 nm/618 nm | LOD = 1.3 nM | Bile Acid | [112] | |
| BLFX-Eu3+-KBrO3-Na2S2O4-SDBS chemiluminescence system | CL excitation/593 nm, 617 nm | LOD = 0.069 nM | Pharma-ceutical formulations and biological fluids | [113] | |
| BLFX-Eu3+-Y3+-SDBS system | 330 nm/618 nm | LOD = 0.83 nM | Pharma-ceutical formulations, human serum, urine | [114] | |
| Ciprofloxacin (CPX) | CPX-Tb.+3-TOPO system pH = 5.5-acetate buffer; micellar solution of CPCl | 333 nm/546 nm | LOD = 1.2 nM | Serum samples | [85] |
| CPX-Tb3+-SLS system; pH = 6- acetic acid- sodium acetate buffer; | 300-325 nm / 549nm | LOD = 30–150 mol/kg | Chicken and trout muscle sample | [115] | |
| CPX-Ad/Tb3+ CPNP nanoparticles; pH = 7.5-HEPES buffer; | 288 nm/545 nm | LOD = 60 nM | Aqueous solution, urine, tablets | [92] | |
| CPX-Tb3+-DO3A NPs; pH = 7.4-HEPES buffer; | 278 nm/542 nm | LOD = 9 nM | Urine | [84] | |
| CPX-Tb3+-TOPO system; pH = 6-piperazine andimidazole buffer; | 320 nm/545 nm | LOD = 100 nM | LC eluent | [116] | |
| CPX-Tb3+ system; pH = 5.7-potassium hydrogen phthalate buffer; (Additionally applied for: enrofloxacin and flumequine) | 337 nm/545 nm | LOD = 9.6 nM | Water samples | [117] | |
| CPX-Tb3+-SDS-Na2SO3 system; pH = 6.5-acetate buffer; | 271 nm/545 nm; | LOD = 1810.7 nM | Serum samples | [108] | |
| CPX-Tb3+-CTAB system; pH = 7.3-Tris buffer; in the presence of tetracycline. | 284 nm/545 nm | LOD = 27162 nM | Serum and urine samples | [118] | |
| CPX-Tb3+ -SDS system; pH = 5.7-sodium acetate buffer; | 278 nm/545 nm. | LOD = 422.5 nM; LOQ = 1508.9 nM. | Milk samples | [86] | |
| CPX-Tb3+-CeIV–SO32- chemiluminescence system | CL excitation/ 490 nm, 545 nm, 585 nm 620 nm | LOD = 0.31 nM | Capsules, human serum and urine samples | [119] | |
| CPX- [Tb(bpy)2]3+–K2S2O8; pH = 5.5- acetic acid– sodium acetate buffer | ECL excitation/ 490 nm, 545 nm, 585 nm 620 nm | LOD = 1.4 nM | Pharma-ceutical tablets | [120] | |
| CPX- Tb3+-calf thymus DNA; pH = 5.5- acetic acid– sodium acetate buffer | 272 nm/545 nm | LOD = 37.9 nM | eye-ear pharma-ceutical dosage forms | [121] | |
| CPX-Eu3+-phen system; pH = 9-acetate-ammonia buffer; micellar solution of DDBS. | 330 nm/615 nm | LOD = 230 nM | Pharma-ceutical tablets, blood serum | [91] | |
| CPX-Eu3+-GMP NPs; pH = 7.4-HEPES buffer; | 276 nm/615 nm | LOD = 780 nM | Pharma-ceutical tablets | [94] | |
| CPX-Eu3+ system in acetonitrile; pH = 6-borate buffer; | 365 nm/615 nm | LOD = 15 nM;LOQ = 45 nM | Pharma-ceutical tablets, serum | [122] | |
| CPX-Eu3+-Ag NPs; pH = 9.4-tris-HCl buffer | 373 nm/614 nm | LOD = 0.057 nM | Pharma-ceutical tablets, serum | [123] | |
| CPX-Eu3+ functionalized Ga(OH)(btec)·0.5H2O, where H4btec = 1,2,4,5-benzenetetra-carboxylic acid | 370 nm/614 nm | LOD = 7243 nM | Urine samples | [124] | |
| Danofloxacin (DAN) | DAN-Tb3+-SDS-Na2SO3 system; pH = 6.5-acetate buffer; | 271 nm/545 nm; | LOD = 1810.7 nM | Serum samples | [108] |
| Enoxacin (ENX) | ENX-Dy3+-Ce4+-S2O32--H2SO4 system; (Additionally applied for: lomefloxacin, ofloxacin, norfloxacin, gatifloxacin) | CL excitation/482 nm, 578 nm | LOD = 624.3 nM | Biological fluids | [109] |
| ENX- Dy3+- MnO4-- S2O32- -HNO3 system; | CL excitation / 482 nm, 578 nm | LOD = 686.8 nM | Biological fluids | [125] | |
| ENX-Tb3+-Na2SO3 system; pH = 7 | ECL excitation /490 nm, 545 nm, 585 nm, 620 nm. | LOD = 0.054 nM | Dosage forms, urine samples. | [106] | |
| ENX-Tb3+-KMnO4-Na2SO3 (Additionally applied for ofloxacin) | CL excitation /490 nm, 545 nm, 585 nm, 620 nm. | LOD = 0.24 nM | Dosage forms, urine sample | [126] | |
| ENX-Tb3+- acetylacetone NPs pH = 7.2, phosphate buffer | 345 nm/559 nm (FRET method) | LOD = 30 nM | Dosage forms, urine sample | [127] | |
| Enrofloxacin (EF) | EF-Eu3+-phen system; pH = 9- acetate-ammonia buffer; micellar solution of DDBS. | 330 nm/615 nm | LOD = 230 nM | Blood sample | [91] |
| EF-Eu3+-optical sensor pH = 7.8 | 395 nm/617 nm | LOD = 75 nM | Pharma-ceutical tablets and serum samples | [128] | |
| EF-Eu3+-polymer nanofilament-based optical sensor pH = 7.5, HEPES buffer | 340 nm/612 nm | LOD = 580 nM | Water-rich media | [129] | |
| EF-Tb3+-SLS system; pH = 6-acetic acid-sodium acetate buffer; | 300 nm- 325 nm/549 nm | LOD = 5.56 nM | Chicken and trout muscle sample | [115] | |
| EF-Tb3+-Ag NPs pH = 6, Tris–HCl buffer | 327 nm/545 nm | LOD = 58.4 nM LOQ = 191.9 nM | Milk samples | [130] | |
| EF-Tb3+-Fe(II)/(III)-H2O2 EF-Tb3+-KNO2-H2O2 (Additionally applied for levofloxacin and pefloxacin) | CL excitation/440 nm 515 nm | LOD = 120 nM LOD = 240 nM | Pharma-ceutical forms and urine samples | [131] | |
| EF-Tb3+-calf thymus DNA pH = 6.5-hexamethylenetetramine buffer | 324 nm/ 546 nm | LOD = 6.95 nM | Beef serum | [132] | |
| Fleroxacin (FLX) | FLX-Dy3+-MnO4−-S2O32−-H6P4O13 system; (Additionally applied for pefloxacin and pipemidic acid) | CL excitation/482 nm, 578 nm | LOD = 0.812 nM | Foods, biological samples | [104] |
| FLX-Dy3+-KMnO4-Na2S2O3-H6P4O13 | CL excitation/482 nm, 578 nm | LOD = 0.3 nM | Injections and urine sample | [133] | |
| FLX-Eu3+-[(Me)4N]2[Pb6K6(m-BDC)9(OH)2]·H2O, where 1,3-H2BDC = 1,3-benzenedicarboxylic acid) | 377 nm/612 nm | LOD = 118.8 nM | Human serum and urine | [134] | |
| FLX-Tb3+ photochemical fluorimetric system pH = 5.7, acetate buffer | 320 nm/545 nm | LOD = 12 nM | Human urine samples | [135] | |
| Flumequine (FLU) | FLU-Tb3+ Ce4+-Na2SO3-H2SO4 system; | Excitation by redox reaction/- | LOD = 382.7 nM; LOQ = 1148.1 nM; | Waste water samples | [105] |
| FLU-Tb3+-SDS micelles system; pH = 7.6-Tris buffer; | 340 nm/545 nm | LOD = 210.5 nM | Chicken muscle and liver, whole milk. | [136] | |
| FLU-Tb3+-1,10-phenanthroline in micellar SDS solutions pH = 7.5-acetate-ammonia buffer | 330 nm/545 nm | LOD = 4.9 nM | Chicken meat | [137] | |
| FLU-Eu3+-Tb3+-nanocomposites pH = 6.9 acetic acid buffered with ammonia | 255 nm/360 nm | LOD = 4.6 nM | Meat samples | [138] | |
| Garenoxacin (GAR) | GAR-Tb3+-SDS micelles-Na2SO3 system; pH = 4.1-acetate buffer; | 281 nm/546 nm | LOD = 46.9 nM; LOQ = 152.4 nM | Serum and urine samples | [89] |
| Gatifloxacin (GFLX) | GFLX- Eu3+ system in acetonitrile; pH = 3.5- acetate buffer; | 395 nm/617 nm | LOD = 16 nM; LOQ = 28 nM | Pharma- ceutical tablets, serum. | [122] |
| GFLX-Eu3+-SDBS system pH = 7.5-Tris–HCl buffer | 338 nm/617 nm | LOD = 1 nM | Injections and human urine/serum samples | [139] | |
| GFLX-Eu3+ system in sol- gel matrix; pH = 6 - borate buffer; | 370 nm/617 nm; | LOD = 0.16 nM | Pharma- ceutical and serum samples. | [140] | |
| GFLX-Tb3+ system in sol-gel matrix; pH = 3.5-acetate buffer; | 350 nm/545 nm; | LOD = 20 nM | Pharma- ceutical and serum samples. | [140] | |
| Grepafloxacin (GREP) | GREP-Tb3+-SDS micellar solution system; pH = 6-acetate buffer; | 275 nm/546 nm | LOD = 27824.5 nM; LOQ = 83473.5 nM | Human serum and urine | [88] |
| GREP-Tb3+-Ce(IV)–Na2SO3 pH = 4-15mM sodium dodecyl sulphate solution | 280 nm/450 nm | LOD = 27.8 nM | Pharmaceutical tablets and human urine | [141] | |
| Levofloxacin (LEVO) | LEVO-Tb3+ system; micellar solution of SDS; pH = 6-acetate buffer; | 292 nm/546 nm | LOD = 27672.6 nM; LOQ = 83017.8 nM | Pharma- ceutical tablets, human serum, urine. | [87] |
| LEVO-Tb3+ system; colloidal silver NPs; pH = 9-tris-HCl buffer; | 284 nm/545 nm | LOD = 7.19x10-9 nM | Serum samples, urine | [95] | |
| LEVO-Tb3+-gold NPs system; pH = 6.8 NaH2PO4/Na2HPO4 buffer; | 373 nm/545 nm | LOD = 0.21 nM; LOQ = 0.72 nM | Powder, tablets. | [99] | |
| LEVO-Sm3+ system; pH = 6-acetate buffer; | 312 nm/553 nm | LOD = 52.5 nM; LOQ = 154.7 nM | Powder, tablets | [142] | |
| LEVO-Eu3+-Ce4+-SO32−-H2SO4 system; (Additionally applied for moxifloxacin and trovafloxacin) | - | LOD = 276.7 nM; LOQ = 968.45 nM | Tablets | [110] | |
| LEVO-Eu3+-covalent organic framework-based hybrid material | 380 nm/613 nm | LOD = 200 nM | Serum and urine samples | [143] | |
| LEVO-Ce(IV) in micellar solutions of of cetyltrimethyl ammonium bromide | 250 nm/355 nm (FRET method) | - | Injections | [144] | |
| Lomefloxacin (LOM) | LOM-Sm3+ system; pH = 6- acetate buffer; | 310 nm/556 nm | LOD = 65.4 nM; LOQ = 199 nM | Powder, tablets. | [142] |
| LOM-Tb3+-silver NPs system; pH = 6-acetate buffer; | 274 nm/545 nm | LOD = 0.11 nM; | Tablets, serum, urine samples. | [98] | |
| LOM-Tb3+-Ce(IV)-Na2SO3 pH = 2-H2SO4 | CL excitation/490 nm, 545 nm, 585 nm, 620 nm | LOD = 1.1 nM | Pharmaceutical tablets, urine and serum samples | [145] | |
| Marbofloxacin (MAR) | MAR-Tb3+ system; pH = 6.0-acetic acid/ ammonium acetate buffer; (Additionally applied for: ciprofloxacin, danofloxacin, enrofloxacin, sarafloxacin, difloxacin, oxolinic acid, flumequine) | FL: 340 nm/545 nm; TR: 281 nm/545 nm | LOD = 165.56 nM (FL); LOD = 262.17 nM (TR); | Whole, semi-skimmed, skimmed milk. | [146] |
| MAR-Tb4O7 NPs-TOPO-SDS-hexamine system; pH = 7.0–7.5-Tris buffer; (Applied also for: ciprofloxacin, danofloxacin, enrofloxacin, sarafloxacin, oxolinic acid, fumequine). | 340 nm/545 nm. | LOD = 96.59 nM | Skimmed, semi- skimmed, whole milk samples. | [97] | |
| Moxifloxacin (MOXI) | MOXI-Tb3+ system; colloidal silver NPs; pH = 9-Tris-HCl buffer; | 284 nm/545 nm | LOD = 8.47 × 10−9 nM | Serum samples, urine | [95] |
| Norfloxacin (NOR) | NOR-Tb3+ -TOPO system pH = 5.5- acetate buffer; micellar solution of CPCl | 333 nm/490 nm | LOD = 1.7 nM | Serum samples | [85] |
| Marbofloxacin (MAR) | MAR-Tb3+ system; pH = 6.0-acetic acid/ ammonium acetate buffer; (Additionally applied for: ciprofloxacin, danofloxacin, enrofloxacin, sarafloxacin, difloxacin, oxolinic acid, flumequine) | FL: 340 nm/545 nm; TR: 281 nm/ 545 nm | LOD = 165.56 nM (FL); LOD = 262.17 nM (TR); | Whole, semi-skimmed, skimmed milk. | [146] |
| MAR-Tb4O7 NPs-TOPO-SDS-hexamine system; pH = 7.0–7.5-Tris buffer; (Applied also for: ciprofloxacin, danofloxacin, enrofloxacin, sarafloxacin, oxolinic acid, fumequine). | 340 nm/545 nm. | LOD = 96.59 nM | Skimmed, semi-skimmed, whole milk samples. | [97] | |
| Moxifloxacin (MOXI) | MOXI-Tb3+ system; colloidal silver NPs; pH = 9-Tris-HCl buffer; | 284 nm/545 nm | LOD = 8.47 × 10−9 nM | Serum samples, urine | [95] |
| Norfloxacin (NOR) | NOR-Tb3+-TOPO system pH = 5.5-acetate buffer; micellar solution of CPCl | 333 nm/490 nm | LOD = 1.7 nM | Serum samples | [85] |
| NOR-Tb3+-calf thymus DNA; pH = 5.5-acetic acid–sodium acetate buffer | 272 nm/545 nm | LOD = 35.8 nM | Eye-ear pharma-ceutical dosage forms | [121] | |
| NOR-Tb3+-zeolite of the CaA-type pH = 7-buffered with a 40% aqueous solution of urothropine (Additionally applied for ciprofloxacin) | 365 nm/545 nm | LOD = 3131 nM | Urine and human plasma samples | [147] | |
| NOR-Tb3+-Na2SO3 system pH = 7 | ECL excitation/490 nm, 545 nm, 585 nm, 620 nm | LOD = 0.028 nM | Pharmaceutical capsules, urine samples | [148] | |
| NOR-Sm3+ system; pH = 6-acetate buffer; | 314 nm/553 nm | LOD = 84.5 nM; LOQ = 253.6 nM; | Powder, tablets | [142] | |
| NOR-Tb3+ sodium tetradecylsulfate system; pH = 7–8; | 337 nm/545 nm | LOD = 0.0031 nM | Used in HELC | [149] | |
| NOR-Ce4+ system; Acidic pH-perchloric acid 0.02-0.08 mol/L; | -/550 nm | LOD = 31.3 nM; LOQ = 93.9 nM | Pharma-ceutical capsules, eye drops, urine | [100] | |
| NOR-Tb3+(bipy)2 (DPA)-K2S2O8 system; pH = 10-NaBO4-NaOH buffer; | ECL excitation /485 nm, 545 nm, 582 nm, 621 nm | LOD = 0.69 nM | Urine samples | [101] | |
| NOR- PEG coated Tb3+ doped ZnS NPs system; pH = 7.6-tris-HCl buffer; | 334 nm/491 nm, 545 nm | LOD = 0.05 nM; LOQ = 0.17 nM | Eye drops, urine samples | [96] | |
| NOR-Tb3+ system flow through solid phase system; pH = 5.6-acetate buffer; | 273 nm/545 nm. | LOD = 4.7 nM; LOQ = 15.7 nM | Serum and urine samples. | [150] | |
| NOR-Tb3+-KMnO4 -Na2SO3 system-optical flow-through sensor; pH = 3-KH phthalate-HCl buffer; | CL excitation/545 nm | LOD = 8.7 nM | Pharma-ceutical samples | [102] | |
| NOR-AMP-Tb3+ CPNSs system; pH = 7.5; (Additionally applied for: pefloxacin, sparfloxacin, fleroxacin, nalidixic acid as well.) | 280 nm/545 nm | LOD = 10 nM | Milk | [93] | |
| NOR-Tb3+-SDS-Na2SO3 system; pH = 6.5-acetate buffer; | 271 nm/545 nm; | LOD = 1.88 nM | Serum samples. | [108] | |
| NOR-Tb3+ system in sol-gel matrix; pH = 3.5-acetate buffer; | 395 nm/545 nm; | LOD = 10 nM | Pharma-ceutical and serum samples. | [140] | |
| NOR- Eu3+ system in sol-gel matrix; pH = 6- borate buffer; | 340 nm/617 nm; | LOD = 3.0 nM | Pharma-ceutical and serum samples. | [140] | |
| NOR fluorescence immunoassay coating-antigen-modified polystyrene particles - anti-norfloxacin monoclonal antibody conjugated with carboxyl-functionalized NaYF4:Yb,Er upconversion NPs pH = 7.4-phosphate-buffered saline | 980 nm/542 nm | LOD = 0.03 nM | Milk, chicken, pork kidney samples | [151] | |
| Ofloxacin (OFLO) | OFLO-Tb3+-TTDC; pH = 7–8; | 337 nm/545 nm | LOD = 0.0000276 nM | Used in HELC | [149] |
| Ce4+-Na2SO3-OFLO-Tb3+ system; Acidic pH- HCl. Flow-injection coupled with CL detection | - | LOD = 20.75 nM | Plasma samples | [152] | |
| OFLO-Ru(bipy)2(CIP)2+ Ce4+ system; Acidic pH -HNO3 | CL excitation / 617 nm, 624 nm | LOD = 4.2 nM | Pharma-ceutical samples, urine samples | [103] | |
| OFLO-Eu3+ system; pH = 5.1- acetate buffer. | 365 nm/617 nm | LOD = 3 nM; LOQ = 9 nM | Pharma-ceutical and serum samples | [153] | |
| OFLO-Tb3+-Na2SO3 system; pH = 7; | ECL excitation /490 nm, 545 nm, 585 nm, 620 nm. | LOD = 0.16 nM | Dosage forms, urine samples. | [106] | |
| Orbifloxacin (ORBI) | ORBI-Tb3+ system; pH = 6-acetate buffer; | 275 nm/545 nm | LOD = 8.35 nM; LOQ = 25.3 nM | Tablets, urine samples | [154] |
| Pazufloxacin (PAZ) | PAZ- Tb3+ system; pH = 6.3- acetate buffer; | 330 nm/545 nm | LOD = 6.2 nM | Serum and urine samples | [155] |
| PAZ-Eu3+-KMnO4- Na2S2O4 system; pH = 6. | CL excitation/592 nm, 617 nm, 695 nm. | LOD = 2.6 nM | Serum and urine samples. | [156] | |
| PAZ- Ce4+- Na2SO3- H2SO4 system; | -/ 247 nm | LOD = 2.2 nM | Urine samples | [157] | |
| Pefloxacin (PEF) | PEF-Tb3+-TOPO system; pH = 5.5-acetate buffer; micellar solution of CPCl | 333 nm/590 nm | LOD = 4.4 nM | Serum samples | [85] |
| PEF-Tb3+- KMnO4/ H2SO3 system; Acidic pH-H2SO4; | CL excitation /543 nm | LOD = 41.1 nM | Semi-skimmed milk samples | [107] | |
| PEF-Tb3+-Ag NPs pH = 6-acetate buffer | 273 nm/545 nm | LOD = 25 nM | Pharmaceutical capsules and serum samples | [158] | |
| Pipemidic acid (PPA) | PPA-Tb3+-Ag+ NPs system; pH = 6-acetate buffer; | 320 nm/545 nm. | LOD = 0.047 nM | Tablets, serum, urine samples. | [98] |
| PPA-Tb3+ system; pH = 5.6- acetate buffer; | 320 nm/545 nm. | LOD = 5.9 nM | Urine and serum samples | [159] | |
| Prulifloxacin (PUFX) | PUFX-Tb3+-KMnO4-Na2S2O4 system; pH = 5.8; (Additionally applied for ulifloxacin –pH = 5.4) | CL excitation/490 nm, 545 nm, 585 nm, 620 nm. | LOD = 7 nM | Tablets | [160] |
| PUFX-Tb3+-KMnO4-Na2SO3 KH2PO4-NaOH buffer | 275 nm/423 nm | LOD = 8 nM | Pharmaceutical tablets, serum, and urine samples | [161] | |
| Ulifloxacin (UFX) | UFX-Eu3+-SDBS system; pH = 8.6-NH4Cl/NH3x H2O buffer; | 276 nm/616 nm. | LOD = 0.2 nM | Human serum and urine | [162] |
| Ion | Quinolone | Conditions | λabsorption | LOD/LOQ | Ref |
|---|---|---|---|---|---|
| Er3+ | Norfloxacin | -Cetylpyridinium chloride; -pH = 9.35-NH4Cl/NH3 buffer; | 515 (+) nm, 517 (−) nm (second derivative spectrum) | LOD = 6.6 × 10−6 mol/L | [164] |
| Eu3+ | Gatifloxacin | -SDBS micelle solution, La3+; -pH = 7-NH4Ac-NH3*H2O buffer; | λex = 336 nm/ λem = 616 nm | LOD = 7 × 10−14 mol/L | [165] |
| Ho3+ | Norfloxacin | -Cetylpyridinium chloride; -pH = 9.35-NH4Cl/ NH3 buffer; | 444 (+) nm, 446 (−) nm (second derivative spectrum) | LOD = 6.6 × 10-6 mol/L | [164] |
| Nd3+ | Norfloxacin | -Cetylpyridinium chloride; -pH = 9.35-NH4Cl/ NH3 buffer; | 570 (+) nm, 568 (−) nm (second derivative spectrum) | LOD = 6.7 × 10−6 mol/L | [164] |
| Cu2+ | 1-methyl-4-hydroxy- 3-(N-2-ethyl-5- aminothiadazolyl-)- carbamoyl- quinoline-2-one | -pH = 7-MOPS buffer; | λex = 320 nm/ λem = 547 nm | LOD = 4.3 × 10−9 mol/L | [166] |
| Sc3+ | Nalidixic acid | -Method used for the extraction of Sc3+; -pH = 3.4; -dicholoromethane; | SF = 1.4 × 104 (from Eu3+); SF = 3.6 × 103 (from Nd3+) | [167] | |
| H2PO4− HPO42− | 4-Hydroxy-1-methyl-2-oxo-1,2-dihydro-quinoline-3-carboxylic acid-ethyl-[1,3,4]thiadiazol-2-yl) amide | -pH = 7.4-HEPES buffer; -Tb3+ complex-Triton X-100 system; | λex = 320 nm/ λem = 548 nm | LOD = 110 × 10−9 mol/L | [168] |
| Type of Nucleic Acid | Ligand (L) | Lanthanide Ion (Ln3+) | Conditions (pH, L:Ln3+) | λex/λem | LOD/LOQ | Ref |
|---|---|---|---|---|---|---|
| fs-DNA | Quinolone derivative | Tb3+ | -pH = 9-Tris- HCl buffer; -1:1; | 340 nm/545 nm | 12 ng/mL; | [172] |
| ct-DNA | Quinolone derivative | Tb3+ | -pH = 9-Tris-HCl buffer; -1:1; | 340 nm/545 nm | 10 ng/mL | [172] |
| ct-DNA | Danoflaxacin | Tb3+ | -pH = 7.8-Tris- HCl buffer; | 347 nm/545 nm | 8 ng/mL | [180] |
| hs-DNA | Prulifloxacin | Tb3+ | -pH = 6.3-Tris-HCl buffer; -2:1; | 345 nm/545 nm | 2.1 ng/mL | [171] |
| cf-DNA | Gatifloxacin | Eu3+ | -pH = 6.5-HMA-HCl buffer; -3.1:1; | 331 nm/617 nm | 6 × 10−9 g/mL | [174] |
| ds-DNA | Difloxacin | Tb3+ | -pH = 7.4-MOPS buffer; -1:1; | 340 nm/545 nm | 0.5 ng/mL | [175] |
| ss-DNA | Difloxacin | Tb3+ | -pH = 7.4-MOPS buffer; -1:1; | 340 nm/545 nm | 2 ng/mL | [175] |
| Oligo-nucleotides | Cs124-TPA- Tb NPs | Tb3+ | -pH = 7.5; | 328 nm/546 nm | 8 × 10−11 mol/L | [179] |
| Oligo- nucleotides | DTPA-cs214-CF3-NCS; DTPA-cs124-NCS | Tb3+, Eu3+, Dy3+, Sm3+ | 0.5–1 × 10−12 mol/L | [177] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Măciucă, A.-M.; Munteanu, A.-C.; Uivarosi, V. Quinolone Complexes with Lanthanide Ions: An Insight into their Analytical Applications and Biological Activity. Molecules 2020, 25, 1347. https://doi.org/10.3390/molecules25061347
Măciucă A-M, Munteanu A-C, Uivarosi V. Quinolone Complexes with Lanthanide Ions: An Insight into their Analytical Applications and Biological Activity. Molecules. 2020; 25(6):1347. https://doi.org/10.3390/molecules25061347
Chicago/Turabian StyleMăciucă, Ana-Mădălina, Alexandra-Cristina Munteanu, and Valentina Uivarosi. 2020. "Quinolone Complexes with Lanthanide Ions: An Insight into their Analytical Applications and Biological Activity" Molecules 25, no. 6: 1347. https://doi.org/10.3390/molecules25061347
APA StyleMăciucă, A.-M., Munteanu, A.-C., & Uivarosi, V. (2020). Quinolone Complexes with Lanthanide Ions: An Insight into their Analytical Applications and Biological Activity. Molecules, 25(6), 1347. https://doi.org/10.3390/molecules25061347








